Herpesvirus telomeric repeats facilitate virus integration into host telomeres, a process which is required for the establishment of virus latency.
Abstract
Some herpesviruses, particularly lymphotropic viruses such as Marek’s disease virus (MDV) and human herpesvirus 6 (HHV-6), integrate their DNA into host chromosomes. MDV and HHV-6, among other herpesviruses, harbor telomeric repeats (TMRs) identical to host telomeres at either end of their linear genomes. Using MDV as a natural virus-host model, we show that herpesvirus TMRs facilitate viral genome integration into host telomeres and that integration is important for establishment of latency and lymphoma formation. Integration into host telomeres also aids in reactivation from the quiescent state of infection. Our results and the presence of TMRs in many herpesviruses suggest that integration mediated by viral TMRs is a conserved mechanism, which ensures faithful virus genome maintenance in host cells during cell division and allows efficient mobilization of dormant viral genomes. This finding is of particular importance as reactivation is critical for virus spread between susceptible individuals and is necessary for continued herpesvirus evolution and survival.
Several DNA viruses and retroviruses are capable of integrating their genetic material into the host genome, which ensures viral genome maintenance and replication during cell division (Li et al., 2006; Hall et al., 2008). Among the DNA viruses, members of the Herpesviridae, which infect animals as diverse as crustaceans and humans, are known to be capable of genomic integration. Although EBV is found integrated in only a small fraction of latently infected cells, human herpesvirus 6 (HHV-6) and Marek’s disease virus (MDV) are found exclusively in an integrated state during the quiescent stage of the viral life cycle (Gulley et al., 1992; Delecluse and Hammerschmidt, 1993; Luppi et al., 1993; Hall et al., 2008). Other members of this group of viruses maintain their genomes exclusively in an episomal state during this phase of infection (Mitchell et al., 2003). Establishment of latency is a unifying and important principle for all herpesviruses that is used for prolonged, usually life-long, maintenance of the genetic material of this group of pathogens in once-infected hosts (Cohrs and Gilden, 2001). From the latent state, virus is reactivated intermittently to allow virus spread from infected to uninfected individuals in a population (Jones, 1998). Latency is a poorly understood series of events, which will uniformly result in the expression of only very few viral genes with the sole purpose of genome maintenance and the avoidance of a fully lytic replicative cycle resulting in cellular death (Cohrs and Gilden, 2001). Dependent on the cell populations targeted by individual herpesviruses, latency may only require genome maintenance when nonproliferative cells such as neurons are infected. However, in other cell types, for example, lymphocytes, faithful and continued replication of viral DNA during each mitosis must be ensured (Stevens, 1989; Cesarman and Mesri, 2007).
Although herpesvirus integration is a well known phenomenon, the mechanistic principles underlying the process have remained entirely elusive. Previously, HHV-6 and MDV, two lymphotropic viruses which can cause lymphoma, have been found integrated and their genomes are present at the distal ends of host chromosomes during latent infection (Delecluse et al., 1993; Luppi et al., 1994; Arbuckle et al., 2010). A recent study suggested that HHV-6 integrates at the proximal end of telomeres, protein-associated repeat sequences which protect chromosomes from damage and extensive shortening during DNA replication, and that integration in telomeres facilitates vertical transmission of the virus’ genetic material (Arbuckle et al., 2010). In addition, HHV-6A and B, HHV-7, MDV, and numerous other herpesviruses harbor telomeric repeats (TMRs) identical to host telomere sequences (TTAGGG)n at either end of their linear genomes (Kishi et al., 1988; Secchiero et al., 1995).
In the case of MDV, genomic integration and efficient induction of the latent state of infection is considered an important prelude to transformation and tumor formation, because the efficiency of lymphoma formation has been shown to correlate directly with the number of cells harboring latent genomes. Out of many latently infected and initially transformed cells, one out-competes all others with the consequence that tumors within one animal are almost invariably monoclonal (Delecluse and Hammerschmidt, 1993; Delecluse et al., 1993). During the latent state of MDV infection, few proteins and RNAs are produced, which ensure the establishment and maintenance of the quiescent state of infection and the continued survival of the host cell. Arguably the most important latency- and tumor-associated MDV protein is Meq, which serves as a repressor of lytic viral gene products, a prerequisite for latency. However, Meq is also a potent transcriptional activator in a heterodimeric complex with the proto-oncoprotein c-jun and can interact with p53 as well as RB, thereby enhancing transformation and T cell proliferation (Liu and Kung, 2000; Lupiani et al., 2004; Brown et al., 2006; Osterrieder et al., 2006). Another factor shown to be important for MDV-induced tumorigenesis is viral telomerase RNA (vTR), a homologue of cellular telomerase RNA (TR). As part of the telomerase complex, vTR ensures increased telomerase activity, at early stages of lymphomagenesis, but also promotes tumor formation independently of its presence in the telomerase complex (Kaufer et al., 2010b). The functions of Meq and vTR contribute to the rapid onset of lymphomas and death of infected animals within a few weeks of infection. These properties make MDV and the chicken an optimal natural virus-host model for studying herpesvirus integration and the effects on tumorigenesis.
Based on the observations of the presence of TMRs in viral genomes and integration into or near host telomeres, we hypothesized that herpesviruses use their TMR sequences for integration, thereby ensuring faithful replication and maintenance of the viral genome in rapidly dividing cells. Using the natural MDV-chicken model, we show that herpesvirus TMRs indeed facilitate directed integration into host telomeres, which is required for efficient tumor formation but also virus reactivation from latency. In the absence of the viral TMRs, herpesvirus integration is severely impaired, occurs in the form of concatemers, and is found at single sites within chromosomes. Our observation that integration occurs even in the absence of viral TMRs suggests that herpesvirus integration into the host genome is not a corollary of MDV replication but rather an important step in the establishment of latency, transformation, and tumorigenesis, as well as reactivation and spread to susceptible hosts.
RESULTS
MDV integrates into host telomeres
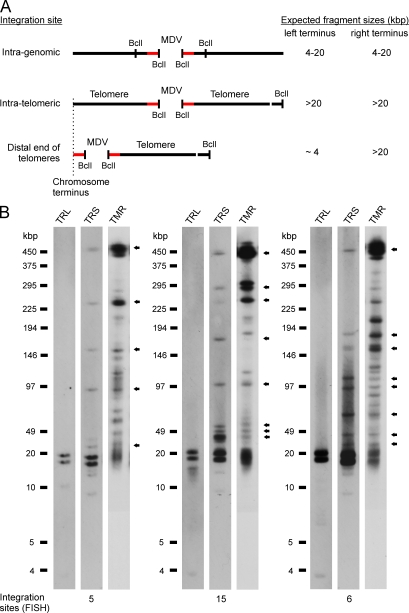
MDV is a highly oncogenic animal herpesvirus (Osterrieder et al., 2006), for which integration of viral DNA into the host genome was shown to be highly efficient and to target the distal end of multiple chromosomes (Delecluse et al., 1993). To elucidate whether MDV is indeed integrated into host telomeres in MDV-transformed cells, we performed pulsed field gel electrophoresis (PFGE) experiments. DNA from MDV-induced lymphoblastoid cell lines (LCLs) were digested with BclI, a restriction enzyme frequently cutting in both the host and viral genome but not in TMRs (Fig. 1 A). Southern blot analysis using probes specific for the terminal fragment of the TRS of the viral genome revealed that MDV is integrated into large fragments of up to 450 kbp that were indigestible with BclI. The sizes of the reactive fragments were found to correlate well with those reported for chicken telomeres, which range in size between 10 and 2,000 kbp (Fig. 1 B; Delany et al., 2003). The presence of telomere repeat sequences in fragments containing the viral genome was confirmed using a telomere-specific probe (TMR), which demonstrated that fragments reactive with the short repeat region of the genome (TRS; Fig. S1 A) coincided with bands positive for telomeres (Fig. 1 B). These results suggested that MDV does indeed integrate into or in very close proximity to the telomeres. To further determine whether MDV integrates into telomeres or at telomere termini, we probed for sequences specific for the other terminal genome region, the terminal repeat long sequences (TRL; Fig. S1 A). Hybridization with TRL sequences resulted in very short reactive fragments, indicating that MDV integrates into telomeres, which are located at either the extreme proximal (subtelomeric) or distal end (Fig. 1 B). Our results for lymphotropic MDV, therefore, are in agreement with a recent study showing that integration of a human lymphotropic herpesvirus, HHV-6, occurs at the internal end of the telomeres (Arbuckle et al., 2010).
Figure 1.
Identification of MDV integration sites. (A) Schematic representation of potential integration sites of the MDV genome within the host chromosome. Fragments resulting from digestion with BclI are depicted and expected fragment sizes are given. Terminal MDV genome fragments (red bars), telomeres, and BclI restriction sites are indicated. (B) PFGE analysis of LCL using BclI. PFGE patterns of three representative cell lines analyzed by Southern blotting, probing for the left MDV terminus (TRL), right MDV terminus (TRS), or telomere sequences (TMR), are shown. The 22.4- and 27.0-kbp fragments correspond to uncleaved termini and internal repeat fragments of the viral DNA. Arrows indicate colocalization of fragments that contain the TRS and high molecular mass telomere sequences. Results are representative of three independent experiments giving identical results. The number of integration sites detected by FISH is given at the bottom of the blot images.
Integrity of viral TMRs is dispensable for virus growth in vitro
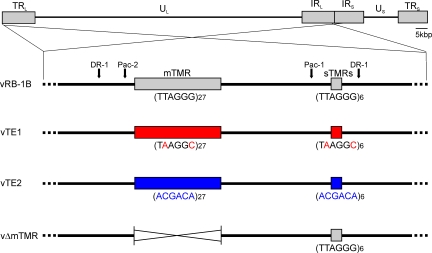
MDV DNA replication results in the formation of four isomers, where its two unique segments (unique-long, UL; and unique-short, US) can invert relative to the long or short inverted repeat sequences (TRL and TRS). MDV DNA harbors an additional copy of TMRs that ensures the presence of telomeres at all possible termini of the four DNA isomers (Fig. S1 A). We addressed the question of whether the viral TMRs mediate integration and if integration is important for establishment of latency and oncogenesis of MDV. Therefore, we replaced the repeats present in pRB-1B, an infectious bacterial artificial chromosome (BAC) clone of the highly oncogenic RB-1B MDV strain, with either structurally similar repeats harboring only two nucleotide exchanges (pTE1) relative to the telomere sequences or with completely scrambled repeats (pTE2; Fig. 2). We targeted both the terminal and internal TMR regions that each contain multiple (m) and short (s) TMRs. The mTMRs have 27 copies of TMRs in the case of RB-1B but up to 100 copies in other MDV stains, and sTMRs constantly specify 6 copies (Fig. 2). In an additional mutant, we deleted both copies of the multiple TMR region (pΔmTMR) to define the role of the longer repeats (Fig. 2). Based on the pTE1, pTE2, and pΔmTMR mutants, we also engineered revertant infectious clones in which the original telomere sequences were restored in both loci (pTE1rev, pTE2rev, and pΔmTMRrev). All infectious clones were confirmed by PCR, DNA sequencing of the targeted regions, and multiple restriction fragment length polymorphism analyses to ensure the integrity of the genome and to exclude fortuitous mutations that might have occurred elsewhere in the genome through the genetic manipulations. Southern blot analysis confirmed that the mutant and restored telomeric repeats were as designed in the recombinant viral genomes (Fig. S1 B). Recombinant viruses were reconstituted from BAC DNA in chicken embryo cells (CECs), and growth properties were evaluated using multistep growth kinetics (unpublished data) and plaque size assays. The viruses derived from pTE1 (vTE1), pTE2 (vTE2), and pΔmTMR (vΔmTMR) showed growth properties that were virtually indistinguishable from those of parental or revertant viruses (Fig. S2, A–C), confirming that neither the exact TMR sequences nor the mTMR region is important for lytic replication in vitro.
Figure 2.
MDV TMR mutants. Schematic representation of the MDV genome with a focus on viral a-like sequences containing the mTMR and sTMR regions present in the MDV genome. Recombinant BAC constructs in which the TMRs (TTAGGG)n were replaced by TE1 (TAAGGC)n or TE2 (ACGACA)n, as well as a BAC construct in which the mTMR region was deleted, are shown.
Absence of herpesvirus TMRs significantly reduces disease incidence and tumor development
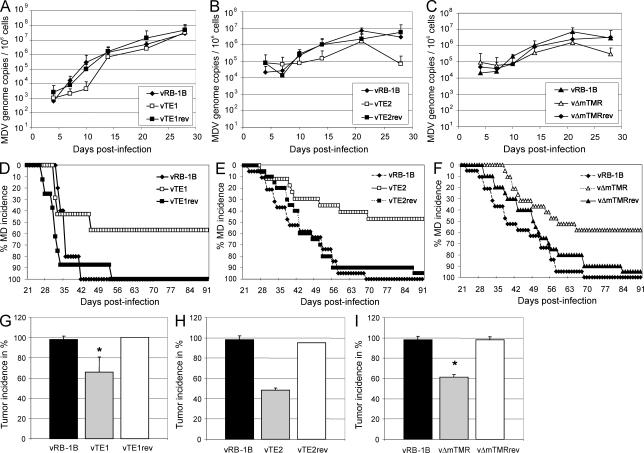
To address the role of the herpesvirus TMRs in disease and tumor development in vivo, 1-d-old P2a chickens highly susceptibly to MDV infection (Cole, 1968) were experimentally infected with parental vRB-1B, mutant vTE1, vTE2, or vΔTMR viruses, or the respective revertant viruses. After infection, we monitored viral levels in peripheral blood as well as disease and tumor development. Viral loads in the blood were only mildly affected in animals infected with vTE1, vTE2, or vΔmTMR (Fig. 3, A–C), indicating that herpesvirus TMRs do not play a prominent role in lytic virus replication in vivo, similar to the situation in cultured cells. In stark contrast, disease and tumor development were severely impaired in the telomere mutant viruses. Only 57.1, 47.1, and 57.9% of chickens, infected with vTE1, vTE2, or vΔmTMR, respectively, developed clinical disease, whereas 95–100% of animals infected with either parental or revertant viruses presented with clinical disease (Fig. 3, D–F). Tumor development was also significantly decreased in animals infected with vTE1 (65.7%, P = 0.021), vTE2 (48.6%), and vΔmTMR (61.4%, P = 1.98 × 10−6) when compared with parental vRB-1B or repair viruses (95–100%; Fig. 3, G–I). Collectively, the mean tumor incidence of 60% observed after intraabdominal infection with either of the three telomere mutants viruses (vMut) in nine independent experiments was significantly reduced when compared with parental vRB-1B (97.5%; P = 1.26 × 10−8) or revertant viruses (vRev; 98%; P = 3.68 × 10−6; Fig. 4 G).
Figure 3.
Mutation of herpesvirus TMRs mildly affects lytic replication but severely impairs disease and tumor development in vivo. (A–C) qPCR analysis of the viral ICP4 gene and host iNOS gene. Blood samples of animals infected with wild-type (vRB-1B), telomere mutant (TE1, TE2, and vΔmTMR), or revertant (TE1rev, TE2rev, and vΔmTMRrev) viruses were taken at 4, 7, 10, 14, 21, and 28 days after infection, and total DNA was extracted. Mean MDV genome copies per 1 × 106 cells of eight infected chickens per group are shown with standard deviations (error bars) determined in one independent experiment for the indicated viruses. (D–F) Marek’s disease (MD) incidence in percentage of chickens infected by the intraabdominal route with wild-type (vRB-1B; D, n = 5; E, n = 19; F, n = 19), telomere mutant (TE1, n = 7; TE2, n = 17; vΔmTMR, n = 19), or revertant (TE1rev, n = 8; TE2rev, n = 20; vΔmTMRrev, n = 20) viruses was monitored during the indicated time period after infection. (G–I) Tumor incidence in P2a chickens infected with indicated viruses. Results are shown as mean tumor incidences in two (G), three (H), or four (I) independent experiments with standard deviations (error bars). The mean tumor incidences in chickens infected with vTE1 and vΔmTMR were significantly decreased compared with those infected with vRB-1B, which is indicated by asterisks (G, P = 0.021; I, P = 1.98 × 10−6). Each group contained between 5 and 20 animals with a mean group size of n = 13.6. vRB-1B, TTAGGG; TE1, TTAGGG→TAAGGC; TE2, TTAGGG→ACGACA; vΔmTMR, mTMR deletion.
Figure 4.
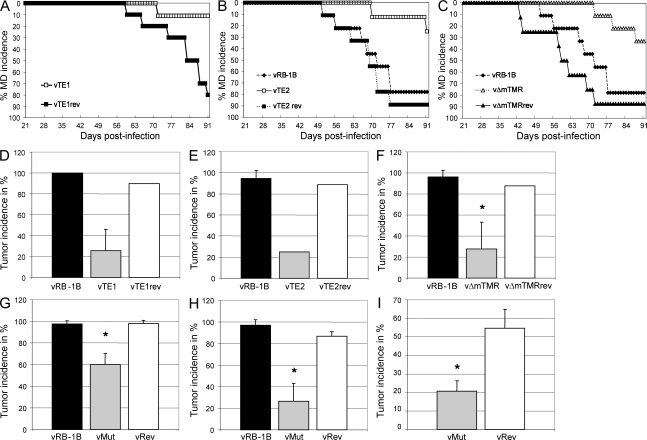
Disease and tumor development are severely impaired in the absence of the viral TMRs in animals infected via the natural route of infection. (A–C) Marek’s disease (MD) incidence in the percentage determined for in-contact animals housed with chickens infected with vRB-1B (n = 9), vTE1 (n = 10), or vTE1rev (n = 10; A), vTE2 (n = 8) or vTE2rev (n = 9; B), and vΔmTMR (n = 8) or vΔmTMRrev (n = 8; C). Chickens were monitored during the indicated time period and MD was recorded after necropsy and gross pathological examination. (D and E) Tumor incidence in percentage of contact animals housed with animals infected with wild-type (vRB-1B), telomere mutant (TE1, TE2, and vΔmTMR), or revertant (TE1rev, TE2rev, and vΔmTMRrev) viruses as indicated. Results are shown as mean tumor incidences of two (D and E) or three (F) independent experiments with standard deviations (error bars). The mean tumor incidences in chickens infected with vΔmTMR (F) were significantly decreased compared with incidences in animals infected with vRB-1B as indicated by the asterisk (P = 0.011). Each group contained between 2 and 10 animals with a mean group size of n = 6.7. (G–I) Mean tumor incidence in highly susceptible P2a chickens infected by either the intraabdominal (G) or the natural (H) route of infection or in more resistant N2a chickens (I) with parental vRB-1B, vMut (vTE1, vTE2, or vΔmTMR), or vRev (vTE1rev, vTE2rev, or vΔmTMRrev) viruses. Results are shown as mean tumor incidences of nine (G), seven (H), or four (I) independent experiments with standard deviations (error bars). The mean group sizes were n = 13.6 (G), n = 6.7 (H), and n = 19.5 (I). A significant decrease of mean tumor incidence compared with vRB-1B (G, P = 1.26 × 10−8; H, P = 2.17 × 10−7) or vRev (G, P = 3.68 × 10−6; H, P = 7.72 × 10−5; I, P = 0.001) is indicated with an asterisk. vRB-1B, TTAGGG; TE1, TTAGGG→TAAGGC; TE2, TTAGGG→ACGACA; vΔmTMR, mTMR deletion.
To determine whether mutation of the TMRs affected interindividual spread within a population, we housed chickens infected by the intraabdominal route together with uninfected chickens. Although mutant viruses were fully able to spread to uninfected animals, as indicated by quantitative (q) PCR detection of viral DNA in chicken blood (not depicted), disease incidence in animals infected with vTE1 (11.1%), vTE2 (25.0%), or vΔmTMR (33.3%) was dramatically diminished when compared with parental or repair viruses (80–100%; Fig. 4, A–C). Tumor development was also significantly decreased in chickens infected with vTE1 (25.5%), vTE2 (25.0%), or vΔmTMR (27.8%; P = 0.01) when compared with those infected with parental vRB-1B or repair viruses (83.8–100%; Fig. 4, D–F). After infection by the aerosol route, mean tumor incidence was significantly reduced after infection with telomere mutant viruses (vMut; 26.3%) in seven independent experiments when compared with parental vRB-1B (96.8%; P = 2.17 × 10−7) or revertant viruses (vRev; 86.6%; P = 7.72 × 10−5; Fig. 4 H). The results suggested an even more drastic defect in tumor development of vTE1, vTE2, and vΔmTMR in chickens exposed by the natural route of infection.
To confirm that the defect in disease and tumor development was not only restricted to chickens that are highly susceptible to MDV, we also infected N2a chickens, which exhibit increased resistance to MDV infection (Cole, 1968). Similar to the situation in P2a animals, tumor incidences were markedly reduced after infection of N2a chickens with vTE1 (25.6%), vTE2 (15.8%), or vΔmTMR (15.8%) when compared with the incidence after infection with the corresponding revertant viruses (42.9–63.1%; Fig. S3, A–C). Overall, the mean tumor incidence of 21% observed after infection of N2a chickens with either of the telomere mutants viruses (vMut) in four independent experiments was significantly reduced when compared with revertant viruses (vRev; 54.75%; P = 0.001; Fig. 4 I).
Integration defects of viruses harboring mutant TMRs
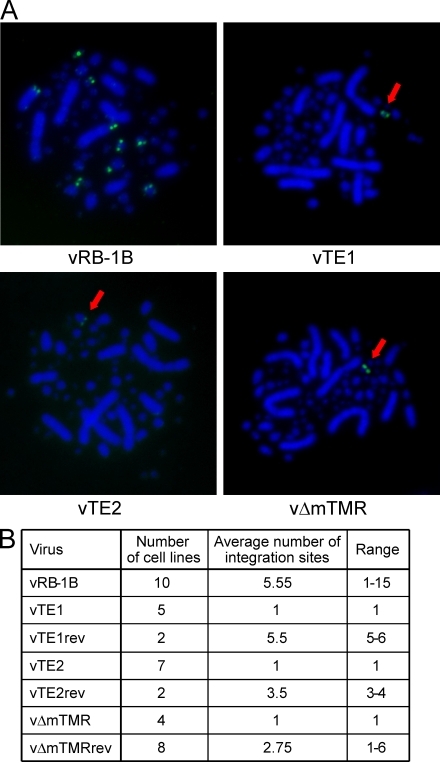
In the next series of experiments, we addressed the question of whether the telomere mutant viruses were still able to integrate their viral DNA into the host genome and performed metaphase fluorescent in situ hybridization (FISH). Analysis of LCL derived from animals infected with parental vRB-1B, TMR mutants, or revertant viruses revealed that vTE1, vTE2, and vΔmTMR had severe integration defects. Although integration of wild-type MDV DNA was identified in the telomeres of up to 15 different chromosomes in individual LCL derived from chickens infected with vRB-1B or revertant viruses, vTE1, vTE2, and vΔmTMR DNA was found integrated invariably in only one single chromosomal locus of each cell line (Fig. 5, A and B).
Figure 5.
Integration is severely impaired in the absence of the viral TMRs. (A) FISH analysis detecting MDV integration sites (anti-DIG FITC, green) in metaphase chromosomes (DAPI stain, blue) of representative cell lines of vRB-1B, vTE1, vTE2, and vΔmTMR. In the case of vTE1, vTE2, and vΔTMR, single integration sites are highlighted by arrows. (B) Mean number of integration sites in vRB-1B, mutant, and revertant cell lines. The number of analyzed cell lines and the frequencies of integration events are given. At least eight independent metaphase spread images were evaluated to determine the number of integration sites for each independent cell line.
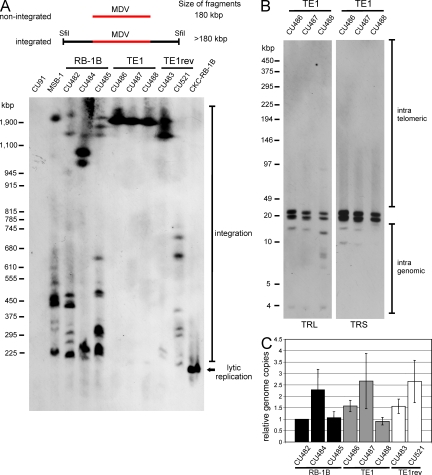
We then applied PFGE and Southern blot analyses to confirm whether integration of wild-type MDV DNA, as detected by FISH, truly represents integration, or rather, tethering, of the viral episome to host chromosomes as was shown, for example, in the case of EBV (Sears et al., 2004). This approach also provided to us the opportunity to investigate the status of viral DNA after integration of viruses with mutant TMRs. Digestion with SfiI, a restriction enzyme which does not cut within the MDV genome, allowed the distinction between integrated and nonintegrated/tethered MDV DNA (Fig. 6 A). Extended proteinase K digestion of LCL DNA embedded in agarose was applied to exclude the possibility of tethering of the viral genome to the host chromosome by a proteinaceous structure. Consistent with the FISH results, multiple integration sites with varying sizes of reactive fragments were identified in DNA of LCL from animals infected with vRB-1B or vTE1rev. In contrast, in the case of LCL derived from vTE1-infected animals, integration was restricted to a single large DNA fragment of ∼1.9 Mbp in size. To address whether the integration sites found in vTE1 LCL also mapped to telomeres, we performed PFGE after BclI digestion. Southern blotting using either the TRS and TRL probe confirmed that vTE1 genomes were only detected in low molecular mass fragments of 4–20 kbp in size, with both the probes derived from either terminal viral DNA fragment. We did not detect, however, any colocalization with telomere-containing fragments (Fig. 6 B; and Fig. S4, A–C). This data and identical results with LCL derived from vTE2-infected animals (unpublished data) demonstrated that only minimal changes in MDV TMR sequences abrogated specific integration into host telomeres and suggests that mutant viral DNA integrates into intrachromosomal sites. Intrachromosomal integration was detected in various chromosomes by FISH (unpublished data), suggesting that there is likely no preferred locus for integration site or that nontelomeric integration sites are present in multiple chromosomes.
Figure 6.
Integration does not occur in host telomeres in the absence of viral TMRs. (A) Schematic representation and corresponding PFGE and Southern blot analysis of LCL DNA digested with SfiI. Fragment sizes generated by SfiI digestion of integrated and nonintegrated MDV genomes are depicted and sizes are given. The size of the linear MDV genome observed during lytic replication is indicated by an arrow. Results are representative of three independent Southern blot analyses. (B) Southern blotting of DNA of LCL derived from animals infected with vTE1 and digested with BclI. Potential intragenomic and telomeric integration sites are indicated. Results are representative of three independent Southern blot analyses. (C) Quantification of MDV copies in tumor cells. Results are shown as mean herpesvirus genome copies detected by the TRL probe relative to B2M in three independent experiments. The data are shown relative to LCL CU482 derived from a vRB-1B–infected chicken with standard deviations (error bars).
In the subsequent experiments, we determined if the vTE1 mutant integrates as a single viral genome or as concatemers and analyzed MDV genome copy numbers in LCL. Large genome fragments with high intensities of reactive bands (Fig. 6 A) had already suggested that vTE1 DNA might be integrated not as a single copy but as multiples in the form of head-to-head or head-to-tail concatemers. Viral copy numbers in LCL were determined by Southern blot analysis using normalization to copies of host β2 microglobulin (B2M; Fig. S4, A-D). Although integrated at only one site, MDV genome copy numbers in all vTE1 cell lines analyzed were comparable to those present in CU482, a cell line with 11 integration sites, clearly indicating that vTE1 genomes were integrated as concatemers (Fig. 6 C and Fig. S4 E). In contrast, wild-type and revertant viruses can apparently also integrate as a single viral genome (Fig. 6 A). These findings were also confirmed by qPCR analysis of numbers of viral DNA copies of individual LCLs (unpublished data). To address whether the site of integration has an effect on expression levels of known latency-associated genes, we examined vTR transcription and Meq protein levels in LCLs (Trapp et al., 2006; Kaufer et al., 2010b). vTR expression, as quantified by qRT-PCR, moderately varied between cell lines (up to 3.3-fold) but levels were not dependent on whether wild-type, vTE1, or revertant cell lines were analyzed (Fig. S5 A). Similarly, Western blot analysis confirmed that Meq expression was also variable between cell lines; however, as shown for vTR transcription, there was no difference between vTE1 cell lines when compared with wild-type and revertant cell lines (Fig. S5 B).
Reactivation of viruses with mutant TMRs is severely impaired
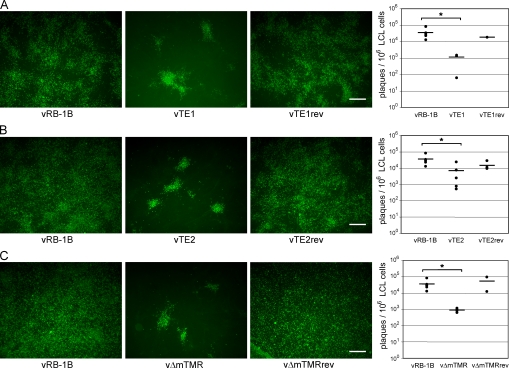
Finally, we determined whether integration of viral genomes into telomeres had a measurable effect on virus fitness, more specifically on reactivation of virus from the latent quiescent state of infection. Although genome maintenance during the latent phase of the virus life cycle is an important transitory phase, reactivation and spread to new hosts are essential for continued virus survival. Because integration into host telomeres could also provide an efficient mechanism for virus genome mobilization through homologous recombination between virus and host telomeres, we analyzed the efficiency of virus reactivation from LCL. Reactivation assays that rely on induction of the lytic replication cycle from latent genomes revealed that vTE1, vTE2, and vΔmTMR were severely impaired with respect to genome mobilization and their reactivation from latently infected cells (Fig. 7, A–C). The mean number of plaques per 1 × 106 tumor cells induced on CECs was significantly (5–39-fold) decreased in the case of cell lines derived from chickens infected with vTE1 (1.13 × 104), vTE2 (7.06 × 104), and vΔmTMR (0.91 × 104) when compared with those from birds infected with parental vRB-1B (3.53 × 105; Fig. 7, A–C). We concluded that the presence of viral TMRs and the resulting integration of MDV DNA into host telomeres serves a purpose beyond genome maintenance during latency by allowing efficient and rapid mobilization of virus genomes in response to external cues for initiation of the lytic phase of infection.
Figure 7.
Reactivation is significantly impaired in the absence of viral TMRs. (A–C) Reactivation assay using LCL derived from animals infected with wild-type (vRB-1B), telomere mutant (TE1, TE2, and vΔmTMR), or revertant (TE1rev, TE2rev, and vΔmTMRrev) viruses as indicated. Representative images of virus reactivation in CECs are shown. Bars, 500 µm. Quantification of reactivation assays is shown as the mean (horizontal bars) number of plaques per 1 × 106 tumor cells for each individual cell line (A–C, right), performed in triplicate for each of three independent experiments. Reactivation of lytic virus from vTE1-, vTE2-, and vΔmTMR-derived LCL was significantly reduced compared with that from vRB-1B–derived LCL, as indicated by the asterisks (A, P = 0.013; B, P = 0.039; C, P = 0.025). vRB-1B, TTAGGG; TE1, TTAGGG→TAAGGC; TE2, TTAGGG→ACGACA; vΔmTMR, mTMR deletion.
DISCUSSION
Herpesvirus integration into the host chromosome, as reported for several lymphotropic herpesviruses, including EBV, HHV-6, and MDV, was viewed as an unintended consequence and by-product of viral replication (Delecluse and Hammerschmidt, 1993; Luppi et al., 1993; Hall et al., 2008). In this paper, we provide the first evidence that telomere sequences present in the MDV genome and the genome of several other herpesvirus genomes facilitate directed integration into host telomeres (Delecluse and Hammerschmidt, 1993; Arbuckle et al., 2010). We show that integration occurs very efficiently if TMRs in the virus genome are present. If, however, TMRs are absent or mutated, integration frequency is greatly reduced and occurs elsewhere intrachromosomally.
It is well known that host telomere sequences prevent the induction of DNA damage responses (Blasco, 2005; Murnane, 2006). We report in this paper that deletion or mutation of viral TMRs did not affect lytic replication in vitro and in vivo suggesting that viral telomeres do not seem to aid the evasion of DNA damage responses. However, we cannot exclude the tumor-promoting functions of MDV’s TMRs beyond facilitating integration into host telomeres. These potential functions will have to be the scope of future studies.
We can confirm in this paper that MDV integration is indeed, as previously shown for another herpesvirus, HHV-6, at telomere termini. PFGE analysis demonstrated that the inverted repeat short terminus of the genome, which contains the mTMR region in the linear MDV genome, is covalently linked to telomeres. Deletion of the mTMR region, which harbors a large but variable number of the hexameric repeats, confirmed that this region is involved in integration. Collectively, we concluded that our data provide initial evidence that integration efficiency is directly correlated with tumor development and disease. It is conceivable that the integration event itself could influence disease development because epigenetic regulation of subtelomeric host genes could also have an effect on the expression of viral genes. It still remains unknown, however, what exact role in the process of integration is served by sTMR, which constantly specifies six copies of the TTAGGG repeats.
Efficient induction of the latent state of infection, likely initiated by efficient integration of the viral into the host genome, leads to lymphoma formation and therefore seems to play an important role in tumor formation in the model chosen here. Integration into telomeres, as shown for MDV and HHV-6, could provide several advantages. The first advantage is the highly recombinogenic nature of TMRs, which could help facilitate efficient entry into the host genome. Homologous recombination between telomeres is known to ensure telomere maintenance via the so-called alternative telomere elongation (ALT) mechanism (Murnane, 2006). We presently surmise that ALT, or a very similar mechanism, is capable of mediating recombination between host and viral telomeres. It is notable that ALT is independent of the action of TR and TERT (telomerase reverse transcription), the two main components of telomerase usually involved in the maintenance of protective telomere structures at the end of linear chromosomes (Murnane, 2006). ALT is commonly seen in telomerase-deficient cancer cells but is also thought to occur in certain somatic cell types (Perrem et al., 2001; Reddel, 2003). In addition, telomere recombination can be achieved by homology-directed repair (HDR) in the absence of an intact telomeric shelterin complex (Sfeir et al., 2010). We propose that the relatively short viral TMRs, consisting of only between 12 and 100 repeats (Spatz et al., 2007), will preclude assembly of the shelterin complex and the structure may, as such, be a potent inducer of HDR. It would be desirable to specifically interfere with HDR after infection of chickens with MDV to directly assess its role in integration and development of latency; however, to our knowledge, HDR inhibitors for in vivo use are currently not available.
The second advantage of telomere integration is the heterochromatic nature of telomeres and adjacent regions (Blasco, 2005), which could aid in silencing of the viral genome during the latent state of the viral life cycle. Our findings of higher frequencies of viral DNA integration and lymphomas in the presence of intact viral telomeres is fully consistent with these theoretical deliberations, as is the fact that mutation of only two nucleotides in the TTAGGG repeat precludes efficient integration into host telomeres.
The third advantage, again dependent on the highly recombinogenic nature of the telomeres, could contribute to the efficient mobilization of the virus genome from the latent, quiescent, and nonproductive state of infection. As observed for the frequency of integration, we determined significantly reduced reactivation levels if the virus does not harbor TMRs. Reduced reactivation was shown for two independent TMR mutant viruses and a virus in which the TMRs were deleted, manipulations which made integration into telomeres impossible. Mobilization of the virus genome is the first and most important step to be taken for the production of fully replicating virus, which is absolutely required for interindividual spread and virus maintenance on a population level. It is worthwhile pointing out that efficiency of mobilization of viral genomes from the integrated state was independent on the number of integration sites. Even in LCLs that presented with low copy numbers of latent viral DNA integrated into telomeres, mobilization and induction of lytic replication was significantly more efficient when compared with LCLs in which viral genomes did not harbor TMRs and, therefore, integrated elsewhere in the chromosome.
Finally, reduced reactivation levels likely cannot be attributed to the loss of terminal sequences that could occur during a nontelomeric integration event. Mutant virus genomes were integrated in the form of concatemers as demonstrated in Fig. 6 (A–C). Integration in the form of concatemers could provide an explanation of how viral genomes are mobilized without the loss of sequence information, namely through HDR between repeat sequences of directly adjacent genomes.
In conclusion, we demonstrated that herpesvirus TMRs, which are present in several herpesviruses including HHV-6 and MDV, facilitate directed integration into the host genome. Our results, therefore, provide the first conclusive evidence not only that herpesvirus TMRs mediate chromosomal integration but that these repeat structures are crucial for efficient tumor formation and reactivation of latent virus from the quiescent state of infection. Mutation or deletion of the TMRs resulted in decreased integration levels with only a single nontelomeric integration site within host chromosomes where viral genomes were present as concatemers. Our observation that integration occurs even in the absence of virus TMRs suggests that herpesvirus integration into the host genome is not simply a consequence of virus replication but, more likely, a prerequisite for transformation and tumorigenesis as well as for reactivation and spread to new susceptible hosts.
MATERIALS AND METHODS
Cells and viruses.
CECs were prepared from specific pathogen-free embryos and maintained as described previously (Osterrieder, 1999). Recombinant viruses were reconstituted in CEC by CaPO4 transfection of purified BAC DNA as described previously (Schumacher et al., 2000; Jarosinski et al., 2007b). The mini-F sequences flanked by loxP sites within the infectious clones were removed by cotransfection with a Cre recombinase expression vector (pCAGGS-NLS/Cre; Jarosinski et al., 2007b). Removal of mini-F sequences was ensured by analyzing recombinant virus stocks via PCR as described previously (Jarosinski et al., 2007b). Virus propagation and determination of virus growth kinetics and plaque sizes were performed exactly as described previously (Schumacher et al., 2005). Viruses used for in vitro and in vivo experiments were passaged no more than five times in vitro. All virus stocks, as well as virus derived from tumor cells, were analyzed by DNA sequencing to ensure that no unexpected changes occurred in the loci targeted by mutagenesis.
Generation of mutant MDV.
The MDV telomere region (Fig. 2) was amplified from pRB-1B and cloned into the pCR2.1 Topo vector (Invitrogen), resulting in pTMR. Plasmids containing a synthetic telomere region in which viral TMRs (TTAGGG)n were replaced by telomere exchange repeats 1 (TAAGGC)n or telomere exchange repeats 2 (ACGACA)n were obtained from Celtec and GenScript, respectively, and designated pTE1 and pTE2. Transfer plasmids for mutagenesis were generated as described previously (Tischer et al., 2006). In brief, the aphAI-I–SceI cassette was amplified from pEPkan-SII and inserted into pTMR, pTE1, or pTE2 using a unique SacI or NheI restriction site within the viral telomere region, resulting in the plasmids pTMR transfer, pTE1 transfer, and pTE2 transfer, respectively. Corresponding transfer plasmids and oligonucleotides (Table S1) were used for mutagenesis as described previously (Tischer et al., 2006). All clones were confirmed by PCR, DNA sequencing, RFLP using different restriction enzymes, and Southern blot analysis to ensure integrity of the genomes and to exclude fortuitous mutations elsewhere in the genome as described previously (Tischer et al., 2006; Kaufer et al., 2010b). Primers used for transfer plasmids and mutagenesis are given in Table S1.
In vivo experiments.
Specific pathogen-free P2a (MHC haplotype B19B19) or N2a (MHC haplotype B21B21) chickens were experimentally infected by intraabdominal inoculation with 500–2,000 plaque-forming units at day 1 of age or via the natural route of infection by cohousing of uninfected with infected animals. All experimental procedures were conducted in compliance with approved Institutional Animal Care and Use Committee protocols under the internal approval number 2002-0085. Chickens were evaluated for symptoms of MDV-induced disease on a daily basis and examined for tumorous lesions when clinical symptoms were evident or at termination of the experiments.
DNA, RNA extraction, and qPCR analysis.
DNA was extracted from whole chicken blood using the EZ 96 blood DNA kit (Omega Bio-Tek) and MDV genomic copies were determined by qPCR (Jarosinski et al., 2005, 2007a). MDV DNA copy numbers were detected using primers and probes specific for the ICP4 locus, and normalization was achieved by determining copies of the chicken iNOS (inducible nitric oxide synthase) gene (Jarosinski et al., 2007b). RNA was extracted from LCLs using the RNeasy Plus Mini kit (QIAGEN), cDNA was synthesized using the Enhanced Avian HS RT-PCR kit (Sigma-Aldrich), and vTR copies were determined by qRT-PCR exactly as described previously (Chbab et al., 2010).
LCLs.
LCLs were generated as described previously (Calnek et al., 1978, 1981). Tumor tissue from MDV-infected animals was harvested and sieved through a cell strainer before lymphocytes were isolated using Histopaque 1119 (Sigma-Aldrich) density gradient centrifugation. Purified cells were cultivated in a 1:1 mixture of Leibovitz-McCoy medium (both obtained from Mediatech, Inc.) and modified according to Hahn as detailed previously (Calnek et al., 1981). The medium was supplemented with 10% FBS and 8% chicken serum at 41°C under a 5% CO2 atmosphere.
FISH.
Metaphase chromosome preparations and FISH analysis were performed as described previously (Rens et al., 2006). A digoxigenin (DIG)-labeled MDV whole genome probe was generated via the DIG High Prime kit (Roche) and used to detect MDV integration sites that were visualized using an FITC-conjugated anti-DIG antibody (Sigma-Aldrich). Metaphase FISH preparations were analyzed using the Axio Imager M1 system and the AxioVision software (Carl Zeiss, Inc.).
PFGE analysis.
PFGE analysis was performed as described previously (Herschleb et al., 2007). 1 × 107/ml LCLs were embedded in 1% agarose and digested at 50°C for 48 h hours in lysis buffer (0.5 M EDTA and 1% wt/vol N-laurylsarcosine) containing 1 mg/ml proteinase K. Proteinase K was inactivated with 0.01 mM phenylmethanesulfonyl fluoride, and agarose plugs were digested with either SfiI or BclI (New England Biolabs) overnight according to the manufacturer’s instructions. DNA fragments were resolved via PFGE using the CHEF-DR III system (Bio-Rad Laboratories). SfiI fragments were resolved with 50–100-s and BclI fragments with 1–25-s switch time gradients at 6 V/cm at 14°C, with a 120° angle and run times of 36 and 24 h, respectively.
Southern blot analysis.
Southern blots were performed after DNA transfer onto a positively charged nylon membrane (GE Healthcare; Sambrook et al., 1989). Fragments containing the TRL or TRS segment of the MDV genome were identified with specific probes generated via the PCR DIG Probe Synthesis kit (Roche). TMRs or mutant repeats were detected using DIG-labeled oligonucleotide probes (Eurofins MWG; Table S1).
Western blot analysis.
Western blots were performed as described previously (Kaufer et al., 2010a). The oncoprotein Meq was detected using a monoclonal mouse anti-Meq antibody (Brown et al., 2006; gift from V. Nair, Institute for Animal Health, Compton, England, UK), and β-actin was probed with a monoclonal rabbit antibody (Cell Signaling Technology). The Meq antibody was used at dilution of 1:100, and the actin antibody was used at a 1:1,000 dilution.
Reactivation assays.
Established LCLs were purified via Histopaque 1119 gradient centrifugation (Sigma-Aldrich; Parcells et al., 1999), washed with PBS, and serial 10-fold dilutions (103, 104, 105, or 106) of LCLs were seeded on fresh CEC cultures. MDV reactivation was stimulated by cocultivation of LCLs with CEC at low serum concentrations (0.1% FBS) and a temperature shift to 37°C as described previously (Calnek et al., 1981). LCLs were completely removed 24 h after seeding through extensive washing. 4 d after infection, CEC cultures were fixed, stained, and analyzed via immunofluorescence analysis using a chicken anti-MDV antiserum and visualized via an Alexa Fluor 488–conjugated anti–chicken antibody.
Statistical analysis.
Significant differences in means of tumor incidences (Fig. 3, G–I; and Fig. 4, F–I) and LCL reactivation assays (Fig. 7, A–C) were determined using Student’s t test.
Online supplemental material.
Fig. S1 depicts the four isomeric forms of the MDV genome and a Southern blot to confirm the mutation of TMRs in recombinant viruses. Fig. S2 presents plaque area determinations of telomere mutant viruses. Fig. S3 shows tumor incidences in N2a chickens infected with telomere mutant viruses. Fig. S4 gives a quantitative analysis of viral copy numbers in LCLs using Southern blot analysis. Fig. S5 shows vTR and Meq expression levels in LCLs derived from chicken infected with telomere mutant viruses. Online supplemental material is available at http://www.jem.org/cgi/content/full/jem.20101402/DC1.
Acknowledgments
We thank Kerstin Osterrieder for help with the statistical calculations and advice. We also thank Annemarie Engel for her assistance with some of the experiments and V. Nair for providing the Meq antibody. We are grateful to Joel Baines and Colin Parrish for editing the manuscript.
This study was supported by PHS grant 5R01CA127238 to N. Osterrieder and K.W. Jarosinski and an unrestricted grant from the Freie Universität Berlin to N. Osterrieder.
The authors have no conflicting financial interests.
Author contributions: B.B. Kaufer and N. Osterrieder conceived and designed the study. B.B. Kaufer and K.W. Jarosinski performed the experiments. B.B. Kaufer and N. Osterrieder wrote the manuscript.
Footnotes
Abbreviations:
- B2M
- β2 microglobulin
- BAC
- bacterial artificial chromosome
- CEC
- chicken embryo cell
- DIG
- digoxigenin
- FISH
- fluorescent in situ hybridization
- HHV-6
- human herpesvirus 6
- LCL
- lymphoblastoid cell line
- MDV
- Marek’s disease virus
- PFGE
- pulsed field gel electrophoresis
- TMR
- telomeric repeat
- vTR
- viral telomerase RNA
References
- Arbuckle J.H., Medveczky M.M., Luka J., Hadley S.H., Luegmayr A., Ablashi D., Lund T.C., Tolar J., De Meirleir K., Montoya J.G., et al. 2010. The latent human herpesvirus-6A genome specifically integrates in telomeres of human chromosomes in vivo and in vitro. Proc. Natl. Acad. Sci. USA. 107:5563–5568 10.1073/pnas.0913586107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blasco M.A. 2005. Telomeres and human disease: ageing, cancer and beyond. Nat. Rev. Genet. 6:611–622 10.1038/nrg1656 [DOI] [PubMed] [Google Scholar]
- Brown A.C., Baigent S.J., Smith L.P., Chattoo J.P., Petherbridge L.J., Hawes P., Allday M.J., Nair V. 2006. Interaction of MEQ protein and C-terminal-binding protein is critical for induction of lymphomas by Marek’s disease virus. Proc. Natl. Acad. Sci. USA. 103:1687–1692 10.1073/pnas.0507595103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calnek B.W., Murthy K.K., Schat K.A. 1978. Establishment of Marek’s disease lymphoblastoid cell lines from transplantable versus primary lymphomas. Int. J. Cancer. 21:100–107 10.1002/ijc.2910210117 [DOI] [PubMed] [Google Scholar]
- Calnek B.W., Shek W.R., Schat K.A. 1981. Spontaneous and induced herpesvirus genome expression in Marek’s disease tumor cell lines. Infect. Immun. 34:483–491 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cesarman E., Mesri E.A. 2007. Kaposi sarcoma-associated herpesvirus and other viruses in human lymphomagenesis. Curr. Top. Microbiol. Immunol. 312:263–287 10.1007/978-3-540-34344-8_10 [DOI] [PubMed] [Google Scholar]
- Chbab N., Egerer A., Veiga I., Jarosinski K.W., Osterrieder N. 2010. Viral control of vTR expression is critical for efficient formation and dissemination of lymphoma induced by Marek’s disease virus (MDV). Vet. Res. 41:56 10.1051/vetres/2010026 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cohrs R.J., Gilden D.H. 2001. Human herpesvirus latency. Brain Pathol. 11:465–474 10.1111/j.1750-3639.2001.tb00415.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cole R.K. 1968. Studies on genetic resistance to Marek’s disease. Avian Dis. 12:9–28 10.2307/1588081 [DOI] [PubMed] [Google Scholar]
- Delany M.E., Daniels L.M., Swanberg S.E., Taylor H.A. 2003. Telomeres in the chicken: genome stability and chromosome ends. Poult. Sci. 82:917–926 [DOI] [PubMed] [Google Scholar]
- Delecluse H.J., Hammerschmidt W. 1993. Status of Marek’s disease virus in established lymphoma cell lines: herpesvirus integration is common. J. Virol. 67:82–92 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Delecluse H.J., Schüller S., Hammerschmidt W. 1993. Latent Marek’s disease virus can be activated from its chromosomally integrated state in herpesvirus-transformed lymphoma cells. EMBO J. 12:3277–3286 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gulley M.L., Raphael M., Lutz C.T., Ross D.W., Raab-Traub N. 1992. Epstein-Barr virus integration in human lymphomas and lymphoid cell lines. Cancer. 70:185–191 10.1002/1097-0142(19920701)70:1<185::AID-CNCR2820700129>3.0.CO;2-J [DOI] [PubMed] [Google Scholar]
- Hall C.B., Caserta M.T., Schnabel K., Shelley L.M., Marino A.S., Carnahan J.A., Yoo C., Lofthus G.K., McDermott M.P. 2008. Chromosomal integration of human herpesvirus 6 is the major mode of congenital human herpesvirus 6 infection. Pediatrics. 122:513–520 10.1542/peds.2007-2838 [DOI] [PubMed] [Google Scholar]
- Herschleb J., Ananiev G., Schwartz D.C. 2007. Pulsed-field gel electrophoresis. Nat. Protoc. 2:677–684 10.1038/nprot.2007.94 [DOI] [PubMed] [Google Scholar]
- Jarosinski K.W., Osterrieder N., Nair V.K., Schat K.A. 2005. Attenuation of Marek’s disease virus by deletion of open reading frame RLORF4 but not RLORF5a. J. Virol. 79:11647–11659 10.1128/JVI.79.18.11647-11659.2005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jarosinski K., Kattenhorn L., Kaufer B., Ploegh H., Osterrieder N. 2007a. A herpesvirus ubiquitin-specific protease is critical for efficient T cell lymphoma formation. Proc. Natl. Acad. Sci. USA. 104:20025–20030 10.1073/pnas.0706295104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jarosinski K.W., Margulis N.G., Kamil J.P., Spatz S.J., Nair V.K., Osterrieder N. 2007b. Horizontal transmission of Marek’s disease virus requires US2, the UL13 protein kinase, and gC. J. Virol. 81:10575–10587 10.1128/JVI.01065-07 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones C. 1998. Alphaherpesvirus latency: its role in disease and survival of the virus in nature. Adv. Virus Res. 51:81–133 10.1016/S0065-3527(08)60784-8 [DOI] [PubMed] [Google Scholar]
- Kaufer B.B., Smejkal B., Osterrieder N. 2010a. The varicella-zoster virus ORFS/L (ORF0) gene is required for efficient viral replication and contains an element involved in DNA cleavage. J. Virol. 84:11661–11669 10.1128/JVI.00878-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaufer B.B., Trapp S., Jarosinski K.W., Osterrieder N. 2010b. Herpesvirus telomerase RNA(vTR)-dependent lymphoma formation does not require interaction of vTR with telomerase reverse transcriptase (TERT). PLoS Pathog. 6:e1001073 10.1371/journal.ppat.1001073 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kishi M., Harada H., Takahashi M., Tanaka A., Hayashi M., Nonoyama M., Josephs S.F., Buchbinder A., Schachter F., Ablashi D.V., et al. 1988. A repeat sequence, GGGTTA, is shared by DNA of human herpesvirus 6 and Marek’s disease virus. J. Virol. 62:4824–4827 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li M., Mizuuchi M., Burke T.R., Jr, Craigie R. 2006. Retroviral DNA integration: reaction pathway and critical intermediates. EMBO J. 25:1295–1304 10.1038/sj.emboj.7601005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu J.L., Kung H.J. 2000. Marek’s disease herpesvirus transforming protein MEQ: a c-Jun analogue with an alternative life style. Virus Genes. 21:51–64 10.1023/A:1008132313289 [DOI] [PubMed] [Google Scholar]
- Lupiani B., Lee L.F., Cui X., Gimeno I., Anderson A., Morgan R.W., Silva R.F., Witter R.L., Kung H.J., Reddy S.M. 2004. Marek’s disease virus-encoded Meq gene is involved in transformation of lymphocytes but is dispensable for replication. Proc. Natl. Acad. Sci. USA. 101:11815–11820 10.1073/pnas.0404508101 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luppi M., Marasca R., Barozzi P., Ferrari S., Ceccherini-Nelli L., Batoni G., Merelli E., Torelli G. 1993. Three cases of human herpesvirus-6 latent infection: integration of viral genome in peripheral blood mononuclear cell DNA. J. Med. Virol. 40:44–52 10.1002/jmv.1890400110 [DOI] [PubMed] [Google Scholar]
- Luppi M., Barozzi P., Marasca R., Torelli G. 1994. Integration of human herpesvirus-6 (HHV-6) genome in chromosome 17 in two lymphoma patients. Leukemia. 8:S41–S45 [PubMed] [Google Scholar]
- Mitchell B.M., Bloom D.C., Cohrs R.J., Gilden D.H., Kennedy P.G. 2003. Herpes simplex virus-1 and varicella-zoster virus latency in ganglia. J. Neurovirol. 9:194–204 [DOI] [PubMed] [Google Scholar]
- Murnane J.P. 2006. Telomeres and chromosome instability. DNA Repair (Amst.). 5:1082–1092 10.1016/j.dnarep.2006.05.030 [DOI] [PubMed] [Google Scholar]
- Osterrieder N. 1999. Sequence and initial characterization of the U(L)10 (glycoprotein M) and U(L)11 homologous genes of serotype 1 Marek’s Disease Virus. Arch. Virol. 144:1853–1863 10.1007/s007050050710 [DOI] [PubMed] [Google Scholar]
- Osterrieder N., Kamil J.P., Schumacher D., Tischer B.K., Trapp S. 2006. Marek’s disease virus: from miasma to model. Nat. Rev. Microbiol. 4:283–294 10.1038/nrmicro1382 [DOI] [PubMed] [Google Scholar]
- Parcells M.S., Dienglewicz R.L., Anderson A.S., Morgan R.W. 1999. Recombinant Marek’s disease virus (MDV)-derived lymphoblastoid cell lines: regulation of a marker gene within the context of the MDV genome. J. Virol. 73:1362–1373 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perrem K., Colgin L.M., Neumann A.A., Yeager T.R., Reddel R.R. 2001. Coexistence of alternative lengthening of telomeres and telomerase in hTERT-transfected GM847 cells. Mol. Cell. Biol. 21:3862–3875 10.1128/MCB.21.12.3862-3875.2001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reddel R.R. 2003. Alternative lengthening of telomeres, telomerase, and cancer. Cancer Lett. 194:155–162 10.1016/S0304-3835(02)00702-4 [DOI] [PubMed] [Google Scholar]
- Rens W., Fu B., O’Brien P.C., Ferguson-Smith M. 2006. Cross-species chromosome painting. Nat. Protoc. 1:783–790 10.1038/nprot.2006.91 [DOI] [PubMed] [Google Scholar]
- Sambrook J., Fritsch D.F., Maniatis T. 1989. Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratories, Cold Spring Harbor, NY: 1659 pp [Google Scholar]
- Schumacher D., Tischer B.K., Fuchs W., Osterrieder N. 2000. Reconstitution of Marek’s disease virus serotype 1 (MDV-1) from DNA cloned as a bacterial artificial chromosome and characterization of a glycoprotein B-negative MDV-1 mutant. J. Virol. 74:11088–11098 10.1128/JVI.74.23.11088-11098.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schumacher D., Tischer B.K., Trapp S., Osterrieder N. 2005. The protein encoded by the US3 orthologue of Marek’s disease virus is required for efficient de-envelopment of perinuclear virions and involved in actin stress fiber breakdown. J. Virol. 79:3987–3997 10.1128/JVI.79.7.3987-3997.2005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sears J., Ujihara M., Wong S., Ott C., Middeldorp J., Aiyar A. 2004. The amino terminus of Epstein-Barr Virus (EBV) nuclear antigen 1 contains AT hooks that facilitate the replication and partitioning of latent EBV genomes by tethering them to cellular chromosomes. J. Virol. 78:11487–11505 10.1128/JVI.78.21.11487-11505.2004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Secchiero P., Nicholas J., Deng H., Xiaopeng T., van Loon N., Ruvolo V.R., Berneman Z.N., Reitz M.S., Jr, Dewhurst S. 1995. Identification of human telomeric repeat motifs at the genome termini of human herpesvirus 7: structural analysis and heterogeneity. J. Virol. 69:8041–8045 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sfeir A., Kabir S., van Overbeek M., Celli G.B., de Lange T. 2010. Loss of Rap1 induces telomere recombination in the absence of NHEJ or a DNA damage signal. Science. 327:1657–1661 10.1126/science.1185100 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spatz S.J., Petherbridge L., Zhao Y., Nair V. 2007. Comparative full-length sequence analysis of oncogenic and vaccine (Rispens) strains of Marek’s disease virus. J. Gen. Virol. 88:1080–1096 10.1099/vir.0.82600-0 [DOI] [PubMed] [Google Scholar]
- Stevens J.G. 1989. Human herpesviruses: a consideration of the latent state. Microbiol. Rev. 53:318–332 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tischer B.K., von Einem J., Kaufer B., Osterrieder N. 2006. Two-step red-mediated recombination for versatile high-efficiency markerless DNA manipulation in Escherichia coli. Biotechniques. 40:191–197 [DOI] [PubMed] [Google Scholar]
- Trapp S., Parcells M.S., Kamil J.P., Schumacher D., Tischer B.K., Kumar P.M., Nair V.K., Osterrieder N. 2006. A virus-encoded telomerase RNA promotes malignant T cell lymphomagenesis. J. Exp. Med. 203:1307–1317 10.1084/jem.20052240 [DOI] [PMC free article] [PubMed] [Google Scholar]