Abstract
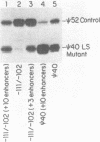
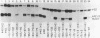
We have assayed a series of linker scanner mutants which cover the Xenopus laevis ribosomal gene promoter at approximately ten base pair intervals. All of these mutations adversely affect promoter activity with the exception of one mutation which stimulates activity. Thus, none are neutral. We show that most of the mutations can be partially rescued by ligating a block of enhancer elements upstream of the promoter. In addition, we have made extracts from liver nuclei which produce DNaseI protection footprints over the promoter. Analysis of both strands reveals a prominent footprinting domain from about -5 to -30. However, lesser changes in the digestion pattern are detected over most of the promoter. Previously published analyses have suggested that this promoter might be composed of three functional domains. The experiments presented here suggest that either 1) the three putative domains are so closely arranged that the boundaries are difficult to discern, or 2) the situation is more complex.
Full text
PDF












Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Dignam J. D., Lebovitz R. M., Roeder R. G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 1983 Mar 11;11(5):1475–1489. doi: 10.1093/nar/11.5.1475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunaway M., Reeder R. H. DNase I footprinting shows three protected regions in the promoter of the rRNA genes of Xenopus laevis. Mol Cell Biol. 1985 Feb;5(2):313–319. doi: 10.1128/mcb.5.2.313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorski K., Carneiro M., Schibler U. Tissue-specific in vitro transcription from the mouse albumin promoter. Cell. 1986 Dec 5;47(5):767–776. doi: 10.1016/0092-8674(86)90519-2. [DOI] [PubMed] [Google Scholar]
- Grummt I. Nucleotide sequence requirements for specific initiation of transcription by RNA polymerase I. Proc Natl Acad Sci U S A. 1982 Nov;79(22):6908–6911. doi: 10.1073/pnas.79.22.6908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haltiner M. M., Smale S. T., Tjian R. Two distinct promoter elements in the human rRNA gene identified by linker scanning mutagenesis. Mol Cell Biol. 1986 Jan;6(1):227–235. doi: 10.1128/mcb.6.1.227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henikoff S. Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene. 1984 Jun;28(3):351–359. doi: 10.1016/0378-1119(84)90153-7. [DOI] [PubMed] [Google Scholar]
- Iida C. T., Kownin P., Paule M. R. Ribosomal RNA transcription: proteins and DNA sequences involved in preinitiation complex formation. Proc Natl Acad Sci U S A. 1985 Mar;82(6):1668–1672. doi: 10.1073/pnas.82.6.1668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohorn B. D., Rae P. M. Localization of DNA sequences promoting RNA polymerase I activity in Drosophila. Proc Natl Acad Sci U S A. 1983 Jun;80(11):3265–3268. doi: 10.1073/pnas.80.11.3265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Labhart P., Reeder R. H. Characterization of three sites of RNA 3' end formation in the Xenopus ribosomal gene spacer. Cell. 1986 May 9;45(3):431–443. doi: 10.1016/0092-8674(86)90329-6. [DOI] [PubMed] [Google Scholar]
- Labhart P., Reeder R. H. Enhancer-like properties of the 60/81 bp elements in the ribosomal gene spacer of Xenopus laevis. Cell. 1984 May;37(1):285–289. doi: 10.1016/0092-8674(84)90324-6. [DOI] [PubMed] [Google Scholar]
- Learned R. M., Smale S. T., Haltiner M. M., Tjian R. Regulation of human ribosomal RNA transcription. Proc Natl Acad Sci U S A. 1983 Jun;80(12):3558–3562. doi: 10.1073/pnas.80.12.3558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKnight S. L., Kingsbury R. Transcriptional control signals of a eukaryotic protein-coding gene. Science. 1982 Jul 23;217(4557):316–324. doi: 10.1126/science.6283634. [DOI] [PubMed] [Google Scholar]
- McStay B., Reeder R. H. A termination site for Xenopus RNA polymerase I also acts as an element of an adjacent promoter. Cell. 1986 Dec 26;47(6):913–920. doi: 10.1016/0092-8674(86)90806-8. [DOI] [PubMed] [Google Scholar]
- Miller K. G., Tower J., Sollner-Webb B. A complex control region of the mouse rRNA gene directs accurate initiation by RNA polymerase I. Mol Cell Biol. 1985 Mar;5(3):554–562. doi: 10.1128/mcb.5.3.554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moss T. Transcription of cloned Xenopus laevis ribosomal DNA microinjected into Xenopus oocytes, and the identification of an RNA polymerase I promoter. Cell. 1982 Oct;30(3):835–842. doi: 10.1016/0092-8674(82)90288-4. [DOI] [PubMed] [Google Scholar]
- Norrander J., Kempe T., Messing J. Construction of improved M13 vectors using oligodeoxynucleotide-directed mutagenesis. Gene. 1983 Dec;26(1):101–106. doi: 10.1016/0378-1119(83)90040-9. [DOI] [PubMed] [Google Scholar]
- Ohlsson H., Edlund T. Sequence-specific interactions of nuclear factors with the insulin gene enhancer. Cell. 1986 Apr 11;45(1):35–44. doi: 10.1016/0092-8674(86)90535-0. [DOI] [PubMed] [Google Scholar]
- Smale S. T., Tjian R. Transcription of herpes simplex virus tk sequences under the control of wild-type and mutant human RNA polymerase I promoters. Mol Cell Biol. 1985 Feb;5(2):352–362. doi: 10.1128/mcb.5.2.352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sollner-Webb B., Wilkinson J. A., Roan J., Reeder R. H. Nested control regions promote Xenopus ribosomal RNA synthesis by RNA polymerase I. Cell. 1983 Nov;35(1):199–206. doi: 10.1016/0092-8674(83)90222-2. [DOI] [PubMed] [Google Scholar]
- Windle J. J., Sollner-Webb B. Two distant and precisely positioned domains promote transcription of Xenopus laevis rRNA genes: analysis with linker-scanning mutants. Mol Cell Biol. 1986 Dec;6(12):4585–4593. doi: 10.1128/mcb.6.12.4585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Windle J., Sollner-Webb B. Upstream domains of the Xenopus laevis rDNA promoter are revealed in microinjected oocytes. Mol Cell Biol. 1986 Apr;6(4):1228–1234. doi: 10.1128/mcb.6.4.1228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamoto O., Takakusa N., Mishima Y., Kominami R., Muramatsu M. Determination of the promoter region of mouse ribosomal RNA gene by an in vitro transcription system. Proc Natl Acad Sci U S A. 1984 Jan;81(2):299–303. doi: 10.1073/pnas.81.2.299. [DOI] [PMC free article] [PubMed] [Google Scholar]