Abstract
Cronobacter spp. (formerly Enterobacter sakazakii) and Salmonella spp. are increasingly implicated internationally as important microbiological contaminants in low-moisture food products, including powdered infant formula. Estimates indicate that 40 to 80% of infants infected with Cronobacter sakazakii and/or Salmonella in the United States may not survive the illness. A systematic approach, combining literature-based data mining, comparative genome analysis, and the direct sequencing of PCR products of specific biomarker genes, was used to construct an initial collection of genes to be targeted. These targeted genes, particularly genes encoding virulence factors and genes responsible for unique phenotypes, have the potential to function as biomarker genes for the identification and differentiation of Cronobacter spp. and Salmonella from other food-borne pathogens in low-moisture food products. In this paper, a total of 58 unique Salmonella gene clusters and 126 unique potential Cronobacter biomarkers and putative virulence factors were identified. A chitinase gene, a well-studied virulence factor in fungi, plants, and bacteria, was used to confirm this approach. We found that the chitinase gene has very low sequence variability and/or polymorphism among Cronobacter, Citrobacter, and Salmonella, while differing significantly in other food-borne pathogens, either by sequence blasting or experimental testing, including PCR amplification and direct sequencing. This computational analysis for Cronobacter and Salmonella biomarker identification and the preliminary laboratory studies are only a starting point; thus, PCR and array-based biomarker verification studies of these and other food-borne pathogens are currently being conducted.
Cronobacter spp. and Salmonella spp. are recognized as food-borne pathogens that cause serious human illness, and in infants, these pathogens are considered to be of great health concern (3, 6). In addition to Salmonella, Cronobacter spp. have been isolated not only from low-moisture food products such as powdered infant milk but also from fresh lettuce, frozen shellfish, ready-to-eat meat, and fermented and cooked food products (5, 17). In 2008, Enterobacter sakazakii was reclassified into the new genus Cronobacter (23, 24). Within the genus Cronobacter, there are five species: C. sakazakii, C. malonaticus, C. turicensis, C. muytjensii, and C. dublinensis. Further, three subspecies currently exist within the species C. dublinensis, including dublinensis, lausannensis, and lactaridi. A wide variety of other bacteria and pathogens, including Pantoea agglomerans, Enterobacter cloacae, Staphylococcus aureus, Hafnia alvei, Citrobacter, Klebsiella pneumoniae, Klebsiella oxytoca, Escherichia vulneris (2, 4), and Listeria monocytogenes (42), have also been found in powdered infant formula (PIF) and other low-moisture products (1-4, 7-9, 13, 26), as well as in foods of animal origin (28, 32).
In recent years, the use of molecular methods, such as multiplex PCR (27), real-time PCR (21, 35), DNA microarrays (7, 34), automated ribotyping (33), amplified fragment length polymorphisms (AFLP) (18), full-length 16S rRNA gene sequencing (11, 12), and immunoassays (20, 44), for the detection and identification of the aforementioned pathogens has been intensively researched. The majority of these methods are heavily dependent on species-specific biomarker genes. One such gene is gluA (α-1,4-glucosidase), which was identified as a biomarker gene in Cronobacter spp. and was not found in any other Enterobacter spp. (22, 39). Additionally, DNase (14), arginine dihydrolase (22), 16S-23S rRNA genes, internal transcribed spacer (ITS) regions (34), outer membrane protein A (ompA) (37, 45), ornithine decarboxylase (14), recN, thdF, and rpoA have been utilized as species-specific biomarkers via multilocus sequence analysis (30).
During the past 2 decades, over 1,000 microbial genomes, including 1 genome from Cronobacter spp. (29) and 31 genomes from Salmonella strains, have been sequenced completely, and over 1,500 microbial genome sequences are in the process of being completed (http://www.ncbi.nlm.nih.gov/genomes/lproks.cgi?view=1). Genome sequence data have shown that bacterial DNA is highly dynamic, and the process of bacterial genome evolution demonstrates substantial differences, even within strains of the same genus. The size of the chromosome may also vary among strains from clinical isolates of Cronobacter (36), Salmonella, and other pathogens. Therefore, finding clinically useful biomarkers that can be used to specifically distinguish Cronobacter spp. and Salmonella spp. from the other food-borne pathogens in mixed bacterial populations is very challenging. A major obstacle for the development of genetic-based detection methods for specific pathogens is the identification of suitable target sequences. Additionally, many methods for detection and isolation of these pathogens in foods are labor intensive and time-consuming. Therefore, the aim of this study was to systematically collect and verify potential genetic biomarker genes through literature-based data mining, comparative genomic comparisons, and verification of specific selected biomarkers to enable the detection and differentiation of Cronobacter and Salmonella spp. from other food-borne pathogens in a direct, single-step, rapid PCR-based method for potential application to food samples and clinical specimens.
MATERIALS AND METHODS
Bacterial strains, DNA isolation, and target gene amplification.
Seventeen Cronobacter strains were obtained from Larry Beuchat (University of Georgia, Center for Food Safety, Griffin, GA) and Dong-Hyun Kang (Washington State University, School of Food Science, Pullman, WA). All of these strains are potentially pathogenic and were isolated from a wide range of food samples and environmental and clinical sources. Strains were grown overnight in Luria-Bertani broth at 37°C, and genomic DNA was isolated using the Qiagen DNeasy kit following the manufacturer's recommendations. The degenerate primer set for chitinase and a nondegenerate primer set for the ompA gene, listed in Table 1, were novelly designed so as to amplify unique bands under standard PCR amplification conditions. PCR amplification was performed in a total volume of 50 μl, containing 5 μl of 10× reaction buffer, 1 μl of deoxynucleoside triphosphates (dNTPs), 5 μl of each of the primers (10 μM), 1 μl of template DNA (50 to 100 ng/μl), 0.25 μl of Taq DNA polymerase (5 U/μl), and 32.75 μl of PCR water to make up the final volume. The amplification was performed using an iCycler thermocycler (Bio-Rad, Hercules, CA). The PCR conditions used were 95°C for 2 min, followed by 30 cycles of 95°C for 10 s, 55°C for 30 s, 72°C for 2 min, and a final extension for 10 min at 72°C. A portion of these amplified PCR products were verified by agarose gel electrophoresis (Fig. 1).
TABLE 1.
Primers used in this study
| Primer name | Expected size | Primer sequencea | Gene |
|---|---|---|---|
| sak_Chi_F | 2.2 kb | ATGGCTACMAGYAAAYTRATYCAGGG | Chitinase |
| sak_chi_R | CACCTGRTAGTTRTGVCCTTTCCAGC | ||
| ompA_F | 469 bp | GGATTTAACCGTGAACTTTTCC | ompA |
| ompA_R | CGCCAGCGATGTTAGAAGA |
M, A or C; Y, C or T; R, A or G; V, A or C or G.
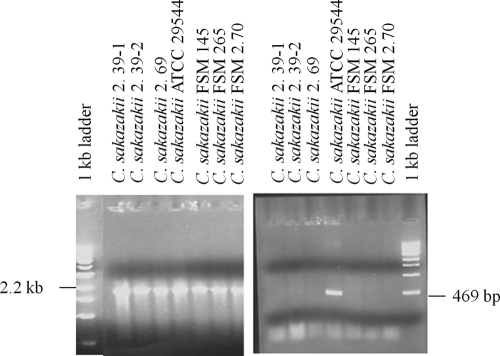
FIG. 1.
Agarose gel electrophoresis analysis of PCR products targeting the chitinase (2.2-kb) and ompA (469-bp) genes.
DNA sequencing and phylogenetic analysis.
In order to explore the evolutionary relationship obtained from an analysis of DNA polymorphisms of the chitinase gene among the tested strains of Cronobacter spp., Salmonella spp., and the other food-borne pathogens, the PCR products of the chitinase gene from various E. sakazakii strains were purified using QIAquick PCR purification columns and sequenced using the BigDye Terminator v3.1 cycle sequencing kit and a 3730 DNA analyzer (Applied Biosystems, Foster City, CA). Sequencher, version 4.9 (Gene Codes, Ann Arbor, MI), was used to trim, combine, and assemble the sequence data to form contiguous stretches. Phylogenetic trees were generated by comparing the nucleotide sequences using ClustalW (43), Phylip (http://evolution.genetics.washington.edu/phylip/general.html), and tree viewer software.
Extraction of unique genes and consensus genes of each genome.
Representative strains (110 in total) of the 15 pathogens listed in Table 2, which are most commonly reported as the cause of food-borne illnesses from low-moisture foods such as dry milk, fruits, peanut butter, cheeses, and chocolate, were analyzed. The complete gene sequence of each bacterial genome was subjected to the BLAST search engine (using default parameters) and compared against each of the sequences of the remaining 14 genomes. If the sequence length (i.e., the total number of bases) of gene A was l and the total number of sequence identities (i.e., the number of identical bases) of gene A with gene B was n, then the identity ratio (similarity) between gene A and gene B was defined as n/l. The identity ratio measures the percentage of similarity between a target gene sequence and a query gene sequence. If the identity ratio between gene A in the query genome and a gene in 1 of 14 subject genomes was greater than or equal to a user-defined threshold, t, then gene A was said to have a hit in the subject genomes. For each query genome, genes that did not have any hits in the other 14 genomes were collected, and these genes were designated unique genes in the query genome. Various t values from 0.5 to 0.05, with a step size of 0.05, were tested, and results were manually analyzed. Manual verification of these computationally generated data showed that a t value between 0.05 and 0.1 would provide the best reasonable data representation. A threshold t value of 0.05 was chosen because it generated a reasonable amount of unique genes. The number of unique genes is shown in Table 2, third column. If gene A in the query genome had hits (various t values from 0.1 to 0.5 were tested, and a t value of 0.1 was chosen) in each of the remaining 14 genomes, then gene A was called a consensus gene. The number of consensus genes in each genome is shown in Table 2, fourth column. To find the unique genes that existed only in Cronobacter spp. and/or Salmonella spp. but not in the other 13 bacterial species, we performed BLAST search analyses of these two genomes against the other 13 genomes. The same threshold (t = 0.05) was used for the genome comparisons, and a subset of the genes unique to Cronobacter spp. and Salmonella spp. is listed in Table 3.
TABLE 2.
Comparative genetic characterization of major pathogens from food-borne illnesses associated with low-moisture food productsa
| Species/strain (GenBank accession no.) | Total no. of: |
GC content (%) | Length (bp) | ||
|---|---|---|---|---|---|
| Genes | Unique genesb | Consensus genesc | |||
| Cronobacter sakazakii(CP000783) | 4,392 | 268 | 105 | 56 | 4,368,373 |
| Salmonella Enteritidis (CP001127) | 4,707 | 401 | 231 | 52 | 4,809,037 |
| Shigella boydii(CP000036) | 4,463 | 123 | 20 | 51 | 4,519,823 |
| Enterobacter sp. strain 638 (CP000653) | 4,230 | 232 | 15 | 52 | 4,518,712 |
| Citrobacter koseri(CP000822) | 5,123 | 243 | 207 | 53 | 4,720,462 |
| Escherichia coli O157:H7 (AE005174) | 5,371 | 366 | 121 | 50 | 5,498,450 |
| Enterobacter cloacae(CP001918) | 5,241 | 391 | 134 | 54 | 5,314,581 |
| Pantoea ananatis(CP001875) | 4,341 | 551 | 21 | 53 | 4,690,298 |
| Klebsiella pneumoniae(CP000964) | 5,567 | 651 | 224 | 57 | 5,641,239 |
| Yersinia pestis (CP000901)d | 4,224 | 696 | 231 | 47 | 4,504,254 |
| Campylobacter jejuni(AL111168) | 1,699 | 804 | 19 | 30 | 1,641,481 |
| Staphylococcus aureus(CP001844) | 2,664 | 902 | 43 | 32 | 2,814,816 |
| Listeria monocytogenes(AL591824) | 2,940 | 1,034 | 5 | 37 | 2,944,528 |
| Clostridium difficile(AM180355) | 3,970 | 1,747 | 29 | 29 | 4,290,252 |
| Bacillus cereus 03BB102 (CP001407) | 5,566 | 2,033 | 201 | 35 | 5,269,628 |
See references 1-4, 7, 9, 13, and 26. Data from the second, fifth, and sixth columns were obtained from NCBI's Entrez database (http://www.ncbi.nlm.nih.gov/genomes/lproks.cgi).
t = 0.05.
t = 0.1.
For more information, visit http://www.iit.edu/ncfst/resources/pdfs/2010poster_iafp_neupane_yerseniabovinemilk.pdf.
TABLE 3.
Select unique genes/regions of Cronobacter and Salmonella spp. based on comparative computational analysis
| Species | GenBank accession no. | Positions | Gene function (unique gene locus tags) |
|---|---|---|---|
| Salmonella spp. | CP001125.1a | 20447-23029 | EstP, putative pesticide-degrading enzyme; esterase |
| CP001125.1a | 85623-83905 | Histidine kinase | |
| CP001125.1a | 88002-87088 | Response regulator receiver protein | |
| CP001125.1a | 92586-93302 | TriD protein | |
| CP001125.1a | 93314-93598 | Putative entry exclusion protein | |
| CP001125.1a | 93616-94623 | TriE protein | |
| CP001125.1a | 82199-82840 | Chloramphenicol acetyltransferase 2 | |
| NC_014476a | 42152-48198 | spvDCBAR gene cluster, virulence gene | |
| CP001127.1 | 17869-19968 | Exochitinase | |
| CP001127.1 | 25762-28428 | Outer membrane usher protein FimD | |
| CP001127.1 | 35340-37058 | Arylsulfotransferase (asst) superfamily protein | |
| CP001127.1 | 213571-212303 | Putative fimbrial-like adhesin protein | |
| CP001127.1 | 249550-246581 | Viral enhancin protein | |
| CP001127.1 | 250369-251670 | Shikimate transporter | |
| CP001127.1 | 343747-344484 | Gram-negative pili assembly chaperone | |
| CP001127.1 | 344508-347018 | Outer membrane fimbrial usher protein | |
| CP001127.1 | 347040-347510 | Putative fimbrial structural subunit | |
| CP001127.1 | 392509-391751 | Fimbrial chaperone protein | |
| CP001127.1 | 397279-396350 | Fimbrial chaperone protein | |
| CP001127.1 | 406659-405202 | Outer membrane protein OprM | |
| CP001127.1 | 566030-565194 | Probable secreted protein | |
| CP001127.1 | 664550-665164 | Lytic enzyme | |
| CP001127.1 | 699362-697905 | O-antigen conversion protein | |
| CP001127.1 | 785621-786787 | Hydrolase, UxaA family | |
| CP001127.1 | 853136-853906 | O-antigen export system, permease protein | |
| CP001127.1 | 986043-985153 | Transcriptional regulator, LysR family | |
| CP001127.1 | 1179341-1180033 | Oligogalacturonate-specific porin | |
| CP001127.1 | 1242120-1241122 | Putative fimbrial protein | |
| CP001127.1 | 1292378-1293106 | Pertussis toxin, subunit 1 subfamily | |
| CP001127.1 | 1295248-1294439 | Cytolethal distending toxin B | |
| CP001127.1 | 1433385-1434401 | Tetrathionate reductase gene cluster | |
| CP001127.1 | 1435347-1437095 | Sensor kinase | |
| CP001127.1 | 1444369-1445862 | Type III secretion outer membrane pore, YscC/HrcC family | |
| CP001127.1 | 1445855-1447054 | Type III secretion apparatus protein, YscD/HrpQ family | |
| CP001127.1 | 1679794-1678562 | l-Lactate oxidase | |
| CP001127.1 | 1692014-1693240 | Secreted effector protein | |
| CP001127.1 | 2187749-2186574 | Wzy | |
| CP001127.1 | 2191447-2190167 | Putative O-antigen transporter | |
| CP001127.1 | 2264284-2263262 | Putative fimbrial protein | |
| CP001127.1 | 2828458-2829702 | Enterochelin esterase | |
| CP001127.1 | 2933896-2931884 | Cell invasion protein SipA | |
| CP001127.1 | 2934982-2933960 | Type III effector protein IpaD/SipD/SspD | |
| CP001127.1 | 2938091-2936310 | Cell invasion protein SipB | |
| CP001127.1 | 2943452-2942442 | Antigen presentation protein SpaN | |
| CP001127.1 | 2945168-2943873 | Flagellum-specific ATP synthase | |
| CP001127.1 | 2945572-2945165 | Surface presentation of antigens protein Spak | |
| CP001127.1 | 2948796-2947678 | Invasion protein InvE | |
| CP001127.1 | 2951227-2950478 | Invasion protein | |
| CP001127.1 | 2999371-2998574 | Beta-lactamase domain protein | |
| CP001127.1 | 3095222-3092733 | Fimbrial usher protein | |
| CP001127.1 | 3207119-3206064 | Putative methyl-accepting chemotaxis protein | |
| CP001127.1 | 3783645-3784859 | O-antigen ligase | |
| CP001127.1 | 4335277-4336125 | ClpP protease | |
| CP001127.1 | 4341875-4343800 | Tail protein | |
| CP001127.1 | 4455678-4458116 | CshB porin | |
| CP001127.1 | 4459680-4460462 | CshE pilin | |
| CP001127.1 | 4700538-4699453 | Putative major fimbrial subunit | |
| CP001127.1 | 4703584-4701155 | Outer membrane usher protein SfmD | |
| Cronobacter spp. | CP000783.1 | 268708-272424 | Hypothetical protein (ESA_00298-ESA_00300) |
| CP000783.1 | 273598-287078 | Hypothetical protein (ESA_00304-ESA_00310, except alginate O-acetyltransferase AlgI [ESA_00303], putative lipoprotein [ESA_00305], and the alpha-2-macroglobulin family region [ESA_00308]) | |
| CP000783.1 | 578742-591835 | Hypothetical protein (ESA_00611, ESA_00612, ESA_00615, ESA_00616, and ESA_00618) | |
| CP000783.1 | 957212-969835 | Hypothetical protein (ESA_00981-ESA_00990, except putative invasin [ESA_00987] and phage integrase [ESA_00990]) | |
| CP000783.1 | 991095-994496 | Hypothetical protein (ESA_01026), peptidase S14 ClpP (ESA_01027), phage major capsid protein, HK97 family (ESA_01028) | |
| CP000783.1 | 1152718-1158275 | O-antigen cluster (ESA_01181-ESA_01185) | |
| CP000783.1 | 1186371-1190015 | Hypothetical protein (ESA_01216-ESA_01218) | |
| CP000783.1 | 1391367-1394671 | Putative fatty acid hydroxylase (ESA_01448), putative fatty acid desaturase (ESA_01449), putative membrane protein (ESA_01450) | |
| CP000783.1 | 2137914-2140588 | Hypothetical protein (ESA_02201), putative esterase/lipase/thioesterase (ESA_02202), transcriptional regulator, LysR family (ESA_02203) | |
| CP000783.1 | 2256328-2258819 | Putative caudovirus prohead protease (ESA_02319), putative phage portal protein, lambda family (ESA_02320), putative phage terminase large subunit (ESA_02321) | |
| CP000783.1 | 3289362-3297665 | Capsular polysaccharide biosynthesis gene cluster (ESA_03352-ESA_03357) | |
| CP000783.1 | 3762752-3765513 | Fimbrial gene cluster (ESA_03812-ESA_03814) | |
| CP000783.1 | 3861832-3864117 | Hypothetical protein (ESA_03913-ESA_03916) | |
| CP000783.1 | 4039090-4042971 | Hypothetical protein (ESA_04084-ESA_04086) | |
| CP000783.1 | 4348939-4352405 | Putative phosphoribosylpyrophosphate synthetase (ESA_04383), putative nicotinamide phosphoribosyl transferase (ESA_04384), hypothetical protein (ESA_04385), putative tellurite resistance protein (ESA_04386) |
GenBank accession number for a plasmid-borne gene.
Literature-based data mining and comparative analysis of unique gene clusters, putative virulence factors, and biomarker genes.
Every unique gene cluster, putative virulence factor, and biomarker gene that we identified, shown in Tables 4 and 5, were based on scientific publications or protein sequence similarity searches (BLASTp) against the GenBank nonredundant protein database or were found by relying on keyword-based analysis of text-mined data from publically available databases such as NCBI (http://www.ncbi.nlm.nih.gov/gene) and ENA (http://www.ebi.ac.uk/ena/). The stand-alone BLAST program was downloaded from NCBI (http://blast.ncbi.nlm.nih.gov/Blast.cgi?CMD=Web&PAGE_TYPE=BlastDocs&DOC_TYPE=Download).
TABLE 4.
Identification of potential biomarker genes that might be used to distinguish Cronobacter and Salmonella spp. from other food-borne pathogens based on literature-based data mining
| C. sakazakiilocus tag | C. sakazakiiprotein accession no. (GenBank) | Gene/protein name (symbol) | Presence of homology |
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Salmonella | Citrobacter | E. coli | Klebsiella | Clostridium | Pantoea | Yersinia | Shigella | E. cloacae | Campylobacter/Listeria | |||
| ESA_01183 | YP_001437286.1 | Wzx | Yes | No | Yes | No | No | No | No | No | No | No |
| ESA_01185 | YP_001437288.1 | Wzy | No | No | No | No | No | No | No | No | No | No |
| ESA_03317 | YP_001439374.1 | Chitinase | Yes | Yes | No | No | No | No | No | No | No | No |
| ESA_02201 | YP_001438286.1 | Hypothetical protein | No | No | No | No | No | No | No | No | No | No |
| ESA_02709 | YP_001438777.1 | α-1,4-Glucosidase gene (gluA) | No | No | No | No | Yes | No | No | No | No | No |
| ESA_02516 | YP_001438597.1 | Putative hemolysin/hemagglutinin | No | No | No | No | No | No | No | No | No | No |
| ESA_02084 | YP_001438170.1 | Putative adhesin | No | No | No | No | No | No | No | No | No | No |
| ESA_00341 | YP_001436476.1 | Beta-carotene hydroxylase pigment (crtZ) | No | No | No | No | No | Yes | No | No | No | No |
| ESA_00341 | YP_001436477.1 | Phytoene/squalene synthetase (crtB) | No | No | No | No | No | Yes | No | No | No | No |
| ESA_00343 | YP_001436478.1 | Phytoene dehydrogenase (crtI) | No | No | No | No | No | Yes | No | No | No | No |
| ESA_00344 | YP_001436479.1 | Lycopene cyclase (crtL) | No | No | No | No | No | Yes | No | No | No | No |
| ESA_00345 | YP_001436480.1 | Glycosyl transferases (crtX) | No | No | No | No | No | Yes | No | No | No | No |
| ESA_00346 | YP_001436481.1 | Isopentenyl pyrophosphate isomerase | No | No | No | No | No | Yes | No | No | No | No |
| ESA_00347 | YP_001436482.1 | Geranylgeranyl pyrophosphate synthase (crtE) | No | No | No | No | No | Yes | No | No | No | No |
| ESA_03721 | YP_001439754.1 | DNase (TatD) | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_02154 | YP_001438239.1 | Succinylarginine dihydrolase | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_02391 | YP_001438473.1 | Outer membrane protein A (ompA) | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_00314 | YP_001436449.1 | Ornithine decarboxylase | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_01872 | YP_001437962.1 | Catalase | Yes | Yes | Yes | Yes | No | No | No | Yes | No | No |
| ESA_01203 | YP_001437307.1 | 4-Aminobutyrate aminotransferase | No | Yes | Yes | Yes | No | No | No | Yes | No | No |
| ESA_01574 | YP_001437664.1 | Aconitate hydratase | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_04127 | YP_001440144.1 | 6-Phosphofructokinase | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_01954 | YP_001438044.1 | Fumarate/nitrate reduction transcriptional regulator | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_04154 | YP_001440171.1 | Alpha-xylosidase (YicI) | Yes | Yes | Yes | Yes | Yes | No | No | Yes | Yes | No |
| ESA_00357 | YP_001436490.1 | DNA primase (dnaG) | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_03853 | YP_001439875.1 | Galactoside permease | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_04206 | YP_001440222.1 | Endo-1,4-d-glucanase (BscZ) | Yes | Yes | Yes | Yes | No | No | Yes | Yes | Yes | No |
| ESA_00523 | YP_001436650.1 | Phosphopyruvate hydratase | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_03753 | YP_001439786.1 | Porphobilinogen deaminase | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_00373 | YP_001436507.1 | Outer membrane channel protein | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_02807 | YP_001438873.1 | Acriflavin resistance protein A precursor (AcrA) | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_03723 | YP_001439756.1 | sec-independent translocase (TatB) | Yes | Yes | Yes | Yes | No | No | Yes | Yes | Yes | No |
| ESA_02187 | YP_001438272.1 | Virulence protein (VirK) | Yes | Yes | Yes | Yes | No | No | Yes | Yes | No | No |
| ESA_02251 | YP_001438336.1 | Acyl carrier protein | Yes | No | Yes | No | No | Yes | Yes | No | No | No |
| ESA_00752 | YP_001436865.1 | Extracellular metalloprotease Prt1 | Yes | Yes | No | No | No | Yes | No | No | No | No |
| ESA_00690 | YP_001436805.1 | GTP-binding protein (lepA) | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_04401 | YP_001440417.1 | Elongation factor G | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_03312 | YP_001439369.1 | Isoleucyl-tRNA synthetase (ileS) | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_03973 | YP_001439995.1 | DNA gyrase subunit B (gyrB) | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_03690 | YP_001439730.1 | DNA-directed RNA polymerase subunit beta (rpoB) | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| GU122171 | GU122171 | 16S rRNA | No | No | No | No | No | No | No | No | No | No |
| ACE74909 | ACE74909 | ATPase (RecN) | Yes | Yes | Yes | Yes | No | No | No | Yes | Yes | No |
| ESA_00031 | YP_001436176 | DNA-directed RNA polymerase subunit alpha (rpoA) | Yes | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | No |
| ESA_03979 | YP_001440001.1 | tRNA modification GTPase TrmE (thdF) | Yes | Yes | Yes | Yes | No | No | No | Yes | Yes | No |
TABLE 5.
Putative virulence factors and unique biomarker genes for distinguishing Cronobacter, Salmonella, and E. coli from other food-borne pathogens based on literature-based data mining combined with comparative genome analysis
| GI no. | GenBank accession no. | Locus tag | Gene/gene function | Present in pathogen: |
||
|---|---|---|---|---|---|---|
| Salmonella | E. coli | Other major pathogensa | ||||
| 156530628 | ABU75454.1 | ESA_00150 | Putative anaerobic decarboxylate transporter | + | + | ± |
| 156530827 | ABU75653.1 | ESA_00355 | rpoS/sigma S (sigma 38) factor of RNA polymerase, major sigma factor during stationary phase | + | + | ± |
| 156530851 | ABU75677.1 | ESA_00379 | katB/catalase-peroxidase KatB | + | + | ± |
| 156530862 | ABU75688.1 | ESA_00390 | pilT/twitching motility protein PilT | − | − | − |
| 156531119 | ABU75945.1 | ESA_00662 | clpE/ATP-dependent protease | + | + | ± |
| 156531139 | ABU75965.1 | ESA_00686 | algU/alginate biosynthesis protein | + | + | ± |
| 156531159 | ABU75985.1 | ESA_00708 | pilR/two-component response regulator | + | + | ± |
| 156531248 | ABU76074.1 | ESA_00797 | Enterobactin synthetase component E | − | − | − |
| 156531242 | ABU76068.1 | ESA_00791 | Putative iron-siderophore transport system, ATP-binding component | + | + | ± |
| 156531430 | ABU76256.1 | ESA_00987 | Putative eae/intimin | − | − | − |
| 156531432 | ABU76258.1 | ESA_00989 | Similar to plasmid virulence: regulation of spv operon, lysR family | − | − | − |
| 156531610 | ABU76436.1 | ESA_01169 | Mannose-1-phosphate guanylyltransferase 1 | + | + | ± |
| 156531686 | ABU76512.1 | ESA_01250 | bscR/putative type III secretion protein | + | + | ± |
| 156531689 | ABU76515.1 | ESA_01253 | fliM/flagellar motor switch protein | + | + | ± |
| 156531693 | ABU76519.1 | ESA_01257 | bscN/putative ATP synthase in type III secretion system | + | + | ± |
| 156531695 | ABU76521.1 | ESA_01259 | fliG/flagellar motor switch protein | + | + | ± |
| 156531696 | ABU76522.1 | ESA_01260 | fliF/flagellar M-ring protein | − | − | − |
| 156531723 | ABU76549.1 | ESA_01287 | fliD/putative flagellar hook-associated protein | − | − | − |
| 156531724 | ABU76550.1 | ESA_01288 | flaA/flagellin | − | − | ± |
| 156531745 | ABU76571.1 | ESA_01309 | bvgA/virulence factors transcription regulator | + | + | ± |
| 156531788 | ABU76614.1 | ESA_01354 | flhB/flagellar biosynthetic protein | − | − | − |
| 156531789 | ABU76615.1 | ESA_01355 | pcrD/type III secretory apparatus protein | + | + | ± |
| 156531820 | ABU76646.1 | ESA_01386 | Lipid A biosynthesis (KDO)2-(lauroyl)-lipid IVA acyltransferase | − | − | − |
| 156531952 | ABU76778.1 | ESA_01520 | Nitrate reductase 1, alpha subunit | + | + | ± |
| 156531953 | ABU76779.1 | ESA_01521 | Nitrate reductase 1, beta subunit | + | + | ± |
| 156531980 | ABU76806.1 | ESA_01552 | Outer membrane receptor FepA | − | − | − |
| 156532312 | ABU77138.1 | ESA_01884 | Putative receptor | − | − | − |
| 156532401 | ABU77227.1 | ESA_01974 | lpfC/long polar fimbrial outer membrane usher protein | − | − | − |
| 156532403 | ABU77229.1 | ESA_01976 | sfaD/SfaD protein | − | − | − |
| 156532510 | ABU77336.1 | ESA_02086 | pykF/pyruvate kinase I | + | + | ± |
| 156532639 | ABU77465.1 | ESA_02216 | phoP/response regulator in two-component regulatory system with PhoQ, transcribes genes expressed under low Mg+ concn (OmpR family) | + | + | ± |
| 156532640 | ABU77466.1 | ESA_02217 | phoQ/sensory kinase protein in two-component regulatory system with PhoP, ligand is Mg+ | − | − | − |
| 156532687 | ABU77513.1 | ESA_02264 | Flagellar hook-associated protein type 3 | − | − | − |
| 156532688 | ABU77514.1 | ESA_02265 | flgK/flagellar hook-filament junction protein | − | − | − |
| 156532690 | ABU77516.1 | ESA_02267 | flgI/flagellar P-ring protein | + | + | ± |
| 156532691 | ABU77517.1 | ESA_02268 | flgH/flagellar L-ring protein | + | + | ± |
| 156532692 | ABU77518.1 | ESA_02269 | flgG/flagellar basal body rod protein | + | + | ± |
| 156532693 | ABU77519.1 | ESA_02270 | flgF/flagellar basal body rod protein | − | − | − |
| 156532694 | ABU77520.1 | ESA_02271 | flgE/flagellar hook protein FlgE | − | − | − |
| 156532695 | ABU77521.1 | ESA_02272 | flgD/flagellar basal body rod modification protein FlgD | − | − | − |
| 156532696 | ABU77522.1 | ESA_02273 | flgC/flagellar basal-body rod protein | + | + | ± |
| 156532767 | ABU77593.1 | ESA_02344 | sfaE/SfaE protein | − | − | − |
| 156532769 | ABU77595.1 | ESA_02347 | Putative oxidoreductase | + | + | ± |
| 156532833 | ABU77659.1 | ESA_02413 | ompF/outer membrane protein F | − | − | − |
| 156532850 | ABU77676.1 | ESA_02430 | Lipid transporter ATP-binding/permease protein | + | + | ± |
| 156532935 | ABU77761.1 | ESA_02516 | fhaB/filamentous hemagglutinin/adhesin | − | − | − |
| 156532957 | ABU77783.1 | ESA_02538 | fimD/fimbrial adhesin | − | − | − |
| 156532958 | ABU77784.1 | ESA_02539 | lpfB/long polar fimbrial chaperone | − | − | − |
| 156532959 | ABU77785.1 | ESA_02540 | Outer membrane usher protein LpfC | − | − | − |
| 156533053 | ABU77879.1 | ESA_02639 | Putative inner membrane protein | − | − | − |
| 156533067 | ABU77893.1 | ESA_02653 | fur/transcriptional repressor of iron-responsive genes | + | + | ± |
| 156533133 | ABU77959.1 | ESA_02727 | Enterobactin synthase subunit F | − | − | ± |
| 156533264 | ABU78090.1 | ESA_02861 | clpP/ATP-dependent Clp protease proteolytic subunit | + | + | ± |
| 156533628 | ABU78454.1 | ESA_03232 | pilB/(type IV) pilus assembly protein | − | − | − |
| 156533629 | ABU78455.1 | ESA_03233 | pilC/(type IV) pilus assembly protein | − | − | − |
| 156533744 | ABU78570.1 | ESA_03349 | kpsE/putative capsule polysaccharide export system inner membrane protein | − | − | − |
| 156533745 | ABU78571.1 | ESA_03350 | kpsD/polysialic acid capsule transport protein | − | − | − |
| 156533747 | ABU78573.1 | ESA_03352 | kpsC/possible polysaccharide modification protein | − | − | − |
| 156533748 | ABU78574.1 | ESA_03353 | kpsS/possible polysaccharide modification protein | − | − | − |
| 156533753 | ABU78579.1 | ESA_03358 | kpsT/putative capsule polysaccharide export ATP-binding protein | − | − | − |
| 156533960 | ABU78786.1 | ESA_03575 | basS/sensory kinase in two-component regulatory system with BasR | − | − | − |
| 156534135 | ABU78961.1 | ESA_03769 | bplF/lipopolysaccharide biosynthesis protein | + | + | ± |
| 156534173 | ABU78999.1 | ESA_03813 | fimC/outer membrane usher protein precursor | − | − | − |
| 156534174 | ABU79000.1 | ESA_03814 | fimB/chaperone protein | − | − | − |
| 156534462 | ABU79288.1 | ESA_04107 | rfaC/lipopolysaccharide heptosyltransferase I | − | + | ± |
| 156534650 | ABU79476.1 | ESA_04296 | dep/capD gamma-glutamyltranspeptidase | + | + | ± |
| 156534760 | ABU79586.1 | ESA_04407 | pilD/type 4 (IV) prepilin-like protein | − | − | − |
Twelve other genomes used in this study.
RESULTS
Analysis of genetic similarities.
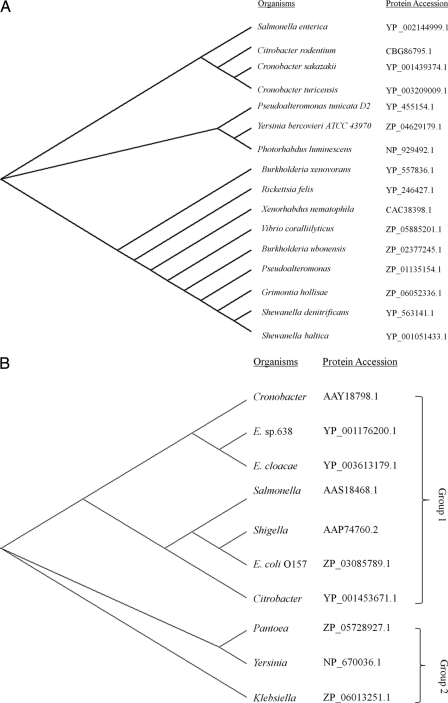
The complete gene data set of the C. sakazakii (CP000783) genome was used to make a gene-by-gene comparison with the representative strains from 13 other bacterial genera. Salmonella enterica serovar Enteritidis (CP001127), Enterobacter cloacae (CP001918), Citrobacter koseri (CP000822), Enterobacter sp. strain 638 (CP000653), Klebsiella pneumoniae (CP000964), Shigella boydii (CP000036), Escherichia coli O157:H7 (AE005174), Pantoea ananatis (CP001875), Yersinia pestis (CP000901), Listeria monocytogenes (AL591824), Bacillus cereus 03BB102 (CP001407), Staphylococcus aureus (CP001844), Clostridium difficile (AM180355), and Campylobacter jejuni (AL111168) were the representative strains used for comparing food-borne pathogens that may be isolated from PIF and other low-moisture food products. The number of genes, the number of unique genes, the total number of consensus genes, the percent GC content, and the size of the genomes (in no. of bp) of the different bacteria are presented in Table 2. We categorized these pathogens into different groups based on the number of unique genes, the GC content obtained from a genome-wide comparative analysis (Table 2), and the phylogenetic analysis of the ompA gene shown in Fig. 2 B. The evolutionary implication of phylogenetic analyses of the chitinase and ompA genes is shown in Fig. 2. Among these genomes are the following groups: group 1, C. sakazakii, Salmonella, Citrobacter, E. cloacae, Enterobacter, Shigella, and E. coli O157:H7; group 2 (no chitinase gene), Pantoea, Klebsiella, and Yersinia; and group 3 (not shown in Fig. 2 due to no significant sequence similarity to ompA and chitinase genes in this group), Listeria, B. cereus, Staphylococcus, C. difficile, and Campylobacter. All pathogens in groups 1 and 2 have 50% or higher GC contents and similar genome sizes, except for Y. pestis, which has a 47% GC content.
FIG. 2.
Phylogenetic analysis of the chitinase (A) and ompA (B) genes shows 2 distinct groups. Protein accession numbers correspond to GenBank.
Identification of unique gene clusters, putative virulence factors, and biomarker genes of Cronobacter spp., Salmonella spp., and the other food-borne pathogens.
The number of unique genes was 268 in C. sakazakii and 401 in Salmonella. Some of these unique genes in C. sakazakii are clustered and are listed as hypothetical genes, with the original annotation provided in GenBank format. Table 3 lists 58 unique clusters of genes, including 8 plasmid-borne genes whose protein products are involved in bacterial pathogenesis and/or have functions or form structures that are important to Salmonella. Likewise, Table 3 also lists 15 unique gene clusters (each gene cluster has a minimum of 3 genes) from Cronobacter spp. whose protein products are also hypothesized to have important functional or structure roles or are involved in pathogenesis. These gene clusters could be used to design a series of unique primers for PCR assays potentially useful for the detection and identification of Cronobacter and Salmonella strains. As an example, there are unique multicopy plasmid-borne genes or clusters listed in Table 3, including estP, a putative esterase or pesticide-degrading enzyme (10) that may exist only in Salmonella. Therefore, these genes have potential application in designing a rapid and sensitive diagnostic test to distinguish Salmonella spp. from related organisms due to the genes' high copy numbers and unique nature; however, comprehensive studies to determine the universal presence of biomarkers, including those that are plasmid encoded in a broad array of Salmonella isolates, would need to be conducted.
Table 4 lists potential biomarkers identified based on scientific publications and protein sequence similarity searches (BLASTp) against the GenBank nonredundant protein database or by relying on keyword-based analysis of text-mined data from publically available databases such as NCBI (http://www.ncbi.nlm.nih.gov/gene) and ENA (http://www.ebi.ac.uk/ena/). In Table 5, the protein homology using the stand-alone BLAST program downloaded from NCBI (http://blast.ncbi.nlm.nih.gov/Blast.cgi?CMD=Web&PAGE_TYPE=BlastDocs&DOC_TYPE=Download) is shown for putative virulence factors and unique biomarker gene discovery. Any protein having an identity of 80% or greater and greater than or equal to 70% query coverage by using the BLASTp program of the BLAST package was considered to have similarity.
Cronobacter yellow pigment gene cluster.
E. sakazakii (now C. sakazakii) was designated a distinct species in 1980 by Farmer et al. (14) and was named in honor of the Japanese bacterial taxonomist/microbiologist Riichi Sakazaki (1920 to 2002), who discovered a distinct yellow-pigmented variant of E. cloacae; however, today non-pigment-producing strains are known to exist within the genus Cronobacter. In the present study, genes responsible for the production of the yellow pigment were used as one of the three targets for the identification of C. sakazakii. The crt operon in C. sakazakii, which contributes to the formation of the yellow pigment (25), was not found in other bacteria shown in Table 4, except for P. ananatis, a plant pathogen, which also produces a yellow pigment. The carotenoid pigment biosynthesis enzymes are encoded by multiple genes within the crt operon. There are 7 genes in this operon, including beta-carotene hydroxylase (crtZ), phytoene synthase (crtB), phytoene dehydrogenase (crtI), phytoene desaturase (crtL), zeaxanthin glucosyl transferase (crtX), isopentenyl pyrophosphate isomerase (idi), and geranylgeranyl diphosphate synthase (crtE). Although these crt operon genes theoretically can be used as targets for bacterial detection, recent studies have shown that approximately 5 to 7% of strains are nonpigmented variants (22; http://www.foodmicrobe.com/food%20poisoning%20microorganisms.htm). Recently, yellow pigment gene clusters from three Cronobacter strains, C. sakazakii ATCC BAA-894 (GenBank accession no. CP000783.1), C. sakazakii BAC 9E10 (GenBank accession no. AM384990.1), and C. turicensis z3032 (GenBank accession no. FN543093.1), have been sequenced. Based on the sequence alignment (data not shown), their DNA sequences are very similar, with only a few nucleotide polymorphisms scattered over the entire 7.6-kb region sequenced; however, there are two significant sequence variabilities in the crtZ gene region. Although the genomic method for the identification of all Cronobacter spp. based on yellow pigment genes as the sole biomarker for clinical diagnosis and early detection could be hampered by these restrictions, since there is limited sequence information to determine if these nonpigmented strains are due to a mutated gene(s), the implication of a consensus sequence in this operon in the genus Cronobacter demonstrates that there is an excellent opportunity to design universal biomarker gene targets for the identification and detection of Cronobacter spp. if also combined with the other suitable biomarkers for identifying nonpigmented Cronobacter strains.
Cronobacter O-antigen gene cluster.
Another potential strain- or serogroup-specific gene cluster, the O-antigen gene cluster, was investigated to determine its suitability as a biomarker (15, 16, 38, 41). The O-antigen gene cluster contains genes with distinctly different functions, including genes involved in the processing and assembly of the O antigen. The genes that encode the O antigen flippase (wzx) and the O antigen polymerase (wzy) are extensively used as biomarker genes for Salmonella and E. coli serotyping (15, 16) and may be suitable biomarkers for Cronobacter spp. as well (Table 4). Both the wzx and wzy genes show limited homology to the same genes in other food-borne pathogens, as confirmed by computational comparative analysis among genomes, making them ideal identification targets for our purposes. It is not clear, however, how much genetic variability of the wzx and wzy genes exists within Cronobacter spp., Salmonella spp., and the other food-borne pathogens due to the limited availability of genomic sequences in the public databases. As shown in Table 4, the wzx gene may not be a suitable biomarker due to significant genetic similarity among Cronobacter spp., Salmonella spp., and E. coli; however, the wzy gene may be suitable.
Cronobacter- and Salmonella-specific virulence factors.
In order to more fully evaluate strain-specific biomarker genes for the differentiation of Cronobacter and Salmonella spp. from the other food-borne pathogens, a systematic approach, combining literature-based data mining (Table 4) and comparative genome analysis (Table 5), was conducted. In Tables 4 and 5, we present an analysis of components of bacterial pathogenesis, including genes that play a role in infection, colonization, and adhesion. In this study, a total of 44 biomarker genes listed in Table 4 were collected by literature-based data mining, and the 14 genomes listed in Table 2 were compared by using the sequences of individual C. sakazakii genes as reference sequences. In Table 4, two important virulence biomarker genes in Cronobacter spp., which are absent in all other pathogens, were analyzed. One gene that was analyzed is a putative hemolysin/hemagglutinin (ESA_02516; GenBank accession no. YP_001438597.1), while another is a putative adhesin (ESA_02084; GenBank accession no. YP_001438170.1). Another putative virulence factor, chitinase, which is of significant clinical importance, was also found to be a suitable biomarker gene for Cronobacter spp. Virulence genes listed in Table 4, such as a gene encoding the hydrolytic enzyme extracellular metalloprotease, prt1, a secretion and transport gene, tatB, and virK, were also analyzed and compared to other pathogens, as well. For the purpose of serotyping, it is necessary to generate a group of species- and serotype-specific biomarker genes. In this study, we found a series of consensus genes (species-specific biomarkers), including 16S rRNA, gyrB, rpoB, recN, thdF, rpoA, ileS, elongation factor G (EF-G), and lepA, through literature data mining and computational analysis suitable for our purposes. In Table 4, we listed only 3 (Clostridium, Campylobacter, and Listeria) out of 5 bacterial species in group 3 food-borne pathogens, due to their importance and the frequencies of their appearance in low-moisture food products.
Verification of analysis of chitinase and ompA genes as potential biomarkers.
As stated above, to verify these virulence factors and unique genes as potential biomarkers (Tables 3 to 5), we amplified and sequenced the PCR products generated from the targeted genes. For this process, we chose two target genes with good potential as biomarkers. One is the chitinase gene, which was identified by the comprehensive sequence comparisons, and the other is the ompA gene, identified by literature-based data mining. Chitinase is a putative virulence factor, and there are various chitinase enzymes present in many organisms, including bacteria. The sequence divergence of the chitinase gene has been used to identify and characterize stage-specific and/or organism-specific genes in several species (9, 31, 40). OmpA is the outer membrane protein A, previously shown to play a role in the invasion of various mammalian host cells. The length of the chitinase gene is ca. 2.2 kb. The primers (Table 1) used for the amplification of the chitinase gene are based on the consensus sequences identified in Salmonella and C. sakazakii through sequence alignments (data not shown). We found that the chitinase gene was present in all 17 tested Cronobacter strains, which originated from three different sources (i.e., environmental, food, and clinical origins). This primer set also amplified the chitinase gene from Salmonella enterica (data not shown). The primers targeting the ompA gene yielded a 469-bp fragment. However, the ompA gene was found only in 11 out of 17 Cronobacter strains tested (Table 6 ) and in 10 out of 15 genomes listed in Table 2. Some of these PCR results are shown in Fig. 1. Partial sequences of the chitinase gene in Cronobacter spp. from various strains and the ompA gene sequences of Cronobacter spp. were analyzed via construction of phylogenetic trees. C. sakazakii was demonstrated to be close to Salmonella, Citrobacter, and Cronobacter turicensis with regard to sequence similarity of the chitinase gene (Fig. 2A). The phylogenetic analysis of the ompA gene (Fig. 2B) was consistent with the genome-level comparison shown in Table 2. Among the food-borne pathogens tested and shown in Table 2, only Salmonella, Citrobacter, and Cronobacter spp. have similar chitinase or chitinase-like genes, as determined by computational blasting and experimental detection of PCR products; however, the chitinase gene is also present in other bacterial pathogens (Fig. 2A), such as Vibrio coralliilyticus, Yersinia bercovieri ATCC 43970, and others.
TABLE 6.
Strains used in this study
| Species (strain/isolate no.) | Presence of: |
Source | |
|---|---|---|---|
| Chitinase | ompA gene | ||
| C. sakazakii (2.39-1 and LCDC 674) | Yes | No | Clinical |
| C. sakazakii (2.39-2) | Yes | No | Clinical |
| C. sakazakii (2.69 and CD1A7[1]) | Yes | No | Shell of hen's egga |
| C. sakazakii (FSM 145) | Yes | No | Environmental, from infant formula manufacturing plant |
| C. sakazakii (FSM 265) | Yes | No | Environmental, from infant formula manufacturing plant |
| C. sakazakii (ATCC 12868) | Yes | Yes | Not available |
| C. sakazakii (ATCC 29004)b | Yes | Yes | Clinical |
| C. sakazakii (ATCC 29544) | Yes | Yes | Clinical (child's throat), type strain |
| C. sakazakii (ATCC 51329)c | Yes | Yes | Not available, type strain |
| C. sakazakii (1) | Yes | Yes | Not available |
| C. sakazakii (2) | Yes | Yes | Not available |
| C. sakazakii (3) | Yes | Yes | Not available |
| C. sakazakii (5) | Yes | Yes | Not available |
| C. sakazakii (6) | Yes | Yes | Not available |
| C. sakazakii (7) | Yes | Yes | Not available |
| C. sakazakii (2.70) | Yes | No | Shell of hen's egga |
| C. sakazakii (2.47 and Gd. St. 8 [HPB 2878]) | Yes | Yes | Environmental |
Originally provided by Michael Musgrove, USDA, ARS.
Now C. sakazakii.
Now C. muytjensii.
DISCUSSION
The discovery of reliable candidate biomarkers for accurate, rapid, and sensitive detection of Cronobacter and Salmonella spp. in complex food matrices and clinical samples remains difficult. This is likely due to several reasons. First, sequence availability detailing genetic differences among food-borne pathogens is very limited. As an example, information as to DNA sequence divergence between Salmonella and E. coli with respect to their O-antigen gene clusters, a common genus- and strain-specific biomarker for differentiation, is incomplete (Table 4). Second, the lack of a comprehensive understanding of phenotypic differences among food-borne pathogens and the absence of particular traditional phenotypic markers may lead to a false-positive diagnosis, as may occur with the absence of the yellow pigment in some strains of Cronobacter spp. (Table 4). Third, the ability to define a set of biomarkers with a role in bacterial pathogenesis or bacterial metabolism that also function as genus-specific markers will provide a more complete and novel picture of the biological activity of food-borne pathogens of interest rather than relying on a single biomarker for identification.
In order to identify potential biomarkers for distinguishing Cronobacter, Salmonella, and the other food-borne pathogens and for developing sensitive and accurate DNA-based detection and identification methods, we investigated a number of candidate biomarker genes. These included genes involved in yellow pigment formation in C. sakazakii (metabolites), the O-antigen cluster genes wzx and wzy, unique genes, and virulence factors common to major food-borne pathogens. The data shown in Tables 3, 4, and 5 in this report are consistent with those shown in a prior report by Healy et al. (19), which demonstrate that many biomarkers were genes related to cell wall/membrane biogenesis/degradation, secretion, and extracellular structures such as fimbriae and flagella. This approach for identifying these putative biomarkers was validated by amplifying and sequencing a particular chitinase gene that showed greater sequence similarity between Cronobacter and Salmonella but was absent in other major food-borne pathogens most frequently found in PIF. As presented in Results, the data obtained in this study demonstrate that chitinase could be a suitable candidate biomarker. In this research, we tested only 17 individual Cronobacter strains and 4 Salmonella strains; therefore, researchers should be cautious when selecting chitinase as the sole biomarker for detection and diagnostic purposes until more comprehensive screening for this gene in Cronobacter and Salmonella spp. is performed. The potential biomarkers listed in Tables 3, 4, and 5 will provide a complete and solid foundation for scientists to draw reliable conclusions based on multiple biomarker genes rather than relying on an individual gene, particular data set, or single experiment.
In this study, we also performed a computational analysis of unique and consensus genes of 15 major food-borne and human pathogens. In total, we identified 292 and 425 unique genes for Cronobacter and Salmonella spp., respectively, that are distinguished from similar genes in other pathogens by a threshold, t, equal to 0.05. Because of the similarity of the genome sequences, the numbers of both the unique and consensus genes suggest that Cronobacter, Salmonella, Shigella, and Enterobacter are genetically more closely related to each other than to other food-borne pathogens, which is not a surprising finding. Further, for detection and diagnostic purposes, this approach could be applied to the use of microarray or target-enrichment strategies for next-generation sequencing technologies when the choice of putative biomarkers can be categorized by functional activities and/or if the number of selected biomarker genes is not a limiting factor. Most importantly, by focusing on a limited number of candidate biomarkers presented in this paper rather than on the whole genome or single biomarkers, appropriate “first tier” and “second tier” of biomarkers are provided for researchers to analyze samples in a complex food matrix or for clinical diagnostic applications. The quantification and characterization of a set of unique biomarkers for various food-borne and human pathogens can be reasonably obtained by either real-time PCR (RT-PCR) or microarray assays, allowing for the detection and identification of important food-borne pathogens based on their genetic differences using currently available technology.
The approaches described above represents our first attempt to describe a systematic approach to identify biomarkers for various food-borne pathogens. Currently, no conventional laboratory method can definitively detect and identify all six of the newly defined species of Cronobacter spp. Recent advances in the areas of genomics and high-throughput studies, as well as the development of new technologies, are improving our understanding of the molecular mechanisms of Cronobacter and Salmonella pathogenesis and are helping to develop effective biomarker identification computational pipelines, an important step toward identifying highly pathogenic Cronobacter spp. and differentiating the other major food-borne pathogens in a timely manner. However, our computational analysis for Cronobacter and Salmonella biomarker identification and the studies described in this report are only a preliminary step toward accomplishing this goal. The fundamental laboratory research, applying PCR and array-based high throughput verification of all major food-borne pathogen biomarkers, is an ongoing project in our laboratory.
Acknowledgments
We express our appreciation to James Smith for his critical review of the manuscript. We also thank Larry Beuchat (University of Georgia, Center for Food Safety, Griffin, GA) and Dong-Hyun Kang (Washington State University, School of Food Science, Pullman, WA) for providing the Cronobacter isolates used in this study.
The mention of trade names or commercial products is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture.
Footnotes
Published ahead of print on 14 January 2011.
REFERENCES
- 1.Anonymous. 2002. Isolation and enumeration of Enterobacter sakazakii from dehydrated powdered infant formula. U.S. Food and Drug Administration, Rockville, MD. http://www.cfsan.fda.gov/comm/mmesakaz.html.
- 2.Anonymous. 2004. Enterobacter sakazakii and other microorganisms in powdered infant formula: meeting report, MRA series 6. FAO/WHO, Geneva, Switzerland.
- 3.Anonymous. 2006. Enterobacter sakazakii and Salmonella in powdered infant formula. Second Risk Assessment Workshop, 16 to 20 January 2006. FAO/WHO, Rome, Italy.
- 4.Anonymous. 2006. Milk and milk products—detection of Enterobacter sakazakii. Technical specification ISO/TS 22964. ISO/TS 22964:2006(E) and IDF/RM 210:2006(E), 1st ed. International Organization for Standardization, Geneva, Switzerland.
- 5.Beuchat, L. R., et al. 2009. Cronobacter sakazakii in foods and factors affecting its survival, growth, and inactivation. Int. J. Food Microbiol. 136:204-213. [DOI] [PubMed] [Google Scholar]
- 6.Bowen, A. B., and C. R. Braden. 2006. Invasive Enterobacter sakazakii disease in infants. Emerg. Infect. Dis. 12:1185-1189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Call, D. R. 2005. Challenges and opportunities for pathogen detection using DNA microarrays. Crit. Rev. Microbiol. 31:91-99. [DOI] [PubMed] [Google Scholar]
- 8.Caubilla-Barron, J., and S. Forsythe. 2007. Dry stress and survival time of Enterobacter sakazakii and other Enterobacteriaceae in dehydrated infant formula. J. Food Prot. 13:467-472. [DOI] [PubMed] [Google Scholar]
- 9.Chaudhuri, S., et al. 2010. Contribution of chitinases to Listeria monocytogenes pathogenesis. Appl. Environ. Microbiol. 76:7302-7305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Choi, Y. J., C. B. Miguez, and B. H. Lee. 2004. Characterization and heterologous gene expression of a novel esterase from Lactobacillus casei CL96. Appl. Environ. Microbiol. 70:3213-3221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Clarridge, J. E., III. 2004. Impact of 16S rRNA gene sequence analysis for identification of bacteria on clinical microbiology and infectious diseases. Clin. Microbiol. Rev. 17:840-862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Drancourt, M., et al. 2000. 16S ribosomal DNA sequence analysis of a large collection of environmental and clinical unidentifiable bacterial isolates. J. Clin. Microbiol. 38:3623-3630. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Drudy, D., et al. 2006. Characterization of a collection of Enterobacter sakazakii isolates from environmental and food sources. Int. J. Food Microbiol. 110:127-134. [DOI] [PubMed] [Google Scholar]
- 14.Farmer, J. J., III., M. A. Asbury, F. W. Hickman, and D. J. Brenner. 1980. The Enterobacteriaceae Study Group (U. S. A.)—Enterobacter sakazakii: a new species of “Enterobacteriaceae” isolated from clinical specimens. Int. J. Syst. Bacteriol. 30:569-584. [Google Scholar]
- 15.Fitzgerald, C., R. Sherwood, L. L. Gheesling, F. W. Brenner, and P. I. Fields. 2003. Molecular analysis of the rfb O antigen gene cluster of Salmonella enterica serogroup O:6,14 and development of a serogroup-specific PCR assay. Appl. Environ. Microbiol. 69:6099-6105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Fratamico, P. M., et al. 2010. Escherichia coli serogroup O2 and O28ac O-antigen gene cluster sequences and detection of pathogenic E. coli O2 and O28ac by PCR. Can. J. Microbiol. 56:308-316. [DOI] [PubMed] [Google Scholar]
- 17.Friedemann, M. 2007. Enterobacter sakazakii in food and beverages (other than infant formula and milk powder). Int. J. Food Microbiol. 116:1-10. [DOI] [PubMed] [Google Scholar]
- 18.Guerra, M. M., F. Bernardo, and J. McLauchlin. 2002. Amplified fragment length polymorphism (AFLP) analysis of Listeria monocytogenes. Syst. Appl. Microbiol. 25:456-461. [DOI] [PubMed] [Google Scholar]
- 19.Healy, B., et al. 2009. Microarray-based comparative genomic indexing of the Cronobacter genus (Enterobacter sakazakii). Int. J. Food Microbiol. 136:159-164. [DOI] [PubMed] [Google Scholar]
- 20.Ho, J. A., and H. W. Hsu. 2003. Procedures for preparing Escherichia coli O157:H7 immunoliposome and its application in liposome immunoassay. Anal. Chem. 75:4330-4334. [DOI] [PubMed] [Google Scholar]
- 21.Iijima, Y., N. T. Asako, M. Aihara, and K. Hayashi. 2004. Improvement in the detection rate of diarrhoeagenic bacteria in human stool specimens by a rapid real-time PCR assay. J. Med. Microbiol. 53:617-622. [DOI] [PubMed] [Google Scholar]
- 22.Iversen, C., et al. 2007. Identification of “Cronobacter” spp. (Enterobacter sakazakii). J. Clin. Microbiol. 45:3814-3816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Iversen, C., et al. 2007. The taxonomy of Enterobacter sakazakii: proposal of a new genus Cronobacter gen. nov. and descriptions of Cronobacter sakazakii comb. nov., Cronobacter sakazakii subsp. Sakazakii comb. nov., Cronobacter sakazakii subsp. malonaticus subsp. nov., Cronobacter turicensis sp. nov., Cronobacter muytjensii sp. nov., Cronobacter dublinensis sp. nov. and Cronobacter genomospecies 1. BMC Evol. Biol. 7:64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Iversen, C., et al. 2008. Cronobacter gen. nov., a new genus to accommodate the biogroups of Enterobacter sakazakii, and proposal of Cronobacter sakazakii gen. nov., comb. nov., Cronobacter malonaticus sp. nov., Cronobacter turicensis sp. nov., Cronobacter muytjensii sp. nov., Cronobacter dublinensis sp. nov., Cronobacter genomospecies 1, and of three subspecies, Cronobacter dublinensis subsp. dublinensis subsp. nov., Cronobacter dublinensis subsp. lausannensis subsp. nov. and Cronobacter dublinensis subsp. lactaridi subsp. nov. Int. J. Syst. Evol. Microbiol. 58:1442-1447. [DOI] [PubMed] [Google Scholar]
- 25.Johler, S., R. Stephan, I. Hartmann, K. A. Kuehner, and A. Lehner. 2010. Genes involved in yellow pigmentation of Cronobacter sakazakii ES5 and influence of pigmentation on persistence and growth under environmental stress. Appl. Environ. Microbiol. 76:1053-1061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kandhai, M. C., M. W. Reij, L. G. M. Gorris, O. Guillaume-Gentil, and M. van Schothorst. 2004. Occurrence of Enterobacter sakazakii in food production environments and households. Lancet 363:39-40. [DOI] [PubMed] [Google Scholar]
- 27.Kim, J. S., et al. 2007. A novel multiplex PCR assay for rapid and simultaneous detection of five pathogenic bacteria: Escherichia coli O157:H7, Salmonella, Staphylococcus aureus, Listeria monocytogenes, and Vibrio parahaemolyticus. J. Food Prot. 70:1656-1662. [DOI] [PubMed] [Google Scholar]
- 28.Kimura, B., et al. 1999. Evaluation of TaqMan PCR assay for detecting Salmonella in raw meat and shrimp. J. Food Prot. 62:329-335. [DOI] [PubMed] [Google Scholar]
- 29.Kucerova, E., et al. 2010. Genome sequence of Cronobacter sakazakii BAA-894 and comparative genomic hybridization analysis with other Cronobacter species. PLoS One 5:e9556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kuhnert, P., B. M. Korczak, R. Stephan, H. Joosten, and C. Iversen. 2009. Phylogeny and prediction of genetic similarity of Cronobacter and related taxa by multilocus sequence analysis (MLSA). Int. J. Food Microbiol. 136:152-158. [DOI] [PubMed] [Google Scholar]
- 31.LeCleir, G. R., A. Buchan, and J. T. Hollibaugh. 2004. Chitinase gene sequences from diverse aquatic habitats reveal complex patterns of diversity. Appl. Environ. Microbiol. 70:6977-6983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Leclercq, A., C. Wanegue, and P. Baylac. 2002. Comparison of fecal coliform agar and violet red bile lactose agar for fecal coliform enumeration in foods. Appl. Environ. Microbiol. 68:1631-1638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lehner, A., T. Tasara, and R. Stephan. 2004. 16S rRNA gene based analysis of Enterobacter sakazakii strains from different sources and development of a PCR assay for identification. BMC Microbiol. 4:43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Liu, Y., et al. 2006. PCR and oligonucleotide array for detection of Enterobacter sakazakii in infant formula. Mol. Cell. Probes 20:11-17. [DOI] [PubMed] [Google Scholar]
- 35.Mackay, I. M. 2004. Real-time PCR in the microbiology laboratory. Clin. Microbiol. Infect. 10:190-212. [DOI] [PubMed] [Google Scholar]
- 36.Miled-Bennour, R., et al. 2010. Genotypic and phenotypic characterisation of a collection of Cronobacter (Enterobacter sakazakii) isolates. Int. J. Food Microbiol. 139:116-125. [DOI] [PubMed] [Google Scholar]
- 37.Mohan Nair, M. K., and K. S. Venkitanarayanan. 2006. Cloning and sequencing of the ompA gene of Enterobacter sakazakii and development of an ompA-targeted PCR for rapid detection of Enterobacter sakazakii in infant formula. Appl. Environ. Microbiol. 72:2539-2546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Mullane, N., et al. 2008. Molecular analysis of the Enterobacter sakazakii O-antigen gene locus. Appl. Environ. Microbiol. 74:3783-3794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Muytjens, H. L., J. van der Ros-van de Repe, and H. A. M. van Druten. 1984. Enzymatic profiles of Enterobacter sakazakii and related species with special reference to the alpha glucosidase reaction and reproducibility of the test system. J. Clin. Microbiol. 20:684-686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Rehli, M., S. W. Krause, and R. Andreesen. 1997. Molecular characterization of the gene for human cartilage gp-39(CHI3L1), a member of the chitinase protein family and marker for late stages of macrophage differentiation. Genomics 43:221-225. [DOI] [PubMed] [Google Scholar]
- 41.Samuel, G., and P. Reeves. 2003. Biosynthesis of O-antigens: genes and pathways involved in nucleotide sugar precursor synthesis and O-antigen assembly. Carbohydr. Res. 338:2503-2519. [DOI] [PubMed] [Google Scholar]
- 42.Steele, M. L., et al. 1997. Survey of Ontario bulk tank raw milk for food-borne pathogens. J. Food Prot. 60:1341-1346. [DOI] [PubMed] [Google Scholar]
- 43.Thompson, J. D., D. G. Higgins, and T. J. Gibson. 1994. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22:4673-4680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Yang, H., L. W. Qu, A. N. Wimbrow, X. P. Jiang, and Y. P. Sun. 2007. Rapid detection of Listeria monocytogenes by nanoparticle-based immunomagnetic separation and real-time PCR. Int. J. Food Microbiol. 118:132-138. [DOI] [PubMed] [Google Scholar]
- 45.Ye, Y.-W., Q.-P. Wu, W.-P. Guo, J.-M. Zhang, and X.-H. Dong. 2007. Rapid detection for Enterobacer sakazakii based on species-specific PCR in powdered milks. Microbiology 34:1192-1197. [Google Scholar]