Abstract
The diagnosis of urinary tract infection (UTI) by urine culture is time-consuming and can produce up to 60 to 80% negative results. Fast screening methods that can reduce the necessity for urine cultures will have a large impact on overall turnaround time and laboratory economics. We have evaluated the detection of bacteria and leukocytes by a new urine analyzer, the UF-1000i, to identify negative urine samples that can be excluded from urine culture. In total, 1,577 urine samples were analyzed and compared to urine culture. Urine culture showed growth of ≥103 CFU/ml in 939 samples (60%). Receiver operating characteristics (ROC) curves and ROC decision plots were been prepared at three different gold standard definitions of a negative urine culture: no growth, growth of bacteria at <104 CFU/ml, and growth of bacteria at <105 CFU/ml. Also, the reduction in urine cultures and the percentage of false negatives were calculated. At the most stringent gold standard definition of no growth, a chosen sensitivity of 95% resulted in a cutoff value of 26 bacteria/μl, a specificity of 43% and a reduction in urine cultures of only 20%, of which 14% were false negatives. However, at a gold standard definition of <105 CFU/ml and a sensitivity of 95%, the UF-1000i cutoff value was 230 bacteria/μl, the specificity was 80%, and the reduction in urine cultures was 52%, of which 0.3% were false negatives. The applicability of the UF-1000i to screen for negative urine samples strongly depends on population characteristics and the definition of a negative urine culture. In our setting, however, the low workload savings and the high percentage of false-negative results do not warrant the UF-1000i to be used as a screening analyzer.
INTRODUCTION
Urinary tract infections (UTIs) are the most common infections in both hospitalized and community patients (16, 19). Although for most patients the burden of disease is minimal, certain subpopulations like children, pregnant women, and the elderly may develop serious complications (8). Symptoms like dysuria, increased urination frequency, hematuria, and back pain are considered specific enough to reliably diagnose a UTI (1). A positive urine culture may confirm the diagnosis and is considered the gold standard in scientific studies. Moreover, susceptibility testing may be performed on cultured bacteria and may guide antibiotic therapy. Considering these facts, it is no surprise that urine samples are the most frequently received specimens in many microbiology laboratories. Nevertheless, a high number of these urine cultures will not yield any bacteria at all, and percentages for negative urine cultures up to 80% have been reported (2, 15). Since urine culture is a laborious procedure, a more cost-effective approach would have a large impact on laboratory economics. A screening method that is able to identify negative urine samples in order to exclude them from costly and laborious culture procedures could reduce the overall turnaround time of analysis, workload, and costs. To prevent positive urine samples from erroneously being classified as negative and not being cultured, a high sensitivity and negative predictive value are prerequisites.
Screening methods, such as dipstick testing for nitrite and leukocytes in urine as well as microscopic sediment analysis for bacteria and white blood cells, are fast but lack sensitivity (5, 21). Moreover, microscopic sediment analysis suffers from interobserver variation and is also labor-intensive. Automated methods for urine sediment analysis, like the Sysmex UF-100, have been developed. The Sysmex UF-100 is an automated urine flow cytometer able to detect particles in urine, including leukocytes and bacteria, quickly by staining the particles with fluorescent dyes and with subsequent identification by impedance, scattering, and fluorescence. Several groups have compared the results of the Sysmex UF-100 to microscopic sediment analysis and reported an adequate performance of the analyzer (3, 7, 17, 18). However, the results of studies that compare the bacterial and leukocyte counts with the UF-100 with urine culture for the diagnosis of UTI vary widely. While some studies have reported an adequate performance in relation to urine culture (6, 9, 10, 13), others have found a relatively large number of false negatives, which decreases sensitivity and makes the UF-100 unsuitable as a screening method to detect negative culture samples (2, 15, 22). Comparing studies is difficult though, since reported sensitivities and specificities depend on the definition used for gold standard positive and negative urines, and these definitions vary among laboratories. For example, some authors consider a urine sample positive if it contains more than 103 CFU/ml (2, 9), whereas others consider 105 or fewer CFU/ml still negative (13).
The Sysmex UF-100 has recently been improved to the Sysmex UF-1000i. One major adaptation to the previous model is that bacteria are stained in a separate bacterial chamber, which prevents interference with red blood cells and improves the detection of bacteria (20). This progress may result in better performance in identifying negative urine samples. In this study we compared the detection of bacteria and leukocytes of the Sysmex UF-1000i to the gold standard, urine culture. The results were analyzed using different gold standard definitions, which made it possible to compare the results to a large number of other studies.
MATERIALS AND METHODS
Urine samples, collected in sterile containers, and submitted for bacterial culture from January to March 2009 to the laboratory for medical microbiology of the PAMM Foundation were enrolled in the study. At arrival the sample was evenly divided over two aliquots. One aliquot was used to prepare a Gram stain, which was analyzed by a trained technician who scored the presence of epithelial cells, leukocytes, yeasts, and bacteria, with the morphology on a scale rating from absent to 4+. From the same aliquot, a chromogenic medium (ChromID CPS3; bioMérieux) and a selective blood agar plate containing 5 μg/ml colisitin and 2 μg/ml aztreonam were inoculated with 10 μl of the sample. Both plates were investigated for growth after 18 to 24 h of aerobic incubation at 37°C. Based on preset validated threshold values, the amount of growth was scored as no growth, 103 to 104 CFU/ml growth, 104 to 105 CFU/ml growth, and ≥105 CFU/ml growth. Grown colonies were identified by their color and a simple additional test in the case of Escherichia coli (brown to burgundy and indole test positive), Proteus mirabilis (clear brown and indole test negative), and Enterococcus spp. (turquoise and growth on CAP blood agar). Other colonies were Gram stained and identified by standard methods when relevant (Gram-negative rods and Gram-positive cocci in groups by the Vitek-2 system [biomeriéux], hemolytic streptococci by Lancefield typing, and viridans streptococci by API-STREP [biomeriéux]). Results of the Gram stain and urine culture procedures were considered the gold standard. Three gold standard definitions for negative urine samples were defined as follows: (i) a negative Gram stain and no growth on the culture plate (here referred to as gold standard no growth); (ii) growth of bacteria at less than 104 CFU/ml (here referred to as gold standard <104 CFU/ml); (iii) growth of bacteria at less than 105 CFU/ml (here referred to as gold standard <105 CFU/ml). Within 2 h after Gram staining and culture plate inoculation, the second aliquots were transported batch-wise to the clinical chemistry laboratory and analyzed for bacteria and leukocytes with the Sysmex UF-1000i according to the manufacturer's guidelines using 800 μl of each sample. The Sysmex UF-1000i performs an analysis of particles in urine by flow cytometry (4, 11, 12, 14, 20). A urine sample is diluted and stained in two different reaction chambers, one for bacteria and one for all other urine particles. The staining agent is a fluorescent polymethine dye that binds to DNA. After staining, the particles are transported to a flow cell and are irradiated by a semiconducting laser (λ = 635 nm). Forward scatter, side scatter, and fluorescence intensities of the individual particles are detected and give information about particle size and structure, which is used to identify and count the particles. The results are presented in histograms and scattergrams.
Using the three gold standard definitions, sensitivity and specificity at different cutoff values were calculated for the UF-1000i. Receiver operating characteristics (ROC) curves and ROC decision plots were made using Analyze-it for Microsoft Excel, version 2.11 (Analyze-it Software, Leeds, United Kingdom).
Cost reduction as a result of omitting cultures for urine samples that were negative by the UF-1000i was calculated by extrapolating data on technician hands-on time and costs of the used materials from a single sample. Hands-on time during culturing included inoculation of the media, transporting the media to the appropriate incubator, recollecting the media the next day, analyzing the media for growth, and entering the data in the laboratory information system. Costs of materials used included a ChromID CPS3 plate (bioMérieux), a blood agar plate containing 5 μg/ml colisitin and 2 μg/ml aztreonam, and a disposable loop. Hands-on time during the UF-1000i procedure included transferring the urine sample to a standardized UF-1000i test tube, placing the test tube in the UF-1000i for analysis, and entering the data in the laboratory information system. Costs comprised the standardized test tube and reagents used.
RESULTS
In total, 1,577 urine samples were included, 681 from outpatients (196 from males) and 896 from hospitalized patients (403 from males). Urine culture showed no growth or bacteria seen in the Gram stain in 638 samples (40%), growth of <104 CFU/ml in 785 samples (50%), and growth of <105 CFU/ml in 981 samples (62%). In 596 samples (38%), bacterial growth of ≥105 CFU/ml was observed. Of all samples in which bacterial growth was observed, 11% contained multiple species and were considered contaminated. Although UTI is unlikely in these patients, these samples were considered gold standard positive when the growth of bacteria exceeded the gold standard definitions of a negative urine culture. The most common microorganisms identified were Escherichia coli (37%), Enterococcus faecalis (14%), Klebsiella spp. (7%), group B streptococci (6%), Proteus mirabilis (4%), Staphylococcus aureus (3%), Pseudomonas aeruginosa (2%), coagulase-negative Staphylococcus (2%), Candida albicans (2%), Gram-negative rods not specified due to small amount of growth (2%), and Aerococcus urinae (1%).
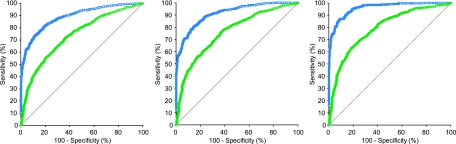
The ROC curves for bacteria and leukocytes at the different gold standard definitions are shown in Fig. 1. In all three cases, the area under the curve (AUC) for bacteria was greater than for leukocytes. The AUCs for bacterial counts were 0.89 (95% confidence interval [CI], 0.87 to 0.90) at no growth, 0.91 (95% CI, 0.90 to 0.93) at <104 CFU/ml, and 0.96 (95% CI, 0.95 to 0.97) at <105 CFU/ml; the AUCs for leukocyte counts were 0.74 (95% CI, 0.71 to 0.76) at no growth, 0.76 (95% CI, 0.74 to 0.78) at <104 CFU/ml, and 0.79 (95% CI, 0.79 to 0.82) at <105 CFU/ml. Furthermore, at any specificity, the corresponding sensitivity was higher for bacteria than for leukocytes.
Fig. 1.
ROC curves for bacterial detection (blue) and leukocyte detection (green) by the UF-1000i when no growth (left), <104 CFU/ml (middle), and <105 CFU/ml (right) were chosen as the gold standard definitions for negative cultures.
In Fig. 2, we analyzed the bacterial detection of the UF-1000i by preparing ROC decision plots. In these graphs, sensitivity and specificity are set against the cutoff value of the UF-1000i, so when a certain UF-1000i cutoff value is chosen, sensitivity and specificity are determined from the curves. For example, when urine cultures without growth are considered negative (Fig. 2, left graph), a cutoff value of 200 bacteria/μl for the UF-1000i results in a sensitivity of 74% and a specificity of 87%. When the gold standard definition for a negative culture value is changed to <104 or <105 CFU/ml (Fig. 2, middle and right graphs), the sensitivity at the same UF-1000i cutoff increases to 82% and 96%, while the specificity decreases to 83% and 78%, respectively. Screening for negative urine samples requires a high sensitivity in order to minimize the number of false-negative screening results. When an arbitrary sensitivity of 95% is chosen, the most stringent gold standard definition would result in a cutoff of 26 bacteria/μl counted by the UF-1000i and a specificity of 43% (Fig. 2, left graph). More liberal definitions of <104 or <105 CFU/ml for negative urine cultures would increase the UF-1000i cutoff values to 39 and 230 bacteria/μl, resulting in specificities of 52% and 80%, respectively.
Fig. 2.
ROC decision plots for the detection of bacteria by the UF-1000i, depicting sensitivity (purple) and specificity (cyan) as a function of the cutoff value chosen for the UF-1000i when no growth (left), <104 CFU/ml (middle), and <105 CFU/ml (right) were chosen as gold standard definitions for negative cultures.
We investigated whether a combination of cutoff values for leukocytes and bacteria might improve the combination of sensitivity and specificity of detecting a urinary tract infection by the UF-1000i. However, we did not find a combination of leukocytes and bacteria that provided higher sensitivities and specificities than when only the bacterial counts were used.
In Fig. 3, the percentage of negative UF-1000i results are shown (dark blue curves) at different UF-1000i cutoff values. This value represents the percentage of cultures saved when the UF-1000i would have been used as a screening test. Not all results were truly negative. The percentage of false-negative tests of the UF-1000i-negative tests is shown by the red line. The graphs in Fig. 3 represent both plots for the different gold standard definitions.
Fig. 3.
The percentage of negative UF-1000i test results at different cutoff values (dark blue) and the percentage of false-negative UF-1000i test results among the total UF-1000i negative test results (red) when no growth (left), <104 CFU/ml (middle), and <105 CFU/ml (right) were chosen as gold standard definitions for negative cultures.
The costs for screening all urine samples by the UF-1000i were €1,183, with 158 min hands-on time. Assuming a desired sensitivity for the UF-1000i of 95% at a gold standard definition of no growth, the UF-1000i would identify 319 negative urine samples. Omitting urine cultures in these negative samples would save €389 and 558 min hands-on time. In total, screening all urine samples with the UF-1000i would increase costs by €794 but would save 400 min hands-on time. Similarly, using the less stringent gold standard definitions of <104 and <105 CFU/ml but maintaining the desired sensitivity of 95% for the UF-1000i, total costs would increase by €641 and €184 but save 619 min and 1,275 min of hands-on time, respectively.
DISCUSSION
The number of gold standard-negative urine samples depends on the definition used. Our study showed 40%, 50%, and 62% negative urines at definitions for negative cultures of no growth, <104 CFU/ml, and <105 CFU/ml, respectively. Although these numbers are comparable to another recent study evaluating the UF-1000i (11), most studies have reported a percentage of negative culture samples around 70 to 80% (2, 4, 15, 22). The organisms identified in our study correspond well with earlier published results (2, 4, 9).
The ROC curves (Fig. 1) indicate that the number of bacteria reported by the UF-1000i can discriminate better between negative and positive urine cultures than the leukocyte count. Interestingly, when increasing the definition of a negative urine culture, the AUC for the bacteria count increases more than the AUC for the detection of leukocytes. Moreover, the differences between the curves are so big that the leukocyte count does not have an additional effect to the bacteria count. This is in contrast to all previous studies showing optimal combinations of sensitivity and specificity when a combination of bacteria and leukocyte counts were used. A possible explanation for this finding might be that the receptacles used for urine sampling in our region do not contain boric acid fluids, a compound known to stabilize leukocytes and inhibit further growth of bacteria. Nevertheless, in our data set, the bacteria count was the only parameter that needed to be monitored while screening for negative urine samples.
The ROC decision plots (Fig. 2) illustrate the influence of the gold standard definition of a negative urine culture on sensitivity and specificity. Loosening the gold standard definition by accepting an increasing number of bacteria grown in urine samples still being considered negative increases the sensitivity and decreases the specificity of the UF-1000i. Screening for negative urine samples requires a high negative predictive value and therefore a high sensitivity, since a false-negative urine test will prohibit the urine from being cultured. In our study high sensitivities are accompanied by lower specificities, compared to the specificities found in other recent studies that evaluated the bacterial detection of the Sysmex UF-1000i. For example, Manoni et al. reported a specificity of 94% at a sensitivity of 97% when a definition of a negative urine culture of <105 CFU/ml was used (11). De Rosa et al. found a specificity of 85% and a sensitivity of 95% at a definition of <104 CFU/ml for a negative urine culture (4). The population described by Manoni et al. is quite similar in percentage of culture negatives and cultured microorganisms to our population, but our study showed many more samples with a high UF-1000i bacteria count that appeared negative when a similar gold standard definition was used. For example, in 981 of our samples that were culture negative (at a gold standard definition of <105 CFU/ml), 281 samples showed a bacteria count of >125 bacteria/μl by the UF-1000i. This number is much higher than the number of high-count false negatives reported by Manoni et al. Several explanations may account for this discrepancy. Antibiotic use prior to urine analysis may result in the detection of nonviable bacteria by the UF-1000i. Other aspects that might play a role are different culture procedures, culture media, or transportation times, and the use of boric acid compounds. The patient population in the study of De Rosa et al. was also similar in number to our data set but contained a much higher number of culture-negative samples (74% versus 50% in our study, at a comparable gold standard definition of <104 CFU/ml). Again, this might reflect culture procedures and prior-to-analysis antibiotic use, but more likely the time point at which a urine sample is collected for microbiological analysis plays a role. In the Netherlands general practitioners are trained to treat patients with uncomplicated UTI empirically with two courses of antibiotics before collecting a urine sample for culture. This might result in a higher a priori chance of a positive urine sample.
Figures 3 illustrates the reduction in urine cultures when the UF-1000i is used to screen urine samples. For example, at the gold standard definition of no growth, a UF-1000i cutoff of 26 bacteria/μl, corresponding to a sensitivity of 95% (Fig. 2, left graph), would result in a reduction of urine cultures of 20% (Fig. 3, left graph). At a gold standard definition for a negative culture of <104 CFU/ml, the same sensitivity would be reached at a UF-1000i cutoff of 39 bacteria/μl and a reduction of urine cultures of 28%. For the gold standard definition of <105 CFU/ml, the same sensitivity was reached at a cutoff of 230 bacteria/μl and would result in a reduction of 52% of urine cultures. The dark blue curves depicted in Fig. 3 illustrate the reduction in urine cultures at different cutoff values of the UF-1000i and are therefore independent of the chosen gold standard definition, and are thus identical in all three figures. As expected, loosening the gold standard definition will result in a decrease of false-negative UF-1000i test results (red lines). For example, at a UF-1000i cutoff of 26 bacteria/μl, the negative predictive value of the UF-1000i will appear to be 86% at the gold standard definition of no growth, 95% at a gold standard definition of <104 CFU/ml, and 99.7% at a gold standard definition of <105 CFU/ml. In other words, a less stringent gold standard definition will result in a lower number of false-negative UF-1000i results, a higher negative predictive value, and a higher number of urine samples not being cultured. It has to be kept in mind, though, that at a gold standard definition for a negative culture of <105 CFU/ml, the presence of very large amount of bacteria is considered not significant, which is questionable, especially for those patients most at risk: children, pregnant women, and the elderly.
Total savings are considerable when using the UF-1000i for screening urine samples. Although reagent costs might be higher for the UF-1000i than material costs for the culture procedure, the most costly factor, hands-on time, is minimal for the UF-1000i. Moreover, the UF-1000i may even be operated by less-skilled technicians than the culture procedure. Finally, savings increase with a less stringent gold standard definition or a higher a priori change of a negative urine sample.
Application of the UF-1000i as a screening analyzer to reduce the number of negative urine culture samples is only reliable when a high sensitivity and a high negative predictive value are obtained. However, this depends on the definition of positive and negative urine cultures used in the laboratory. In our opinion, it is therefore not possible to decide on a single, general cutoff value of the bacteria count for the UF-1000i. In our laboratory, urine samples with no growth of bacteria are normally considered negative, which results in an a priori chance of 40% for a negative urine sample. In this setting, the UF-1000i analyzer would only identify 20% of the urines as negative and thus save 20% of the workload; moreover, 14% of these urine samples would be false negative. In our opinion the low reduction in workload and the high percentage of false-negative results do not warrant the use of the UF-1000i as a screening analyzer. However, in laboratories with a higher a priori chance of a negative urine sample, the UF-1000i might be helpful in significantly reducing workload and costs.
ACKNOWLEDGMENTS
We acknowledge Truus Buijssen, Kim van Nieuwenhuijzen, and Gerard Coppens for their help in analyzing the urine specimens.
Footnotes
Published ahead of print on 19 January 2011.
REFERENCES
- 1. Bent S., Nallamothu B. K., Simel D. L., Fihn S. D., Saint S. 2002. Does this woman have an acute uncomplicated urinary tract infection? JAMA 287:2701–2710 [DOI] [PubMed] [Google Scholar]
- 2. Brilha S., Proenca H., Cristino J. M., Hanscheid T. 2010. Use of flow cytometry (Sysmex UF-100) to screen for positive urine cultures: in search for the ideal cut-off. Clin. Chem. Lab. Med. 48:289–292 [DOI] [PubMed] [Google Scholar]
- 3. Delanghe J. R., et al. 2000. The role of automated urine particle flow cytometry in clinical practice. Clin. Chim. Acta 301:1–18 [DOI] [PubMed] [Google Scholar]
- 4. De Rosa R., et al. 2010. Evaluation of the Sysmex UF1000i flow cytometer for ruling out bacterial urinary tract infection. Clin. Chim. Acta 411:1137–1142 [DOI] [PubMed] [Google Scholar]
- 5. Deville W. L., et al. 2004. The urine dipstick test useful to rule out infections. A meta-analysis of the accuracy. BMC Urol. 4:4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Evans R., Davidson M. M., Sim L. R., Hay A. J. 2006. Testing by Sysmex UF-100 flow cytometer and with bacterial culture in a diagnostic laboratory: a comparison. J. Clin. Pathol. 59:661–662 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Fenili D., Pirovano B. 1998. The automation of sediment urinalysis using a new urine flow cytometer (UF-100). Clin. Chem. Lab. Med. 36:909–917 [DOI] [PubMed] [Google Scholar]
- 8. Foxman B. 2002. Epidemiology of urinary tract infections: incidence, morbidity, and economic costs. Am. J. Med. 113(Suppl. 1A):5S–13S [DOI] [PubMed] [Google Scholar]
- 9. Kim S. Y., et al. 2007. Evaluation of the Sysmex UF-100 urine cell analyzer as a screening test to reduce the need for urine cultures for community-acquired urinary tract infection. Am. J. Clin. Pathol. 128:922–925 [DOI] [PubMed] [Google Scholar]
- 10. Koken T., et al. 2002. Determination of cut-off values for leucocytes and bacteria for urine flow cytometer (UF-100) in urinary tract infections. Int. Urol. Nephrol. 34:175–178 [DOI] [PubMed] [Google Scholar]
- 11. Manoni F., et al. 2009. Cutoff values for bacteria and leukocytes for urine flow cytometer Sysmex UF-1000i in urinary tract infections. Diagn. Microbiol. Infect. Dis. 65:103–107 [DOI] [PubMed] [Google Scholar]
- 12. Manoni F., et al. 2010. Urine particle evaluation: a comparison between the UF-1000i and quantitative microscopy. Clin. Chem. Lab. Med. 48:1107–1111 [DOI] [PubMed] [Google Scholar]
- 13. Manoni F., et al. 2002. Field evaluation of a second-generation cytometer UF-100 in diagnosis of acute urinary tract infections in adult patients. Clin. Microbiol. Infect. 8:662–668 [DOI] [PubMed] [Google Scholar]
- 14. Nanos N. E., Delanghe J. R. 2008. Evaluation of Sysmex UF-1000i for use in cerebrospinal fluid analysis. Clin. Chim. Acta 392:30–33 [DOI] [PubMed] [Google Scholar]
- 15. Okada H., et al. 2000. Detection of significant bacteriuria by automated urinalysis using flow cytometry. J. Clin. Microbiol. 38:2870–2872 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Peleg A. Y., Hooper D. C. 2010. Hospital-acquired infections due to gram-negative bacteria. N. Engl. J. Med. 362:1804–1813 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Regeniter A., et al. 2001. Urine analysis performed by flow cytometry: reference range determination and comparison to morphological findings, dipstick chemistry and bacterial culture results. A multicenter study. Clin. Nephrol. 55:384–392 [PubMed] [Google Scholar]
- 18. Shayanfar N., Tobler U., von Eckardstein A., Bestmann L. 2007. Automated urinalysis: first experiences and a comparison between the Iris iQ200 urine microscopy system, the Sysmex UF-100 flow cytometer and manual microscopic particle counting. Clin. Chem. Lab. Med. 45:1251–1256 [DOI] [PubMed] [Google Scholar]
- 19. Stamm W. E., Hooton T. M. 1993. Management of urinary tract infections in adults. N. Engl. J. Med. 329:1328–1334 [DOI] [PubMed] [Google Scholar]
- 20. Wang J., Zhang Y., Xu D., Shao W., Lu Y. 2010. Evaluation of the Sysmex UF-1000i for the diagnosis of urinary tract infection. Am. J. Clin. Pathol. 133:577–582 [DOI] [PubMed] [Google Scholar]
- 21. Williams G. J., et al. 2010. Absolute and relative accuracy of rapid urine tests for urinary tract infection in children: a meta-analysis. Lancet Infect. Dis. 10:240–250 [DOI] [PubMed] [Google Scholar]
- 22. Zaman Z., Roggeman S., Verhaegen J. 2001. Unsatisfactory performance of flow cytometer UF-100 and urine strips in predicting outcome of urine cultures. J. Clin. Microbiol. 39:4169–4171 [DOI] [PMC free article] [PubMed] [Google Scholar]