Abstract
A study recently published in EMBO reports solves the solution structure of E. coli BamE, thus providing the basis for a better understanding of the mechanism of β-barrel assembly in bacterial and mitochondrial outer membranes.
EMBO Rep (2011) advance online publication. doi: 10.1038/embor.2010.202
β-barrel membrane proteins are found exclusively in the outer membrane of Gram-negative bacteria and the outer membranes of eukaryotic organelles of prokaryotic origin, mitochondria and chloroplasts. In contrast to the inner membrane, the bacterial outer membrane is an asymmetrical bilayer that consists mainly of lipopolysaccharides in the outer leaflet and phospholipids in the inner leaflet. Bacterial β-barrel outer membrane proteins (OMPs) mediate many cellular functions, for example, passive or selective diffusion of small molecules through the β-barrel pores across the outer membrane. By contrast, only a few mitochondrial β-barrel outer membrane proteins (MBOMPs) have been identified so far. The central machineries that mediate insertion and assembly of OMPs/MBOMPs are the β-barrel assembly machine (BAM) complex in the bacterial outer membrane and the topogenesis of outer-membrane β-barrel proteins (TOB)/sorting and assembly machinery (SAM) complex in the mitochondrial outer membrane (Knowles et al, 2009; Endo & Yamano, 2010; Stroud et al, 2010; Fig 1). However, the molecular mechanisms of β-barrel protein topogenesis in bacterial and mitochondrial outer membranes remain poorly understood.
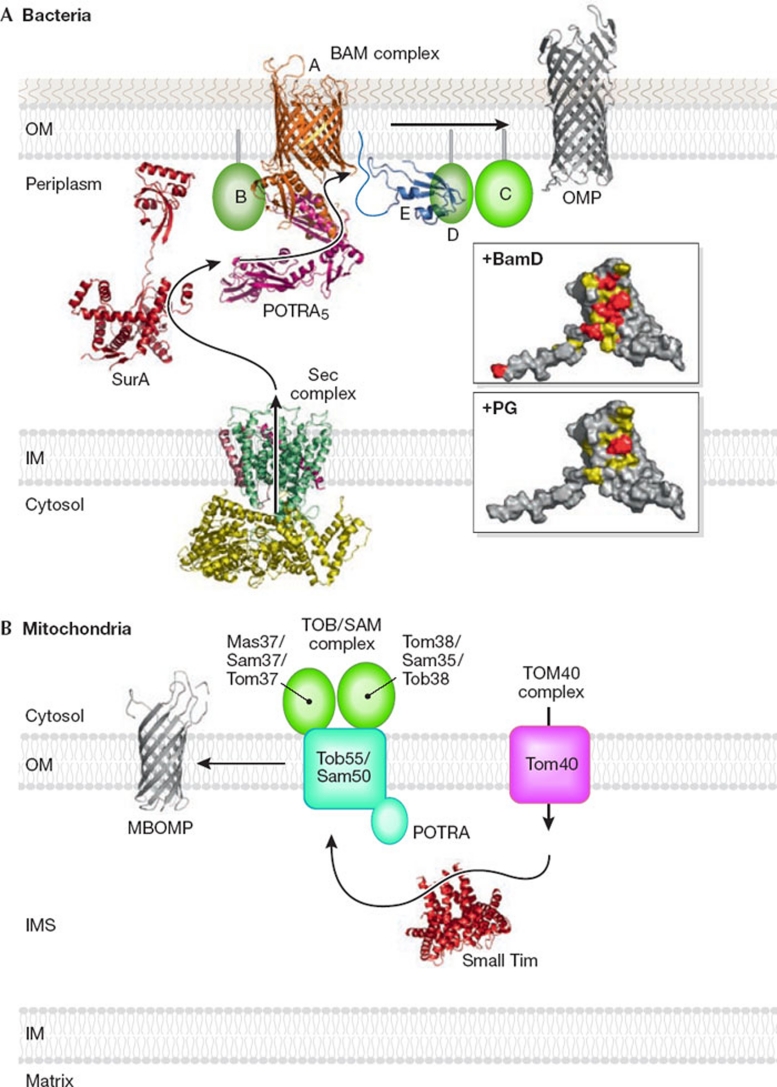
Figure 1.
β-barrel protein assembly in bacterial and mitochondrial outer membranes. (A) Bacteria. Ribbon models of the structures of the Sec complex, SurA, BamA (Clantin et al, 2007; Kim et al, 2007), BamE and OMP. The upper and lower inserts show the surface of BamE (residues 20–108; viewed after approximately 90° rotation of the ribbon model around the horizontal axis toward the reader). Residues important for BamD binding are shown in red and residues with NMR signals that were perturbed by BamD binding are shown in yellow. The residue (Phe 74) important for PG binding is shown in red and the residues with NMR signals that were perturbed by PG binding are shown in yellow. (B) Mitochondria. Ribbon models were drawn for the structures of small Tim and MBOMP. IM, inner membrane; IMS, intermembrane space; MBOMP, mitochondrial β-barrel outer membrane protein; OM, outer membrane; OMP, outer membrane protein; PG, phosphatidylglycerol; POTRA, polypeptide transport-associated domain.
Bacterial OMPs are synthesized in the cytosol as precursor proteins with an amino-terminal signal sequence that guides the proteins to the Sec machinery for crossing the inner membrane and is cleaved off in the periplasm. Periplasmic chaperones then escort OMPs through the aqueous periplasmic space in a partly unfolded state. On reaching the outer membrane, OMPs assemble into a β-barrel structure and insert into the outer membrane with the help of the BAM complex. The bacterial OMP insertion pathway can be compared to the assembly pathway of MBOMPs from the mitochondrial intermembrane space into the outer membrane. MBOMPs are synthesized in the cytosol and imported into the intermembrane space by the outer membrane translocator TOM40. The subsequent chaperone-mediated escort across the intermembrane space and insertion into the outer membrane by the TOB complex is similar to the OMP assembly process. Notably, the BAM and TOB complexes share the homologous β-barrel proteins BamA and Tob55/Sam50, respectively, as the central components of their insertion machineries. The BAM complex in Escherichia coli consists of BamA (YaeT/Omp85) and four accessory lipoproteins: BamB (YfgL), BamC (NlpB), BamD (YfiO) and BamE (SmpA). BamA and BamD are essential for cell growth, yet deletion of dispensable BamB, BamC or BamE leads to outer membrane defects manifested in hypersensitivity to antibiotics. Although BamAB and BamCDE can form distinct subcomplexes, they become functional only after formation of the entire BAM complex with all five subunits (Hagan et al, 2010).
In this issue of EMBO reports, Knowles et al (2011) solve the nuclear magnetic resonance (NMR) solution structure of E. coli BamE, which sheds light on the roles of one of the Bam subunits in β-barrel protein assembly. The structure of BamE consists of a three-stranded antiparallel β-sheet packed against a pair of α-helices (Fig 1).
As the ΔbamE mutant cannot grow in the presence of vancomycin, the authors identify functionally important residues of BamE by testing the effects of amino-acid substitutions in BamE on its inability to complement the growth defects of ΔbamE, without destabilizing BamE itself. Many of the identified residues are conserved among BamE proteins from different organisms and map to a single surface area on BamE. Interestingly, NMR signals of the residues around this region are sensitive to the addition of micelles containing the lipid phosphatidylglycerol, but not phosphatidylethanolamine or cardiolipin. In parallel, the authors analyse perturbation of the NMR spectra of BamE after the addition of purified BamB, C and D proteins. Only BamD affects the NMR spectra of BamE, and the BamD interacting region of BamE is found to overlap partly with the residues involved in phosphatidylglycerol binding. As the addition of BamD and phosphatidylglycerol have different effects on the NMR spectra of BamE, the binding of BamD and phosphatidylglycerol to BamE seem to take place simultaneously. What is the biological relevance of the observed interactions of BamE with both BamD and phosphatidylglycerol? As phosphatidylglycerol was found to help the insertion of OMPs into lipid liposomes (Patel et al, 2009), BamE might recruit the BAM complex through BamD to phosphatidylglycerol-rich regions in the outer membrane, or might directly recruit phosphatidylglycerol to form assembly points for OMP insertion and folding.
What are the roles of other subunits of the BAM complex in β-barrel protein assembly? The essential subunit of the E. coli BAM complex BamA consists of two domains: the N-terminal polypeptide transport-associated (POTRA) domain repeat in the periplasm and the carboxy-terminal β-barrel domain, embedded in the outer membrane. The number of POTRA domains ranges from one to five in BamA homologues from different organisms. Of these POTRA domains, the one nearest to the C-terminal that is most connected to the β-barrel domain is essential for cell viability and its deletion leads to disassembly of the BAM complex (Kim et al, 2007). Structural studies of the E. coli BamA POTRA domains suggest that each POTRA domain has a common fold, whereas conformational rigidity might differ between inter-domain linkers (Gatzeva-Topalova et al, 2010; Fig 1). As individual POTRA domains have some affinity for unfolded substrate proteins, the periplasmic tandem POTRA repeat probably provides several substrate binding sites that slide the substrate progressively towards the BamA β-barrel domain. The β-barrel domain of BamA probably functions as a scaffold to facilitate the formation of β-strands, possibly through β-augmentation and subsequent spontaneous membrane insertion of the β-barrel. Yet, it is not clear whether this cradle for β-strand formation is provided by the pore formed within the monomer or oligomeric forms of the BamA β-barrel domain. Alternatively, membrane insertion and folding of OMPs might take place at the interface between BamA and the outer membrane lipid bilayer.
How much of the β-barrel assembly process is conserved during the evolution of mitochondria from Gram-negative bacteria? Although the central subunits BamA and Tob55 of the BAM and TOB complexes are conserved, other subunits of these complexes are unrelated to each other. The POTRA domains of BamA are essential for recognition and assembly of bacterial OMPs, whereas that of Tob55 is dispensable for MBOMP assembly in the mitochondrial outer membrane. Nevertheless, the mitochondrial TOB complex facilitates assembly of bacterial OMPs at low efficiency (Walther et al, 2009) and, in turn, the bacterial BAM complex can mediate assembly of mitochondrial porin. Therefore, the basic mechanism of β-barrel assembly in the outer membranes of bacteria and mitochondria seems to be conserved. High-resolution structures of each component of the BAM and TOB complexes—including that of BamE in this study—will thus provide the basis for a better understanding of the mechanism of β-barrel assembly in evolutionarily related bacterial and mitochondrial outer membranes.
References
- Clantin B et al. (2007) Science 317: 957–961 [DOI] [PubMed] [Google Scholar]
- Endo T, Yamano K (2010) Biochim Biophys Acta 1803: 706–714 [DOI] [PubMed] [Google Scholar]
- Gatzeva-Topalova PZ et al. (2010) Structure 18: 1492–1501 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hagan CL et al. (2010) Science 327: 890–892 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim S et al. (2007) Science 317: 961–964 [DOI] [PubMed] [Google Scholar]
- Knowles TJ et al. (2009) Nat Rev Microbiol 7: 206–214 [DOI] [PubMed] [Google Scholar]
- Knowles TJ et al. (2011) EMBO Rep [Epub 7 January 2011] doi:10.1038/embor.2010.202 [Google Scholar]
- Patel GJ et al. (2009) Biochemistry 48: 10235–10245 [DOI] [PubMed] [Google Scholar]
- Stroud DA et al. (2010) Science 328: 831–832 [DOI] [PubMed] [Google Scholar]
- Walther DM et al. (2009) Proc Natl Acad Sci USA 106: 2531–2536 [DOI] [PMC free article] [PubMed] [Google Scholar]