Abstract
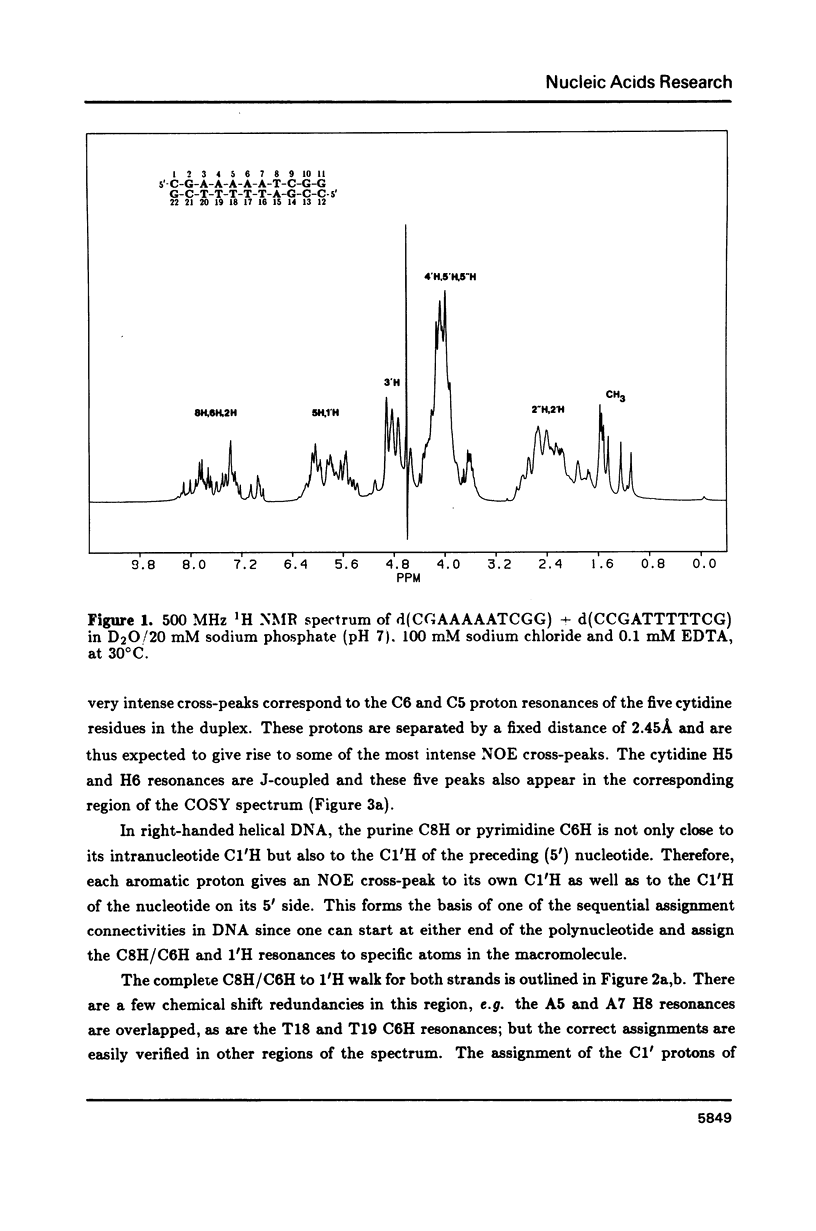
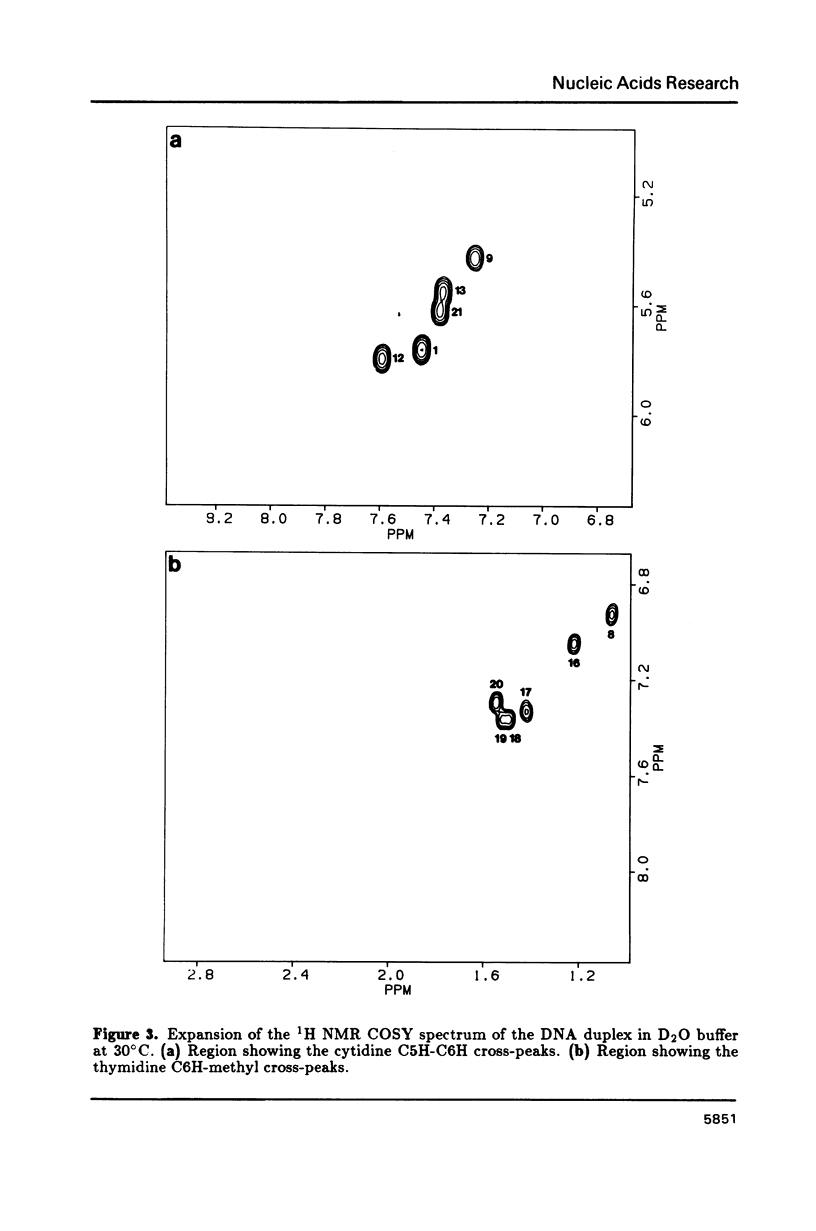
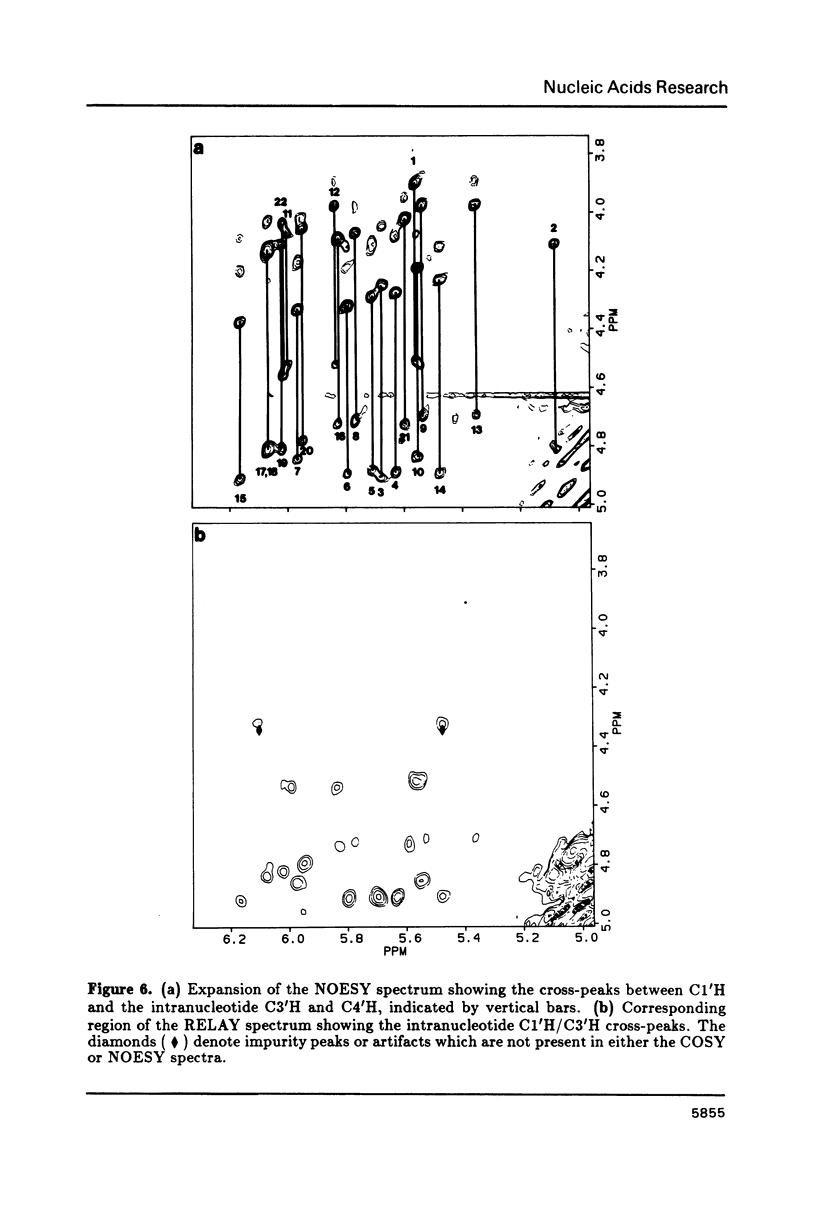
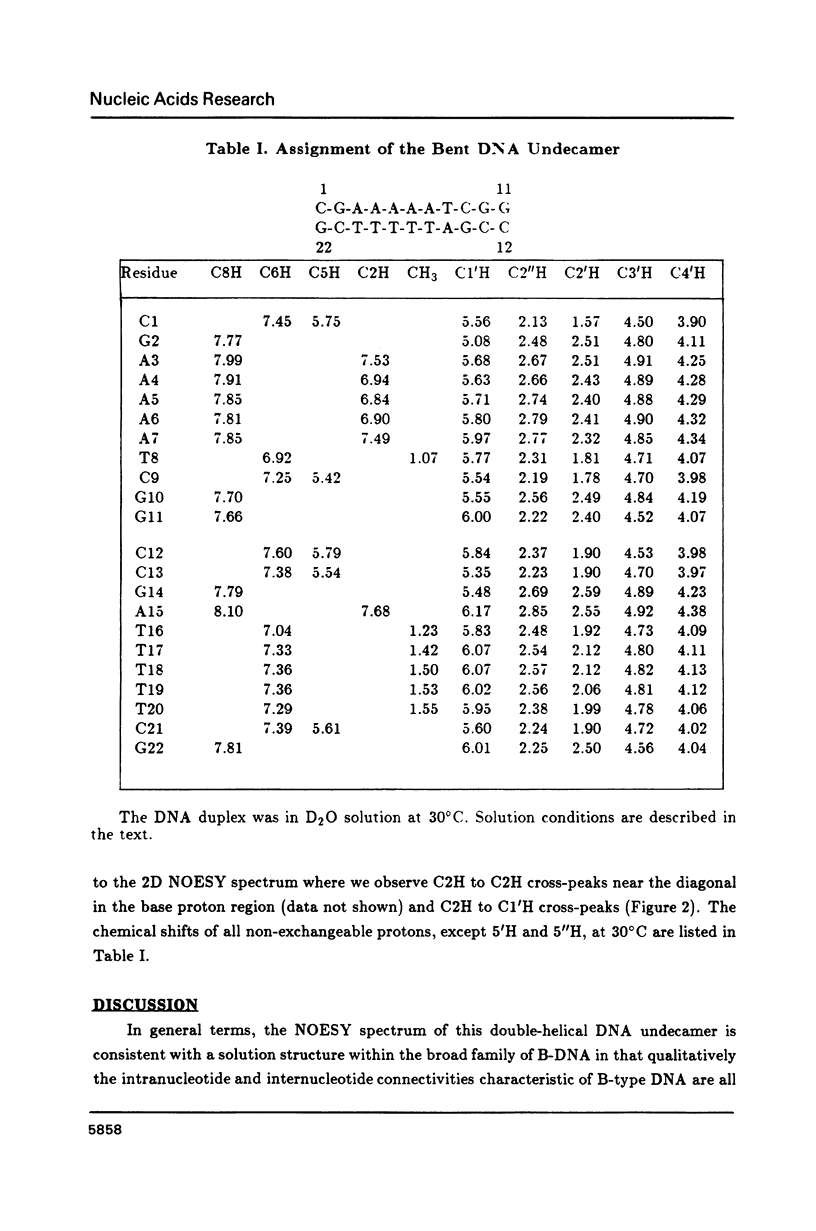
The resonances of all the non-exchangeable protons (except 5'H and 5"H) of d(CGAAAAATCGG) + d(CCGATTTTTCG), a putatively bent DNA duplex, have been assigned using 1H two-dimensional nuclear magnetic resonance methods. The nuclear Overhauser effect data indicate an overall B-form structure for this double-helical DNA undecamer. However, several features of the NMR data such as some unusually weak C8/C6 proton to C1' proton NOE cross-peaks, the presence of relatively intense C2H to C1'H NOE cross-peaks, and unusual chemical shifts of some 2", 2', and 1' protons suggest a substantial perturbation of the helix structure at the junctions and along the length of the tract of A residues. These structural deviations are considered in terms of models of DNA bending.
Full text
PDF













Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arnott S., Hukins D. W. Optimised parameters for A-DNA and B-DNA. Biochem Biophys Res Commun. 1972 Jun 28;47(6):1504–1509. doi: 10.1016/0006-291X(72)90243-4. [DOI] [PubMed] [Google Scholar]
- Behling R. W., Kearns D. R. 1H two-dimensional nuclear Overhauser effect and relaxation studies of poly(dA).poly(dT) Biochemistry. 1986 Jun 3;25(11):3335–3346. doi: 10.1021/bi00359a037. [DOI] [PubMed] [Google Scholar]
- Chou S. H., Hare D. R., Wemmer D. E., Reid B. R. Sequence-specific recognition of deoxyribonucleic acid. Chemical synthesis and nuclear magnetic resonance assignment of the imino protons of lambda OR3 operator deoxyribonucleic acid. Biochemistry. 1983 Jun 21;22(13):3037–3041. doi: 10.1021/bi00282a002. [DOI] [PubMed] [Google Scholar]
- Dickerson R. E., Drew H. R. Structure of a B-DNA dodecamer. II. Influence of base sequence on helix structure. J Mol Biol. 1981 Jul 15;149(4):761–786. doi: 10.1016/0022-2836(81)90357-0. [DOI] [PubMed] [Google Scholar]
- Feigon J., Leupin W., Denny W. A., Kearns D. R. Two-dimensional proton nuclear magnetic resonance investigation of the synthetic deoxyribonucleic acid decamer d(ATATCGATAT)2. Biochemistry. 1983 Dec 6;22(25):5943–5951. doi: 10.1021/bi00294a038. [DOI] [PubMed] [Google Scholar]
- Hagerman P. J. Evidence for the existence of stable curvature of DNA in solution. Proc Natl Acad Sci U S A. 1984 Aug;81(15):4632–4636. doi: 10.1073/pnas.81.15.4632. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hagerman P. J. Sequence dependence of the curvature of DNA: a test of the phasing hypothesis. Biochemistry. 1985 Dec 3;24(25):7033–7037. doi: 10.1021/bi00346a001. [DOI] [PubMed] [Google Scholar]
- Hare D. R., Reid B. R. Direct assignment of the dihydrouridine-helix imino proton resonances in transfer ribonucleic acid nuclear magnetic resonance spectra by means of the nuclear Overhauser effect. Biochemistry. 1982 Apr 13;21(8):1835–1842. doi: 10.1021/bi00537a020. [DOI] [PubMed] [Google Scholar]
- Hare D. R., Reid B. R. Three-dimensional structure of a DNA hairpin in solution: two-dimensional NMR studies and distance geometry calculations on d(CGCGTTTTCGCG). Biochemistry. 1986 Sep 9;25(18):5341–5350. doi: 10.1021/bi00366a053. [DOI] [PubMed] [Google Scholar]
- Hare D. R., Wemmer D. E., Chou S. H., Drobny G., Reid B. R. Assignment of the non-exchangeable proton resonances of d(C-G-C-G-A-A-T-T-C-G-C-G) using two-dimensional nuclear magnetic resonance methods. J Mol Biol. 1983 Dec 15;171(3):319–336. doi: 10.1016/0022-2836(83)90096-7. [DOI] [PubMed] [Google Scholar]
- Hare D., Shapiro L., Patel D. J. Extrahelical adenosine stacks into right-handed DNA: solution conformation of the d(C-G-C-A-G-A-G-C-T-C-G-C-G) duplex deduced from distance geometry analysis of nuclear Overhauser effect spectra. Biochemistry. 1986 Nov 18;25(23):7456–7464. doi: 10.1021/bi00371a030. [DOI] [PubMed] [Google Scholar]
- Hare D., Shapiro L., Patel D. J. Wobble dG X dT pairing in right-handed DNA: solution conformation of the d(C-G-T-G-A-A-T-T-C-G-C-G) duplex deduced from distance geometry analysis of nuclear Overhauser effect spectra. Biochemistry. 1986 Nov 18;25(23):7445–7456. doi: 10.1021/bi00371a029. [DOI] [PubMed] [Google Scholar]
- Havel T. F., Wüthrich K. An evaluation of the combined use of nuclear magnetic resonance and distance geometry for the determination of protein conformations in solution. J Mol Biol. 1985 Mar 20;182(2):281–294. doi: 10.1016/0022-2836(85)90346-8. [DOI] [PubMed] [Google Scholar]
- Koo H. S., Wu H. M., Crothers D. M. DNA bending at adenine . thymine tracts. Nature. 1986 Apr 10;320(6062):501–506. doi: 10.1038/320501a0. [DOI] [PubMed] [Google Scholar]
- Marini J. C., Levene S. D., Crothers D. M., Englund P. T. Bent helical structure in kinetoplast DNA. Proc Natl Acad Sci U S A. 1982 Dec;79(24):7664–7668. doi: 10.1073/pnas.79.24.7664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patel D. J., Kozlowski S. A., Marky L. A., Broka C., Rice J. A., Itakura K., Breslauer K. J. Premelting and melting transitions in the d(CGCGAATTCGCG) self-complementary duplex in solution. Biochemistry. 1982 Feb 2;21(3):428–436. doi: 10.1021/bi00532a002. [DOI] [PubMed] [Google Scholar]
- Patel D. J., Shapiro L., Kozlowski S. A., Gaffney B. L., Jones R. A. Covalent carcinogenic O6-methylguanosine lesions in DNA. Structural studies of the O6 meG X A and O6meG X G interactions in dodecanucleotide duplexes. J Mol Biol. 1986 Apr 20;188(4):677–692. doi: 10.1016/s0022-2836(86)80014-6. [DOI] [PubMed] [Google Scholar]
- Simpson L. Isolation of maxicircle component of kinetoplast DNA from hemoflagellate protozoa. Proc Natl Acad Sci U S A. 1979 Apr;76(4):1585–1588. doi: 10.1073/pnas.76.4.1585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss M. A., Patel D. J., Sauer R. T., Karplus M. 1H-NMR study of the lambda operator site OL1: assignment of the imino and adenine H2 resonances. Nucleic Acids Res. 1984 May 11;12(9):4035–4047. doi: 10.1093/nar/12.9.4035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss M. A., Patel D. J., Sauer R. T., Karplus M. Two-dimensional 1H NMR study of the lambda operator site OL1: a sequential assignment strategy and its application. Proc Natl Acad Sci U S A. 1984 Jan;81(1):130–134. doi: 10.1073/pnas.81.1.130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wemmer D. E., Chou S. H., Reid B. R. Sequence-specific recognition of DNA. Nuclear magnetic resonance assignments and structural comparison of wild-type and mutant lambda OR3 operator DNA. J Mol Biol. 1984 Nov 25;180(1):41–60. doi: 10.1016/0022-2836(84)90429-7. [DOI] [PubMed] [Google Scholar]
- Wu H. M., Crothers D. M. The locus of sequence-directed and protein-induced DNA bending. Nature. 1984 Apr 5;308(5959):509–513. doi: 10.1038/308509a0. [DOI] [PubMed] [Google Scholar]
- Zahn K., Blattner F. R. Sequence-induced DNA curvature at the bacteriophage lambda origin of replication. Nature. 1985 Oct 3;317(6036):451–453. doi: 10.1038/317451a0. [DOI] [PubMed] [Google Scholar]



