Abstract
In the title zwitterionic compound, C15H19NO2S, the six-membered heterocycle adopts a sofa conformation. The negative charge is delocalized along the carbonyl and enolate system on the ring and the positive charge is localized on the S atom. Two intermolecular C—H⋯O interactions help to establish the packing.
Related literature
For background to the synthesis of chiral non-racemic zwitterionic 4-alkoxy-3-sulfonium ylide pyridine-2-ones, see: Zang et al. (2008 ▶); Kappe et al. (1983 ▶); Palillero et al. (2009 ▶). For the biological activity of related structures, see: Basco et al. (1994 ▶); Koruzňjak et al., 2003 ▶). For ring conformation analysis, see: Cremer & Pople (1975 ▶).
Experimental
Crystal data
C15H19NO2S
M r = 277.37
Orthorhombic,

a = 5.9860 (17) Å
b = 7.4050 (14) Å
c = 31.589 (5) Å
V = 1400.2 (5) Å3
Z = 4
Mo Kα radiation
μ = 0.23 mm−1
T = 293 K
0.5 × 0.4 × 0.2 mm
Data collection
Siemens P4 diffractometer
Absorption correction: ψ scan (North et al., 1968 ▶) T min = 0.728, T max = 0.846
3016 measured reflections
2683 independent reflections
1928 reflections with I > 2σ(I)
R int = 0.045
3 standard reflections every 97 reflections intensity decay: 3%
Refinement
R[F 2 > 2σ(F 2)] = 0.061
wR(F 2) = 0.153
S = 1.03
2683 reflections
172 parameters
H-atom parameters constrained
Δρmax = 0.63 e Å−3
Δρmin = −0.39 e Å−3
Absolute structure: Flack (1983 ▶), 532 Friedel pairs
Flack parameter: −0.01 (16)
Data collection: XSCANS (Siemens, 1994 ▶); cell refinement: XSCANS; data reduction: XSCANS; program(s) used to solve structure: SIR2004 (Burla et al., 2005 ▶); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008 ▶); molecular graphics: ORTEP-3 for Windows (Farrugia, 1997 ▶); software used to prepare material for publication: WinGX (Farrugia, 1999 ▶).
Supplementary Material
Crystal structure: contains datablocks global, I. DOI: 10.1107/S1600536810052955/bt5438sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S1600536810052955/bt5438Isup2.hkl
Additional supplementary materials: crystallographic information; 3D view; checkCIF report
Table 1. Hydrogen-bond geometry (Å, °).
| D—H⋯A | D—H | H⋯A | D⋯A | D—H⋯A |
|---|---|---|---|---|
| C7—H7C⋯O2i | 0.96 | 2.36 | 3.315 (6) | 172 |
| C15—H15A⋯O1ii | 0.96 | 2.38 | 3.167 (5) | 138 |
Symmetry codes: (i)  ; (ii)
; (ii)  .
.
Acknowledgments
We are grateful to CONACyT (Project 83185) for financial support. PGG also thanks CONACyT for a scholarship (169011). Special thanks go to Dr Marcos Flores (USAI-FQ-UNAM) for useful comments.
supplementary crystallographic information
Comment
The synthesis of chiral non racemic zwitterionic 4-alkoxy-3-sulfonium ylide pyridine-2-ones is an original area of interest in organic chemistry (Zang et al., 2008; Kappe et al., 1983) because they are useful for the synthesis of piperidine-2,4-dione and pyridine-2-one (Palillero et al., 2009) compounds and because of their interesting biological properties (Basco et al., 1994; Koruzňjak et al., 2003).
The title compound I, features a zwitterionic molecule. The chiral centre shows an R configuration with [α]D= +70.5. The six member ring N1/C1/C2/C3/C4/C5 shows an sofa conformation with puckering parameters (Cremer & Pople, 1975) Q = 0.465 (4) Å, θ2 = 119.7 (5)°, φ2 = 103.2 (6)°, q2 = 0.404 (4) Å and q3 = -0.231 (4) Å. The bond distances of N1—C1, N1—C5, C5—C4 and C4—C3 show typical values, so that C2—C3 distance shows a single double bond (1.415 (5) Å), while C1—C2 (1.455 (5) Å) distance is shorter than common sp3—sp3 bonds, furthermore C3—O2 (1.244 (5) Å) and C1—O1 (1.250 (4) Å) distances are longer than related enolates and amide groups respectively these values suggest that negative charge is delocalized over O1/C1/C2/C3/O2 system and in the sulfonium group is located the positive charge in the zwitterion. Crystal packing is stabilized by two weak intermolecular C —H···O interactions.
Experimental
The title compound, was obtained by an intramolecular cyclization reaction of (1'R)-(+)-{[(2-methoxycarbonyl-ethyl)-(1'-phenyl-ethyl)-carbamoyl]-methyl}-dimethyl-sulfonium; bromide (1 mmol), which was dissolved in CH3CN (10 mL), treated with KOH (1.2 mmol) and stirred for two hours at room temperature. The resulting mixture was concentrated in vacuum and dissolved in ethyl acetate, filtered and concentrated giving the desired compound in 95%. Crystals were obtained from an ethyl acetate/diethylether solution; m.p. 139–140°C, [α]D= +70.5 (c 1.1, MeOH). IR (KBr) 1656 cm-1. 1H NMR (400 MHz, CDCl3) δ (p.p.m., J Hz): 1.51 (d, J = 7.2, 3H, Me), 2.32 (m, 2H), 2.90 (m, 1H), 2.98 (s, 3H, SMe), 3.00 (s, 3H, SMe), 3.16 (m, 1H), 5.94 (q, J =7.2, 1H), 7.27–7.40 (m, 5H). 13C NMR (100 MHz, CDCl3) δ p.p.m. 15.4, 26.0, 26.3, 36.3, 37.6, 48.9, 74.3, 126.5–141.0, 166.2, 187.5. HRMS (FAB): Calcd for C15H19NO2S: 277.1124. Found: 277.1103.
Refinement
H atoms linked to C atoms were placed in geometrical idealized positions and refined as riding on their parent atoms, with C—H = 0.93–0.98 Å and with Uiso(H) = 1.2 Ueq(C) or Ueq(H) = 1.5 Ueq(C) for methyl groups.
Figures
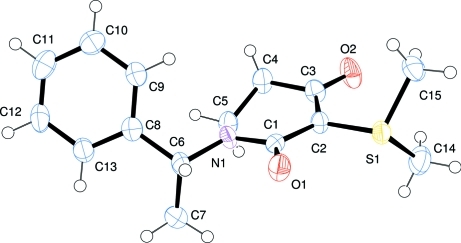
Fig. 1.
The molecular structure of the title compound with 50% probability displacement ellipsoids for non-H atoms.
Crystal data
| C15H19NO2S | F(000) = 592 |
| Mr = 277.37 | Dx = 1.316 Mg m−3 |
| Orthorhombic, P212121 | Mo Kα radiation, λ = 0.71073 Å |
| Hall symbol: P 2ac 2ab | Cell parameters from 46 reflections |
| a = 5.9860 (17) Å | θ = 21.3–35.1° |
| b = 7.4050 (14) Å | µ = 0.23 mm−1 |
| c = 31.589 (5) Å | T = 293 K |
| V = 1400.2 (5) Å3 | Prism, colorless |
| Z = 4 | 0.5 × 0.4 × 0.2 mm |
Data collection
| Siemens P4 diffractometer | Rint = 0.045 |
| graphite | θmax = 29.0°, θmin = 2.6° |
| 2θ/ω scans | h = −1→8 |
| Absorption correction: ψ scan (North et al., 1968) | k = −1→10 |
| Tmin = 0.728, Tmax = 0.846 | l = −43→1 |
| 3016 measured reflections | 3 standard reflections every 97 reflections |
| 2683 independent reflections | intensity decay: 3% |
| 1928 reflections with I > 2σ(I) |
Refinement
| Refinement on F2 | Secondary atom site location: difference Fourier map |
| Least-squares matrix: full | Hydrogen site location: inferred from neighbouring sites |
| R[F2 > 2σ(F2)] = 0.061 | H-atom parameters constrained |
| wR(F2) = 0.153 | w = 1/[σ2(Fo2) + (0.0631P)2 + 1.1965P] where P = (Fo2 + 2Fc2)/3 |
| S = 1.03 | (Δ/σ)max < 0.001 |
| 2683 reflections | Δρmax = 0.63 e Å−3 |
| 172 parameters | Δρmin = −0.39 e Å−3 |
| 0 restraints | Absolute structure: Flack (1983), 532 Friedel pairs |
| Primary atom site location: structure-invariant direct methods | Flack parameter: −0.01 (16) |
Special details
| Geometry. All s.u.'s (except the s.u. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell s.u.'s are taken into account individually in the estimation of s.u.'s in distances, angles and torsion angles; correlations between s.u.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell s.u.'s is used for estimating s.u.'s involving l.s. planes. |
| Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2, conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > 2σ(F2) is used only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2 are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
| x | y | z | Uiso*/Ueq | ||
| S1 | 1.10027 (19) | 0.87732 (14) | 0.03372 (3) | 0.0323 (2) | |
| O1 | 0.8859 (6) | 0.5383 (4) | 0.05505 (8) | 0.0393 (7) | |
| N1 | 0.7630 (6) | 0.6122 (4) | 0.12114 (10) | 0.0300 (7) | |
| O2 | 1.0851 (7) | 1.1013 (4) | 0.11356 (10) | 0.0511 (9) | |
| C9 | 0.8558 (7) | 0.3759 (6) | 0.19114 (12) | 0.0351 (9) | |
| H9 | 0.9723 | 0.4434 | 0.1798 | 0.042* | |
| C1 | 0.8724 (7) | 0.6511 (5) | 0.08437 (10) | 0.0278 (8) | |
| C6 | 0.6341 (8) | 0.4423 (5) | 0.12418 (12) | 0.0345 (10) | |
| H6 | 0.7011 | 0.3572 | 0.1041 | 0.041* | |
| C4 | 0.8971 (9) | 0.8843 (6) | 0.15731 (11) | 0.0373 (9) | |
| H4A | 1.0124 | 0.8199 | 0.1727 | 0.045* | |
| H4B | 0.8501 | 0.9863 | 0.1744 | 0.045* | |
| C3 | 0.9911 (8) | 0.9518 (6) | 0.11576 (12) | 0.0340 (9) | |
| C8 | 0.6557 (7) | 0.3579 (5) | 0.16847 (12) | 0.0305 (9) | |
| C10 | 0.8816 (9) | 0.2937 (6) | 0.23044 (13) | 0.0404 (10) | |
| H10 | 1.0143 | 0.3077 | 0.2454 | 0.049* | |
| C15 | 1.3821 (8) | 0.9447 (7) | 0.04512 (13) | 0.0427 (11) | |
| H15A | 1.4586 | 0.9711 | 0.0191 | 0.064* | |
| H15B | 1.4578 | 0.8487 | 0.0596 | 0.064* | |
| H15C | 1.3805 | 1.0505 | 0.0627 | 0.064* | |
| C2 | 0.9708 (7) | 0.8304 (5) | 0.08140 (11) | 0.0285 (9) | |
| C5 | 0.7006 (8) | 0.7604 (6) | 0.14990 (12) | 0.0345 (10) | |
| H5A | 0.6507 | 0.7108 | 0.1767 | 0.041* | |
| H5B | 0.578 | 0.8286 | 0.1377 | 0.041* | |
| C13 | 0.4867 (8) | 0.2558 (6) | 0.18589 (14) | 0.0396 (10) | |
| H13 | 0.3523 | 0.2426 | 0.1714 | 0.048* | |
| C12 | 0.5156 (9) | 0.1720 (6) | 0.22507 (14) | 0.0445 (11) | |
| H12 | 0.401 | 0.1021 | 0.2363 | 0.053* | |
| C11 | 0.7107 (9) | 0.1916 (6) | 0.24725 (14) | 0.0443 (12) | |
| H11 | 0.7278 | 0.1363 | 0.2735 | 0.053* | |
| C14 | 0.9890 (9) | 1.0887 (7) | 0.01517 (15) | 0.0544 (13) | |
| H14A | 1.0569 | 1.1193 | −0.0114 | 0.082* | |
| H14B | 1.0205 | 1.1817 | 0.0355 | 0.082* | |
| H14C | 0.8304 | 1.078 | 0.0115 | 0.082* | |
| C7 | 0.3927 (9) | 0.4733 (7) | 0.10987 (16) | 0.0524 (13) | |
| H7A | 0.3922 | 0.5252 | 0.082 | 0.079* | |
| H7B | 0.3199 | 0.5539 | 0.1293 | 0.079* | |
| H7C | 0.3147 | 0.36 | 0.1094 | 0.079* |
Atomic displacement parameters (Å2)
| U11 | U22 | U33 | U12 | U13 | U23 | |
| S1 | 0.0354 (5) | 0.0314 (4) | 0.0300 (4) | −0.0059 (5) | 0.0034 (5) | 0.0035 (4) |
| O1 | 0.056 (2) | 0.0317 (14) | 0.0304 (13) | −0.0091 (19) | 0.0042 (15) | −0.0028 (11) |
| N1 | 0.0319 (17) | 0.0246 (15) | 0.0334 (15) | −0.0086 (18) | 0.0047 (14) | 0.0014 (14) |
| O2 | 0.066 (2) | 0.0325 (15) | 0.0543 (17) | −0.023 (2) | 0.0150 (19) | −0.0100 (13) |
| C9 | 0.034 (2) | 0.0334 (19) | 0.0379 (19) | −0.002 (2) | 0.0054 (17) | 0.0015 (18) |
| C1 | 0.030 (2) | 0.0288 (18) | 0.0245 (15) | −0.003 (2) | −0.0033 (16) | 0.0029 (14) |
| C6 | 0.040 (3) | 0.0282 (18) | 0.0352 (19) | −0.010 (2) | 0.0001 (19) | 0.0041 (16) |
| C4 | 0.052 (2) | 0.0315 (18) | 0.0287 (17) | −0.007 (3) | 0.009 (2) | −0.0067 (16) |
| C3 | 0.037 (2) | 0.0305 (19) | 0.0348 (19) | −0.003 (2) | 0.0052 (19) | −0.0037 (17) |
| C8 | 0.036 (2) | 0.0214 (17) | 0.0341 (18) | 0.0030 (19) | 0.0052 (17) | −0.0042 (15) |
| C10 | 0.042 (2) | 0.041 (2) | 0.039 (2) | 0.006 (3) | −0.001 (2) | −0.0008 (18) |
| C15 | 0.031 (2) | 0.055 (3) | 0.042 (2) | −0.006 (3) | 0.008 (2) | 0.001 (2) |
| C2 | 0.033 (2) | 0.0234 (17) | 0.0292 (17) | −0.0035 (18) | 0.0036 (16) | 0.0020 (14) |
| C5 | 0.042 (2) | 0.030 (2) | 0.0321 (19) | −0.002 (2) | 0.0077 (19) | −0.0005 (16) |
| C13 | 0.034 (2) | 0.037 (2) | 0.048 (2) | −0.003 (2) | 0.001 (2) | 0.0091 (19) |
| C12 | 0.046 (3) | 0.040 (2) | 0.048 (3) | −0.006 (3) | 0.012 (2) | 0.013 (2) |
| C11 | 0.057 (3) | 0.041 (2) | 0.035 (2) | 0.013 (3) | 0.009 (2) | 0.0050 (19) |
| C14 | 0.051 (3) | 0.057 (3) | 0.055 (3) | 0.004 (3) | 0.004 (2) | 0.026 (2) |
| C7 | 0.042 (3) | 0.057 (3) | 0.058 (3) | −0.016 (3) | −0.015 (3) | 0.019 (2) |
Geometric parameters (Å, °)
| S1—C2 | 1.729 (4) | C8—C13 | 1.378 (6) |
| S1—C15 | 1.796 (5) | C10—C11 | 1.379 (7) |
| S1—C14 | 1.799 (5) | C10—H10 | 0.93 |
| O1—C1 | 1.250 (4) | C15—H15A | 0.96 |
| N1—C1 | 1.364 (5) | C15—H15B | 0.96 |
| N1—C5 | 1.473 (5) | C15—H15C | 0.96 |
| N1—C6 | 1.479 (5) | C5—H5A | 0.97 |
| O2—C3 | 1.244 (5) | C5—H5B | 0.97 |
| C9—C10 | 1.391 (6) | C13—C12 | 1.395 (6) |
| C9—C8 | 1.402 (6) | C13—H13 | 0.93 |
| C9—H9 | 0.93 | C12—C11 | 1.369 (7) |
| C1—C2 | 1.455 (5) | C12—H12 | 0.93 |
| C6—C7 | 1.531 (7) | C11—H11 | 0.93 |
| C6—C8 | 1.538 (5) | C14—H14A | 0.96 |
| C6—H6 | 0.98 | C14—H14B | 0.96 |
| C4—C5 | 1.510 (6) | C14—H14C | 0.96 |
| C4—C3 | 1.513 (5) | C7—H7A | 0.96 |
| C4—H4A | 0.97 | C7—H7B | 0.96 |
| C4—H4B | 0.97 | C7—H7C | 0.96 |
| C3—C2 | 1.415 (5) | ||
| C2—S1—C15 | 107.6 (2) | H15A—C15—H15B | 109.5 |
| C2—S1—C14 | 107.0 (2) | S1—C15—H15C | 109.5 |
| C15—S1—C14 | 99.9 (2) | H15A—C15—H15C | 109.5 |
| C1—N1—C5 | 119.4 (3) | H15B—C15—H15C | 109.5 |
| C1—N1—C6 | 119.0 (3) | C3—C2—C1 | 124.4 (3) |
| C5—N1—C6 | 117.5 (3) | C3—C2—S1 | 120.1 (3) |
| C10—C9—C8 | 120.6 (4) | C1—C2—S1 | 114.9 (3) |
| C10—C9—H9 | 119.7 | N1—C5—C4 | 110.5 (3) |
| C8—C9—H9 | 119.7 | N1—C5—H5A | 109.5 |
| O1—C1—N1 | 121.4 (3) | C4—C5—H5A | 109.5 |
| O1—C1—C2 | 122.4 (3) | N1—C5—H5B | 109.5 |
| N1—C1—C2 | 116.2 (3) | C4—C5—H5B | 109.5 |
| N1—C6—C7 | 110.2 (4) | H5A—C5—H5B | 108.1 |
| N1—C6—C8 | 111.2 (3) | C8—C13—C12 | 120.5 (4) |
| C7—C6—C8 | 114.1 (4) | C8—C13—H13 | 119.8 |
| N1—C6—H6 | 107 | C12—C13—H13 | 119.8 |
| C7—C6—H6 | 107 | C11—C12—C13 | 120.9 (4) |
| C8—C6—H6 | 107 | C11—C12—H12 | 119.6 |
| C5—C4—C3 | 110.8 (3) | C13—C12—H12 | 119.6 |
| C5—C4—H4A | 109.5 | C12—C11—C10 | 119.6 (4) |
| C3—C4—H4A | 109.5 | C12—C11—H11 | 120.2 |
| C5—C4—H4B | 109.5 | C10—C11—H11 | 120.2 |
| C3—C4—H4B | 109.5 | S1—C14—H14A | 109.5 |
| H4A—C4—H4B | 108.1 | S1—C14—H14B | 109.5 |
| O2—C3—C2 | 124.2 (4) | H14A—C14—H14B | 109.5 |
| O2—C3—C4 | 120.7 (4) | S1—C14—H14C | 109.5 |
| C2—C3—C4 | 115.1 (3) | H14A—C14—H14C | 109.5 |
| C13—C8—C9 | 118.4 (4) | H14B—C14—H14C | 109.5 |
| C13—C8—C6 | 121.6 (4) | C6—C7—H7A | 109.5 |
| C9—C8—C6 | 119.9 (4) | C6—C7—H7B | 109.5 |
| C11—C10—C9 | 120.1 (5) | H7A—C7—H7B | 109.5 |
| C11—C10—H10 | 120 | C6—C7—H7C | 109.5 |
| C9—C10—H10 | 120 | H7A—C7—H7C | 109.5 |
| S1—C15—H15A | 109.5 | H7B—C7—H7C | 109.5 |
| S1—C15—H15B | 109.5 | ||
| C5—N1—C1—O1 | −164.3 (4) | O2—C3—C2—S1 | 5.4 (7) |
| C6—N1—C1—O1 | −7.7 (6) | C4—C3—C2—S1 | −172.1 (3) |
| C5—N1—C1—C2 | 15.5 (5) | O1—C1—C2—C3 | −169.5 (4) |
| C6—N1—C1—C2 | 172.0 (3) | N1—C1—C2—C3 | 10.7 (6) |
| C1—N1—C6—C7 | −90.2 (5) | O1—C1—C2—S1 | 1.8 (5) |
| C5—N1—C6—C7 | 66.8 (5) | N1—C1—C2—S1 | −178.0 (3) |
| C1—N1—C6—C8 | 142.2 (4) | C15—S1—C2—C3 | 46.9 (4) |
| C5—N1—C6—C8 | −60.8 (5) | C14—S1—C2—C3 | −59.6 (4) |
| C5—C4—C3—O2 | 151.0 (4) | C15—S1—C2—C1 | −124.8 (3) |
| C5—C4—C3—C2 | −31.4 (6) | C14—S1—C2—C1 | 128.7 (3) |
| C10—C9—C8—C13 | −0.6 (6) | C1—N1—C5—C4 | −48.4 (5) |
| C10—C9—C8—C6 | −177.2 (4) | C6—N1—C5—C4 | 154.7 (3) |
| N1—C6—C8—C13 | 149.8 (4) | C3—C4—C5—N1 | 54.5 (5) |
| C7—C6—C8—C13 | 24.4 (6) | C9—C8—C13—C12 | −0.3 (6) |
| N1—C6—C8—C9 | −33.6 (5) | C6—C8—C13—C12 | 176.3 (4) |
| C7—C6—C8—C9 | −159.1 (4) | C8—C13—C12—C11 | 0.9 (7) |
| C8—C9—C10—C11 | 0.8 (6) | C13—C12—C11—C10 | −0.7 (7) |
| O2—C3—C2—C1 | 176.3 (4) | C9—C10—C11—C12 | −0.2 (7) |
| C4—C3—C2—C1 | −1.2 (6) |
Hydrogen-bond geometry (Å, °)
| D—H···A | D—H | H···A | D···A | D—H···A |
| C7—H7C···O2i | 0.96 | 2.36 | 3.315 (6) | 172 |
| C15—H15A···O1ii | 0.96 | 2.38 | 3.167 (5) | 138 |
Symmetry codes: (i) x−1, y−1, z; (ii) x+1/2, −y+3/2, −z.
Footnotes
Supplementary data and figures for this paper are available from the IUCr electronic archives (Reference: BT5438).
References
- Basco, L. K., Mitaku, S., Skaltsounis, A. L., Ravelomanantsoa, N., Tillequin, F., Koch, M. & Le Bras, J. (1994). J. Antimicrob. Agents Chemother 38, 1169–1171. [DOI] [PMC free article] [PubMed]
- Burla, M. C., Caliandro, R., Camalli, M., Carrozzini, B., Cascarano, G. L., De Caro, L., Giacovazzo, C., Polidori, G. & Spagna, R. (2005). J. Appl. Cryst. 38, 381–388.
- Cremer, D. & Pople, J. A. (1975). J. Am. Chem. Soc. 97, 1354–1358.
- Farrugia, L. J. (1997). J. Appl. Cryst. 30, 565.
- Farrugia, L. J. (1999). J. Appl. Cryst. 32, 837–838.
- Flack, H. D. (1983). Acta Cryst. A39, 876–881.
- Kappe, T., Korbuuly, G. & Pongratz, E. (1983). Monatsh. Chem 114, 303–315.
- Koruzňjak, J. D., Grdiša, M., Slade, N., Zamola, B., Pavelic, K. & Karminski-Zamola, G. (2003). J. Med. Chem 46, 4516–4524. [DOI] [PubMed]
- North, A. C. T., Phillips, D. C. & Mathews, F. S. (1968). Acta Cryst. A24, 351–359.
- Palillero, A., Terán, J. L., Gnecco, D., Juárez, J. R., Orea, M. L. & Castro, A. (2009). Tetrahedron Lett. 50, 4208–4211.
- Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. [DOI] [PubMed]
- Siemens (1994). XSCANS Siemens Analytical X-ray Instruments Inc., Madison, Wisconsin, USA.
- Zang, S. L., Huang, Z. S., Li, Y. M., Chan, A. S. C. & Gu, L. Q. (2008). Tetrahedron, 64, 4403–4407.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crystal structure: contains datablocks global, I. DOI: 10.1107/S1600536810052955/bt5438sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S1600536810052955/bt5438Isup2.hkl
Additional supplementary materials: crystallographic information; 3D view; checkCIF report