Abstract
In the title compound, C21H20ClN5·H2O, the 1H-imidazo[4,5-c]quinoline ring is approximately planar, with a maximum deviation of 0.0795 (7) Å, and it forms a dihedral angle of 7.65 (3)° with the chlorophenyl ring. In the crystal, the components are linked into chains along the a axis via intermolecular N—H⋯O, O—H⋯N and C—H⋯O hydrogen bonds. One of the H atoms of the water molecule is disordered over two positions with a site-occupancy ratio of 0.80 (4):0.20 (4).
Related literature
For background to quinolines and their microbial activity, see: El-Subbagh et al. (2000 ▶); Atwell et al. (1989 ▶); Kuo et al. (1993 ▶); Xia et al. (1998 ▶). For the biological activity of Schiff base hydrazones, see: Colins & Lyne (1970 ▶); Ochiai (1977 ▶). For bond-length data, see: Allen et al. (1987 ▶). For related structures, see: Loh et al. (2011a ▶,b
▶). For the stability of the temperature controller used in the data collection, see: Cosier & Glazer (1986 ▶).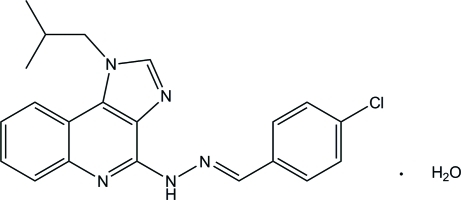
Experimental
Crystal data
C21H20ClN5·H2O
M r = 395.89
Monoclinic,

a = 10.4117 (3) Å
b = 18.2365 (6) Å
c = 11.9019 (3) Å
β = 117.809 (2)°
V = 1998.85 (10) Å3
Z = 4
Mo Kα radiation
μ = 0.21 mm−1
T = 100 K
0.49 × 0.45 × 0.18 mm
Data collection
Bruker SMART APEXII DUO CCD area-detector diffractometer
Absorption correction: multi-scan (SADABS; Bruker, 2009 ▶) T min = 0.904, T max = 0.963
39468 measured reflections
10411 independent reflections
8351 reflections with I > 2σ(I)
R int = 0.032
Refinement
R[F 2 > 2σ(F 2)] = 0.045
wR(F 2) = 0.135
S = 1.04
10411 reflections
260 parameters
H atoms treated by a mixture of independent and constrained refinement
Δρmax = 1.19 e Å−3
Δρmin = −0.47 e Å−3
Data collection: APEX2 (Bruker, 2009 ▶); cell refinement: SAINT (Bruker, 2009 ▶); data reduction: SAINT; program(s) used to solve structure: SHELXTL (Sheldrick, 2008 ▶); program(s) used to refine structure: SHELXTL; molecular graphics: SHELXTL; software used to prepare material for publication: SHELXTL and PLATON (Spek, 2009 ▶).
Supplementary Material
Crystal structure: contains datablocks global, I. DOI: 10.1107/S1600536811001577/is2658sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S1600536811001577/is2658Isup2.hkl
Additional supplementary materials: crystallographic information; 3D view; checkCIF report
Table 1. Hydrogen-bond geometry (Å, °).
| D—H⋯A | D—H | H⋯A | D⋯A | D—H⋯A |
|---|---|---|---|---|
| N4—H1N4⋯O1Wi | 0.874 (19) | 2.559 (18) | 3.2789 (13) | 140.2 (14) |
| O1W—H1W1⋯N1ii | 0.83 | 2.09 | 2.9178 (14) | 173 |
| C10—H10A⋯O1Wiii | 0.93 | 2.52 | 3.3513 (16) | 149 |
| C18—H18B⋯O1Wiii | 0.97 | 2.59 | 3.4776 (14) | 153 |
Symmetry codes: (i)  ; (ii)
; (ii)  ; (iii)
; (iii)  .
.
Acknowledgments
HKF and WSL thank Universiti Sains Malaysia (USM) for the Research University Grant (1001/PFIZIK/811160). WSL also thanks the Malaysian Government and USM for the award of a Research Fellowship.
supplementary crystallographic information
Comment
Quinolines and their derivatives are important constituents of pharmacologically active synthetic compounds as these systems have been associated with a wide spectrum of biological properties (El-Subbagh et al., 2000) such as DNA binding capability (Atwell et al., 1989) and antitumor activities (Kuo et al., 1993; Xia et al., 1998). The study of Schiff base hydrazones has been growing because of their antimicrobial, anti-tuberculosis and anti-tumour activities (Colins & Lyne, 1970; Ochiai, 1977).
The asymmetric unit of the title compound, (Fig. 1), consists of one 4-chlorobenzaldehyde(1-isobutyl-1H-imidazo[4,5-c]quinolin-4-yl) hydrazone molecule and one water molecule. One of the H atoms attached to the water molecule is disordered over two positions with the site occupancy ratio of 0.80 (4):0.20 (4). The 1H-imidazo[4,5-c]quinoline ring (C1–C6/N1/C7/C8/N3/C10/N2/C9) is approximately planar with a maximum deviation of 0.0795 (7) Å at atom C2 and it forms a dihedral angle of 7.65 (3)° with the chlorophenyl ring (Cl1/C11–C16) with maximum deviation of 0.0286 (3) Å at atom Cl1. Bond lengths (Allen et al., 1987) and angles are within the normal ranges and are comparable to the related structures (Loh et al., 2011a,b).
In the crystal packing (Fig. 2), the molecules are linked into chains along the a axis by the water molecules via intermolecular N4—H1N4···O1W, O1W—H1W1···N1, C10—H10A···O1W and C18—H18B···O1W hydrogen bonds (Table 1).
Experimental
A mixture of 4-hydrazino-1-isobutyl-1H-imidazo[4,5-c]quinoline (2.5 g, 0.0098 mole) and 4-chlorobenzaldehyde (1.38 g, 0.0098 mole) in absolute ethanol was refluxed for 4 h in the presence of acetic acid (1 ml). The product, 4-chlorobenzaldehyde (1-isobutyl-1H-imidazo[4,5-c]quinolin-4-yl)hydrazone, was obtained after cooling and it was crystallized from absolute ethanol. Yield: 3.4 g (80%). Crystals suitable for X-ray analysis were obtained from ethanol by slow evaporation.
Refinement
O- and N-bound H atoms were located from a difference Fourier map. O-bound H atoms were then fixed at their found positions (O—H = 0.8330 to 0.8554 Å), with Uiso(H) = 1.5Ueq(O), whereas N-bound H atoms was refined freely [N—H = 0.875 (18) Å]. The remaining H atoms were positioned geometrically with the bond lengths of C—H = 0.93 to 0.98 Å and were refined using a riding model, with Uiso(H) = 1.2 or 1.5Ueq(C). A rotating group model was applied to the methyl groups. The heighest residual electron density peak is located 1.01 Å from atom O1W.
Figures
Fig. 1.
The molecular structure of the title compound, showing 50% probability displacement ellipsoids and the atom-numbering scheme. The open bond indicates the minor component.
Fig. 2.
The crystal packing of the title compound, showing the chains along the a axis. H atoms not involved in the intermolecular interactions (dashed lines) have been omitted for clarity. Only major component is shown.
Crystal data
| C21H20ClN5·H2O | F(000) = 832 |
| Mr = 395.89 | Dx = 1.316 Mg m−3 |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.71073 Å |
| Hall symbol: -P 2ybc | Cell parameters from 9978 reflections |
| a = 10.4117 (3) Å | θ = 4.0–37.5° |
| b = 18.2365 (6) Å | µ = 0.21 mm−1 |
| c = 11.9019 (3) Å | T = 100 K |
| β = 117.809 (2)° | Plate, yellow |
| V = 1998.85 (10) Å3 | 0.49 × 0.45 × 0.18 mm |
| Z = 4 |
Data collection
| Bruker SMART APEXII DUO CCD area-detector diffractometer | 10411 independent reflections |
| Radiation source: fine-focus sealed tube | 8351 reflections with I > 2σ(I) |
| graphite | Rint = 0.032 |
| φ and ω scans | θmax = 37.6°, θmin = 4.0° |
| Absorption correction: multi-scan (SADABS; Bruker, 2009) | h = −17→17 |
| Tmin = 0.904, Tmax = 0.963 | k = −31→30 |
| 39468 measured reflections | l = −20→20 |
Refinement
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.045 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.135 | H atoms treated by a mixture of independent and constrained refinement |
| S = 1.04 | w = 1/[σ2(Fo2) + (0.0715P)2 + 0.4845P] where P = (Fo2 + 2Fc2)/3 |
| 10411 reflections | (Δ/σ)max = 0.001 |
| 260 parameters | Δρmax = 1.19 e Å−3 |
| 0 restraints | Δρmin = −0.47 e Å−3 |
Special details
| Experimental. The crystal was placed in the cold stream of an Oxford Cryosystems Cobra open-flow nitrogen cryostat (Cosier & Glazer, 1986) operating at 100.0 (1) K. |
| Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
| Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2, conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2 are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
| x | y | z | Uiso*/Ueq | Occ. (<1) | |
| Cl1 | −0.37283 (3) | −0.053039 (16) | 0.46655 (3) | 0.03357 (7) | |
| N1 | 0.00417 (7) | 0.13498 (4) | 0.01185 (6) | 0.01373 (11) | |
| N2 | 0.33923 (7) | 0.14996 (4) | −0.08204 (7) | 0.01514 (11) | |
| N3 | 0.35384 (7) | 0.06015 (4) | 0.05254 (7) | 0.01630 (12) | |
| N4 | 0.13161 (8) | 0.03567 (4) | 0.13737 (7) | 0.01564 (11) | |
| N5 | 0.02778 (7) | 0.02713 (4) | 0.17545 (6) | 0.01482 (11) | |
| C1 | 0.09304 (8) | 0.20371 (4) | −0.11917 (7) | 0.01242 (11) | |
| C2 | 0.06369 (8) | 0.25958 (4) | −0.21028 (7) | 0.01464 (12) | |
| H2A | 0.1260 | 0.2664 | −0.2455 | 0.018* | |
| C3 | −0.05646 (8) | 0.30406 (4) | −0.24743 (7) | 0.01589 (12) | |
| H3A | −0.0749 | 0.3407 | −0.3075 | 0.019* | |
| C4 | −0.15108 (8) | 0.29408 (4) | −0.19448 (8) | 0.01637 (13) | |
| H4A | −0.2297 | 0.3255 | −0.2171 | 0.020* | |
| C5 | −0.12834 (8) | 0.23813 (4) | −0.10935 (8) | 0.01560 (12) | |
| H5A | −0.1932 | 0.2314 | −0.0768 | 0.019* | |
| C6 | −0.00740 (8) | 0.19089 (4) | −0.07104 (7) | 0.01287 (11) | |
| C7 | 0.11700 (8) | 0.09093 (4) | 0.05390 (7) | 0.01293 (11) | |
| C8 | 0.22740 (8) | 0.09992 (4) | 0.01609 (7) | 0.01314 (11) | |
| C9 | 0.21572 (8) | 0.15557 (4) | −0.06761 (7) | 0.01280 (11) | |
| C10 | 0.41677 (9) | 0.09246 (4) | −0.00837 (8) | 0.01736 (13) | |
| H10A | 0.5054 | 0.0774 | −0.0017 | 0.021* | |
| C11 | 0.04705 (9) | −0.02385 (4) | 0.25628 (7) | 0.01614 (13) | |
| H11A | 0.1279 | −0.0543 | 0.2850 | 0.019* | |
| C12 | −0.05978 (9) | −0.03357 (4) | 0.30283 (7) | 0.01541 (12) | |
| C13 | −0.18721 (9) | 0.00855 (4) | 0.25432 (7) | 0.01694 (13) | |
| H13A | −0.2067 | 0.0415 | 0.1887 | 0.020* | |
| C14 | −0.28457 (10) | 0.00168 (5) | 0.30303 (8) | 0.01947 (14) | |
| H14A | −0.3690 | 0.0296 | 0.2704 | 0.023* | |
| C15 | −0.25383 (10) | −0.04775 (5) | 0.40152 (8) | 0.02080 (15) | |
| C16 | −0.13059 (11) | −0.09132 (5) | 0.44958 (8) | 0.02182 (15) | |
| H16A | −0.1126 | −0.1248 | 0.5142 | 0.026* | |
| C17 | −0.03391 (10) | −0.08415 (5) | 0.39943 (8) | 0.01941 (14) | |
| H17A | 0.0488 | −0.1134 | 0.4306 | 0.023* | |
| C18 | 0.38977 (9) | 0.19795 (4) | −0.15243 (8) | 0.01655 (13) | |
| H18A | 0.3124 | 0.2040 | −0.2385 | 0.020* | |
| H18B | 0.4711 | 0.1749 | −0.1570 | 0.020* | |
| C19 | 0.43656 (8) | 0.27358 (4) | −0.09030 (8) | 0.01657 (13) | |
| H19A | 0.3546 | 0.2959 | −0.0839 | 0.020* | |
| C20 | 0.56378 (11) | 0.26719 (6) | 0.04269 (9) | 0.02498 (17) | |
| H20A | 0.5932 | 0.3153 | 0.0784 | 0.037* | |
| H20B | 0.6435 | 0.2433 | 0.0383 | 0.037* | |
| H20C | 0.5349 | 0.2390 | 0.0952 | 0.037* | |
| C21 | 0.47505 (11) | 0.32187 (6) | −0.17470 (10) | 0.02624 (18) | |
| H21A | 0.5008 | 0.3700 | −0.1382 | 0.039* | |
| H21B | 0.3929 | 0.3253 | −0.2576 | 0.039* | |
| H21C | 0.5556 | 0.3008 | −0.1815 | 0.039* | |
| H1N4 | 0.2039 (19) | 0.0049 (9) | 0.1601 (16) | 0.035 (4)* | |
| O1W | 0.70320 (9) | 0.09494 (4) | 0.93489 (12) | 0.0421 (3) | |
| H1W1 | 0.7912 | 0.1025 | 0.9595 | 0.063* | |
| H2WA | 0.6633 | 0.0764 | 0.8603 | 0.063* | 0.80 (4) |
| H2WB | 0.6915 | 0.0492 | 0.9405 | 0.063* | 0.20 (4) |
Atomic displacement parameters (Å2)
| U11 | U22 | U33 | U12 | U13 | U23 | |
| Cl1 | 0.03921 (14) | 0.03795 (14) | 0.03987 (14) | −0.00160 (10) | 0.03211 (12) | 0.00615 (10) |
| N1 | 0.0138 (2) | 0.0145 (2) | 0.0160 (2) | 0.00128 (19) | 0.0096 (2) | 0.0020 (2) |
| N2 | 0.0138 (2) | 0.0149 (3) | 0.0216 (3) | 0.0012 (2) | 0.0124 (2) | 0.0016 (2) |
| N3 | 0.0137 (3) | 0.0151 (3) | 0.0227 (3) | 0.0020 (2) | 0.0107 (2) | 0.0021 (2) |
| N4 | 0.0154 (3) | 0.0166 (3) | 0.0188 (3) | 0.0029 (2) | 0.0112 (2) | 0.0044 (2) |
| N5 | 0.0165 (3) | 0.0153 (3) | 0.0161 (3) | −0.0005 (2) | 0.0104 (2) | 0.0009 (2) |
| C1 | 0.0114 (3) | 0.0139 (3) | 0.0136 (3) | 0.0000 (2) | 0.0072 (2) | 0.0001 (2) |
| C2 | 0.0139 (3) | 0.0168 (3) | 0.0152 (3) | 0.0000 (2) | 0.0085 (2) | 0.0017 (2) |
| C3 | 0.0142 (3) | 0.0177 (3) | 0.0162 (3) | 0.0010 (2) | 0.0074 (2) | 0.0034 (2) |
| C4 | 0.0134 (3) | 0.0168 (3) | 0.0196 (3) | 0.0019 (2) | 0.0083 (2) | 0.0032 (2) |
| C5 | 0.0134 (3) | 0.0169 (3) | 0.0197 (3) | 0.0020 (2) | 0.0103 (2) | 0.0029 (2) |
| C6 | 0.0122 (3) | 0.0142 (3) | 0.0147 (3) | 0.0002 (2) | 0.0084 (2) | 0.0004 (2) |
| C7 | 0.0132 (3) | 0.0136 (3) | 0.0141 (3) | −0.0003 (2) | 0.0081 (2) | −0.0001 (2) |
| C8 | 0.0121 (3) | 0.0132 (3) | 0.0163 (3) | 0.0004 (2) | 0.0084 (2) | 0.0003 (2) |
| C9 | 0.0120 (3) | 0.0138 (3) | 0.0154 (3) | −0.0003 (2) | 0.0087 (2) | −0.0003 (2) |
| C10 | 0.0149 (3) | 0.0157 (3) | 0.0258 (3) | 0.0025 (2) | 0.0132 (3) | 0.0025 (3) |
| C11 | 0.0175 (3) | 0.0161 (3) | 0.0162 (3) | 0.0004 (2) | 0.0090 (2) | 0.0025 (2) |
| C12 | 0.0187 (3) | 0.0150 (3) | 0.0142 (3) | −0.0020 (2) | 0.0091 (2) | 0.0007 (2) |
| C13 | 0.0193 (3) | 0.0173 (3) | 0.0167 (3) | −0.0005 (2) | 0.0105 (3) | 0.0020 (2) |
| C14 | 0.0211 (3) | 0.0198 (3) | 0.0218 (3) | −0.0012 (3) | 0.0136 (3) | 0.0011 (3) |
| C15 | 0.0261 (4) | 0.0216 (3) | 0.0209 (3) | −0.0049 (3) | 0.0162 (3) | −0.0001 (3) |
| C16 | 0.0277 (4) | 0.0224 (4) | 0.0189 (3) | −0.0022 (3) | 0.0138 (3) | 0.0045 (3) |
| C17 | 0.0228 (4) | 0.0192 (3) | 0.0175 (3) | 0.0001 (3) | 0.0104 (3) | 0.0043 (3) |
| C18 | 0.0162 (3) | 0.0188 (3) | 0.0200 (3) | 0.0003 (2) | 0.0130 (3) | 0.0017 (2) |
| C19 | 0.0139 (3) | 0.0177 (3) | 0.0198 (3) | 0.0002 (2) | 0.0094 (3) | 0.0031 (2) |
| C20 | 0.0216 (4) | 0.0287 (4) | 0.0210 (4) | −0.0013 (3) | 0.0070 (3) | 0.0022 (3) |
| C21 | 0.0255 (4) | 0.0260 (4) | 0.0297 (4) | −0.0023 (3) | 0.0150 (4) | 0.0094 (3) |
| O1W | 0.0217 (3) | 0.0222 (3) | 0.0865 (8) | 0.0064 (3) | 0.0287 (4) | 0.0188 (4) |
Geometric parameters (Å, °)
| Cl1—C15 | 1.7430 (9) | C11—H11A | 0.9300 |
| N1—C7 | 1.3143 (10) | C12—C17 | 1.3985 (11) |
| N1—C6 | 1.3842 (9) | C12—C13 | 1.4032 (12) |
| N2—C10 | 1.3638 (10) | C13—C14 | 1.3884 (11) |
| N2—C9 | 1.3782 (9) | C13—H13A | 0.9300 |
| N2—C18 | 1.4684 (10) | C14—C15 | 1.3924 (12) |
| N3—C10 | 1.3210 (10) | C14—H14A | 0.9300 |
| N3—C8 | 1.3836 (10) | C15—C16 | 1.3857 (14) |
| N4—N5 | 1.3622 (9) | C16—C17 | 1.3955 (12) |
| N4—C7 | 1.3728 (10) | C16—H16A | 0.9300 |
| N4—H1N4 | 0.875 (18) | C17—H17A | 0.9300 |
| N5—C11 | 1.2845 (10) | C18—C19 | 1.5330 (12) |
| C1—C2 | 1.4135 (10) | C18—H18A | 0.9700 |
| C1—C6 | 1.4267 (10) | C18—H18B | 0.9700 |
| C1—C9 | 1.4308 (10) | C19—C20 | 1.5222 (12) |
| C2—C3 | 1.3793 (11) | C19—C21 | 1.5233 (12) |
| C2—H2A | 0.9300 | C19—H19A | 0.9800 |
| C3—C4 | 1.4072 (11) | C20—H20A | 0.9600 |
| C3—H3A | 0.9300 | C20—H20B | 0.9600 |
| C4—C5 | 1.3783 (11) | C20—H20C | 0.9600 |
| C4—H4A | 0.9300 | C21—H21A | 0.9600 |
| C5—C6 | 1.4142 (10) | C21—H21B | 0.9600 |
| C5—H5A | 0.9300 | C21—H21C | 0.9600 |
| C7—C8 | 1.4259 (10) | O1W—H1W1 | 0.8330 |
| C8—C9 | 1.3872 (10) | O1W—H2WA | 0.8554 |
| C10—H10A | 0.9300 | O1W—H2WB | 0.8508 |
| C11—C12 | 1.4665 (11) | ||
| C7—N1—C6 | 119.13 (6) | C17—C12—C11 | 120.07 (7) |
| C10—N2—C9 | 106.49 (6) | C13—C12—C11 | 121.10 (7) |
| C10—N2—C18 | 124.26 (6) | C14—C13—C12 | 120.87 (7) |
| C9—N2—C18 | 129.08 (6) | C14—C13—H13A | 119.6 |
| C10—N3—C8 | 103.72 (6) | C12—C13—H13A | 119.6 |
| N5—N4—C7 | 119.18 (6) | C13—C14—C15 | 118.96 (8) |
| N5—N4—H1N4 | 121.7 (12) | C13—C14—H14A | 120.5 |
| C7—N4—H1N4 | 119.0 (12) | C15—C14—H14A | 120.5 |
| C11—N5—N4 | 117.46 (7) | C16—C15—C14 | 121.57 (8) |
| C2—C1—C6 | 119.31 (6) | C16—C15—Cl1 | 119.88 (6) |
| C2—C1—C9 | 126.97 (6) | C14—C15—Cl1 | 118.54 (7) |
| C6—C1—C9 | 113.71 (6) | C15—C16—C17 | 118.91 (8) |
| C3—C2—C1 | 120.60 (7) | C15—C16—H16A | 120.5 |
| C3—C2—H2A | 119.7 | C17—C16—H16A | 120.5 |
| C1—C2—H2A | 119.7 | C16—C17—C12 | 120.84 (8) |
| C2—C3—C4 | 120.01 (7) | C16—C17—H17A | 119.6 |
| C2—C3—H3A | 120.0 | C12—C17—H17A | 119.6 |
| C4—C3—H3A | 120.0 | N2—C18—C19 | 112.26 (6) |
| C5—C4—C3 | 120.59 (7) | N2—C18—H18A | 109.2 |
| C5—C4—H4A | 119.7 | C19—C18—H18A | 109.2 |
| C3—C4—H4A | 119.7 | N2—C18—H18B | 109.2 |
| C4—C5—C6 | 120.65 (7) | C19—C18—H18B | 109.2 |
| C4—C5—H5A | 119.7 | H18A—C18—H18B | 107.9 |
| C6—C5—H5A | 119.7 | C20—C19—C21 | 111.16 (7) |
| N1—C6—C5 | 116.55 (6) | C20—C19—C18 | 111.05 (7) |
| N1—C6—C1 | 124.78 (6) | C21—C19—C18 | 108.91 (7) |
| C5—C6—C1 | 118.67 (6) | C20—C19—H19A | 108.5 |
| N1—C7—N4 | 120.12 (6) | C21—C19—H19A | 108.5 |
| N1—C7—C8 | 121.23 (7) | C18—C19—H19A | 108.5 |
| N4—C7—C8 | 118.64 (6) | C19—C20—H20A | 109.5 |
| N3—C8—C9 | 111.13 (6) | C19—C20—H20B | 109.5 |
| N3—C8—C7 | 129.05 (7) | H20A—C20—H20B | 109.5 |
| C9—C8—C7 | 119.81 (6) | C19—C20—H20C | 109.5 |
| N2—C9—C8 | 105.12 (6) | H20A—C20—H20C | 109.5 |
| N2—C9—C1 | 133.65 (7) | H20B—C20—H20C | 109.5 |
| C8—C9—C1 | 121.23 (6) | C19—C21—H21A | 109.5 |
| N3—C10—N2 | 113.54 (7) | C19—C21—H21B | 109.5 |
| N3—C10—H10A | 123.2 | H21A—C21—H21B | 109.5 |
| N2—C10—H10A | 123.2 | C19—C21—H21C | 109.5 |
| N5—C11—C12 | 119.30 (7) | H21A—C21—H21C | 109.5 |
| N5—C11—H11A | 120.3 | H21B—C21—H21C | 109.5 |
| C12—C11—H11A | 120.3 | H1W1—O1W—H2WA | 110.7 |
| C17—C12—C13 | 118.81 (7) | H1W1—O1W—H2WB | 107.9 |
| C7—N4—N5—C11 | −178.12 (7) | N3—C8—C9—N2 | −0.42 (9) |
| C6—C1—C2—C3 | 3.55 (11) | C7—C8—C9—N2 | 178.87 (7) |
| C9—C1—C2—C3 | −177.20 (7) | N3—C8—C9—C1 | 179.51 (7) |
| C1—C2—C3—C4 | 0.05 (12) | C7—C8—C9—C1 | −1.21 (11) |
| C2—C3—C4—C5 | −2.71 (12) | C2—C1—C9—N2 | 3.78 (14) |
| C3—C4—C5—C6 | 1.68 (12) | C6—C1—C9—N2 | −176.94 (8) |
| C7—N1—C6—C5 | −177.76 (7) | C2—C1—C9—C8 | −176.12 (7) |
| C7—N1—C6—C1 | 2.14 (11) | C6—C1—C9—C8 | 3.17 (10) |
| C4—C5—C6—N1 | −178.17 (7) | C8—N3—C10—N2 | −0.36 (9) |
| C4—C5—C6—C1 | 1.94 (11) | C9—N2—C10—N3 | 0.12 (10) |
| C2—C1—C6—N1 | 175.61 (7) | C18—N2—C10—N3 | 175.72 (7) |
| C9—C1—C6—N1 | −3.74 (10) | N4—N5—C11—C12 | 178.17 (7) |
| C2—C1—C6—C5 | −4.50 (11) | N5—C11—C12—C17 | −174.05 (8) |
| C9—C1—C6—C5 | 176.15 (7) | N5—C11—C12—C13 | 4.30 (12) |
| C6—N1—C7—N4 | 179.21 (7) | C17—C12—C13—C14 | 1.54 (12) |
| C6—N1—C7—C8 | 0.24 (11) | C11—C12—C13—C14 | −176.82 (8) |
| N5—N4—C7—N1 | 0.18 (11) | C12—C13—C14—C15 | 0.15 (13) |
| N5—N4—C7—C8 | 179.17 (7) | C13—C14—C15—C16 | −1.62 (13) |
| C10—N3—C8—C9 | 0.48 (9) | C13—C14—C15—Cl1 | 177.25 (7) |
| C10—N3—C8—C7 | −178.72 (8) | C14—C15—C16—C17 | 1.32 (14) |
| N1—C7—C8—N3 | 178.48 (7) | Cl1—C15—C16—C17 | −177.53 (7) |
| N4—C7—C8—N3 | −0.50 (12) | C15—C16—C17—C12 | 0.44 (13) |
| N1—C7—C8—C9 | −0.66 (11) | C13—C12—C17—C16 | −1.84 (12) |
| N4—C7—C8—C9 | −179.64 (7) | C11—C12—C17—C16 | 176.54 (8) |
| C10—N2—C9—C8 | 0.18 (8) | C10—N2—C18—C19 | −105.84 (9) |
| C18—N2—C9—C8 | −175.14 (7) | C9—N2—C18—C19 | 68.72 (10) |
| C10—N2—C9—C1 | −179.73 (8) | N2—C18—C19—C20 | 62.01 (9) |
| C18—N2—C9—C1 | 4.95 (14) | N2—C18—C19—C21 | −175.26 (7) |
Hydrogen-bond geometry (Å, °)
| D—H···A | D—H | H···A | D···A | D—H···A |
| N4—H1N4···O1Wi | 0.874 (19) | 2.559 (18) | 3.2789 (13) | 140.2 (14) |
| O1W—H1W1···N1ii | 0.83 | 2.09 | 2.9178 (14) | 173 |
| C10—H10A···O1Wiii | 0.93 | 2.52 | 3.3513 (16) | 149 |
| C18—H18B···O1Wiii | 0.97 | 2.59 | 3.4776 (14) | 153 |
Symmetry codes: (i) −x+1, −y, −z+1; (ii) x+1, y, z+1; (iii) x, y, z−1.
Footnotes
Supplementary data and figures for this paper are available from the IUCr electronic archives (Reference: IS2658).
References
- Allen, F. H., Kennard, O., Watson, D. G., Brammer, L., Orpen, A. G. & Taylor, R. (1987). J. Chem. Soc. Perkin Trans. 2, pp. S1–19.
- Atwell, G. J., Baguley, B. C. & Denny, W. A. (1989). J. Med. Chem. 32, 396–401. [DOI] [PubMed]
- Bruker (2009). APEX2, SAINT and SADABS Bruker AXS Inc., Madison, Wisconsin, USA.
- Colins, C. H. & Lyne, P. M. (1970). Microbiological Methods Baltimore: University Park Press.
- Cosier, J. & Glazer, A. M. (1986). J. Appl. Cryst. 19, 105–107.
- El-Subbagh, H. I., Abu-Zaid, S. M., Mahran, M. A., Badria, F. A. & Al-Obaid, A. M. (2000). J. Med. Chem. 43, 2915–2921. [DOI] [PubMed]
- Kuo, S. C., Lee, H. Z., Juang, J. P., Lin, Y. T., Wu, T. S., Chang, J. J., Lednicer, D., Paull, K. D., Lin, C. M., Hamel, E. & Lee, K. H. (1993). J. Med. Chem. 36, 1146–1156. [DOI] [PubMed]
- Loh, W.-S., Fun, H.-K., Kayarmar, R., Viveka, S. & Nagaraja, G. K. (2011a). Acta Cryst. E67, o405. [DOI] [PMC free article] [PubMed]
- Loh, W.-S., Fun, H.-K., Kayarmar, R., Viveka, S. & Nagaraja, G. K. (2011b). Acta Cryst. E67, o406. [DOI] [PMC free article] [PubMed]
- Ochiai, E. (1977). In Bioinorganic Chemistry Boston: Allyn and Bacon.
- Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. [DOI] [PubMed]
- Spek, A. L. (2009). Acta Cryst. D65, 148–155. [DOI] [PMC free article] [PubMed]
- Xia, Y., Yang, Z. Y., Xia, P., Bastow, K. F., Tachibana, Y., Kuo, S. C., Hamel, E., Hackl, T. & Lee, K. H. (1998). J. Med. Chem. 41, 1155–1162. [DOI] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crystal structure: contains datablocks global, I. DOI: 10.1107/S1600536811001577/is2658sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S1600536811001577/is2658Isup2.hkl
Additional supplementary materials: crystallographic information; 3D view; checkCIF report




