Abstract
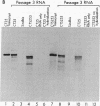
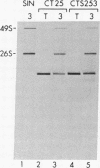
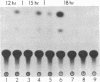
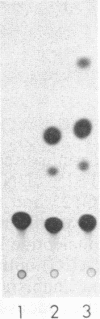
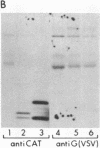
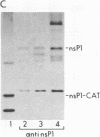
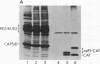
We are investigating the feasibility of using the positive-strand RNA virus Sindbis virus and its defective interfering (DI) particles as vectors for introducing foreign genes into cells. In previous work we showed by deletion mapping of a cloned cDNA derived from one of the DI RNAs that only nucleotides at the 3' and 5' termini of the RNA are essential for the DI RNA to be amplified after it is transfected into cells in the presence of helper virus. As a first step in developing a vector we replaced 75% of the internal nucleotides of this DI cDNA with foreign sequences including the bacterial chloramphenicol acetyltransferase (CAT; EC 2.3.1.28) gene. DI RNAs transcribed from this cDNA were replicated and packaged by helper Sindbis virus and became a major viral RNA species in infected cells by the third passage after transfection. They were also translated to produce enzymatically active CAT. CAT activity was measured at passage 3 but could also be detected in transfected cells. DI RNAs containing the CAT gene were translated in vivo and in vitro to produce two polypeptides immunoprecipitable by anti-CAT antibodies. One polypeptide was identical in size to the authentic CAT polypeptide; the other was the size expected for a protein initiated at an upstream, viral-specific AUG in frame with the CAT AUG. These studies establish that DI genomes of Sindbis virus can tolerate the insertion and direct the expression of at least one foreign gene.
Full text
PDF
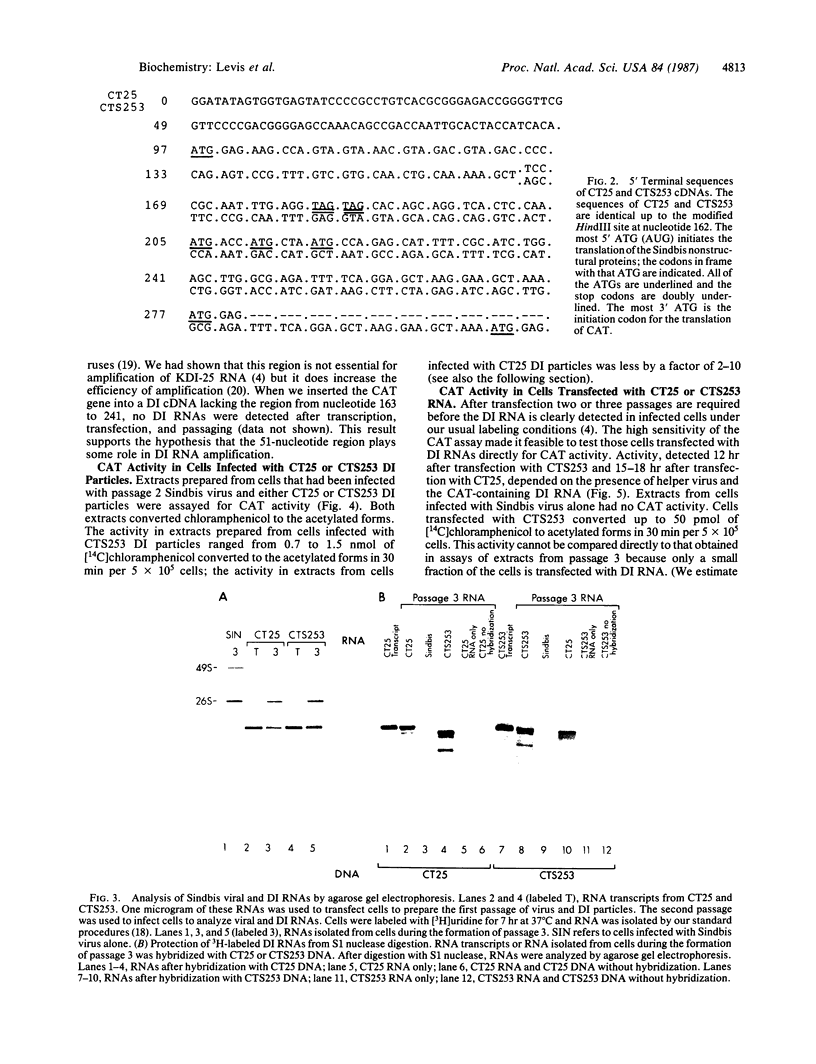


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Anderson D. J., Blobel G. Immunoprecipitation of proteins from cell-free translations. Methods Enzymol. 1983;96:111–120. doi: 10.1016/s0076-6879(83)96012-3. [DOI] [PubMed] [Google Scholar]
- Bujarski J. J., Ahlquist P., Hall T. C., Dreher T. W., Kaesberg P. Modulation of replication, aminoacylation and adenylation in vitro and infectivity in vivo of BMV RNAs containing deletions within the multifunctional 3' end. EMBO J. 1986 Aug;5(8):1769–1774. doi: 10.1002/j.1460-2075.1986.tb04425.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carmichael G. G., McMaster G. K. The analysis of nucleic acids in gels using glyoxal and acridine orange. Methods Enzymol. 1980;65(1):380–391. doi: 10.1016/s0076-6879(80)65049-6. [DOI] [PubMed] [Google Scholar]
- French R., Janda M., Ahlquist P. Bacterial gene inserted in an engineered RNA virus: efficient expression in monocotyledonous plant cells. Science. 1986 Mar 14;231(4743):1294–1297. doi: 10.1126/science.231.4743.1294. [DOI] [PubMed] [Google Scholar]
- Gorman C. M., Moffat L. F., Howard B. H. Recombinant genomes which express chloramphenicol acetyltransferase in mammalian cells. Mol Cell Biol. 1982 Sep;2(9):1044–1051. doi: 10.1128/mcb.2.9.1044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Konarska M. M., Padgett R. A., Sharp P. A. Recognition of cap structure in splicing in vitro of mRNA precursors. Cell. 1984 Oct;38(3):731–736. doi: 10.1016/0092-8674(84)90268-x. [DOI] [PubMed] [Google Scholar]
- Kozak M. Bifunctional messenger RNAs in eukaryotes. Cell. 1986 Nov 21;47(4):481–483. doi: 10.1016/0092-8674(86)90609-4. [DOI] [PubMed] [Google Scholar]
- Kozak M. Point mutations define a sequence flanking the AUG initiator codon that modulates translation by eukaryotic ribosomes. Cell. 1986 Jan 31;44(2):283–292. doi: 10.1016/0092-8674(86)90762-2. [DOI] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Levis R., Weiss B. G., Tsiang M., Huang H., Schlesinger S. Deletion mapping of Sindbis virus DI RNAs derived from cDNAs defines the sequences essential for replication and packaging. Cell. 1986 Jan 17;44(1):137–145. doi: 10.1016/0092-8674(86)90492-7. [DOI] [PubMed] [Google Scholar]
- Melton D. A., Krieg P. A., Rebagliati M. R., Maniatis T., Zinn K., Green M. R. Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter. Nucleic Acids Res. 1984 Sep 25;12(18):7035–7056. doi: 10.1093/nar/12.18.7035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Monroe S. S., Schlesinger S. Common and distinct regions of defective-interfering RNAs of Sindbis virus. J Virol. 1984 Mar;49(3):865–872. doi: 10.1128/jvi.49.3.865-872.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Monroe S. S., Schlesinger S. RNAs from two independently isolated defective interfering particles of Sindbis virus contain a cellular tRNA sequence at their 5' ends. Proc Natl Acad Sci U S A. 1983 Jun;80(11):3279–3283. doi: 10.1073/pnas.80.11.3279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ou J. H., Strauss E. G., Strauss J. H. The 5'-terminal sequences of the genomic RNAs of several alphaviruses. J Mol Biol. 1983 Jul 25;168(1):1–15. doi: 10.1016/s0022-2836(83)80319-2. [DOI] [PubMed] [Google Scholar]
- Perrault J. Origin and replication of defective interfering particles. Curr Top Microbiol Immunol. 1981;93:151–207. doi: 10.1007/978-3-642-68123-3_7. [DOI] [PubMed] [Google Scholar]
- Rigby P. W. Cloning vectors derived from animal viruses. J Gen Virol. 1983 Feb;64(Pt 2):255–266. doi: 10.1099/0022-1317-64-2-255. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strauss E. G., Rice C. M., Strauss J. H. Complete nucleotide sequence of the genomic RNA of Sindbis virus. Virology. 1984 Feb;133(1):92–110. doi: 10.1016/0042-6822(84)90428-8. [DOI] [PubMed] [Google Scholar]
- Tsiang M., Monroe S. S., Schlesinger S. Studies of defective interfering RNAs of Sindbis virus with and without tRNAAsp sequences at their 5' termini. J Virol. 1985 Apr;54(1):38–44. doi: 10.1128/jvi.54.1.38-44.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss B., Rosenthal R., Schlesinger S. Establishment and maintenance of persistent infection by Sindbis virus in BHK cells. J Virol. 1980 Jan;33(1):463–474. doi: 10.1128/jvi.33.1.463-474.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- al-Moslih M. I., Dubes G. R. The kinetics of DEAE-dextran-induced cell sensitization to transfection. J Gen Virol. 1973 Feb;18(2):189–193. doi: 10.1099/0022-1317-18-2-189. [DOI] [PubMed] [Google Scholar]