Abstract
Cellular responses to DNA damage can prevent mutations and death. In this study, we have used high-throughput screens and developed a comparative genomic approach, termed Functionome mapping, to discover conserved responses to UVC-damage. Functionome mapping uses Gene Ontology (GO) information to link proteins with similar biological functions from different organisms, and we have used it to compare 303, 311 and 288 UVC-toxicity modulating proteins from E. coli, S. pombe and S. cerevisiae, respectively. We have demonstrated that all three organisms use DNA repair, translation and aerobic respiration associated processes to modulate the toxicity of UVC, with these last two categories highlighting the importance of ribosomal proteins and electron transport machinery. Our study has demonstrated that comparative genomic approaches can be used to identify conserved responses to damage, and suggest roles for translational machinery and components of energy metabolism in optimizing the DNA damage response.
Keywords: DNA repair, Functional mapping, High-throughput screening, Ribosomal proteins, Translation, UV-damage
Introduction
Ultra-violet (UV) radiation is a major source of DNA damage and can cause the formation of cyclobutane-pyrimidine dimers (CPDs) and 6-4 photoproducts (6-4PPs) [1]. Specifically, CPDs and 6-4PPs can inhibit the progress of RNA polymerases, resulting in transcriptional blocks that can promote cell death. CPDs and 6-4PPs can also block the action of replicative DNA polymerases, which can cause either cell death or promote translesion polymerase associated mutations [2], with the generation of UV-induced mutations dependent on the specific adduct. For example, while thymine-thymine lesions are often replicated correctly, lesions that contain cytosine frequently result in cytosine to thymine transition mutations [3]. These transition mutations are found at a high frequency in the p53 tumor suppressor gene in many skin cancers [4]; thus, the efficient removal of UV-induced lesions in DNA is critical for cancer prevention.
UV-induced DNA lesions can be removed by direct reversal and nucleotide excision repair (NER). Both mechanisms have been extensively characterized in Escherichia coli [1]. Direct reversal, or photo-reactivation, is facilitated by DNA photolyase enzymes using energy from light to split the CPD and 6-4 photoproduct lesions [5]. Photolyases are found in some bacteria and are closely related to the blue light sensing cryptochrome proteins. Although functionally absent in humans, photolyases are found in Bacteria, Archaea and vertebrates [6]. UV photoproducts can also be removed from the genome through the action of proteins participating in nucleotide excision repair (NER), a process in which a short segment of damaged DNA is removed and then re-synthesized. In E. coli, NER is initially facilitated by the binding of dimeric UvrA and UvrB to bulky DNA damage and the subsequent unwinding of the DNA and recruitment of UvrC to incise DNA 3’ and 5’ to the lesion [7, 8]. Cho (UvrC homologue, ydjQ) can also make the 3’ incision downstream from the normal UvrC incision point, with Cho proposed to be an efficient 3’ endonuclease at some bulky lesions [8, 9]. After 5’ and 3’ incisions are made around the lesion, the DNA helicase UvrD displaces the excised fragment and DNA polymerase I and DNA ligase fill and seal the gap, respectively [7].
NER is found in species ranging from bacteria to humans and its mechanism of action is highly conserved [1, 10]. NER in eukaryotes involves the coordinated action of over 30 proteins and, similarly to E. coli, can occur in both a general and transcription dependent mode. In Saccharomyces cerevisiae, members of the Rad3 epistasis group participate in NER and, similarly to bacteria, these activities include proteins that recognize bulky lesions, those that incise the DNA 5’ and 3’ to the lesion and others that remove the DNA fragment containing the lesion [1]. In general, defects in genes belonging to the Rad3 epistasis group confer sensitivity to UV. Importantly, NER has been shown to operate in a global and transcription-coupled manner, with the latter coordinating DNA repair with the action of RNA polymerases [11, 12]. NER defects in humans can lead to Xeroderma Pigmentosum (XP), Cockayne's Syndrome, and Trichothiodystrophy, all of which are associated with varying degrees of increased UV-sensitivity and in some cases, neurodegenerative conditions [1, 13]. XP patients in particular demonstrate UV-induced genome instability and are diagnosed with skin cancer 50 years earlier than the general population [14].
In most organisms studied to date, DNA repair pathways are activated after DNA damage and this activation usually coincides with activation of a global signaling program. For example, UV radiation has been shown to induce the SOS response in E. coli. The SOS response is regulated by the RecA protein, a single stranded DNA binding protein that accumulates at sites of DNA damage. RecA protein bound to DNA will promote cleavage of the repressor protein LexA [15]. LexA cleavage up-regulates the transcription of many genes important for cell survival after DNA damage, including those in the NER and recombination pathways [16, 17]. DNA replication and repair are not the only cellular processes up-regulated by the SOS response. Other transcripts regulated as part of the SOS response include those whose corresponding proteins are involved in transcription (LexA, RpoD, Fis), nucleoside metabolism (NrdA, NrdB, GrxA), translation (ArgS, PrfB, PrfC) and heat shock (YcaH, CorA, GlvB). Similarly to prokaryotic cells, eukaryotic organisms have DNA damage response (DDR) pathways activated by DNA strand breaks. Recognition of DNA strand breaks is facilitated by protein-based damage detection resulting in signaling through the ataxia telangiectasia mutated (ATM) and ATM and Rad3-related (ATR) kinases [18-20]. In S. cerevisiae, the ATM and ATR homologs are named Mec1 and Tel1. Mec1-dependent transcriptional reprogramming occurs after DNA damage and includes hundreds of different transcripts corresponding to a wide range of cellular proteins. Transcripts regulated in a Mec1-dependent manner include those associated with the DDR, as well as transcripts belonging to the environmental stress response (ESR) [21]. The ESR is thought to protect the internal homeostasis of the cell and includes transcripts whose corresponding proteins participate in reactive oxygen species detoxification, protein folding and degradation, carbohydrate metabolism, ribosomal function and translational regulation. The regulation of a broad range of cellular functions after DNA damage is also conserved in humans. For example, ATM and ATR have been shown to phosphorylate over 700 downstream proteins including the DDR associated proteins Chk1, Chk2, p53, Brca1 and Cdc25 in addition to many other targets [22, 23]. The other ATM/ATR targets belong to the cellular processes of nucleic acid metabolism, protein metabolism, cell cycle, signal transduction, cell structure and motility, protein traffic and oncogenesis. Thus, the regulation of many different cellular processes after DNA damage is a theme that is conserved from bacteria to lower and higher eukaryotes. In addition, the diversity of responses regulated by SOS, Mec1 and ATM/ATR signaling suggests that DNA repair is coordinated with other cellular processes.
In an effort to identify proteins and pathways that help cells respond to damaging agents, scientists have screened gene deletion libraries derived from different single celled organisms [24-28]. In these libraries, gene deletion is facilitated by targeted homologous recombination to replace a specific gene with a selection cassette. Gene deletion libraries are made when, in theory, all genes in a genome have been individually removed, and the resulting mutants have been arrayed into separate wells of a multi-well plate. In diploid cells, removal of one allele can result in haploinsufficiency or cell death in some cases [28]. In haploid cells, only 75-95% of the genes can be removed to yield a viable mutant, as many genes encode essential activities that when removed result in cell death. Gene deletion libraries have been made in E. coli, S. cerevisiae, and Schizosaccharomyces pombe [28-30], and all of these resources have been used in organized screens and have proven to be valuable tools for identifying gene products that modulate the toxicity of different DNA damaging agents. For example, after the S. cerevisiae gene deletion library was screened against UVC, we reported that in addition to DNA repair and cell cycle, a number of proteins associated with RNA and protein metabolism, aerobic respiration, and other functional categories were classified as toxicity-modulating [24]. This identification of a broad range of unexpected biological processes in functional screens supports the contention that either the corresponding proteins are linked to the DNA damage response in some way or that other metabolic pathways are coordinated with the repair of UV-induced DNA lesions. Other possibilities exist, though, and the identified UV-toxicity modulating proteins might just be experimental anomalies specific to the S. cerevisiae gene deletion library, as these cells have a distinct physiology.
In the following study, we have utilized comparative functional genomic approaches to identify similar biological processes that modulate the toxicity of UV in three different and evolutionarily distinct cell types: E. coli, S. cerevisiae, and S. pombe. Specifically, we have screened two different species-specific deletion libraries, E. coli and S. pombe, to identify 303 and 311 UV-toxicity modulating proteins, respectively. We have also computationally compared the UV-toxicity modulating proteins identified from E. coli and S. pombe to our previously reported list of 288 UV-toxicity modulating proteins from S. cerevisiae [24]. To do this, we have developed a functional interactome mapping approach to identify GO-specified biological processes that are significantly enriched for UV-toxicity modulating proteins from multiple organisms. We have demonstrated that multiple species use the biological processes associated with DNA repair, translation and aerobic respiration to modulate the toxicity of UV. In addition, we have demonstrated that our functional mapping approach is predictive and can be used to identify UV-toxicity modulating proteins in different cell types. Finally, at the mechanistic level, our results support the idea that cells use translational machinery and ATP levels to optimize DNA repair or coordinate DNA repair with other metabolic processes.
Results and Discussion
303 E. coli gene deletion mutants identified as sensitive to UV
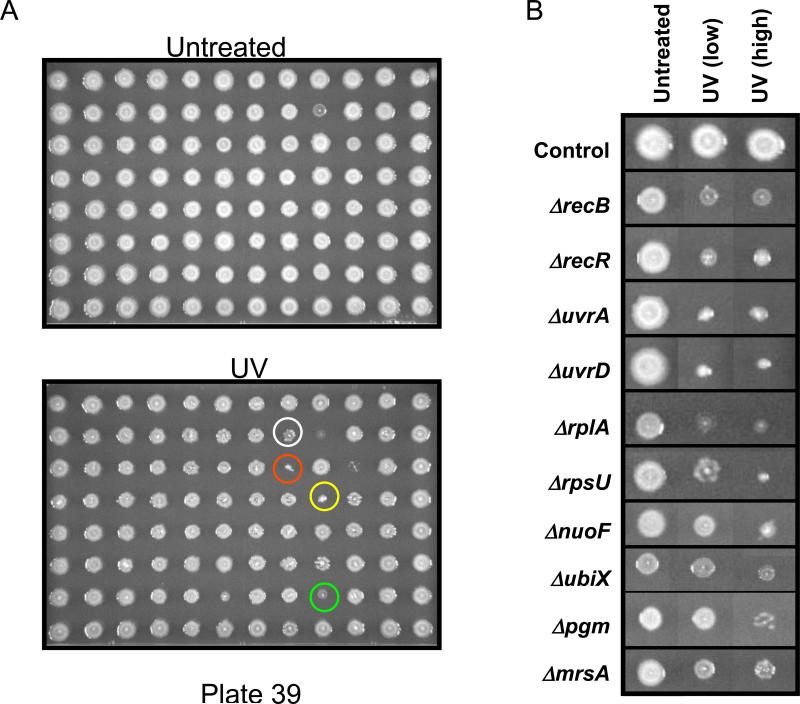
The E. coli deletion set from Keio contains 8,640 mutants specific for 3,968 genes [29]. Mutants were individually spotted onto LB-agar plates using the 96-syringe Matrix Scientific Hydra. Upon drying, the cells were left untreated or exposed to two different doses of UV (10.0 and 12.5 joules) and then allowed to grow for 24 hours (Figure 1A). UV exposure conditions were chosen such that cells deficient in RecA and Cho were consistently identified as UV-sensitive in preliminary experiments. In total, 364 plates containing 34,944 spotted cultures were assayed as described above and the resulting images of these plates were compiled into Supplemental Figure S1. We analyzed our UV screen data using a similar methodology as previously reported [26]. First, a virtual mutant representing at least two isolates of each gene mutant was given a UV-toxicity modulating score, derived from the behavior of the corresponding mutants exposed to different doses of UV. For example, the Keio library contains 2 mutants representing uvrD, and the UV-toxicity modulating score describes the behavior of each of these uvrD mutants after exposure to two different doses of UV. For each exposure condition (low and high), mutants that demonstrated reduced growth were given a score of 4 to 2, depending on the severity of the growth defect (4 = high, 2 = low), and those displaying a color change were scored 1. In theory, the most sensitive virtual mutants scored 16 (4 + 4 + 4 + 4), because two corresponding isolates displayed severely reduced growth (score of 4) at both UV-exposure conditions. We used a cut-off of 3 to identify virtual mutants that were slightly sensitive to UV, as this category of mutants repeatedly displayed a UV-induced reduction in growth and/or a color change (Figure 1B).
Figure 1. Genomic phenotyping of E. coli mutants with UV.
(A) 96 gene deletion mutants were spotted onto agar plates, left untreated or exposed to two doses of UV, incubated at 37° C for 16 hours and imaged. These doses were used because some cells were only sensitive to the higher dose, while others were sensitive to both conditions. White, red, yellow and green circles identify the UV-sensitive mutants ΔruvA, ΔruvC, ΔuvrA and ΔholC, respectively. (B) Images were taken from many different plates and recompiled to demonstrate that varying degrees of UV-sensitivity were observed in the screen. Examples of a color change from white to grey are shown for ΔrecB and ΔmrsA.
Ultimately, we identified 303 E. coli virtual mutants as being sensitive to UV (Table I) and observed a range of sensitivities from high (25 mutants scored from 10 to 16), medium (30 mutants scored from 7 to 9), low (90 mutants scored from 5 to 6), to slightly (157 mutants scored from 3 to 4). Mutants classified as being highly sensitive to UV included uvrAΔ, uvrBΔ, uvrCΔ, and uvrDΔ, all of whose corresponding proteins are components of NER. It is known that uvrAΔ, uvrBΔ, uvrCΔ, and uvrDΔ mutants are sensitive to UV [1, 31] and their identification in our screen validated our methodology. Other mutants highly sensitive to UV included those deficient in the peptide chain release factor PrfB and the 50S ribosomal subunit protein RflA. These results suggest that a defect in ribosome assembly, ribosomal protein deficiency or corrupted protein synthesis disrupts cellular responses to damage. Mutants in the medium sensitivity category included dnaGΔ, parCΔ, and ruvAΔ, all of which encode activities associated with DNA synthesis and DNA repair. Again, these mutants highlight the importance of DNA repair and DNA metabolism after UV-damage and their presence in our list was expected. Other mutants that fall into the medium sensitive category included those cells deficient in the 30S ribosome binding factor (RbfA), ribonuclease RNase T and adenylate cyclase (CycA), highlighting the roles for protein, RNA and small molecule metabolism in response to UV-damage. Mutants in the low sensitivity category included those deficient in the NER activity Cho, the 30S ribosomal subunit protein (RpsT) and NADH ubiquinone oxidoreductase (NuoF). Our mutants classified as having low sensitivity further demonstrated that deficiencies in DNA repair and protein synthesis corrupt cellular viability after UV-exposure. In addition, the identification of NuoF in our UV-sensitive catalog introduced respiration and energy metabolism to the list of cellular process important after damage. The slightly sensitive list is also populated by mutants defective in DNA repair (UmuC), ribosome assembly (RpsO), protein synthesis (PoxA), RNA metabolism (Tgt) and aerobic respiration (NuoH), further highlighting the wide range of cellular processes involved in the response to UV-damage. [We note for the preceding paragraph all annotation information was obtained from EcoGene [32].]
Table I.
Proteins corresponding to the 301 UV-sensitive mutants identified in E. coli
| Protein | Description | Sensitivity |
|---|---|---|
| ArtM | arginine transporter subunit, membrane component of ABC superfamily | High |
| DedD | affects Col V production | High |
| DnaT | DNA biosynthesis protein (primosomal protein I) | High |
| Fis | global DNA-binding transcriptional dual regulator | High |
| FlgH | flagellar protein of basal-body outer-membrane L ring | High |
| FtnA | ferritin iron storage protein (cytoplasmic) | High |
| Mfd | transcription-repair coupling factor | High |
| Pnp | polynucleotide phosphorylase/polyadenylase | High |
| PrfB | peptide chain release factor RF-2 | High |
| PriA | Primosome factor n’ (replication factor Y) | High |
| RecA | DNA strand exchange and recombination protein with protease and nuclease activity | High |
| RecC | exonuclease V (RecBCD complex) | High |
| RecO | gap repair protein | High |
| RecR | gap repair protein | High |
| RuvC | component of RuvABC resolvasome, endonuclease | High |
| UvrA | ATPase and DNA damage recognition protein of nucleotide excision repair excinuclease UvrABC | High |
| UvrB | excinulease of nucleotide excision repair | High |
| UvrC | excinuclease UvrABC, endonuclease subunit | High |
| UvrD | DNA-dependent ATPase I and helicase II | High |
| YohF | predicted oxidoreductase with NAD(P)-binding Rossmann-fold domain | High |
| CyaA | adenylate cyclase | Medium |
| DacD | D-alanyl-D-alanine carboxypeptidase (penicillin-binding protein 6b) | Medium |
| Dam | DNA adenine methylase | Medium |
| DnaG | DNA primase | Medium |
| DnaK | chaperone Hsp70, co-chaperone with DnaJ | Medium |
| Hfq | HF-I, host factor for RNA phage Q beta replication | Medium |
| IntA | CP4-57 prophage; integrase | Medium |
| JW1227.5 | unknown function | Medium |
| JW5460 | predicted protein (pseudogene) | Medium |
| ParC | DNA topoisomerase IV, subunit A | Medium |
| PdxH | pyridoxine 5'-phosphate oxidase | Medium |
| RbfA | 30s ribosome binding factor | Medium |
| RecF | gap repair protein | Medium |
| Rnt | ribonuclease T (RNase T) | Medium |
| RplA | 50S ribosomal subunit protein L1 | Medium |
| RseA | anti-sigma factor | Medium |
| RuvA | component of RuvABC resolvasome, regulatory subunit | Medium |
| Uup | fused predicted transporter subunits of ABC superfamily: ATP-binding components | Medium |
| YbaB | conserved protein | Medium |
| YceB | predicted lipoprotein | Medium |
| YciG | unknown function | Medium |
| YdaE | Rac prophage; conserved protein | Medium |
| YddO | D-ala-D-ala transporter subunit, ATP-binding component of ABC superfamily | Medium |
| YdeH | unknown function | Medium |
| YdiI | unknown function | Medium |
| YehP | unknown function | Medium |
| YeiC | predicted kinase | Medium |
| YgfA | predicted ligase | Medium |
| YibP | protease with a role in cell division | Medium |
| YihX | predicted hydrolase | Medium |
| YjeK | predicted lysine aminomutase | Medium |
| YneF | predicted diguanylate cyclase | Medium |
| YnjI | predicted inner membrane protein | Medium |
| YohG | unknown function | Medium |
| YphB | conserved protein | Medium |
| Cho | endonuclease of nucleotide excision repair | Low |
| CybB | Cytochrome b561 | Low |
| CysA | sulfate/thiosulfate transporter subunit, ATP-binding component of ABC superfamily | Low |
| DedA | conserved inner membrane protein | Low |
| FecA | KpLE2 phage-like element; ferric citrate outer membrane transporter | Low |
| FecC | KpLE2 phage-like element; iron-dicitrate transporter subunit, membrane component of ABC superfamily | Low |
| FimC | chaperone, periplasmic | Low |
| FumC | fumarate hydratase (fumarase C), aerobic Class II | Low |
| GntP | fructuronate transporter | Low |
| Gph | phosphoglycolate phosphatase | Low |
| HolC | DNA polymerase III, chi subunit | Low |
| HolD | DNA polymerase III, psi subunit | Low |
| HtgA | unknown function | Low |
| IlvC | ketol-acid reductoisomerase, NAD(P)-binding | Low |
| IspZ | predicted inner membrane protein | Low |
| JW0258 | predicted IS protein | Low |
| JW5411 | unknown function | Low |
| JW5766 | unknown function | Low |
| MtlA | fused mannitol-specific PTS enzymes: IIA components , IIB components , IIC components | Low |
| NuoF | NADH:ubiquinone oxidoreductase, chain F | Low |
| OppD | oligopeptide transporter subunit, ATP-binding component of ABC superfamily | Low |
| PriB | primosomal protein N | Low |
| PrpB | 2-methylisocitrate lyase | Low |
| RadA | predicted repair protein | Low |
| RecB | exonuclease V (RecBCD complex), beta subunit | Low |
| RpsT | 30S ribosomal subunit protein S20 | Low |
| SgcE | KpLE2 phage-like element; predicted epimerase | Low |
| UbiX | 3-octaprenyl-4-hydroxybenzoate carboxy-lyase | Low |
| YbhD | unknown function | Low |
| YcaM | predicted transporter | Low |
| YceP | cold shock gene | Low |
| YcfS | conserved protein | Low |
| YeaH | conserved protein | Low |
| YebG | conserved protein regulated by LexA | Low |
| yeeL | predicted protein, C-ter fragment (pseudogene) | Low |
| YfaO | predicted NUDIX hydrolase | Low |
| YfbQ | predicted aminotransferase | Low |
| YfdZ | prediected aminotransferase, PLP-dependent | Low |
| YfiM | predicted protein | Low |
| YgaY | predicted transporter (pseudogene) | Low |
| YgfY | conserved protein | Low |
| YnjE | predicted thiosulfate sulfur transferase | Low |
| YodB | predicted cytochrome | Low |
| DicB | Qin prophage; cell division inhibition protein | Slightly |
| FhiA | flagellar system protein, promoterless fragment (pseudogene) | Slightly |
| FimH | minor component of type 1 fimbriae | Slightly |
| FliD | flagellar filament capping protein | Slightly |
| GroL | Cpn60 chaperonin GroEL, large subunit of GroESL | Slightly |
| HisB | fused histidinol-phosphatase, imidazoleglycerol-phosphate dehydratase | Slightly |
| IhfB | integration host factor (IHF), DNA-binding protein, beta subunit | Slightly |
| JW1277 | unknown function | Slightly |
| JW5474 | unknown function | Slightly |
| LuxS | S-ribosylhomocysteinase | Slightly |
| MinC | cell division inhibitor | Slightly |
| NhaA | sodium-proton antiporter | Slightly |
| QueA | S-adenosylmethionine:tRNA ribosyltransferase-isomerase | Slightly |
| Rof | modulator of Rho-dependent transcription termination | Slightly |
| RrmJ | 23S rRNA methyltransferase | Slightly |
| RspB | predicted oxidoreductase, Zn-dependent and NAD(P)-binding | Slightly |
| SapA | predicted antimicrobial peptide transporter subunit, periplasmic-binding component of ABC superfamily | Slightly |
| SotB | predicted arabinose transporter | Slightly |
| SpeG | spermidine N1-acetyltransferase | Slightly |
| SucB | dihydrolipoyltranssuccinase | Slightly |
| ThrL | thr operon leader peptide | Slightly |
| WbbL | lipopolysaccharide biosynthesis protein, N-ter fragment (pseudogene) | Slightly |
| WecG | UDP-N-acetyl-D-mannosaminuronic acid transferase | Slightly |
| YadB | Glutamyl-Q tRNA(Asp) synthase | Slightly |
| YadM | predicted fimbrial-like adhesin protein | Slightly |
| YagT | predicted xanthine dehydrogenase, 2Fe-2S subunit | Slightly |
| YahN | neutral amino-acid efflux system | Slightly |
| YbgI | conserved metal-binding protein | Slightly |
| YcgR | protein involved in flagellar function | Slightly |
| YchE | predicted inner membrane protein | Slightly |
| YciQ | unknown function | Slightly |
| YcjG | L-Ala-D/L-Glu epimerase | Slightly |
| YdbL | conserved protein | Slightly |
| YdfA | Qin prophage; predicted protein | Slightly |
| YdhL | conserved protein | Slightly |
| YdhZ | predicted protein | Slightly |
| YdiK | predicted inner membrane protein | Slightly |
| YeeV | CP4-44 prophage; toxin of the YeeV-YeeU toxin-antitoxin system | Slightly |
| YegV | predicted kinase | Slightly |
| YehK | predicted protein | Slightly |
| YfdI | unknown function | Slightly |
| YfhD | predicted transglycosylase | Slightly |
| YgdB | predicted protein | Slightly |
| YgfZ | predicted folate-dependent regulatory protein | Slightly |
| YhbX | predicted hydrolase, inner membrane | Slightly |
| YmgD | unknown function | Slightly |
| YoaF | conserved outer membrane protein | Slightly |
| YobD | conserved inner membrane protein | Slightly |
| AcrR | DNA-binding transcriptional repressor | Slightly |
| AgaI | galactosamine-6-phosphate isomerase | Slightly |
| AraF | L-arabinose transporter subunit, periplasmic-binding component of ABC superfamily | Slightly |
| Asr | acid shock-inducible periplasmic protein | Slightly |
| BtuE | predicted glutathione peroxidase | Slightly |
| CycA | D-alanine/D-serine/glycine transporter | Slightly |
| DeaD | ATP-dependent RNA helicase | Slightly |
| EamA | cysteine and O-acetyl-L-serine efflux system | Slightly |
| FruA | fused fructose-specific PTS enzymes: IIBcomponent, IIC components | Slightly |
| FruB | fused fructose-specific PTS enzymes: IIA component, HPr component | Slightly |
| HokD | Qin prophage; small toxic polypeptide | Slightly |
| HybA | hydrogenase 2 4Fe-4S ferredoxin-type component | Slightly |
| IhfA | integration host factor (IHF), DNA-binding protein, alpha subunit | Slightly |
| JW5386 | predicted protein | Slightly |
| JW5846 | predicted protein | Slightly |
| Kbl | glycine C-acetyltransferase | Slightly |
| MglC | methyl-galactoside transporter subunit, membrane component of ABC superfamily | Slightly |
| MiaA | delta(2)-isopentenylpyrophosphate tRNA-adenosine transferase | Slightly |
| MotB | protein that enables flagellar motor rotation | Slightly |
| NagA | N-acetylglucosamine-6-phosphate deacetylase | Slightly |
| OtsB | trehalose-6-phosphate phosphatase, biosynthetic | Slightly |
| PotA | polyamine transporter subunit, ATP-binding component of ABC superfamily | Slightly |
| PoxA | predicted lysyl-tRNA synthetase | Slightly |
| PrpE | predicted propionyl-CoA synthetase with ATPase domain | Slightly |
| RhsA | rhsA element core protein RshA | Slightly |
| RpmF | 50S ribosomal subunit protein L32 | Slightly |
| RpsU | 30S ribosomal subunit protein S21 | Slightly |
| SsuB | alkanesulfonate transporter subunit, ATP-binding component of ABC superfamily | Slightly |
| SufB | Complexed with SufC and SufD | Slightly |
| TfaR | Rac prophage; predicted tail fiber assembly protein | Slightly |
| TrpL | trp operon leader peptide | Slightly |
| UbiF | 2-octaprenyl-3-methyl-6-methoxy-1,4-benzoquinol oxygenase | Slightly |
| Upk | undecaprenyl pyrophosphate phosphatase | Slightly |
| XylG | fused D-xylose transporter subunits of ABC superfamily; ATP-binding components | Slightly |
| YagE | CP4-6 prophage; predicted lyase/synthase | Slightly |
| YbaL | predicted transporter with NAD(P)-binding Rossmann-fold domain | Slightly |
| YbdF | unknown function | Slightly |
| YbeQ | unknown function | Slightly |
| YbfO | unknown function | Slightly |
| YbgQ | predicted outer membrane protein | Slightly |
| YbjS | predicted NAD(P)H-binding oxidoreductase with NAD(P)-binding Rossmann-fold domain | Slightly |
| YcaQ | unknown function | Slightly |
| YccA | inner membrane protein | Slightly |
| YccS | predicted inner membrane protein | Slightly |
| YcdL | unknown function | Slightly |
| YceH | unknown function | Slightly |
| YciW | predicted oxidoreductase | Slightly |
| YcjO | predicted sugar transporter subunit: membrane component of ABC superfamily | Slightly |
| YdaM | predicted diguanylate cyclase, GGDEF domain signalling protein | Slightly |
| YdbJ | unknown function | Slightly |
| YddG | predicted methyl viologen efflux pump | Slightly |
| YdhB | predicted DNA-binding transcriptional regulator | Slightly |
| YdhQ | unknown function | Slightly |
| YdiD | Acyl-CoA synthase | Slightly |
| YdjN | predicted transporter | Slightly |
| yedS | predicted protein, middle fragment (pseudogene) | Slightly |
| YeeJ | adhesin | Slightly |
| YeiA | predicted oxidoreductase | Slightly |
| YfbU | unknown function | Slightly |
| YfcU | predicted export usher protein | Slightly |
| YgcR | predicted flavoprotein | Slightly |
| YhcP | p-hydroxybenzoic acid efflux system component | Slightly |
| YhdZ | predicted amino-acid transporter subunit, ATP-binding component of ABC superfamily | Slightly |
| YliA | Glutathione transporter ATP-binding protein | Slightly |
| YmfA | predicted inner membrane protein | Slightly |
| YmfE | e14 prophage; predicted inner membrane protein | Slightly |
| YmfO | e14 prophage; conserved protein | Slightly |
| YnbD | predicted phosphatase, inner membrane protein | Slightly |
| YnhG | unknown function | Slightly |
| YoaG | unknown function | Slightly |
| BtuC | vitamin B12 transporter subunit: membrane component of ABC superfamily | Slightly |
| BtuD | vitamin B12 transporter subunit : ATP-binding component of ABC superfamily | Slightly |
| CinA | unknown function | Slightly |
| CydB | cytochrome d terminal oxidase, subunit II | Slightly |
| DcuB | C4-dicarboxylate antiporter | Slightly |
| GidA | glucose-inhibited cell-division protein | Slightly |
| GrxA | glutaredoxin 1, redox coenzyme for ribonucleotide reductase (RNR1a) | Slightly |
| GutQ | D-arabinose 5-phosphate isomerase | Slightly |
| HisG | ATP phosphoribosyltransferase | Slightly |
| HyfB | hydrogenase 4, membrane subunit | Slightly |
| JW1421 | unknown function | Slightly |
| JW1640 | unknown function | Slightly |
| JW2679 | unknown function | Slightly |
| JW3017 | unknown function | Slightly |
| JW4246 | KpLE2 phage-like element; predicted protein | Slightly |
| JW5029 | unknown function | Slightly |
| LdcC | lysine decarboxylase 2, constitutive | Slightly |
| LeuC | 3-isopropylmalate isomerase subunit, dehydratase component | Slightly |
| ManX | fused mannose-specific PTS enzymes: IIA component, IIB component | Slightly |
| MenC | o-succinylbenzoyl-CoA synthase | Slightly |
| MrsA | phosphoglucosamine mutase | Slightly |
| NhoA | N-hydroxyarylamine O-acetyltransferase | Slightly |
| NuoH | NADH:ubiquinone oxidoreductase, membrane subunit H | Slightly |
| NuoK | NADH:ubiquinone oxidoreductase, membrane subunit K | Slightly |
| PaaC | predicted multicomponent oxygenase/reductase subunit for phenylacetic acid degradation | Slightly |
| PaaX | DNA-binding transcriptional repressor of phenylacetic acid degradation, aryl-CoA responsive | Slightly |
| PdxB | erythronate-4-phosphate dehydrogenase | Slightly |
| Pgm | phosphoglucomutase | Slightly |
| PpiB | peptidyl-prolyl cis-trans isomerase B (rotamase B) | Slightly |
| Rmf | ribosome modulation factor | Slightly |
| RpsO | 30S ribosomal subunit protein S15 | Slightly |
| Rtn | conserved protein | Slightly |
| SanA | predicted protein | Slightly |
| SapB | predicted antimicrobial peptide transporter subunit, membrane component of ABC superfamily | Slightly |
| SfmF | predicted fimbrial-like adhesin protein | Slightly |
| SirB2 | predicted transcriptional regulator | Slightly |
| SufD | component of SufBCD complex | Slightly |
| Tgt | tRNA-guanine transglycosylase | Slightly |
| TruA | pseudouridylate synthase I | Slightly |
| UgpA | glycerol-3-phosphate transporter subunit, membrane component of ABC superfamily | Slightly |
| UmuC | DNA polymerase V, subunit C | Slightly |
| Usg | predicted semialdehyde dehydrogenase | Slightly |
| VacJ | predicted lipoprotein | Slightly |
| YagW | predicted receptor | Slightly |
| YaiT | predicted protein | Slightly |
| ycdN | predicted protein, N-ter fragment (pseudogene) | Slightly |
| YcfF | purine nucleoside phosphoramidase | Slightly |
| YcgK | unknown function | Slightly |
| YciF | unknown function | Slightly |
| YciN | unknown function | Slightly |
| YcjD | unknown function | Slightly |
| YdaY | Rac prophage; predicted protein | Slightly |
| YddP | D-ala-D-ala transporter subunit, ATP-binding component of ABC superfamily | Slightly |
| YdeE | predicted transporter | Slightly |
| YdeP | predicted oxidoreductase | Slightly |
| YdfZ | unknown function | Slightly |
| YdgD | predicted peptidase | Slightly |
| YdjE | predicted transporter | Slightly |
| YecN | unknown function | Slightly |
| YeeY | predicted DNA-binding transcriptional regulator | Slightly |
| YefI | lipopolysaccharide biosynthesis protein | Slightly |
| YegN | multidrug efflux system, subunit B | Slightly |
| YehL | unknown function | Slightly |
| YehM | unknown function | Slightly |
| YeiE | predicted DNA-binding transcriptional regulator | Slightly |
| YejB | predicted oligopeptide transporter subunit, membrane component of ABC superfamily | Slightly |
| YejO | predicted autotransporter outer membrane protein | Slightly |
| yfaS | predicted protein, C-ter fragment (pseudogene) | Slightly |
| YfbK | conserved protein | Slightly |
| YfdQ | CPS-53 (KpLE1) prophage; predicted protein | Slightly |
| YfeH | unknown function | Slightly |
| YfhQ | predicted methyltransferase | Slightly |
| YgbI | predicted DNA-binding transcriptional regulator | Slightly |
| YgcI | unknown function | Slightly |
| YgdI | unknown function | Slightly |
| YgfE | protein that localizes to the cytokinetic ring | Slightly |
| YhhZ | unknown function | Slightly |
| YjgK | unknown function | Slightly |
| YjgM | predicted acetyltransferase | Slightly |
| YkfC | CP4-6 prophage; conserved protein | Slightly |
| YmbA | unknown function | Slightly |
| YmfL | e14 prophage; predicted DNA-binding transcriptional regulator | Slightly |
| YneC | unknown function | Slightly |
| YneE | unknown function | Slightly |
| YojI | fused predicted multidrug transport subunits of ABC superfamily: membrane component, ATP-binding component | Slightly |
| YqcC | unknown function | Slightly |
| YtfF | predicted inner membrane protein | Slightly |
Conserved biological process that modulate UV-toxicity identified by Functionome mapping
After we identified 303 E. coli proteins that modulate the toxicity of UV-damage and linked them to a number of cellular processes, we wanted to determine which of these responses were conserved across species. In most model organisms, DNA repair is essential for survival after UV-damage and we wanted to determine if any other biological processes showed a similar association. Previously, we had performed high throughput screens in S. cerevisiae and identified 288 proteins that modulate the toxicity of UV in this eukaryotic organism [24]. To identify conserved cellular responses to UV-damage, we developed a functional interactome approach to align our list of UV-toxicity modulating proteins from E. coli with those previously identified in S. cerevisiae. Matching toxicity modulating proteins from each organism using similarities in primary amino acid sequence is problematic in that sequence homology is limited between these prokaryotic and eukaryotic organisms. Conversely, functional classifications offered an avenue to perform a cross-species analysis of UV-toxicity modulating data. To compare functional categories associated with UV-toxicity modulating proteins, we took advantage of GO annotations. GO provides a controlled vocabulary describing a protein's biological process and the information has been methodically compiled by annotation teams [33]. Annotations reflect the biological function of each protein, and GO assignments are made using manual and automated approaches. In both cases, annotation assignments are based on an attributable source (literature, another database or computational study) and the annotation is detailed to indicate the type of evidence used to make each assignment. The controlled vocabulary allows for proteins from different organisms to be linked via an identical biological process and we exploited GO associations to generate a cross-species functional interactome that we call a Functionome. Functional associations have previously been used to group species-specific data and analyze high-throughput data sets [22, 34, 35], but our cross-species Functionome expands this approach to efficiently compare data from different organisms. To compile the E. coli-S. cerevisiae Functionome, we identified 511 GO biological processes found in both organisms. Next, we associated 3,120 E. coli and 4,415 S. cerevisiae proteins with these GO identifiers. We note that not all E. coli and S. cerevisiae proteins are found in this Functionome, as some proteins are only associated with species-specific functional information. The 7,535 proteins found in the Functionome averaged 2.4 functional interactions per protein, with a total of 18,250 functional interactions associated with the compiled structure (Figure 2A).
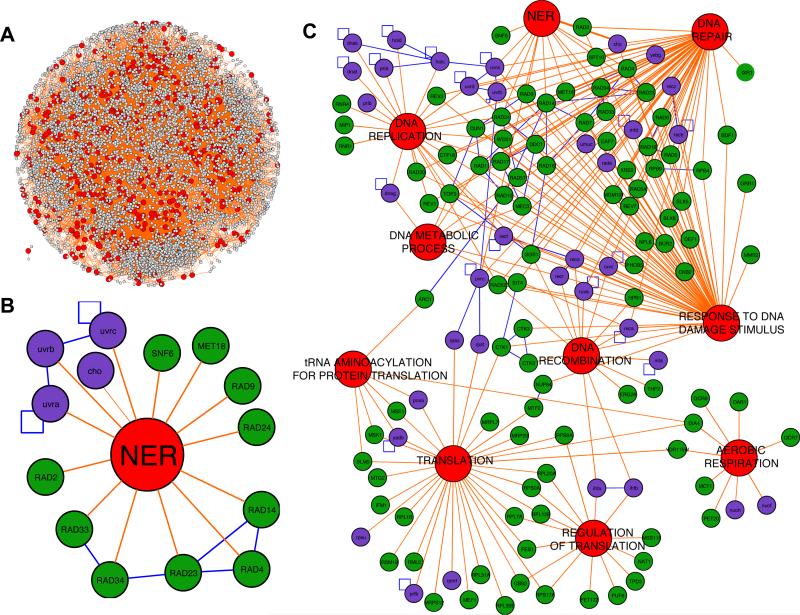
Figure 2. Functional interactome mapping identified multi-species nodes over-represented with UV-toxicity modulating proteins.
(A) The Functionome was compiled using GO identifiers for biological processes specific to 3,120 E. coli and 4,271 S. cerevisiae proteins (small grey spheres). A total of 511 GO identifiers (large red spheres) and 18,254 functional links (orange lines) were used to compile the functional interactome. (B) The Functionome was computationally analyzed to identify nodes over-represented with both E. coli (purple spheres, lower case protein names) and S. cerevisiae (green spheres, upper case protein names) UV-toxicity modulating proteins. One of the top scoring functional nodes was NER (p < 10-12). Blue lines represent protein-protein interactions. (C) All functional nodes that were over-represented (p < 0.06) with UV-toxicity modulating proteins from both E. coli and S. cerevisiae were visualized using Cytoscape.
After we compiled the Functionome, we mapped it with UV-toxicity modulating proteins from E. coli (171) and S. cerevisiae (236). Next, we computationally analyzed this mapped Functionome to identify GO-nodes significantly over-represented with UV-toxicity modulating proteins from both organisms. Significance was verified using two methodologies: random sampling of proteins in the Functionome and network randomizations. Using our computational approaches, we expected to identify the GO biological process of NER as a node that was significantly over-represented with UV-toxicity modulating proteins from both organisms. We specified that in order to be identified in our analysis at least two UV-toxicity modulating proteins from each organism must be associated with the GO biological process. This was done to prevent the identification of GO-nodes predominated by a single organism's UV-toxicity modulating proteins. As expected, our computational analysis identified NER as being over-represented with UV-toxicity modulating proteins from both E. coli and S. cerevisiae. NER was, in fact, one of the top scoring nodes (P < 0.0001) in our Functionome analysis (Figure 2B), helping to validate our methodology. Additionally, we identified 9 other GO biological processes that met our criteria (Figure 2C and Table II).
Table II.
GO biological processes over-represented in both E. coli and S. cerevisiae UV-toxicity modulating proteins, as identified by Functionome mapping
| GO Functional Category | E. coli | Saccharomyces cerevisiae | Combined | |||
|---|---|---|---|---|---|---|
| # UV Sensitive | Genes | # UV Sensitive | Genes | Total UV Sensitive | P- Value | |
| Response to DNA damage stimulus | 16 | cho, mfd, rada, reca, recb, recc, recf, reco, recr, ruva, ruvc, umuc, uvra, uvrb, uvrc, uvrd, yebg | 51 | RAD16, RAD18, SLX5, DUN1, NUP84, RAD57, HPR1, RAD9, RAD34, XRS2, RAD30, RAD23, SLX8, RAD4, RAD24, RAD6, RPB9, MMS2, RAD54, RAD2, WSS1, MET18, REV7, CTK2, RPB4, RAD7, GRR1, MGM101, DEF1, CTK1, RAD5, BUR2, TOP3, MEC3, BDF1, RAD33, RAD52, RAD10, CTK3, CTF18, NPL6, SGS1, RAD14, EAF7, CKB2, REV1, RAD17, RAD1, PHO85, REV3, DDC1 | 67 | 1.00E-04 |
| DNA repair | 17 | cho, mfd, radA, recA, recB, recC, recF, recO, recR, ruvA, ruvC, umuC, uvrA, uvrB, uvrC, uvrD, yebG | 42 | RAD16, RAD18, SLX5, SIT4, DUN1, RAD57, RAD9, RAD34, XRS2, RAD30, RAD23, SLX8, RAD4, RAD24, RAD6, RPB9, RAD54, RAD2, WSS1, MET18, REV7, SPT10, RPB4, RAD7, MGM101, IXR1, RAD5, TOP3, MEC3, BDF1, RAD33, RAD52, RAD10, CTF18, SGS1, RAD14, EAF7, REV1, RAD17, RAD1, REV3, DDC1 | 59 | 1.00E-04 |
| Nucleotide-excision repair | 3 | uvrA, uvrB, uvrC, cho | 10 | RAD9, RAD34, RAD23, RAD4, RAD24, RAD2, SNF6, MET18, RAD33, RAD14 | 13 | 1.00E-04 |
| DNA replication | 9 | dnaG, dnaK, dnaT, holC, holD, priA, priB, recF, uvrD | 14 | RAD9, RAD30, RNR1, RAD24, RNR4, WSS1, TOP3, RAD52, CTF18, SGS1, MIP1, REV1, RAD17, REV3 | 23 | 1.00E-04 |
| DNA metabolic process | 3 | dnaG, recA, recR | 4 | RAD57, TOP3, MEC3, RAD1 | 7 | 1.24E-02 |
| tRNA aminoacylation for protein translation | 2 | poxA, yadB | 5 | SLM5, ARC1, DIA4, MSK1, MSE1 | 7 | 1.24E-02 |
| Aerobic respiration | 2 | nuoF, nuoH | 7 | YDR115W, QCR7, QCR6, DIA4, OAR1, MCT1, PET20 | 9 | 1.64E-02 |
| Translation | 7 | poxA, prfB, rpmF, rpsO, rpsT, rpsU, yadB | 26 | RPS8A, FES1, SLM5, MTF2, RPL31A, RPL35B, YDR115W, CBS2, MRPL7, MRP20, RPL12B, RML2, RPL7A, RPL1B, RPS0A, DIA4, MTG2, CTK1, MEF1, RPS17A, RPL20A, MSK1, MRPS12, RSM19, IFM1, MSE1 | 33 | 2.78E-02 |
| Regulation of translation | 2 | ihfA, ihfB | 13 | TPD3, RPS8A, FES1, NAT1, MSS116, CBS2, RPL12B, PUF6, PET122, RPL7A, RPS0A, RPS17A, RPL20A | 15 | 4.55E-02 |
| DNA recombination | 8 | ihfA, ihfB, intA, recA, recO, recR, ruvA, ruvC | 5 | HPR1, ERG28, THP2, RAD52, SGS1 | 13 | 5.70E-02 |
GO biological processes identified in our analysis included response to DNA damage stimulus, DNA repair, NER, DNA replication, DNA metabolic process, DNA recombination, translation, regulation of translation, tRNA aminoacylation for protein translation and aerobic respiration. In all, these categories spanned three general areas: DNA repair, protein synthesis and energy production. Because DNA repair is a conserved response to UV-damage [1], it was expected to be identified and was highlighted by a number of nodes. The identified DNA repair node serves as the namesake for this group and its inclusion reflects the fact that UV generates DNA lesions, that when left unrepaired can cause cell death. In addition, cells are known to initiate signal transduction cascades and activate cellular repair pathways after DNA damage [16]; thus, response to DNA damage stimulus was also a category that we expected to identify in our computational analysis. The repair of UV-induced DNA lesions can also occur by DNA recombination and as long tracks of DNA are replaced during recombination, our identification of DNA replication as an over-represented functional category was also expected. In all, the identification of these five DNA centric biological processes is consistent with published data and their identification further supports the validity of our computational analysis.
The surprising aspect of our Functionome study was the identification of protein synthesis and energy production categories, as these corresponding biological processes are not routinely associated with modulating UV toxicity. In fact, all of the identified biological processes associated with protein synthesis and energy metabolism were found to be more statistically significant than DNA recombination (P < 0.057), a known and well studied response to DNA damage in both E. coli and S. cerevisiae. Biological processes involving protein synthesis were identified and included translation (P < 0.0278), tRNA aminoacylation for protein translation (P < 0.0124) and regulation of translation (P < 0.0455). The identification of these three biological processes suggests that cells mount an evolutionarily conserved protein synthesis or ribosomal protein associated response to UV-damage. It is tempting to speculate that this protein synthesis based response is specific, as reports have indicated that regulation of translation initiation [36], tRNA modification status [37], tRNA cleavage [38] and tRNA mischarging [39] are cellular strategies used to respond to damage. It is also interesting to note that transcription was not identified in our GO analysis, which is added support for a specific translational program. We note, though, that general protein synthesis will be important for producing enzymes that respond to DNA damage and for maintaining cellular homeostasis. Of note is the predominance of many ribosomal proteins (18) in the protein synthesis categories, suggesting the integrity of the ribosome or ribosomal protein-based responses are essential for repairing the damage.
Additionally, our Functionome analysis demonstrated that the node of aerobic respiration (P < 0.0164) was over-represented with UV-toxicity modulating proteins. Associated E. coli proteins included NuoF and NuoH, part of the complex that shuttles electrons from NADH to quinones in the respiratory chain and acts to couple a redox reaction to proton translocation [32]. The S. cerevisiae proteins identified in the aerobic respiration node (Ydr115w, Qcr7, Qcr6, Dia4, Oar1, Mct1 and Pet20) include two components of ubiquinol cytochrome-c reductase complex involved in the electron transport chain (Qcr6 and Qcr7) and a protein required for respiratory growth (Pet20) [40]. Four of the E. coli and S. cerevisiae UV-toxicity modulating proteins linked to the aerobic respiration node ultimately promote ATP synthesis, suggesting that an intact metabolic response driving energy production is vital to cellular viability after UV-damage. This conclusion was further tested in S. cerevisiae. Ultimately our results suggest that cells with defective mitochondria (rho-), and thus deficient in oxidative phosphorylation, would be sensitive to UVC. Using ethidium bromide induced rho- strains we have shown that these cells are in fact sensitive to UVC (Supplemental Figure S2), thus supporting our Functionome mapping conclusion.
Functionome results predict UV-sensitive themes for S. pombe mutants
Our Functionome analysis has highlighted three biological themes that modulate the toxicity of UV in both E. coli and S. cerevisiae. These results also lead us to predict that DNA repair, protein synthesis and energy production are important UV-toxicity modulating processes for cells from other species. To test this prediction, we performed a high throughput screen of the S. pombe gene deletion library to identify gene products that modulate the toxicity of UV. The S. pombe deletion set from Bioneer (Daejeon, Republic of Korea) contains mutant strains specific to 3,006 genes [28]. Mutants were individually spotted onto YES-agar plates using the 96-syringe Matrix Scientific Hydra. Upon drying, the cells were left untreated or exposed to three different concentrations of UV (10, 15 and 20 joules) and then allowed to grow for 60 hours (Figure 3). UV exposure conditions were chosen based on the behavior of rad17Δ cells, as this DNA repair mutant was consistently identified as UV-sensitive in preliminary experiments. In total, 122 plates containing ~11,468 spotted cultures were assayed as described above and the resulting images of these plates were compiled into Supplemental Figure S3. We classified 310 S. pombe gene products as modulating the toxicity of UV (Table III). In a similar fashion to our results in E. coli and S. cerevisiae, we identified a wide range of S. pombe cellular proteins that modulate the toxicity of UV. In all, we determined that 18 biological processes from S. pombe were over-represented with UV-toxicity modulating proteins (Table IV) and, as predicted by our Functionome study, the categories of DNA repair, protein synthesis and, to some extent, energy production were well represented. Specifically related to our Functionome prediction, we determined that S. pombe proteins associated with the GO biological processes of NER, DNA recombination and response to DNA damage stimulus were over-represented (P < 0.09) in our list of UV-toxicity modulating proteins. It is worth noting that while NER proteins consistently modulate the toxicity of UV damage in lower organisms, our observed trend may not be observed in higher eukaryotes, as the redundancy in DNA repair pathway and differences in cell metabolism may allow for compensatory responses. We also determined that the GO biological process of translation (P < 0.09) was over-represented in our list of S. pombe UV-toxicity modulating proteins, supporting our prediction that components of the protein synthesis machinery play an important yet unknown role after UV-exposure. Significantly for both DNA repair and protein synthesis, we have observed exact identity in four GO terms (NER, DNA recombination, response to DNA damage stimulus and translation), as they were all over-represented with UV-toxicity modulating proteins from E. coli, S. cerevisiae and S. pombe. The identity in DNA repair-associated terms between S. pombe, S. cerevisiae and E. coli UV-toxicity modulating proteins was expected and provided validation of our methodology for comparing high throughput screening results. We note that in S. pombe, we only classified two GO annotated NER proteins as modulators of UV-toxicity (Rad8 and Uve1) and had a third fall just below our sensitivity cut off (Rad13). Analysis of the S. pombe library indicated that only 15 of the 28 mutants corresponding to NER proteins were represented and of these 15, we classified 5 as slow growers; thus, our search space was limited to 10 mutants. Our analysis suggests that in S. pombe NER is a vital cellular process under basal conditions.
Figure 3. Genomic phenotyping of S. pombe mutants with UV.
(A) 93 gene deletion mutants were spotted onto agar plates, left untreated or exposed to UV, incubated at 30° C for 60 hours and imaged. Green, red and yellow circles identify spbc21c3.02cΔ, ubc13Δ and msa1Δ, respectively. (B) Images were taken from many different plates and recompiled to demonstrate that varying degrees of UV sensitivity were observed in the S. pombe screen.
Table III.
Proteins corresponding to the 310 UV-sensitive mutants identified in S. pombe
| Protein | Deescription | UV Sensitivity |
|---|---|---|
| Rad17 | RFC related checkpoint protein Rad17 | High |
| Spcc576.12c | conserved eukaryotic protein | High |
| Rad9 | checkpoint clamp complex protein Rad9 | High |
| Tpp1 | trehalose-6-phosphate phosphatase Tpp1 | High |
| Apm1 | AP-1 adaptor complex subunit Apm1 | High |
| Zds1 | zds family protein Zds1 | High |
| Rhp18 | Rad18 homolog Rhp18 | High |
| Ubc13 | ubiquitin conjugating enzyme Ubc13 | High |
| Spbc16g5.13 | sequence orphan | High |
| Sol1 | SWI/SNF complex subunit Sol1 | High |
| Spbc947.04 | DIPSY family | High |
| Rad22 | DNA repair protein Rad22 | High |
| Spbc9b6.07 | nucleolar protein Nop52 family | High |
| Nmt1 | no message in thiamine Nmt1 | High |
| Spbc21c3.02c | Sds3-like family | High |
| Rip1 | ubiquinol-cytochrome-c reductase complex subunit 5 | High |
| Spp27 | RNA polymerase I upstream activation factor complex subunit Spp27 | High |
| Crb2 | DNA repair protein RAD9 homolog, Rhp9 | High |
| Spac27d7.06 | electron transfer flavoprotein alpha subunit | High |
| Rad50 | DNA repair protein Rad50 | High |
| Spbc1539.02 | sequence orphan | High |
| Spcc794.10 | UTP-glucose-1-phosphate uridylyltransferase | High |
| Kin1 | microtubule affinity-regulating kinase Kin1 | High |
| Spac16c9.01c | carbohydrate kinase | High |
| Rpl2701 | 60S ribosomal protein L27 | High |
| Spcp1e11.10 | ankyrin repeat protein, unknown biological role | High |
| Spbc16a3.10 | membrane bound O-acyltransferase, MBOAT | High |
| Spbc29a10.16c | cytochrome b5 | High |
| Spbc11b10.07c | CDC50 domain protein | High |
| Spac4f10.16c | P-type ATPase | High |
| Spt6 | transcription elongation factor Spt6 | High |
| Elf1 | AAA family ATPase ELf1 | High |
| Sty1 | MAP kinase Sty1 | High |
| Spac4a8.02c | conserved protein (broad species distribution) | High |
| Spbc6b1.03c | Pal1 family protein | High |
| Cds1 | replication checkpoint kinase Cds1 | High |
| Swi3 | replication fork protection complex subunit Swi3 | High |
| Tom7 | mitochondrial TOM complex subunit Tom7 | High |
| Mug42 | sequence orphan | High |
| Gar2 | GAR family | High |
| Ras1 | GTPase Ras1 | High |
| Rpl2002 | 60S ribosomal protein L20 | High |
| Rps1801 | 40S ribosomal protein S18 | High |
| Spbc2a9.05c | DUF846 family protein | High |
| Fsv1 | SNARE Fsv1 | High |
| Mug183 | histone chaperone Rtt106-like | High |
| Spac1f5.03c | FAD-dependent oxidoreductase | High |
| Rhp55 | RecA family ATPase Rhp55 | Medium |
| Str1 | siderophore-iron transporter Str1 | Medium |
| Ilv1 | acetolactate synthase catalytic subunit | Medium |
| Vps5 | retromer complex subunit Vps5 | Medium |
| Alp14 | Mad2-dependent spindle checkpoint component | Medium |
| Mto1 | MT organizer Mto1 | Medium |
| Mug80 | cyclin Clg1 | Medium |
| Spac688.03c | human AMMECR1 homolog | Medium |
| Mfm2 | M-factor precursor Mfm2 | Medium |
| Spbc17d11.08 | WD repeat protein, human WDR68 family | Medium |
| Spac1486.01 | manganese superoxide dismutase (AF069292) | Medium |
| Spbc16d10.08c | heat shock protein Hsp104 | Medium |
| Spac869.04 | formamidase-like protein | Medium |
| Spbp35g2.14 | RNA-binding protein | Medium |
| Mug136 | acetylglucosaminyltransferase | Medium |
| Pku80 | Ku domain protein Pku80 | Medium |
| Ctf1 | mRNA cleavage and polyadenylation specificity factor complex subunit Ctf1 | Medium |
| Cuf1 | Cu metalloregulatory transcription factor Cuf1 | Medium |
| Spac9e9.15 | CIA30 family protein | Medium |
| Spbc1604.12 | sequence orphan | Medium |
| Spcc1020.05 | phosphoprotein phosphatase | Medium |
| Btf3 | nascent polypeptide-associated complex subunit | Medium |
| Spac22e12.18 | conserved fungal protein | Medium |
| Rpl803 | 60S ribosomal protein L8 | Medium |
| Rtt109 | RTT109 family histone lysine acetyltransferase Rtt109 | Medium |
| Rad8 | ubiquitin-protein ligase E3 | Medium |
| Spac631.02 | bromodomain protein | Medium |
| Spcc757.11c | membrane transporter | Medium |
| Dak1 | dihydroxyacetone kinase Dak1 | Medium |
| Spac589.10c | ribomal-ubiquitin fusion protein Ubi5 | Medium |
| Spbp23a10.02 | conserved fungal protein | Medium |
| Spac1851.02 | 1-acylglycerol-3-phosphate O-acyltransferase | Medium |
| Spac1705.02 | human 4F5S homolog | Medium |
| Spcc126.12 | NGG1p interacting factor 3 family | Medium |
| Spbc3h7.07c | phosphoserine phosphatase | Medium |
| Mug4 | sequence orphan | Medium |
| Cwf11 | complexed with Cdc5 protein Cwf11 | Medium |
| Spcc1739.08c | short chain dehydrogenase | Medium |
| Spcc736.13 | short chain dehydrogenase | Medium |
| Spbc2d10.11c | nucleosome assembly protein Nap2 | Medium |
| Rnc1 | RNA-binding protein that suppresses calcineurin deletion Rnc1 | Medium |
| Gms1 | UDP-galactose transporter Gms1 | Medium |
| Spcc663.06c | short chain dehydrogenase | Medium |
| Spcc777.12c | sequence orphan | Medium |
| Sir2 | Sir2 family histone deacetylase Sir2 | Medium |
| Wtf16 | wtf element Wtf16 | Medium |
| Spbc56f2.05c | transcription factor | Medium |
| Mug96 | sequence orphan | Medium |
| Pnu1 | endodeoxyribonuclease Pnu1 | Medium |
| Spac3a12.13c | translation initiation factor eIF3 complex subunit | Medium |
| Spcc11e10.06c | RNA polymerase II elongator complex subunit Elp4 | Medium |
| Spbc651.09c | RNA polymerase II associated Paf1 complex | Medium |
| Mpd2 | GYF domain | Medium |
| Pex1 | AAA family ATPase Pex1 | Medium |
| Spbc418.02 | NatA N-acetyltransferase complex subunit | Medium |
| Spac12g12.10 | WD repeat protein, human WDR21 family | Medium |
| Chr2 | chitin synthase regulatory factor Chr2 | Medium |
| Spcc794.03 | amino acid permease, unknown 13 | Medium |
| Spcc1259.08 | conserved fungal protein | Medium |
| Spac22e12.03c | THIJ/PFPI family peptidase | Medium |
| Rps902 | 40S ribosomal protein S9 | Medium |
| Ain1 | alpha-actinin | Medium |
| Spac20g4.01 | CCR4-Not complex subunit Caf16 | Medium |
| Did2 | vacuolar sorting protein Did2 | Medium |
| Spbc21d10.09c | ubiquitin-protein ligase E3 | Medium |
| Hem2 | porphobilinogen synthase Hem2 | Medium |
| Rds1 | conserved fungal protein | Medium |
| Ish1 | LEA domain protein | Medium |
| Sin1 | stress activated MAP kinase interacting protein Sin1 | Medium |
| Spbc365.07c | TATA element modulatory factor homolog | Medium |
| Air1 | TRAMP complex subunit | Medium |
| Spac15a10.07 | sequence orphan | Medium |
| Sgo2 | shugoshin Sgo2 | Medium |
| Oxa102 | mitochondrial inner membrane translocase Oxa102 | low |
| Rpl702 | 60S ribosomal protein L7 | low |
| Rpl501 | 60S ribosomal protein L5 | low |
| Spac25g10.01 | RNA-binding protein | low |
| Spac1093.01 | PPR repeat protein | low |
| Mug182 | YjeF family protein | low |
| Mug24 | RNA-binding protein | low |
| Exo1 | exonuclease I Exo1 | low |
| Spcc24b10.12 | CGI121 family protein | low |
| Ctu1 | ATP binding protein | low |
| Mph1 | dual specificity protein kinase Mph1 | low |
| Spac31g5.07 | conserved fungal protein | low |
| Rmt3 | type I ribosomal protein arginine N-methytransferase Rmt3 | low |
| Fta5 | Sim4 and Mal2 associated (4 and 2 associated) protein 5 | low |
| Meu32 | sequence orphan | low |
| Cwf21 | complexed with Cdc5 protein Cwf21 | low |
| Rps1602 | 40S ribosomal protein S16 | low |
| Spbc839.03c | neddylation protein Dcn1 | low |
| Spac27e2.11c | sequence orphan | low |
| Vps1302 | chorein homolog | low |
| Mcs4 | two-component response regulator | low |
| Gsk31 | serine/threonine protein kinase Gsk31 | low |
| Spbc543.10 | GET complex subunit | low |
| Spbc1709.14 | peptide N-glycanase | low |
| Spbc1709.09 | mitochondrial translation termination factor | low |
| Spac11d3.14c | oxoprolinase | low |
| Spcc584.13 | amino acid permease, unknown 14 | low |
| Spbc2f12.03c | EST1 family protein | low |
| Meu29 | sequence orphan | low |
| Cys12 | cysteine synthase Cys12 | low |
| Yak3 | aldose reductase YakC | low |
| Ctr5 | copper transporter complex subunit Ctr5 | low |
| Ssb3 | DNA replication factor A subunit Ssb3 | low |
| Spbc1271.07c | N-acetyltransferase | low |
| Spac17a5.08 | COPII-coated vesicle component Erp2/3/4 | low |
| Cdd1 | cytidine deaminase Pcd1 | low |
| Arp42 | SWI/SNF and RSC complex subunit Arp42 | low |
| Brl2 | ubiquitin-protein ligase E3 | low |
| Gaf1 | transcription factor Gaf1 | low |
| Arg4 | carbamoyl-phosphate synthase Arg4 | low |
| Spbc359.01 | amino acid permease, unknown 7 | low |
| Nse5 | Smc5-6 complex non-SMC subunit Nse5 | low |
| Yam8 | calcium transport protein | low |
| Mug165 | sequence orphan | low |
| Spbc21c3.06 | sequence orphan | low |
| Spac1f5.05c | sequence orphan | low |
| Spac29b12.08 | sequence orphan | low |
| Rps1102 | 40S ribosomal protein S11 | low |
| Spac1687.08 | sequence orphan | low |
| Omt2 | 4-alpha-hydroxytetrahydrobiopterin dehydratase | low |
| Spcc320.14 | threo-3-hydroxyaspartate ammonia-lyase | low |
| Spcc191.10 | sequence orphan | low |
| Spbc18h10.16 | amino acid permease, unknown 9 | low |
| Spac212.02 | sequence orphan | low |
| Spcc1322.10 | conserved fungal protein | low |
| Spac26a3.14c | DUF1748 family protein | low |
| Spac27e2.01 | alpha-amylase homolog | slightly |
| Ght7 | hexose transporter Ght7 | slightly |
| Rga9 | RhoGAp, GTPase activator towards Rho/Rac/Cdc42-like small GTPases | slightly |
| Thi5 | transcription factor Thi5 | slightly |
| Spbc3b8.06 | conserved fungal protein | slightly |
| Coq2 | para-hydroxybenzoate--polyprenyltransferase Coq2 | slightly |
| Pof3 | F-box protein Pof3 | slightly |
| Oca2 | serine/threonine protein kinase Oca2 | slightly |
| Lyn1 | sequence orphan | slightly |
| Spac14c4.12c | SWIRM domain protein | slightly |
| Kap1 | chromatin remodeling complex subunit Ngg1 | slightly |
| Ctu2 | conserved eukaryotic protein | slightly |
| Spac22a12.17c | short chain dehydrogenase | slightly |
| Csn71 | COP9/signalosome complex subunit 7a | slightly |
| Spac644.13c | Rab GTPase binding | slightly |
| Spbc543.08 | phosphoinositide biosynthesis protein | slightly |
| Spbc31f10.02 | thioesterase superfamily protein | slightly |
| Spcc1442.02 | DUF1760 family protein | slightly |
| Spbc25b2.01 | elongation factor 1 alpha related protein | slightly |
| Spcc594.07c | sequence orphan | slightly |
| Bqt1 | bouquet formation protein Bqt1 | slightly |
| Spcc70.02c | mitochondrial ATPase inhibitor | slightly |
| Atp15 | F0-ATPase epsilon subunit | slightly |
| Spac9g1.05 | actin cortical patch component Aip1 | slightly |
| Pet127 | mitochondrial membrane protein Pet127 | slightly |
| Spac823.10c | mitochondrial carrier with solute carrier repeats | slightly |
| Nap1 | nucleosome assembly protein Nap1 | slightly |
| Nup124 | nucleoporin Nup124 | slightly |
| Spac1687.07 | conserved fungal protein | slightly |
| Pep3 | ubiquitin-protein ligase E3 | slightly |
| Smd3 | Sm snRNP core protein Smd3 | slightly |
| Spbc776.16 | sequence orphan | slightly |
| Spac1952.17c | GTPase activating protein | slightly |
| Spac1f3.03 | Lgl family protein | slightly |
| Tcg1 | single-stranded telomeric binding protein Tgc1 | slightly |
| Spac1782.01 | proteasome component | slightly |
| Spac959.06c | sequence orphan | slightly |
| Spac1687.19c | queuine tRNA-ribosyltransferase | slightly |
| Spbc25d12.06 | RNA helicase | slightly |
| Rex3 | exonuclease Rex3 | slightly |
| Spbc1685.05 | serine protease | slightly |
| Hus1 | checkpoint clamp complex protein Hus1 | slightly |
| Spcc16a11.16c | ARM1 family | slightly |
| Ats1 | N-acetyltransferase Ats1 | slightly |
| Spac17c9.15c | sequence orphan | slightly |
| Spcc306.02c | Rab GTPase binding | slightly |
| Tsf1 | mitochondrial translation elongation factor EF-Ts Tsf1 | slightly |
| Spac31g5.15 | phosphatidylserine decarboxylase | slightly |
| Spcc1919.07 | sequence orphan | slightly |
| Spac13g7.09c | sequence orphan | slightly |
| Spac30d11.06c | DUF300 family protein | slightly |
| But1 | neddylation pathway protein But1 | slightly |
| Rti1 | Rad22 homolog Rti1 | slightly |
| Rrp16 | rRNA processing protein Rrp16 | slightly |
| Med20 | TATA-box related factor (TRF) | slightly |
| Spbc21b10.03c | ataxin-2 homolog | slightly |
| Spbp22h7.05c | ATPase with bromodomain protein | slightly |
| Gos1 | SNARE Gos1 | slightly |
| Spcc1795.10c | Sed5 Vesicle Protein Svp26 | slightly |
| Spac9.02c | N-acetyltransferase | slightly |
| Rpl1801 | 60S ribosomal protein L18 | slightly |
| Rep2 | transcriptional activator Rep2 | slightly |
| Spbc21c3.17c | conserved fungal protein | slightly |
| Spac8f11.02c | diphthamide biosynthesis protein Dph3 | slightly |
| Sco1 | copper chaperone Sco1 | slightly |
| Spac3h8.07c | prefoldin subunit 3 | slightly |
| Spbc359.03c | amino acid permease, unknown 8 | slightly |
| Spac4g8.03c | RNA-binding protein | slightly |
| Dak2 | dihydroxyacetone kinase Dak2 | slightly |
| Ght1 | hexose transporter Ght1 | slightly |
| 39722 | mitochondrial intermediate peptidase Oct1 | slightly |
| Spcc61.03 | conserved protein (broad species distribution) | slightly |
| Spac13f5.03c | glycerol dehydrogenase | slightly |
| Spac227.06 | Rab GTPase binding | slightly |
| Rps1501 | 40S ribosomal protein S15 | slightly |
| Spac3g9.11c | pyruvate decarboxylase | slightly |
| Met16 | phosphoadenosine phosphosulfate reductase | slightly |
| Spbc13e7.08c | RNA polymerase II associated Paf1 complex | slightly |
| Spbc405.05 | sequence orphan | slightly |
| Mug106 | sequence orphan | slightly |
| Rsc1 | RSC complex subunit Rsc1 | slightly |
| Hsp9 | heat shock protein Hsp9 | slightly |
| Spbc409.08 | spermine family transporter | slightly |
| Dga1 | diacylglycerol O-acyltransferase | slightly |
| Spbc106.13 | conserved eukaryotic protein | slightly |
| Spac1952.03 | cysteine protease | slightly |
| Spac29b12.11c | human WW domain binding protein-2 ortholog | slightly |
| Par1 | protein phosphatase regulatory subunit Par1 | slightly |
| Ppk4 | serine/threonine protein kinase Ppk4 | slightly |
| Srb11 | cyclin Srb11 | slightly |
| Ago1 | argonaute | slightly |
| Vps3 | GTPase regulator Vps3 | slightly |
| Spac3a11.04 | siepin homolog | slightly |
| Cid12 | poly(A) polymerase Cid12 | slightly |
| Pep7 | prevacuole/endosomal FYVE tethering component Pep7 | slightly |
| Eaf1 | RNA polymerase II transcription elongation factor SpEAF | slightly |
| Spcc4b3.08 | C-terminal domain kinase I (CTDK-I) gamma subunit | slightly |
| Gyp1 | GTPase activating protein Gyp1 | slightly |
| Spbc582.08 | alanine aminotransferase | slightly |
| Sce3 | translation initiation factor eIF4B | slightly |
| Cis4 | membrane transporter | slightly |
| Spbc21c3.08c | ornithine aminotransferase | slightly |
| Uve1 | endonuclease Uve1 | slightly |
| Rrg1 | methyltransferase | slightly |
| Tel1 | ATM checkpoint kinase | slightly |
| Spcc162.01c | RNA-binding protein | slightly |
| Spbc29a10.07 | nucleoporin Pom152 | slightly |
| Spcc663.14c | membrane transporter | slightly |
| Spac10f6.11c | kinase activator | slightly |
| Spbc21h7.04 | ATP-dependent RNA helicase Dbp7 | slightly |
| Apl1 | AP-2 adaptor complex subunit Apl1 | slightly |
| But2 | neddylation pathway protein But2 | slightly |
| Wis2 | cyclophilin family peptidyl-prolyl cis-trans isomerase Wis2 | slightly |
| Spcc285.13c | nucleoporin Nup60 | slightly |
| Str3 | siderophore-iron transporter Str3 | slightly |
| Spap27g11.14c | sequence orphan | slightly |
| Spac23a1.14c | cystathionine gamma-synthase | slightly |
| Spac977.14c | aldo/keto reductase, unknown biological role | slightly |
| Spacunk4.15 | 2',3'-cyclic-nucleotide 3'-phosphodiesterase | slightly |
| Spac17c9.14 | Pex19 protein family | slightly |
| Spbc21d10.10 | bromodomain protein | slightly |
| Spac31g5.18c | ubiquitin family, human C1ORF55 related | slightly |
| Spbc16h5.12c | conserved fungal protein | slightly |
| Spcc1840.04 | caspase | slightly |
| Spcc1235.01 | sequence orphan | slightly |
| Spbc27b12.14 | mitochondrial membrane protein complex assembly protein | slightly |
| Spcc548.04 | ubiquitin family protein Urm1 | slightly |
| Spbc776.17 | rRNA processing protein Rrp7 | slightly |
| Wsc1 | transmembrane receptor Wsc1 | slightly |
| Spcc736.07c | cell polarity protein | slightly |
| Spbp8b7.13 | conserved fungal protein | slightly |
| Rif1 | telomere length regulator protein Rif1 | slightly |
| Rpl2301 | 60S ribosomal protein L23 | slightly |
| Mug63 | TLDc domain protein 1 | slightly |
Table IV.
GO biological processes over-represented in the 310 UV-toxicity modulating proteins from S. pombe
| Functional Process | Total Tested | # UV Sensitive | Genes | P-value |
|---|---|---|---|---|
| Protein amino acid acetylation | 5 | 3 | ATS1, SPBC1271.07C, SPBC418.02 | 1.59E-04 |
| Ribosome biogenesis | 50 | 11 | GAR2, RPL501, RPS1102, RPS1602, RPS1801, RPS902, RRP16, SPBC776.17, RMT3, SPAC589.10C, TCG1 | 6.87E-04 |
| RNA processing | 4 | 2 | PET127, REX3 | 2.56E-03 |
| Glycerophospholipid biosynthetic process | 3 | 2 | SPAC1851.02, SPBC16A3.10 | 2.56E-03 |
| mRNA polyadenylation | 4 | 2 | SPBC21B10.03C, CTF1 | 3.47E-03 |
| Protein homooligomerization | 5 | 2 | RAD22, RTI1 | 4.66E-03 |
| Response to arsenic | 6 | 2 | MCS4, STY1 | 4.66E-03 |
| Chromosome segregation | 23 | 5 | AGO1, ALP14, CID12, SIR2, SPCC576.12C | 8.20E-03 |
| Response to hydrogen peroxide | 7 | 2 | MCS4, STY1 | 1.39E-02 |
| Cellular iron ion homeostasis | 10 | 3 | STR1, STR3, CUF1 | 2.28E-02 |
| Copper ion transport | 4 | 2 | CTR5, SCO1 | 2.28E-02 |
| Respiratory chain complex iv assembly | 5 | 2 | OXA102, SCO1 | 2.87E-02 |
| DNA recombination* | 6 | 2 | PNU1, SSB3 | 4.46E-02 |
| Response to DNA damage stimulus* | 27 | 5 | NSE5, RAD8, RHP55, RTT109, SWI3 | 6.68E-02 |
| Nucleotide-excision repair* | 10 | 2 | RAD8, UVE1 | 8.08E-02 |
| Intracellular protein transport | 67 | 10 | APL1, APM1, SPAC4F10.16C, SPAC644.13C, VPS1302, VPS3, VPS5, FSV1, GOS1, SPAC17A5.08 | 8.08E-02 |
| Regulation of catalytic activity | 59 | 9 | PAR1, RGA9, TSF1, ZDS1, SPAC1F3.03, SPAC227.06, SPAC644.13C, SPCC306.02C, VPS3 | 9.68E-02 |
| Translation* | 116 | 14 | RPL1801, RPL2002, RPL2301, RPL2701, RPL702, RPL803, RPS1102, RPS1501, SPAC3A12.13C, RPL501, RPS1602, RPS1801, RPS902, SPBC25B2.01 | 9.68E-02 |
indicates a biological process that was also identified in our Functionome analysis
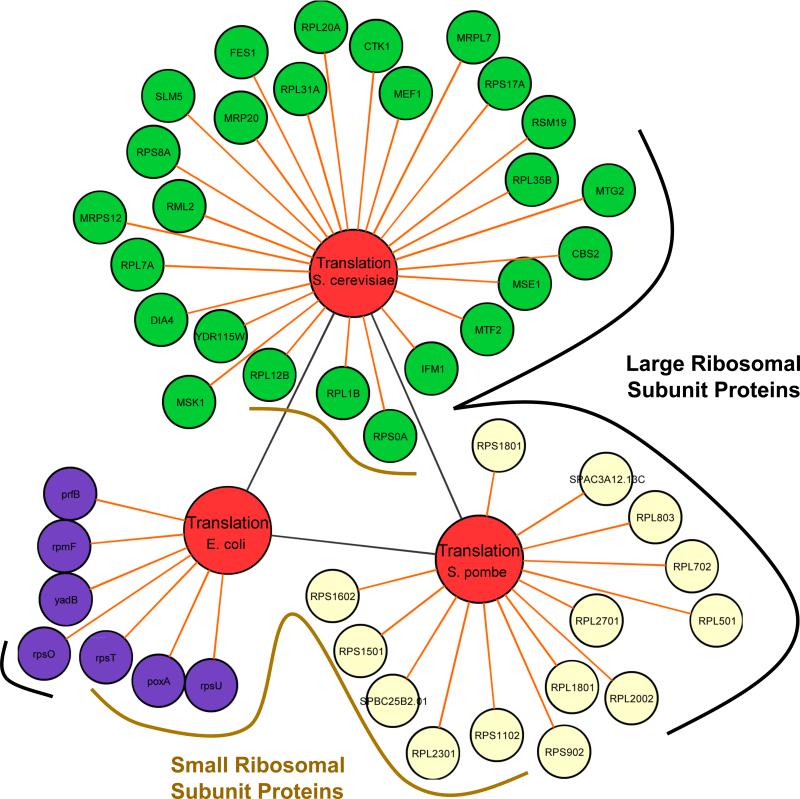
The trend that the GO term of translation was consistently identified in each organism's list of UV-toxicity modulating proteins (Figure 4) further supports our conclusion that protein synthesis or ribosomal protein-based responses to UV-damage are universally important to many cell types. Detailed annotation information on 7 E. coli, 26 S. cerevisiae and 14 S. pombe proteins identified in the biological process of translation indicated that many are associated with the small (11) or large (13) ribosomal subunits. The ribosome is a complicated machine and in both prokaryotic and eukaryotic organisms, it is composed of ribonucleoproteins and divided into large and small subunits. We wanted to determine if there was any amino acid similarity between the individual proteins found associated with the translation node, and using the S. pombe gene database [41] we looked for orthologous proteins between the two yeasts. We determined that in the translation node, two sets of UV-toxicity modulating proteins from S. pombe and S. cerevisiae were orthologs and thus connected to each other: Rpl702 - Rpl7A, and Rpl2002 – Rpl20A. While the exact role for these large ribosomal proteins in preventing UV-toxicity is unknown, we can speculate that a protein synthesis based response to damage is corrupted in these cells. This response could include the use of specific ribosomal proteins to promote the translation of stress response genes, which is akin to a ribosomal code [42, 43]. Other realistic possibilities exist, though, as ribosomal proteins have been demonstrated to perform auxiliary activities in stress signaling [44] and they may directly contribute to DNA repair or cell cycle in some fashion. It is worth noting that ribosomal protein S3 has been shown to nick AP containing DNA, has affinity for abasic sites and 7,8-dihydro-8-oxoguanine DNA, and has been shown to localize to the nucleus in response to genotoxic stress [45, 46]. Thus, there are already published biochemical and damage-response roles for ribosomal proteins in DNA repair. In addition, there is sufficient evidence that ribosomal proteins have extra-ribosomal functions [44]. Another possibility is that perturbations in ribosome assembly promote stress on their own and in conjunction with UV-damage, this may overwhelm the cellular stress response. The role of ribosomal proteins during stress is largely unexplored and the identified sets of orthologs (Rpl702 - Rpl7A, and Rpl2002 – Rpl20A) highlight starting points for focused studies to better understand the role of protein synthesis machinery after UV-exposure.
Figure 4. Tri-species node of translation identified as a conserved biological process that modulated the toxicity of UV.
UV-toxicity modulating proteins from E. coli (purple spheres, lower case protein names), S. cerevisiae (green spheres, upper case protein names) and S. pombe (yellow spheres, upper case protein names) were connected to their GO biological process of translation (orange line) and each protein's basic function was analyzed in a species-specific database (Ecogene, S. pombe GeneDB and SGD). Those proteins belonging to the large and small ribosomal subunit are underlined in black and brown, respectively.
Many of the S. cerevisiae proteins (16 in total) belonging to the translation node play a role in mitochondrial protein synthesis. Defects in mitochondrial translation are associated with corrupted aerobic metabolism [40], which provides a link between the process of translation and energy production. This link is also demonstrated in our Functionome analysis as the dual annotation of specific proteins in both translation and energy metabolism based processes was observed (Figure 2C). The biological process of aerobic respiration was specifically identified in our Functionome analysis of UV-toxicity modulating proteins from E. coli and S. cerevisiae. In S. pombe, we analyzed 8 of 25 mutants corresponding to proteins annotated with the aerobic respiration designation and none of these were sensitive to UV. In general though, the GO annotations for all S. pombe proteins used in this study are limited with ~1.3 annotations per protein as compared to ~2.7 (12130/4415) and ~2.0 (6120/3120) annotations per protein for S. cerevisiae and E. coli, respectively. In addition, the S. pombe deletion library only contained 32% of the mutants corresponding to the aerobic respiration category, suggesting that many are essential genes. Thus the small sample size (eight mutants) limited our search space. We did observe a hint of aerobic respiration in our list of S. pombe UV-toxicity modulating proteins, as the category of respiratory chain complex iv assembly was over-represented (P < 0.03). Corresponding proteins included Oxa102, required for the insertion of integral membrane proteins into the mitochondrial inner membrane and essential for the activity and assembly of cytochrome c oxidase, and Sco1, a copper chaperone used to transport copper to the Cu(A) site on the cytochrome c oxidase subunit II [41]. While respiratory chain complex iv assembly is not an identical category to aerobic respiration, there is a connection via ATP formation. The precise reason for the identification of aerobic respiration in our analysis is unknown, but we speculate that ATP levels optimize stress responses. A deficiency in an energy-associated metabolite, NAD(+), has recently been implicated in the damage-induced death of DNA repair deficient cells [47], supporting a role for ATP synthesis in modulating DNA damage-induced toxicity. We also note that many of the proteins that participate in DNA repair, recombination, and the DNA damage response are ATP-dependent enzymes and their activity could be affected in cells compromised for aerobic respiration. These enzymes include helicases and recombinases that manipulate DNA strands, as well as chromatin remodeling enzymes vital to transcriptional responses and DNA replication; as such deficiencies in ATP levels should thus impede their activity. While the exact reasons for our identification of proteins associated with aerobic respiration as modulating the toxicity of UV are speculative, our results highlight a potential area for future studies.
Conclusions
Cellular responses to DNA damage play an important role in dictating cellular outcomes, and model organisms continue to be an important tool for understanding stress response pathways. High throughput screens using the above described model systems are fast and cost-efficient and when linked to computational analysis allowed for the identification of highly significant themes associated with UV-toxicity modulation. Our identification of over 600 new UV-toxicity modulating proteins in E. coli and S. pombe and their comparison to previously reported proteins from S. cerevisiae has further cemented a universal role for DNA repair after UV-exposure. In addition, our study has highlighted roles for protein synthesis machinery and aerobic respiration components after UV-exposure. Ultimately, we have demonstrated the feasibility of using comparative functional genomics approaches to identify highly conserved biological responses to UV-damage. Our methodology can be easily extended to other stress responses and has the potential to help identify novel damage-response themes and proteins in higher organisms.
Materials and methods
High throughput screening of E. coli gene-deletion mutants against UV
Luria Bertani broth (LB) (BP1426-3, Fisher Scientific, Waltham, MA) was used to culture E. coli. The library of E. coli gene deletion mutants was acquired from the Genome Analysis Project in Japan [29]. High throughput screening of the E. coli gene deletion library was performed as previously described [26]. This procedure is very similar to what we used to screen the S. cerevisiae gene deletion library [24]. Briefly, 96-well plates containing the gene deletion mutants were replicated into liquid medium (LB-kanamycin), grown for 16 hours at 37°C, diluted 10-fold into LB and 1 μl cell suspensions were then robotically (Matrix Hydra) spotted on LB-kanamycin agar plates. Plates were allowed to dry for 30 minutes and UV doses were applied using a Stratalinker (Stragene, Cedar Creek, Texas). Inoculated plates were incubated for 16-hours at 37 °C and then imaged using an AlphaImager (Alpha Innotech Corporation, San Leandro, CA). Reduced growth for a specific gene-deletion mutant was identified relative to other mutants found on the 96-well plate and wild-type BW25113 cells, and was also relative to growth of the mutant on an untreated plate. To identify UV-toxicity modulating proteins, we linked sensitive mutants to their corresponding deleted gene and assumed the protein encoded by the deleted gene was responsible for the observed phenotype.
Functional Interactome Construction, Mapping and Analysis
We constructed a Functionome by using Gene Ontology information for E. coli and S. cerevisiae proteins. Species-specific gene ontology information associated with biological process was downloaded from the Gene Ontology Project website (Feb 1, 2010, www.geneontology.org) [33]. We choose to use an intermediate level of biological process to limit redundancy in our search space and only used 511 biological processes common to both E. coli and S. cerevisiae (Supplemental Table 1). The data files were compiled so that each protein was linked to one or more biological processes and the compiled information was visualized as a functional interactome using the program Cytoscape [48]. Sensitivity data representing 171 E. coli and 236 S. cerevisiae UV-toxicity modulating proteins was mapped onto our functional interactome. Next, we identified GO-nodes that were significantly over-represented with UV-toxicity modulating proteins and contained at least two from E. coli and two from S. cerevisiae. GO-node significance was determined by mapping randomly sampled proteins (171 from E. coli and 236 from S. cerevisiae) onto the Functionome and tracking occurrences in specific nodes over 200 iterations. Significant GO-nodes were first determined based on Z-score calculations and then using Normal curve approximations, similar as described [24, 26]. The protein randomization method utilized a static network and random groups of proteins to determine significance. We also verified the significance of our UV-target lists by mapping our 171 and 236 UV-toxicity modulating proteins from E. coli and S. cerevisiae on 200 randomized Functionomes. This last method employed a static target list and randomly compiled functional interactions, and is similar to a method we have previously reported [49]. Once we identified GO-nodes that met our criteria, significance (p < 0.05) and number of associated proteins (=>4), we utilized publically available protein-protein interaction information [50] to enhance our network visualizations. S. cerevisiae strains with mitochondrial mutations (rho-) were generated as previously described [51] and tested for UVC sensitivity on plating a 10-fold dilution series of cells on YPD plates and treating with increasing doses of UVC.
High throughput screening of S. pombe gene-deletion mutants against UV
Yeast extract with supplements (YES) (Fisher Scientific, Waltham, MA) was used to culture S. pombe. The library of S. pombe gene deletion mutants was acquired from Bioneer Corporation (Daejeon, Republic of Korea). High throughput screening of the S. pombe gene deletion library was performed in a similar fashion to the methodology we have described for E. coli and S. cerevisiae [24, 26]. Briefly, 96-well plates containing the gene deletion mutants were replicated into liquid medium (YES), grown for 24-hours at 30°C, and 1 μl cell suspensions were robotically (Matrix Hydra) spotted onto YES agar plates. Plates were allowed to dry for 30 minutes and UV doses were applied using a Stratalinker (Stragene, Cedar Creek, Texas) with 254 nm bulbs. Inoculated plates were incubated for 60 hours at 30 °C and then imaged using an AlphaImager (Alpha Innotech Corporation, San Leandro, CA). Reduced growth for a specific gene-deletion mutant was identified relative to other mutants found on the 96-well plate and wild-type. This was done for both untreated and UV treated plates. Mutants were initially scored for their ability to grow on untreated plates, with 357 mutants displaying a general slow growth phenotype (Supplemental Table 2). UV-sensitive mutants were scored (4 to 1, high to slightly sensitive) based on their ability to grow normally on untreated plates and on their decreased growth after UV treatment. A cumulative UV-toxicity sensitivity score for each mutant was determined, representing the behavior of each mutant after three increasing UV doses over two biological replicates. In theory, the most sensitive mutant could score a 24 ((4 + 4 + 4) * 2 replicates) and we set a minimum sensitivity score of 4. To identify UV-toxicity modulating proteins, we linked sensitive mutants to their corresponding deleted gene and assumed the protein encoded by the deleted gene was responsible for the observed phenotype. GO functional mapping using the 310 UV-toxicity modulating proteins and corresponding GO-annotations (Supplemental Table 3) was performed similar to as described above. We note that all GO assignments specific to each protein reported are included in Supplemental Table 4 and these tables can be imported into Cytoscape to visualize the Functionomes used in this study. In addition, all GO terms used in this study are reported in Supplemental Table 5.
Supplementary Material
Supplemental Figure Legends
Figure S1. Genomic phenotyping of E. colimutants with UV.
All plates and replicates from the genomic phenotyping analysis of the E. coli deletion library.
Figure S2.UV sensitivity of S. cerevisiaerho- mutants.
By4741 cells were made rho- by ethidium bromide treatment and then analyzed for UV sensitivity on YPD plates. Wild-type and rho- mutants were serially diluted, plated and then exposed to increasing doses of UVC.
Figure S3. Genomic phenotyping of S. pombemutants with UV.
All plates and replicates from the genomic phenotyping analysis of the S. pombe deletion library.
Acknowledgments
We regret the omission of many important references due to space constraints. This work was supported by NIH grants to TJB (ES01225101 and ES015037) and RPC (CA116318, RR015464, and GM46312) and a NYSTAR James Watson Award to TJB. Special thanks to members of the Cancer Research Center and Wadsworth NY State Public Health Laboratories for helpful comments.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Friedberg E, Walker G, Siede W, Wood RD, Schultz RA, Ellenberger T. DNA Repair and Mutagenesis. 2nd Edition ASM Press; Washington D.C.: 2005. [Google Scholar]
- 2.Sinha RP, Hader DP. UV-induced DNA damage and repair: a review. Photochem Photobiol Sci. 2002;1:225–236. doi: 10.1039/b201230h. [DOI] [PubMed] [Google Scholar]
- 3.Johnson RE, Prakash S, Prakash L. Efficient bypass of a thymine-thymine dimer by yeast DNA polymerase, Poleta. Science. 1999;283:1001–1004. doi: 10.1126/science.283.5404.1001. [DOI] [PubMed] [Google Scholar]
- 4.You YH, Szabo PE, Pfeifer GP. Cyclobutane pyrimidine dimers form preferentially at the major p53 mutational hotspot in UVB-induced mouse skin tumors. Carcinogenesis. 2000;21:2113–2117. doi: 10.1093/carcin/21.11.2113. [DOI] [PubMed] [Google Scholar]
- 5.Sancar A. Structure and function of DNA photolyase. Biochemistry. 1994;33:2–9. doi: 10.1021/bi00167a001. [DOI] [PubMed] [Google Scholar]
- 6.Todo T. Functional diversity of the DNA photolyase/blue light receptor family. Mutat Res. 1999;434:89–97. doi: 10.1016/s0921-8777(99)00013-0. [DOI] [PubMed] [Google Scholar]
- 7.Sancar A. DNA excision repair. Annu Rev Biochem. 1996;65:43–81. doi: 10.1146/annurev.bi.65.070196.000355. [DOI] [PubMed] [Google Scholar]
- 8.Van Houten B, Eisen JA, Hanawalt PC. A cut above: Discovery of an alternative excision repair pathway in bacteria. Proc Natl Acad Sci U S A. 2002;99:2581–258. doi: 10.1073/pnas.062062599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Moolenaar GF, van Rossum-Fikkert S, van Kesteren M, Goosen N. Cho, a second endonuclease involved in Escherichia coli nucleotide excision repair. Proc Natl Acad Sci U S A. 2002;99:1467–1472. doi: 10.1073/pnas.032584099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Meyer JN, Boyd WA, Azzam GA, Haugen AC, Freedman JH, Van Houten B. Decline of nucleotide excision repair capacity in aging Caenorhabditis elegans. Genome Biol. 2007;8:R70. doi: 10.1186/gb-2007-8-5-r70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hanawalt PC, Spivak G. Transcription-coupled DNA repair: two decades of progress and surprises. Nat Rev Mol Cell Biol. 2008;9:958–970. doi: 10.1038/nrm2549. [DOI] [PubMed] [Google Scholar]
- 12.Ganesan AK, Hanawalt PC. Transcription-coupled nucleotide excision repair of a gene transcribed by bacteriophage T7 RNA polymerase in Escherichia coli. DNA Repair. 2009 doi: 10.1016/j.dnarep.2010.06.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Chu G, Mayne L. Xeroderma pigmentosum, Cockayne syndrome and Trichothiodystrophy-Do the genes explain the diseases. Trends in Genetics. 1996;12:187–192. doi: 10.1016/0168-9525(96)10021-4. [DOI] [PubMed] [Google Scholar]
- 14.Friedberg EC. How nucleotide excision repair protects against cancer. Nat Rev Cancer. 2001;1:22–33. doi: 10.1038/35094000. [DOI] [PubMed] [Google Scholar]
- 15.Janion C. Inducible SOS response system of DNA repair and mutagenesis in Escherichia coli. Int J Biol Sci. 2008;4:338–344. doi: 10.7150/ijbs.4.338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Courcelle J, Khodursky A, Peter B, Brown PO, Hanawalt PC. Comparative gene expression profiles following UV exposure in wild-type and SOS-deficient Escherichia coli. Genetics. 2001;158:41–64. doi: 10.1093/genetics/158.1.41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Walker GC, Kenyon CJ, Bagg A, Langer PJ, Shanabruch WG. Mutagenesis and cellular responses to DNA damage. Natl Cancer Inst Monogr. 1982;60:257–267. [PubMed] [Google Scholar]
- 18.Cortez D, Guntuku S, Qin J, Elledge SJ. ATR and ATRIP: partners in checkpoint signaling. Science. 2001;294:1713–1716. doi: 10.1126/science.1065521. [DOI] [PubMed] [Google Scholar]
- 19.Kastan MB, Bartek J. Cell-cycle checkpoints and cancer. Nature. 2004;432:316–323. doi: 10.1038/nature03097. [DOI] [PubMed] [Google Scholar]
- 20.Hurley PJ, Wilsker D, Bunz F. Human cancer cells require ATR for cell cycle progression following exposure to ionizing radiation. Oncogene. 2007;26:2535–2542. doi: 10.1038/sj.onc.1210049. [DOI] [PubMed] [Google Scholar]
- 21.Gasch AP, Huang M, Metzner S, Botstein D, Elledge SJ, Brown PO. Genomic expression responses to DNA-damaging agents and the regulatory role of the yeast ATR homolog Mec1p. Mol Biol Cell. 2001;12:2987–3003. doi: 10.1091/mbc.12.10.2987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Matsuoka S, Ballif BA, Smogorzewska A, McDonald ER, 3rd, Hurov KE, Luo J, Bakalarski CE, Zhao Z, Solimini N, Lerenthal Y, et al. ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science. 2007;316:1160–1166. doi: 10.1126/science.1140321. [DOI] [PubMed] [Google Scholar]
- 23.Matsuoka S, Rotman G, Ogawa A, Shiloh Y, Tamai K, Elledge SJ. Ataxia telangiectasia-mutated phosphorylates Chk2 in vivo and in vitro. Proc Natl Acad Sci U S A. 2000;97:10389–10394. doi: 10.1073/pnas.190030497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Begley TJ, Rosenbach AS, Ideker T, Samson LD. Hot spots for modulating toxicity identified by genomic phenotyping and localization mapping. Mol Cell. 2004;16:117–125. doi: 10.1016/j.molcel.2004.09.005. [DOI] [PubMed] [Google Scholar]
- 25.Bennett CB, Lewis LK, Karthikeyan G, Lobachev KS, Jin YH, Sterling JF, Snipe JR, Resnick MA. Genes required for ionizing radiation resistance in yeast. Nat Genet. 2001;29:426–434. doi: 10.1038/ng778. [DOI] [PubMed] [Google Scholar]
- 26.Rooney JP, George AD, Patil A, Begley U, Bessette E, Zappala MR, Huang X, Conklin DS, Cunningham RP, Begley TJ. Systems based mapping demonstrates that recovery from alkylation damage requires DNA repair, RNA processing, and translation associated networks. Genomics. 2008;93:42–51. doi: 10.1016/j.ygeno.2008.09.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Deshpande GP, Hayles J, Hoe KL, Kim DU, Park HO, Hartsuiker E. Screening a genome-wide S. pombe deletion library identifies novel genes and pathways involved in genome stability maintenance. DNA Repair (Amst) 2009;8:672–679. doi: 10.1016/j.dnarep.2009.01.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kim DU, Hayles J, Kim D, Wood V, Park HO, Won M, Yoo HS, Duhig T, Nam M, Palmer G, et al. Analysis of a genome-wide set of gene deletions in the fission yeast Schizosaccharomyces pombe. Nat Biotechnol. 2009;28:617–623. doi: 10.1038/nbt.1628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, Datsenko KA, Tomita M, Wanner BL, Mori H. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol. 2006;2:2006–2008. doi: 10.1038/msb4100050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Winzeler EA, Shoemaker DD, Astromoff A, Liang H, Anderson K, Andre B, Bangham R, Benito R, Boeke JD, Bussey H, et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science. 1999;285:901–906. doi: 10.1126/science.285.5429.901. [DOI] [PubMed] [Google Scholar]
- 31.Hanawalt PC, Cooper PK, Ganesan AK, Smith CA. DNA repair in bacteria and mammalian cells. Annu Rev Biochem. 1979;48:783–836. doi: 10.1146/annurev.bi.48.070179.004031. [DOI] [PubMed] [Google Scholar]
- 32.Rudd KE. EcoGene: a genome sequence database for Escherichia coli K-12. Nucleic Acids Res. 2000;28:60–64. doi: 10.1093/nar/28.1.60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25:25–29. doi: 10.1038/75556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Robinson MD, Grigull J, Mohammad N, Hughes TR. FunSpec: a web-based cluster interpreter for yeast. BMC Bioinformatics. 2002;3:35. doi: 10.1186/1471-2105-3-35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Jelinsky SA, Samson LD. Global response of Saccharomyces cerevisiae to an alkylating agent. Proc Natl Acad Sci U S A. 1999;96:1486–1491. doi: 10.1073/pnas.96.4.1486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Harding HP, Novoa I, Zhang Y, Zeng H, Wek R, Schapira M, Ron D. Regulated Translation Initiation Controls Stress-Induced Gene Expression in Mammalian Cells. Molecular Cell. 2000;6:1099. doi: 10.1016/s1097-2765(00)00108-8. [DOI] [PubMed] [Google Scholar]
- 37.Begley U, Dyavaiah M, Patil A, Rooney JP, Direnzo D, Young CM, Conklin DS, Zitomer RS, Begley TJ. Trm9-Catalyzed tRNA Modifications Link Translation to the DNA Damage Response. Mol Cell. 2007;28:860–870. doi: 10.1016/j.molcel.2007.09.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Thompson DM, Lu C, Green PJ, Parker R. tRNA cleavage is a conserved response to oxidative stress in eukaryotes. Rna. 2008;14:2095–2103. doi: 10.1261/rna.1232808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Netzer N, Goodenbour JM, David A, Dittmar KA, Jones RB, Schneider JR, Boone D, Eves EM, Rosner MR, Gibbs JS, et al. Innate immune and chemically triggered oxidative stress modifies translational fidelity. Nature. 2009;462:522–526. doi: 10.1038/nature08576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Cherry JM, Adler C, Ball C, Chervitz SA, Dwight SS, Hester ET, Jia Y, Juvik G, Roe T, Schroeder M, et al. SGD: Saccharomyces Genome Database. Nucleic Acids Res. 1998;26:73–79. doi: 10.1093/nar/26.1.73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Hertz-Fowler C, Peacock CS, Wood V, Aslett M, Kerhornou A, Mooney P, Tivey A, Berriman M, Hall N, Rutherford K, et al. GeneDB: a resource for prokaryotic and eukaryotic organisms. Nucleic Acids Res. 2004;32:D339–343. doi: 10.1093/nar/gkh007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Komili S, Farny NG, Roth FP, Silver PA. Functional specificity among ribosomal proteins regulates gene expression. Cell. 2007;131:557–571. doi: 10.1016/j.cell.2007.08.037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.McIntosh KB, Warner JR. Yeast ribosomes: variety is the spice of life. Cell. 2007;131:450–451. doi: 10.1016/j.cell.2007.10.028. [DOI] [PubMed] [Google Scholar]
- 44.Warner JR, McIntosh KB. How common are extraribosomal functions of ribosomal proteins? Mol Cell. 2009;34:3–11. doi: 10.1016/j.molcel.2009.03.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Kim J, Chubatsu LS, Admon A, Stahl J, Fellous R, Linn S. Implication of mammalian ribosomal protein S3 in the processing of DNA damage. J Biol Chem. 1995;270:13620–13629. doi: 10.1074/jbc.270.23.13620. [DOI] [PubMed] [Google Scholar]
- 46.Yadavilli S, Hegde V, Deutsch WA. Translocation of human ribosomal protein S3 to sites of DNA damage is dependant on ERK-mediated phosphorylation following genotoxic stress. DNA Repair (Amst) 2007;6:1453–1462. doi: 10.1016/j.dnarep.2007.04.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Tang JB, Goellner EM, Wang XH, Trivedi RN, St Croix CM, Jelezcova E, Svilar D, Brown AR, Sobol RW. Bioenergetic metabolites regulate base excision repair-dependent cell death in response to DNA damage. Mol Cancer Res. 2009;8:67–79. doi: 10.1158/1541-7786.MCR-09-0411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–2504. doi: 10.1101/gr.1239303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Said MR, Begley TJ, Oppenheim AV, Lauffenburger DA, Samson LD. Global network analysis of phenotypic effects: protein networks and toxicity modulation in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 2004;101:18006–18011. doi: 10.1073/pnas.0405996101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Xenarios I, Salwinski L, Duan XJ, Higney P, Kim SM, Eisenberg D. DIP, the Database of Interacting Proteins: a research tool for studying cellular networks of protein interactions. Nucleic Acids Res. 2002;30:303–305. doi: 10.1093/nar/30.1.303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Rasmussen AK, Chatterjee A, Rasmussen LJ, Singh KK. Mitochondria-mediated nuclear mutator phenotype in Saccharomyces cerevisiae. Nucleic Acids Res. 2003;31:3909–3917. doi: 10.1093/nar/gkg446. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental Figure Legends
Figure S1. Genomic phenotyping of E. colimutants with UV.
All plates and replicates from the genomic phenotyping analysis of the E. coli deletion library.
Figure S2.UV sensitivity of S. cerevisiaerho- mutants.
By4741 cells were made rho- by ethidium bromide treatment and then analyzed for UV sensitivity on YPD plates. Wild-type and rho- mutants were serially diluted, plated and then exposed to increasing doses of UVC.
Figure S3. Genomic phenotyping of S. pombemutants with UV.
All plates and replicates from the genomic phenotyping analysis of the S. pombe deletion library.