Abstract
Myxoma virus (MYXV) M062R is a functional homolog of the C7L family of host range genes from orthopoxviruses. We constructed a targeted M062R-knockout-MYXV (vMyxM062-KO) and characterized its properties in vitro and in vivo. In European rabbits, infection by vMyxM062-KO was completely asymptomatic. The surviving rabbits did not gain full protection against the subsequent lethal-dose challenge with wild-type MYXV. We also looked for cellular tropism defects in a variety of cultured cells. In all of the rabbit cells tested, vMyxM062-KO conducts an abortive infection, although it initiates viral DNA replication. In many, but not all, human cancer cells that are permissive for wild-type MYXV, vMyxM062-KO exhibited a profound replication defect. We categorized human cells tested into two groups: (i) type A, which support productive replication for wild-type MYXV but are unable to produce significant levels of progeny virus by vMyxM062-KO, and (ii) type B, which are permissive to infections by both wild-type MYXV and vMyxM062-KO. Furthermore, using proteomic strategies, we identified sterile α motif domain containing 9 (SAMD9), an interferon-regulated cellular protein implicated in human inflammatory disorders, as a unique host binding partner of M062 in human cells. Significantly, knocking down SAMD9 in type A human cancer cells led to a substantial rescue of vMyxM062-KO infection. In summary, M062 is a novel host range factor that controls productive MYXV replication in rabbit cells and in a wide variety of human cells. M062 also binds and antagonizes cellular SAMD9 in human cells, suggesting that SAMD9 is a novel innate antiviral factor against poxviruses.
The poxvirus C7L family represents a group of viral host range genes widely distributed among almost all mammalian poxviruses (16). Unlike some of the other known host range proteins in orthopoxvirus members, such as vaccinia virus (VACV) K1 and cowpox CP77, members of the C7 protein family do not contain well-defined protein consensus motifs or domains other than sequence similarity to each other. Little is known about the biological functions of the members of the C7L host range factor family. In VACV, K1 and C7 can complement their host range functions to rescue viral replication in most cell lines. However, K1L or CP77, but not C7L, can compensate for the defect of a knockout VACV for both K1L and C7L in infected rabbit cells (20). It was suggested that VACV C7 may antagonize the antiviral effects triggered by type 1 interferon (IFN) to allow productive infection of VACV in the absence of K1 (17). However, the molecular mechanism of C7L gene family members in relation to host antiviral pathways remains to be determined. The present study was initiated to study the role of one C7 family member, M062 of MYXV, in its native susceptible rabbit host. Since MYXV is also being currently developed as an oncolytic therapeutics for human cancer (12), we also investigated the potential host range properties of M062 in a broad spectrum of human cancer cells.
M062 was recently shown to be a functional homolog of VACV C7 (16), such that the replacement of the deleted VACV C7L by M062R in a K1L knockout background of VACV restored the host tropism of VACV infection in cultured cells and rescued pathogenesis in vivo (16). Interestingly, the MYXV genome also includes two other related C7L-like genes, M063R and M064R, but neither of these was able to compensate for the host range function of C7L in VACV. It was previously shown that the expression of M063 is essential for the productive MYXV infection in rabbit cells, but M063 is fully dispensable for infecting all human cancer cells tested that were permissive for wild-type MYXV (3, 4).
We focused here on studying the role of M062 in MYXV infection in cultured cells of rabbit and human origin, and on its role as a virulence factor in MYXV-infected European rabbits, by examining for any novel biological phenotype of the targeted M062R-knockout virus in vitro and in vivo. As a part of this study, we identified a novel cellular binding partner of M062, called SAMD9, in MYXV-infected human cells. We also investigated the interaction between M062 and SAMD9 and discovered a role of the viral M063 protein in facilitating this protein-protein interaction. Finally, we studied the role of the human SAMD9 as a novel antiviral restriction factor for MYXV infection in human cells and determined that its antiviral effects are antagonized by M062, which thus explains why M062 is essential for MYXV replication in most human cancer cells.
MATERIALS AND METHODS
Cell culture and virus stock.
Unless stated specifically otherwise, most cell lines were cultured in Dulbecco minimal essential medium (DMEM; Invitrogen) supplemented with 10% fetal bovine serum (FBS; Gibco), 2 mM glutamine (Invitrogen), and 100 μg of penicillin-streptomycin (Pen/Strep; Invitrogen)/ml, including human cells, e.g., U87 (kindly provided by P. Forsyth) (14), Huh7 (17), HeLa (ATCC CCL-2), HepG2 (ATCC CRL-10741), and Hs766T (ATCC HTB-134), monkey cell lines including BSC-40 (ATCC CRL-2761) and BGMK (21), and the rabbit cell line RK-13 (ATCC CCL-37). The RL-5 rabbit CD4+ T cell line (5) and the THP1 human cell line (ATCC TIB-202) are cultured in RPMI 1640 (obtained from Gibco and Lonza-BioWhittaker, respectively) supplemented with 10% FBS, 2 mM glutamine, and 100 μg of Pen/Strep per ml. Human vascular endothelial cells (ATCC PCS-100-010) were maintained in endothelial cell growth media (Lonza-BioWhittaker). CAL-27 (ATCC CRL-2095) and SSC-25 (ATCC CRL-1628 cultured in a 1:1 mixture of DMEM and F-12 medium containing 2 g of sodium bicarbonate/liter, 2.5 mM l-glutamine, 15 mM HEPES, and 0.5 mM sodium pyruvate and supplemented with 400 ng of hydrocortisone/ml and 10% FBS) were kindly provided by A. Jakymiw (Medical University of South Carolina).
MYXV viral stock was prepared using BSC-40 cells as described previously (21).
Construction of recombinant MYXV.
To create a targeted M062R-knockout MYXV construct, vMyxM062-KO, a recombinant plasmid was first constructed using MultiSite Gateway Pro (Invitrogen) system (13, 15). Briefly, 78% of the central region of M062R sequence was replaced by an enhanced green fluorescent protein (eGFP) expression cassette (driven by a poxvirus synthetic early/late promoter) with 51 bp remaining at the 5′ end. Upstream and downstream hybridizing sequences were amplified by PCR to generate entry clones by Gateway BP recombination with appropriate pDONR vectors. Hybridizing sequences were confirmed by sequencing. The final recombinant plasmid was constructed by recombining entry clones with a destination vector in a sequential manner. M062R knockout virus (vMyxM062-KO) was constructed by infecting BSC-40 cells with wild-type MYXV Lausanne strain (vMyx-Lau), followed by transfection of the recombination plasmid (13). Multiple rounds of focus purifications were conducted on BSC-40 cells until a pure stock of knockout virus was achieved, which was confirmed by PCR. The similar method was used to construct vMyxM062V5 with C-terminal V5-tagged M062 that also contains an eGFP expression cassette driven by a VACV p11 late promoter inserted downstream the M062RV5 sequence. A revertant of M062R knockout virus, vMyxM062-Rev, was constructed by infecting BSC-40 cells with vMyxM062-KO, followed by transfection with a “revertant” plasmid containing the intact M061R, M062R, and M063R gene sequences. Focus purification was conducted on BGMK cells until a nonfluorescent M062R containing pure stock was achieved, which was confirmed by PCR using the appropriate primer set.
Cloning and transfection.
DNA sequences encoding the open reading frames (ORFs) for M062R and M063R were individually cloned in plasmid pcDNA3.1-Myc/His, respectively. The expression of corresponding protein is driven by an immediate-early cytomegalovirus promoter and fused with c-terminal Myc and His6. Viral DNA (vMyx-Lau) was used to amplify by PCR the genomic sequence containing the M061R, M062R, and M063R genes which was then inserted into pCR2.1 using TOPO TA cloning kit (Invitrogen). The resulting revertant plasmid was confirmed by sequencing and used to construct the revertant virus vMyxM062-Rev. The transfection of cells was performed using Effectene reagent (Qiagen) according to the manufacturer's protocol.
One-step and multiple-step viral replication growth curves.
One-step growth curves were constructed by quantifying viral yields at various time points from cells infected with virus at a various multiplicities of infection (MOIs) ranging from 3 to 5. After 1 h of incubation with virus inocula, the cells were washed with phosphate-buffered saline (PBS), and the medium was replaced with the appropriate growth medium. Multiple-step growth curves were constructed by tittering virus yield from cell lysates harvested at various time points after infection at an MOI of 0.1. Titration of viral yield was conducted using BSC-40 cells and a protocol that was described previously (11).
Cytosine β-d-arabinofuranoside (AraC; Sigma Life Science) was used to study MYXV gene expression kinetics or a DNA replication profile, and a 0.2 mM concentration was used to pretreat cells for 1 h before viral infection at an MOI of 5 in the presence of AraC. After 1 h of incubation with virus inocula, the infection medium was replaced with fresh growth medium with AraC until the time for harvesting.
Ni-NTA pull-down, coimmunoprecipitation (co-IP), and Western blot analysis.
For the nickel-nitrilotriacetic acid (Ni-NTA) pull-down experiment, HeLa cells were transiently transfected with plasmid DNA as described above. At 72 h after transfection, monolayer cells were washed with ice-cold PBS and lysed with NP-40 lysis buffer with an appropriate concentration of proteinase inhibitor (Roche) and 10 mM imidazole (pH 7). Cell lysates were collected and gently rotated at 4°C for 10 min before spinning them down at 12,000 × g at 4°C. Meanwhile, Ni-NTA agarose (Qiagen) was washed in Tris-HCl buffer (50 mM pH 7.4), followed by two more washes with the lysis buffer. The supernatant containing total protein from cell lysis was added to the washed Ni-NTA beads, rotated at 4°C for 2 h, and then spun down briefly to collect any protein associated with and/or bound to the beads. The beads were washed five times with lysis buffer containing 20 mM imidazole, and any bound proteins were eluted with lysis buffer containing 250 mM imidazole. Eluted proteins were separated on SDS-PAGE gel for Western blot analysis.
Co-IP experiments were conducted with an anti-V5 antibody (Invitrogen, Carlsbad, CA) or agarose-conjugated anti-V5 antibody (Sigma-Aldrich, St. Louis, MO). The anti-V5 antibody (not agarose conjugated) was incubated with Pierce protein A/G-agarose (Thermo Scientific) in washing buffer (50 mM Tris-Cl [pH 7.4], 150 mM sodium chloride, 0.1% Nonidet P-40 or NP-40) for 30 min. The agarose was then washed three times to get rid of the excess antibody. Cell lysates were incubated with antibody bound to agarose and rotated at 4°C for approximately 2 h, followed by washing steps, and the samples were then analyzed by Western blotting.
For Western blot analysis, antibodies targeting the human SAMD9 protein (Sigma-Aldrich), c-Myc (9E10; Santa Cruz Biotechnology), anti-V5 (Invitrogen), anti-β-actin (Ambion), goat anti-mouse IgG conjugated with peroxidase (Thermo Scientific), and goat anti-rabbit IgG conjugated with peroxidase (Santa Cruz Biotechnology) were purchased commercially for this study. Antibodies against MYXV SERP-1 and M063 were previously described (4, 19). Briefly, after the addition of Laemmli loading buffer and heat treatment, 25 to 50 μg of total protein or samples from pull-down or co-IP were separated by SDS-PAGE before transferring to HyBond-P hydrophobic polyvinylidene difluoride (PVDF) membrane (GE Healthcare) using the XCell II Blot Module (Invitrogen). The membranes were blocked with 5% bovine serum albumin (Fisher Scientific) or 5% nonfat milk in TBS (150 mM NaCl, 2.5 mM KCl, 25 mM Tris-HCl [pH 7.4]) containing 0.1% Tween 20 (Fisher Scientific) for 30 min to 2 h. The membranes were then incubated at 4°C overnight with primary antibody at the appropriate dilution, washed five times, and incubated with the secondary antibody conjugated to horseradish peroxidase (HRP) for 1 h. After washing, proteins of interest were detected using Immobilon Western HRP substrate (Millipore) on Kodak BioMax film (Carestream Health, Inc.).
Mass spectrometry of protein samples.
Protein samples from co-IP experiments were electrophoresed by SDS-PAGE and stained with mass spectrometry-compatible Coomassie blue (SimplyBlue SafeStain; Invitrogen) according to the manufacturer's protocol. Unique protein bands compared to proper controls were dissected for trypsin digestion, followed by liquid chromatography-tandem mass spectrometry analysis (proteomic service by the Interdisciplinary Center for Biotechnology Research or ICBR, University of Florida). The results were searched against an appropriate protein database and then analyzed and displayed with matching polypeptide sequences in the identified protein sequence using Scaffold (Proteome Software, Portland, OR).
DNA and RNA extraction and real-time PCR.
Viral infection was conducted at an MOI of 5 and, at given time points, infected or mock treated cells were harvested for DNA extraction using a FlexiGene DNA kit (Qiagen). At the designated time points, RNA extraction using TRIzol (Invitrogen) was conducted on infected cells (at an MOI of 5 or mock treated). Contaminating genomic DNA was cleaned by using a DNA-free kit (Ambion) (11). Reverse transcription was performed using 1 to 2 μg of total RNA and SuperScript III reverse transcriptase (Invitrogen) for cDNA synthesis according to the manufacturer's protocol. Real-time PCR was set up using SYBR green PCR core reagents (Applied Biosystems) as previously described (9), and analysis was performed by using a thermocycler (Opticon II; MJ Research). A melting-curve analysis was conducted to ensure the specificity of the primer sets. A comparative threshold cycle (CT) method was used to calculate and compare DNA or RNA level for the gene of interest.
Pathogenesis study in rabbits.
This animal study was approved by the Institutional Animal Care and Usage Committee at the University of Florida. MYXV pathogenesis in New Zealand White rabbits (Charles River Laboratories International) was compared between four groups: (i) PBS (n = 3): 100 μl of PBS was injected intradermally (ID) on the left flank; (ii) vMyx-Lau (n = 5); (iii) vMyxM062-KO (n = 4); and (iv) vMyxM062-Rev (n = 3). Groups ii to iv received 1,000 focus-forming units (FFU) of the indicated virus in 100 μl of PBS and were inoculated intradermally in the left flank of each rabbit. Daily physical examinations (13) were conducted to evaluate respiration, weight, temperature, heart rate, lung sound, food and water intake, urine and feces output, hydration status, attitude, activity, posture, and indications of primary lesion and secondary lesions, as well as the condition of these lesions. An overall daily clinical score was obtained for each animal that ranged from 0 to 34 (the maximum score). Animals were humanely euthanized when one of the following conditions was observed: (i) a clinical score reaching 26 of 34, (ii) extreme respiratory stress (e.g., open mouth breathing), (iii) orthopnea, (iv) cyanosis, or (v) no food or water intake for 48 h. If animals initially infected with knockout virus survived at 24 days after initial infection, a challenge was performed with a lethal dose of vMyx-Lau (e.g., 1,000 FFU) administered by the intradermal route in the right flank of the animal. One group of naive animals (n = 2) was used as a control for this infection. Daily physical examination as previously described was conducted.
Transfection of siRNAs and cell line establishment using lentivirus packaged with short hairpin RNAs (shRNAs).
Oligofectamine (Invitrogen) was used to transfect cells with small interfering RNAs (siRNAs), including control and siRNAs targeting human SAMD9 (ON-TARGET plus SMART pool and individual siRNA sets [Dharmacon]; target sequences are listed in Table S1 in the supplemental material), respectively, at a final concentration of 100 nM according to the manufacturer's protocol. At 48 h posttransfection, the cells were either harvested for Western blot analysis to check protein knockdown level or infected with MYXV to examine the effect of the protein level of SAMD9 on viral replication.
Control lentivirus packaged with a scrambled shRNA (Santa Cruz Biotechnology, Inc.) or SAMD9 shRNA lentivirus particles (Santa Cruz Biotechnology, Inc.) were used individually to infect cells to construct stable cell lines according to the manufacturer's protocol. Western blot analysis was performed to determine the knockdown level of SAMD9.
Statistic analysis.
GraphPad Prism 5.0 (GraphPad Software, Inc.) or SigmaStat3.1 (Systat Software Asia Pacific, Ltd.) software was used for statistical analysis in the present study. A statistical significance was defined as a P value of <0.05.
RESULTS
M062 is expressed as an early or late protein and is essential to MYXV infection in rabbit cells.
To construct a targeted M062R knockout virus (vMyxM062-KO), the central 78% of the M062R coding sequence was deleted from the parental MYXV genome (vMyx-Lau) and replaced with an eGFP expression cassette driven by a poxvirus synthetic early/late promoter (Fig. 1A and B). The M062 knockout virus was created and amplified in BSC-40 cells but, unexpectedly, did not replicate successfully in BGMK cells, a related but distinct monkey cell line that has also been used routinely in our lab to propagate the parental MYXV (results not shown). In RK-13 rabbit kidney cells (Fig. 2B and D) and in RL-5, a rabbit CD4+ T cell line (Fig. 2A and C), infection by vMyxM062-KO was abortive, with significantly reduced early marker gene expression (GFP expression) and no production of progeny virus. Viral DNA replication in RK-13 cells appeared to be normal up to 8 h postinfection (p.i.) (Fig. 3A) but was significantly reduced compared to wild-type MYXV (vMyx-gfp) from 12 to 24 h p.i. (Fig. 3A). A typical late MYXV gene product (e.g., SERP-1) was not expressed during the course of vMyxM062-KO infection in rabbit cells, even as late as 24 h p.i. Meanwhile, in wild-type MYXV-infected cells, SERP-1 was detected as early as 8 h p.i. (under prolonged blot exposure time [data not shown]) and exhibited robust expression by 12 h p.i. SERP-1 production continued to increase in wild-type MYXV-infected cells reaching a maximum at 24 h p.i. (Fig. 3B). To confirm M062 ablation, after infection of vMyxM062-KO in rabbit kidney cell line RK-13, the expression of M062 mRNA could not be detected by reverse transcription, followed by real-time PCR (Fig. 3C).
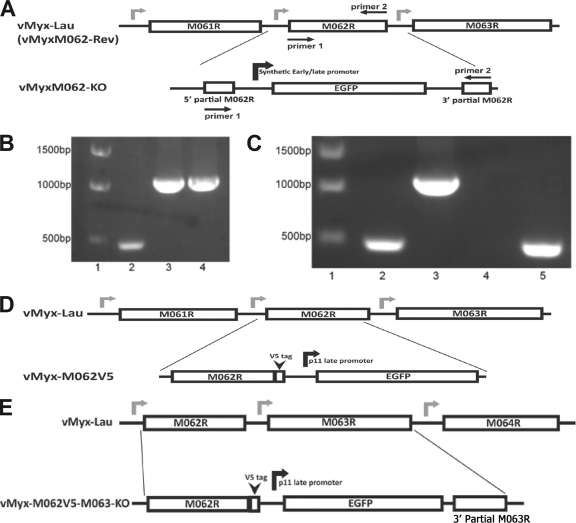
FIG. 1.
Construction and confirmations of recombinant MYXVs. (A) Construction of vMyxM062-KO. The central 78% of MYXV M062R sequence was replaced by an eGFP expression cassette under a viral early/late promoter using the three-element recombination of the MultiSite Gateway system. The revertant virus of vMyxM062-KO was constructed by replacing the eGFP expression cassette in the knockout virus with an intact M062R sequence. (B) Confirmation of purified vMyxM062-KO. PCR analysis using primers 1 and 2 (Table 3) designed to recognize M062R coding sequences (shown in panel A) will generate a 474-bp fragment with wild-type MYXV viral DNA as a template (lane 2) and a 1,077-bp fragment using either the recombination plasmid (lane 3) or purified vMyxM062-KO viral DNA (lane 4) as a template. The molecular markers are in lane 1. (C) Confirmation of the purified vMyxM062-Rev. The set of screening primers used in panel B was used to determine whether vMyxM062-Rev contained any contaminating vMyxM062-KO. Lane 1, molecular marker; lane 2, vMyx-Lau viral DNA; 3, vMyxM062-KO DNA; 4, no template DNA; 5, DNA of vMyxM062-Rev. (D) Construction of vMyx-M062V5. Using the MultiSite Gateway system, a V5 epitope sequence was inserted just before the stop codon of the M062R. In addition, a cassette containing a VACV p11 late promoter-driven eGFP expression was inserted following the V5-tagged M062R and in front of the intact M063R promoter region. Primers 1 and 3 (Table 3) were used for screening during the purification process. (E) The construction of vMyx-M062V5-M063-KO. MultiSite Gateway system was used to insert the V5 sequence before the stop codon of M062R, VACV p11 late promoter-driven eGFP, and 115 bp of the 3′-terminal sequence of M063R in front of an intact M064R promoter sequence. Primers 1 and 3 were used for screening during the process of purification.
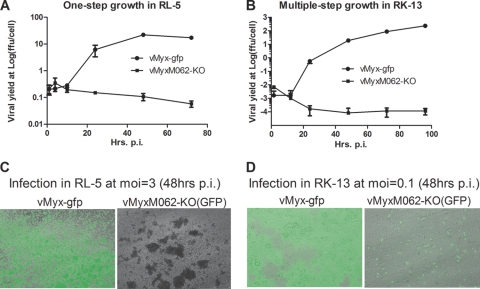
FIG. 2.
Infection by vMyxM062-KO in rabbit cells is abortive. (A) One-step growth curve of MYXV infection in RL-5 rabbit T cells. The rabbit CD4+ T cell line RL-5 was infected with either vMyx-gfp or vMyxM062-KO at an MOI of 3, and total cell lysates were collected at given time points (1, 4, 10, 24, 48, and 72 h p.i.). The viral yield at each time point was estimated by titrating on BSC-40 cells, and titrations were performed in triplicate to calculate the average titer at each time point, while an error bar represents the ± the standard deviation (SD). Shown is a representative one-step growth curve of three independent experiments. (B) Multiple-step growth of MYXV in RK-13 rabbit kidney cells. Either vMyx-gfp or vMyxM062-KO was used to infect RK-13 at an MOI of 0.1, at given time points (1.5, 12, 24, 48, 72, and 96 h p.i.) total cell lysates were harvested for titration as described in panel A. Shown is a representative multiple-step growth curve from three independent experiments. (C) Fluorescence microscope images of infected RL-5 cells. Images of infected cells from panel A were taken using an inverted fluorescence microscope (Leica DMI6000B) using a lens with ×10 magnification at 48 h p.i. The pictures shown are merged images from both GFP and bright fields. Left side, infection by vMyx-gfp; right side, infection by vMyxM062-KO. (D) Fluorescent images of infected RK-13 cells. At 48 h p.i., pictures of infected cells from panel B were taken under both GFP filter and bright field. Merged images are shown. Left side, infection by vMyx-gfp; right side, infection by vMyxM062-KO.
FIG. 3.
M062 is an early/late gene product and is essential for MYXV infection in rabbit cells. (A) Viral DNA is detected in RK-13 cells infected with vMyxM062-KO. RK-13 cells were infected with either vMyx-gfp or vMyxM062-KO at an MOI of 5, and cells were harvested at 2, 4, 8, 12, and 24 h p.i. Total DNA was extracted and real-time PCR was conducted using Sybr green real-time PCR system. Primer sets of MYXV M071L and the 18S ribosomal genes from O. cuniculus were designed to selectively amplify MYXV DNA and rabbit cellular DNA, respectively. Real-time PCR for M071L or 18S at each time point for each infection was conducted in triplicate. The average CT from the triplicate was used for the calculation of the ratio comparison. The comparative CT method or the 2−ΔΔCT method was used to calculate the relative fold increase (RFI) of viral DNA level during infection by vMyx-gfp or vMyxM062-KO. For each infection, the fold increase of sample (viral) DNA level at each time point was measured against the sample DNA level at 2 h p.i., which was set artificially as 1-fold. Curve shown is a summary of results from two independent experiments, and error bars represent ± the SD. (B) Infection of RK-13 cells with vMyxM062-KO led to an abortive infection with no late viral gene expression. RK-13 cells were mock treated (lane 1) or infected with either vMyx-gfp or vMyxM062-KO at an MOI of 5 in the absence or the presence of AraC. At 4, 8, 12, or 24 h p.i., monolayer cells were lysed to measure total protein concentration. Total protein (50 μg) from each sample was separated using 12% SDS-PAGE, transferred to a PVDF membrane, and analyzed by Western blotting. The membrane was first probed for an antibody raised against a late protein of MYXV (SERP-1), stripped, and probed for β-actin as a loading control. Lanes 1 and 11, molecular marker; lane 2, mock infection without AraC; lanes 3, 4, and 5, infection by vMyxM062-KO in the absence of AraC harvested at 4, 8, and 12 h p.i., respectively; lanes 6, 7, and 8, infection by vMyx-gfp in the absence of AraC harvested at 4, 8, 12 h p.i., respectively; lanes 9 and 10, infection by vMyx-gfp in the presence of AraC harvested at 8 and 12 h p.i., respectively; lanes 12 and 13, infection by vMyx-gfp and vMyxM062-KO, respectively, harvested at 24 h in the absence of AraC. (C) M062 mRNA is no longer expressed after vMyxM062-KO infection in rabbit cells. RK-13 cells were infected with either vMyx-gfp or vMyxM062-KO at an MOI of 5. At 1, 2, 4, 8, and 12 h p.i. for vMyx-gfp infection or at 2, 8, and 12 h p.i. for vMyxM062-KO infection or a mock infection sample (as controls for “no M062 cDNA”), cells were harvested to extract total RNA, and reverse transcription was conducted to produce cDNA. Sybr green real-time PCR was conducted to measure the M062 cDNA level using a primer set shown in Table 3, and O. cuniculus 18S cDNA was also measured as an internal control. Comparative CT method was used to calculate the level increase of M062 mRNA during vMyx-gfp infection, while no stable M062 mRNA can be detected at any time point after infection by vMyxM062-KO or in the mock-treated sample. (D) M062 protein is expressed as an early/late gene product during wild-type MYXV infection in rabbit cells. RK-13 cells were infected with vMyxM062V5 or mock infected at an MOI of 5 in the presence or absence of AraC. At 0, 1, 2, 4, 8, 12, and 24 h p.i. in the absence of AraC and at 8, 12, and 24 h p.i. in the presence of AraC, cells were harvested for analysis by Western blot, and 50 μg of total protein from each sample was separated by a 12% SDS-PAGE, transferred to a PVDF membrane, and analyzed by Western blotting with a V5 antibody. The blot was then stripped and probed for M063 (an early/late MYXV protein), M-T7 (an early/late viral protein), and SERP-1 (a late viral protein), respectively, and finally β-actin as loading control. Lanes 1 to 7, infection at 0, 1, 2, 4, 8, 12, and 24 h p.i., respectively, in the absence of AraC; lanes 8, 9, and 10, infection at 8, 12, and 24 h p.i., respectively, in the presence of AraC.
A recombinant M062-tagged MYXV virus, vMyx-M062V5, was constructed by inserting a V5 epitope tag sequence at the C-terminal end of M062R, followed by a cassette expressing eGFP driven by a poxvirus p11 late promoter (Fig. 1D). This virus behaves identically to wild-type vMyx-gfp in cultured cells (data not shown) and was used to determine the expression kinetics of M062 protein during infection of rabbit cells. As shown in Fig. 3D, V5-tagged M062 can be detected as early as 2 h p.i., and the expression persisted throughout the course of infection; the presence of AraC reduced protein levels of V5-tagged M062 at 8, 12, and 24 h p.i. Therefore, we conclude that M062 is an early/late gene product and is essential for productive MYXV infection in cultured rabbit cells.
M062 is essential for pathogenesis of MYXV in European rabbits and vMyxM062-KO does not vaccinate against subsequent lethal challenge with wild-type MYXV.
To determine the role of M062 in MYXV infection in vivo, we tested the pathogenesis of vMyxM062-KO in the European rabbit by the intradermal route of inoculation. For the present study, rabbits that were mock infected (PBS), infected with the wild-type parental MYXV Lausanne strain (vMyx-Lau), or infected with a revertant virus (vMyxM062-Rev) containing a rescued intact M062R gene sequence (Fig. 1A and C) were used as controls. Revertant virus-infected animals showed similar manifestations of myxomatosis (Table 1 and Fig. 4A) and showed the same mortality rate (100%) as that caused by the parental wild-type virus (Fig. 4B). Meanwhile, vMyxM062-KO caused no discernible lesion at the primary inoculation site, nor were any disease indications detected throughout the course of observation, and all animals infected with this virus survived the initial infection with no observable symptoms (Table 1 and Fig. 4A). This suggests an obligatory role for M062 in the infection and pathogenesis of MYXV in its susceptible European rabbit host. We next tested whether the aborted M062 knockout infection might function as a protective vaccine for myxomatosis caused by the wild-type MYXV strain Lausanne. Interestingly, when rabbits previously inoculated with vMyxM062-KO were challenged with the parental vMyx-Lau at 24 days after the initial infection, rabbits were not protected from the development of full-blown myxomatosis (Table 1). Although all animals showed some signs of recovery in primary and secondary lesions at a later time, only two of five rabbits survived the challenge with full recovery without respiratory stress. We conclude that the M062 knockout virus does not have the immune-inducing properties needed for a myxomatosis vaccine, presumably because the virus infection is aborted too quickly to stimulate a robust acquired immune response.
TABLE 1.
Role of M062 during in vivo pathogenesis by MYXV in European rabbits
| Infection group | Observation(s)a |
|
|---|---|---|
| vMyx-Lau (wt) or vMyxM062-Rev | vMyxM062-KO | |
| Initial infection (intradermal inoculation of 1,000 FFU) | At 2 to 3 dpi, there was a light pink area at the infection site; by 5 dpi, secondary lesions were visible; at 7 dpi, there were extensive primary and secondary lesions, animals' attitude was dull; at 10 to 11 dpi, there was severe respiratory stress and nasal discharge with reduced input/output and no or low activity (endpoint) | No signs of primary or secondary lesions were observed, and all animals remained healthy throughout the time of observation |
| Surviving animals from the initial infection were challenged intradermally with 1,000 FFU of vMyx-Lau (wt) | Challenge with vMyx-Lau took place at 24 dpi after the initial infection with vMyxM062KO. A full-blown infection occurred at up to 5 dpc, and there was no obvious protection in reducing primary or secondary lesion size or numbers. At 7 dpc, primary and secondary lesions started healing. At 10 dpc, three animals showed respiratory stress, while the other two animals fully recovered without respiratory complications | |
dpi, days postinfection; dpc, days postchallenge.
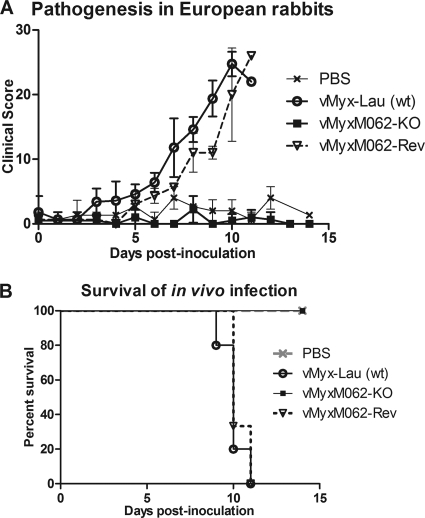
FIG. 4.
M062 is essential for MYXV pathogenesis in European rabbits. (A) Comparison of in vivo pathogenesis showed that vMyxM062-KO can no longer cause productive infection in rabbits. A total of 1,000 FFU of vMyx-Lau (n = 5), vMyxM062-KO (n = 4), or vMyxM062-Rev (n = 3) were diluted in 100 μl of PBS and inoculated intradermally into the left flank of European rabbits. Another group of rabbits (n = 3) were inoculated with PBS only. The overall physical condition and disease progression were evaluated daily in a clinical score system, and the average daily score was calculated. The error bar in the chart represents ± the SD within each group. (B) MYXV without M062 can no longer cause lethal myxomatosis. The results of the in vivo study described in panel A were analyzed for survival rates among the groups. The daily percentage of survival in each group was plotted to generate the survival curve.
M062 is essential for the replication of MYXV in most human cells.
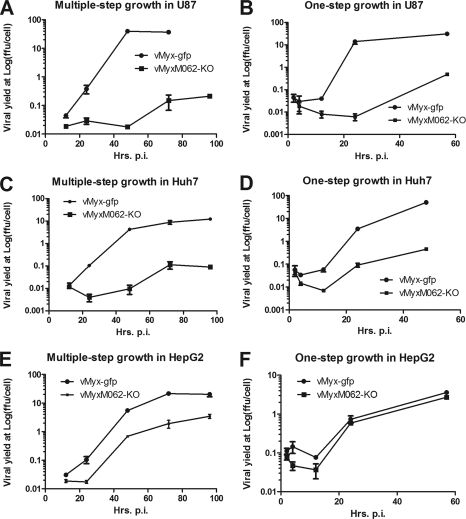
In addition to infecting rabbit cells, wild-type MYXV can also productively infect a broad variety of human cancer cells and is being developed as an oncolytic virotherapeutic to treat cancer (12). M062 has been predicted to be a host range factor belonging to the poxvirus C7L family, and it was previously shown to be a functional homolog of VACV C7, at least in vaccinia virus-infected cells (16). We next examined the function of M062 for MYXV replication in human cells by screening vMyxM062-KO in a spectrum of cancerous and nontumorigenic human cell lines that are permissive for wild-type MYXV infection. In the majority of the human cell lines tested, including cancerous (e.g., U87 glioma in Fig. 5A and B and HeLa cells [not shown]) or nontumorigenic cells, (e.g., MCF-10A [not shown]), M062R knockout virus could not conduct a productive infection. Furthermore, in most of these nonpermissive cell lines (e.g., U87), neither late gene product (see Fig. 8A) nor DNA replication (data not shown) could be detected during vMyxM062-KO infection. In a small subset of cell lines, such as Huh7 cells, although low levels of late gene expression were detected (Fig. 8D) and progeny virus were produced when they were infected with vMyxM062-KO (Fig. 5C and D), infection by the M062 knockout virus was so inefficient that consistently a 2-log-lower viral yield was observed than that of vMyx-gfp infection. Therefore, cells exhibiting no or minimal viral replication of just the M062 knockout virus were designated as type A. In contrast, in a smaller subset of cell lines, e.g., HepG2 cells, vMyxM062-KO replicated very similarly to the wild-type virus (vMyx-gfp) (Fig. 5E and F). Thus, we categorized these human cells into a second group, designated as type B, to denote cells that are permissive for both wild-type and M062R knockout viruses (Table 2). In summary, in type A cells, vMyxM062-KO cannot conduct a productive viral replication (associated with either the complete lack, or reduced levels, of late gene expression and progeny virus production), whereas infection by wild-type MYXV is permissive. Type B cells were characterized by a productive viral replication after either wild-type MYXV or vMyxM062-KO infection (Fig. 5E and F). One possible interpretation of these results is that vMyxM062-KO is antagonized to a greater degree than wild-type MYXV by specific antiviral responses that are operational in many, but not all, cultured human cells.
FIG. 5.
M062 is essential for MYXV infection in many diverse human cells. (A) Multiple-step growth curves of MYXV in U87 glioma cells. U87 cells were infected at an MOI of 0.1 with either vMyx-gfp or vMyxM062-KO. At given time points (12, 24, 48, and 72 h p.i. for both viruses and up to 96 h p.i. for vMyxM062-KO), total cell lysates were harvested for titration. Error bars represent ± the SD generated from titration in triplicate, and the results of a representative growth curve of two independent experiments are shown. (B) One-step growth curves of MYXV infection in U87 cells. U87 cells were infected at an MOI of 3 with vMyx-gfp or vMyxM062-KO. Cell lysates were harvested at 2, 4, 12, 24, and 57 h p.i. for titration, and error bars represent ± the SD generated from titration in triplicate. The results of a representative growth curve from two independent experiments are shown. (C) Multiple-step growth curves of MYXV in Huh7 cells. Experiments were carried out as described for panel A. (D) One-step growth curves of MYXV in Huh7. Experiments were carried out as described for panel B. (E) Multiple-step growth curves in HepG2 cells. Experiments were carried out as described for panel A. (F) One-step growth curves in HepG2 cells. Experiments were carried out as described for panel B.
TABLE 2.
Human cell lines that are permissive for parental wild-type MYXV categorized into two groups according to their permissiveness of infection by vMyxM062-KO
| Categories of human cells for vMyxM062-KO tropism | Description | Human cell types |
|---|---|---|
| Type A | Permissive for vMyx-gfp infection but nonpermissive for vMyxM062-KO | U87, HeLa, HEK293, A549, THP1, MCF-10A, U251, HOS, 786-0, MDA-MB231, PANC-1, CAL-27, SCC-25, HUVEC, HT29, Colo-205, GM02504, Huh7 |
| Type B | Permissive to infection by both vMyx-Lau and vMyxM062-KO | Hs766T, HepG2, HCT116 |
M062 forms a complex with another MYXV protein, M063, and this interaction enhances the binding to a human host protein, SAMD9.
We next investigated whether M062 interacted with any specific target proteins in human cells that might participate in antiviral response pathways and thus help explain why vMyx-M062-KO is uniquely nonpermissive in so many cultured human cells. The recombinant MYXV expressing a V5-tagged M062R was used to infect type A human cells (e.g., HeLa, U87, or Huh7) for the purpose of using co-IP and mass spectrometry to identify any potential protein binding partners of M062. Two distinct protein binding partners were consistently identified in the immunoprecipitates with V5-tagged M062. Mass spectrometry confirmed they were the human sterile alpha motif domain containing protein 9 (SAMD9) and the viral M063 of MYXV (see Fig. S1A and B in the supplemental material). A co-IP experiment was also performed using human anti-SAMD9 antibody after MYXV infection, and the presence of M062 and M063 was detected in the immunoprecipitates (Fig. 6C). The interaction between M062 and M063 in the absence of SAMD9 was also confirmed by infecting murine cells with either vMyxM062V5 or MYXV expressing a V5-tagged M063 (vMyxM063V5 described in supplemental Fig. 2A) (see Fig. S2B in the supplemental material). Murine cells were chosen for this analysis because SAMD9 is not expressed in cells of mouse origin due to evolutionary remodeling of the locus containing SAMD9 in the mouse genome (10). Co-IP comparison of M062 with other functional C7 family members (e.g., VACV C7L and M67R of Yaba-like disease virus [YLDV]) showed that the protein-protein interaction with human SAMD9 is unique to M062 (Fig. 6B), at least in the context of MYXV infection. When these proteins were provided in trans during infections with vMyxM062-KO, although with relatively reduced efficiency judged by smaller foci development (Fig. 6A), both VACV C7 and YLDV M67 could partially compensate the loss of M062 function in human cells. However, neither of them was able to rescue the defect of infection by vMyxM062-KO in RK-13 cells, as evidenced by the absence of focus formation (Fig. 6D, subpanel a) and late gene expression (Fig. 6D, subpanel b) compared to permissive viral replication caused by transfection of a plasmid expressing M062 after vMyxM062-KO infection.
FIG. 6.
M062 cannot be fully compensated for by other functional homologs of C7L family, and M062 has a unique interaction to a human host factor, SAMD9. (A) Transient expression of C7L family members can partially rescue vMyxM062-KO infection in human cells. HEK293 cells were infected with vMyxM062-KO at an MOI of 1 for 1 h, followed by transient transfection of the plasmids pYLDV-67R, pVACV-C7L, and pMYXV-M062R, respectively, or were mock transfected. The construction of these plasmids has been previously described (16). In these plasmids, the sequence of the VACV-C7L native promoter drives the expression of V5 tagged YLDV-67R, VACV-C7L, or MYXV-M062R. At 48 h posttransfection, pictures were taken using a fluorescence microscope with a lens of ×10 magnification under both a GFP filter and a bright field. Merged images are shown. (B) The interaction with human SAMD9 protein is unique and specific to M062 and does not occur with other functional homologs of the C7 family. Cell lysates from the experiment described in panel A were harvested at 72 h posttransfection. Before co-IP using the agarose-conjugated anti-V5 antibody, a 1/50 volume of total protein lysate from each sample was saved to prepare a Western blot to reveal the input human SAMD9 (bottom panel). Proteins associated with V5-tagged proteins were then separated on 10% SDS-PAGE, transferred to a PVDF membrane, and immunoblotted. The blot was first probed for human SAMD9 (top panel), stripped, and then probed for V5-tagged protein expression (middle panel). Lanes 1 to 4 represent cells infected with vMyxM062-KO and treated with mock transfection, transfected with pYLDV-67R, pVACV-C7L, or pMYXV-62R, respectively. (C) Antibody recognizing human SAMD9 for co-IP also confirms the interaction with M062 and M063. Human HCT116 cells were mock infected (lane 1), infected with vMyx-gfp (lane 2), or infected with vMyxM062V5 (lane 3) at an MOI of 1 for 72 h. Before co-IP, 1% volume of each cell lysate was used for Western blotting to show the input SAMD9. Proteins associated with SAMD9 were separated on 10% SDS-PAGE for immunoblotting as previously described for panel B. The blot was first probed for V5-tagged protein, stripped, and probed for M063 and then SAMD9. (D) In RK-13 rabbit cells, the expression of VACV-C7 or YLDV-67 did not compensate for the loss of M062R gene during the infection by vMyxM062-KO. RK-13 cells were infected with vMyxM062-KO at an MOI of 1 for 1 h before they were mock transfected or transfected with pMYXV-M062RV5, pVACV-C7LV5, or pYLDV-67RV5, respectively. At 72 h p.i., the cells were examined by using a fluorescence microscope, and images were taken using a lens of ×10 magnification. Merged green fluorescent and bright-field images are shown in subpanel a. The total protein (50 μg) was separated by 12% SDS-PAGE, transferred to a PVDF membrane, and analyzed by Western blotting. The membrane was probed first using anti-SERP-1 antibody, stripped, and then probed to monitor V5-tagged protein expression (not shown). At last, the membrane was stripped and probed for β-actin expression as a loading control. The results of Western blot analyses are shown in subpanel b. Lane 1, RK-13 cells were infected with vMyxM062-KO and mock transfected; lanes 2 to 4, RK-13 cells were infected with vMyxM062-KO and transfected with pMYXV-M062RV5, pVACV-C7LV5, or pYLDV-67RV5, respectively; lane 5, HeLa cells were infected with vMyxM062-KO at an MOI of 1 for 72 h as a positive control for SERP-1 expression.
In transiently transfected human cells (HeLa) in the absence of MYXV infection, expressed M062 binds to endogenous SAMD9 protein while M063 does not (Fig. 7A). However, when M063R is knocked out from MYXV genome, the ability of M062 in binding to human SAMD9 is significantly reduced but not eliminated (Fig. 7B).
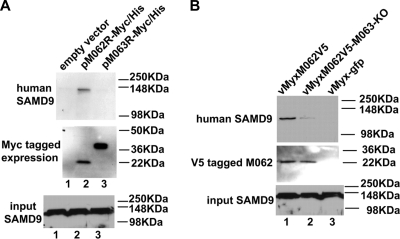
FIG. 7.
In human cells M062, but not M063, binds to endogenous SAMD9 protein, but the presence of M063 during MYXV infection enhances the interaction of M062 and SAMD9. (A) M062 binds to human SAMD9. HeLa cells were transfected with empty vector (pCDNA3.1-Myc/His) (lane 1), pM062R-Myc/His (lane 2), or pM063R-Myc/His (lane 3) for 48 h. Cells were lysed, and before a pull-down experiment using Ni-NTA beads was constructed, 1/50 volume of the cell lysate was saved for Western blotting to show the input SAMD9 (bottom panel). Proteins associated with the indicated transfected His-tagged proteins were pulled down, washed, and eluted from the beads. The eluted protein was separated by 10% SDS-PAGE and transferred to a PVDF membrane for immunoblotting. The membrane was first probed with anti-human SAMD9 antibody, stripped, and then probed with anti-myc antibody for myc-tagged protein expression. The SAMD9 protein was detected at a molecular mass of ∼148 kDa only in cells transfected with pM062R-Myc/His (top panel). The M063-myc tagged protein was detected at the observed molecular mass of 29.2 kDa, while the M062-myc tagged protein was detected at 23.7 kDa (middle panel). (B) The expression of M063 protein enhances the binding between M062 and SAMD9 in human cells during MYXV infection. U87 cells were infected with vMyxM062V5 (lane 1), vMyxM062V5-M063-KO (lane 2), or vMyx-gfp (lane 3) at an MOI of 1 for 48 h. Cells were harvested and, before co-IP using anti-V5 antibody was performed, a 1/50 volume of total cell lysate was saved for Western blotting to show SAMD9 input (bottom panel). Precipitated proteins were separated by 8% SDS-PAGE and transferred for an immunoblot procedure. The blot was first probed for SAMD9 expression (top panel). After stripping, it was then probed with anti-V5 antibody (middle panel).
We conclude that (i) M062 function in MYXV infection in rabbit cells cannot be compensated by other C7 functional homologs; (ii) in human cells, the loss of M062's function can only partially be compensated for by other C7 functional poxviral homologs in the context of MYXV infection; and (iii) binding to human SAMD9 is a unique feature of M062, and this interaction can be increased (but not mediated solely) by M063. Therefore, we hypothesize that M062 is a multifunctional host range factor and that human SAMD9 is the major host factor targeted by M062. We next tested the model that, in human cells (and presumably in rabbit cells as well), M062 functions by antagonizing SAMD9 during MYXV infection.
Knocking down SAMD9 protein expression relieves a block on vMyxM062-KO infection in human cells.
Human SAMD9 protein can be detected in all of the human cells tested, including cancerous, nontumorigenic (Table 2), and primary cells (e.g., primary human macrophages and GM02504) (data not shown). The major protein product detected for SAMD9 here migrated on SDS-PAGE with an observed molecular mass of ∼148 kDa. A minor fraction of the protein product, visible only after longer exposure of the autoradiogram of a Western blot, was also detected with a molecular mass between 148 and 250 kDa. These observations were consistent with SAMD9 predicted protein products of approximately 149 and 184 kDa, respectively, resulting from alternative splicing of the full-length transcript (10). Knocking down endogenous SAMD9 expression by transient transfection of an siRNA pool targeting human SAMD9 resulted in only a partial reduction in the detectable SAMD9 protein level (Fig. 8C, top panel, and see Fig. S4A in the supplemental material). However, even this modest reduction in SAMD9 protein level caused a significant increase in vMyxM062-KO replication in cell lines such as Huh7 (Fig. 8C and D; see also Fig. S4B in the supplemental material), Hs766T (data not shown), and HCT116 (data not shown). In order to achieve higher levels of gene expression knockdown, a lentivirus containing shRNAs targeting SAMD9 was used to construct cell lines with constitutively reduced SAMD9 protein levels (see Fig. S3B in the supplemental material). In SAMD9-knockdown U87 or Huh7 cells, no increase in wild-type MYXV viral replication can be detected (data not shown). However, in SAMD9 shRNA-transduced U87 human glioma cells, relief of the block to late viral gene expression can be observed specifically after vMyxM062-KO infection (Fig. 8A). Ultimately, this constitutive reduction of SAMD9 expression led to the partial rescue of vMyxM062-KO infection in the previously nonpermissive cell line (Fig. 8B). In addition, similar effects on vMyxM062-KO infection in Huh7 cells were observed following comparable constitutive SAMD9 knock down (Fig. 8C). However, in these constructed cell lines with significant but incomplete knockdown in endogenous SAMD9 levels, vMyxM062-KO viral replication was not fully restored to the same level as that of the wild-type MYXV (compare Fig. 5B to Fig. 8B). In conclusion, M062 is essential for permissive MYXV replication in the majority of human cells tested, and one of M062's functions is to antagonize the antiviral effects of human SAMD9 to ensure a productive viral infection.
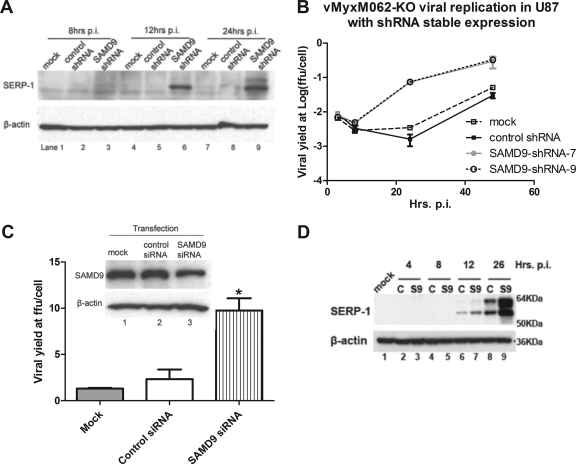
FIG. 8.
Reducing the expression of endogenous human SAMD9 can rescue vMyxM062-KO infection in nonpermissive cells. (A) Knocking down the expression of endogenous SAMD9 in U87 can rescue late viral gene expression following vMyxM062-KO infection. U87 lines with constitutive SAMD9 knockdown were constructed by infecting parental U87 with a lentivirus packaged with shRNAs targeting SAMD9 mRNA and by selecting for transduced clones with the highest knockdown at the protein level. In this experiment, the number 9 clone, designated U87 SAMD9-shRNA-9, was used. A control U87 cell line, designated U87 control-shRNA, was constructed by infecting U87 cells with a lentivirus packaged with control shRNAs and selecting for stably transduced cells. Parental U87, U87 control shRNA, and U87 SAMD9-shRNA-9 were infected with vMyxM062-KO at an MOI of 5. Cell lysates were harvested at 8, 12, and 24 h p.i., and 25 μg of total protein of each sample was separated by 12% SDS-PAGE and then analyzed by Western blotting. The blot was first probed against SERP-1, and after stripping it was probed against β-actin as a loading control. (B) Cell lines with reduced SAMD9 expression showed rescue of productive infection by vMyxM062-KO. One-step growth of vMyxM062-KO was conducted in parental U87, U87 control-shRNA, U87 SAMD9-shRNA-7, and U87 SAMD9-shRNA-9 at an MOI of 5. At 3, 8, 24, and 48 h p.i., total cell lysates were collected for titration. Error bars represent ± the SD calculated from infections performed in triplicate. (C) Transient knockdown of endogenous SAMD9 improved the M062 knockout MYXV yield in Huh7 cells. Huh7 cells were treated in triplicate as follows: (i) transiently transfected with control siRNA (lane 2) or (ii) siRNAs targeting human SAMD9 (final concentration of 100 nM) (lane 3) or (iii) treated with only transfection reagent (mock) (lane 1) for 48 h before infection with vMyxM062-KO at an MOI of 3 for 48 h. Cell lysates were harvested for titration, and error bars represent ± the SD calculated from viral yields in triplicate in each treatment. One-way analysis of variance was used to analyze the statistical significance among the different viral yields in each group (P = 0.0016) and Tukey's multiple-comparison test was conducted to conclude that viral yield from SAMD9 siRNA-transfected group (labeled with an asterisk [*]) is significantly different from the other two groups. (D) Late gene expression of MYXV after infection increased by knocking down SAMD9 expression in Huh7 cells. Huh7 cells were infected with a lentivirus packaged with either a control shRNA or with SAMD9 shRNAs, followed by puromycin selection to enrich for the transduced cells. Significant knockdown of SAMD9 was observed in cells transduced with SAMD9 shRNA (labeled as “S9”), while the SAMD9 protein level is intact in control shRNA transduced cells (labeled as “C”). Infection by vMyxM062-KO was conducted at an MOI of 5 in both Huh7-S9 (SAMD9 knockdown) and Huh7-C (control shRNA) cells. At 4, 8, 12, and 26 h p.i., cell lysates were collected, along with cell lysate from the mock-infected cells (Huh7-C) (lane 1), and 25 μg of total protein was separated by 12% SDS-PAGE and analyzed by Western blotting. The blot was first probed for SERP-1 (top panel), stripped, and then probed for β-actin (bottom panel).
DISCUSSION
Viral infection by vMyxM062-KO in rabbit cells is abortive, which explains the asymptomatic infection of European rabbits at the primary site of inoculation. This abortive infection by vMyxM062-KO in rabbits must have been cleared very rapidly because very little effective acquired immunity was generated. Rabbits that were previously infected with vMyxM062-KO, when challenged with a lethal dose of wild-type MYXV, were provided only a minor protection against the development of full-blown myxomatosis. Although starting at 7 days postchallenge infection, primary and secondary lesions had started to heal in all of the wild-type MYXV-challenged animals, suggesting some suboptimal protective effects from the vaccination, the majority (three of five) of animals developed respiratory stress and had to be sacrificed. The disease progression after challenge is very similar to the vaccination effect by UrΔM063, a M063R-inactivated MYXV constructed in strain Uriarra (1).
An extensive defect in vMyxM062-KO viral replication can be observed not only in rabbit cells but also in a wide spectrum of cultured cell lines from other species. For example, BGMK monkey cells have been a standard cell line in our lab for MYXV stock preparation, because they are so permissive for MYXV infection; however, vMyxM062-KO cannot infect these cells productively and does not even progress to the stage of late gene expression (not shown). In addition, vMyxM062-KO only produces small foci when infecting Vero cells, another standard monkey cell line commonly used for MYXV stock preparations (not shown). The phenotype of vMyxM062-KO infection in human cells is also very distinct from all other previously characterized MYXV targeted gene knockout viruses (8). For example, vMyxM062-KO is dramatically different from the phenotype of the M063R-knockout virus (vMyxM063-KO), which demonstrated no difference in the ability to infect human cancer cells compared to wild-type MYXV (4). Here, we report that M062 and M063 form a heteromeric protein complex during MYXV infection in human and murine cells, as well as in rabbit cells (data not shown). Despite the obvious sequence similarity between these two related host range genes and the fact that M062 and M063 proteins bind to each other, these two host range factors nevertheless play distinct roles during viral replication. When MYXV infects rabbit cells, the expression of both M062 and M063 is required for a productive viral replication. However, when infecting human cells, including a wide spectrum of human cancer cells, only M062 is essential for permissive viral replication. Previously, it was shown that M062 is a functional homolog of VACV C7 (16), at least in the context of a VACV infection, whereas M063 fails to compensate for the C7 function in human or murine cell lines. These observations suggest that the existence of both M062 and M063 to form a complex in the MYXV infection may be an evolutionary consequence of MYXV specifically adapting to the rabbit host.
In the present study, we also identified a human host protein, designated SAMD9, which binds to M062, albeit with higher efficiency in the presence of M063, during MYXV infection of human cells. In chromosome 10 of Oryctolagus cuniculus (Thorbecke inbred), a predicted gene sequence (NCBI reference XR_085081.2) with an identity of 86% similar to human SAMD9 can be found located in between sterile α domain containing 9-like (SAMD9L) and cyclin-dependent kinase 6 (CDK6), an gene layout identical to that of SAMD9 in the human genome. Co-IP of V5-tagged M062 from infected rabbit (RK-13) cells resulted in far more pull-down proteins in addition to M063 by silver staining than those can be detected from human cells; when Western blot was used with the intention of probing rabbit SAMD9 from a similar co-IP experiment using a polyclonal antibody generated by an immunogen of the N-terminal 82 amino acids of human SAMD9, a negative result was observed. The sequence alignment of the 82-amino-acid polypeptide showed only 75% similarity to the predicted SAMD9 sequence of Oryctolagus cuniculus breed Thorbecke inbred. Due to the difficulty of investigating SAMD9 in the rabbit background, we focused on human SAMD9 here.
Human SAMD9 was reported to play a key role in a human inflammatory disease called normophosphatemic familial tumoral calcinosis (NFTC) (6, 23). In these studies, a missense mutation responsible for the degradation of SAMD9 and a nonsense mutation leading to a truncation of the protein were identified to be the causative genetic sources of NFTC among patients from families in a well-isolated ethnic group. Patients with NFTC showed prominent inflammatory manifestations accompanied with dystrophic calcinosis (18), suggesting that SAMD9 may play a role in innate inflammatory responses. Human SAMD9 is a cytoplasmic protein ubiquitously expressed in various tissues, and its expression can be further upregulated either by tumor necrosis factor (TNF) signaling through p38 (6) or by interferons (IFNs) (7, 22). In addition, human SAMD9 was shown to regulate cell proliferation and to suppress tumor growth; interestingly, the SAMD9 gene has been lost in the mouse genome due to an evolutionary rearrangement of the chromosomes (10). However, little is known about the physiological function of this protein in human cells. A recent study suggested a cloned short fragment of the N-terminal sequence of human SAMD9 may interact with human RGL2 (7). In the present study, we first determined the interaction between a MYXV host range factor, M062, and the full-length human SAMD9 and suggest a potential antiviral role for this host protein. When we compared the co-IP products from cells nonpermissive to vMyxM062-KO with either infection of vMyxM062V5 only or infection with vMyxM062-KO, followed by transient transfection of tagged M062 at 1 h p.i., we observed an intact slower migrating band of SAMD9 in the wild-type virus-infected cells (see Fig. S3A, top panel, in the supplemental material) that coimmunoprecipitated with M062 expressed from the viral genome. However, an extensive degradation pattern of SAMD9 protein was detected in cells infected with M062 knockout virus and subsequently transfected with ectopic V5-tagged M062 (Fig. 6B, top panel). This suggests a putative role of M062 in sequestering full-length unprocessed SAMD9 and preventing downstream modifications and/or cleavages that would otherwise participate in virus-induced responses that function to antagonize MYXV infection. Further proteomic analysis of the SAMD9 species that interact with M062 will help to more definitively address these questions. The full-length human SAMD9 protein (1,589 amino acids) contains an amino-terminal SAM domain (65 amino acids) but lacks sequence similarity to any other known protein families or protein motifs. The protein-protein interaction between the viral M062 and host SAMD9 provides a model system to further investigate the functional role of this cellular protein as an innate or intrinsic antiviral defense factor in human cells. The fact that SAMD9 can be upregulated by both TNF and IFNs also further suggests a role for this protein in antiviral defenses of human cells.
In modified VACV Ankara (MVA) infection of human cells, C7 was suggested to inhibit protein kinase R-mediated phosphorylation of eIF2α (2); however, wild-type MYXV infection in human cells triggered the phosphorylation of eIF2α and there was no significant difference in the presence or absence of M062 (data not shown). This suggests that very different cell signaling interactions are mediated by MVA versus MYXV infection in human cells. It was noted that although M062 can compensate for the loss of VACV C7 in VACV-infected cells, C7, or another functional homolog, YLDV-67, only has the ability to partially rescue infection by vMyxM062-KO in human cells but does so at much lower efficiency. The fact that neither C7 nor YLDV-67 binds to human SAMD9 in the context of MYXV infection suggests a novel antiviral signaling pathway where SAMD9 is but one player in a virus-induced response cascade. Our observations also suggest that multiple functions are possessed by M062: an essential role in initiating later stages of the viral life cycle and the host range function which antagonizes the host antiviral defense mechanism mediated by SAMD9. When SAMD9 expression was knocked down, this only produced a partial rescue of vMyxM062-KO infection in several type A human cell lines tested. However, we could never achieve full SAMD9 knockdown, either transiently with siRNA or constitutively in shRNA-transduced cell lines, and thus could not fully restore the replication efficiency of the M062 knockout virus to that of wild-type levels. We observed that the constructed cell lines with constitutive SAMD9 knockdown provided no obvious benefit in improving viral replication by either wild-type MYXV or wild-type VACV (data not shown). It is possible that the antiviral effects of SAMD9, or the antiviral pathway it presumably belongs to, functions in response to a variety of indirect triggering stimuli instead of just the basal level of viral replication. Furthermore, in type B cells, siRNA knockdown of SAMD9 had different impacts on vMyxM062-KO viral replication: in HCT116 and Hs766T, this knockdown increased vMyxM062-KO replication, while in HepG2 a comparable level of knockdown in SAMD9 protein level showed no significant effect on vMyxM062-KO replication. We therefore hypothesize that the antiviral pathway, in which SAMD9 plays a role, has inherent defects in these type B cells such that the pathway is no longer functional in antagonizing vMyxM062-KO infection. However, the very defect in these cells may well be different from each other, which caused the different outcome from HepG2 to HCT116 and Hs766T when the SAMD9 protein level is reduced. We are further investigating the difference in cellular signaling in these three cell lines to understand the molecular antiviral functions of SAMD9.
At this point, we presume that SAMD9 will also play a role in battling infections by other pathogens. Therefore, we are further studying the SAMD9 pathway functions in the presence of various cellular stress signals, while screening for the role of SAMD9 in cellular responses to other pathogens as well.
Supplementary Material
TABLE 3.
Real-time PCR and screening primer sequences
| Primer | Sequence (5′-3′) |
|---|---|
| Screening primer | |
| Primer 1-M062R forward | ATGGGCGTGCAACACAAATTGGA |
| Primer 2-M062R reverse | CGAATGTGTTCTATCCTCGTACCA |
| Primer 3-M063R reverse | ATCTTCTTCCTCCTCCGTGTCTT |
| Real-time PCR | |
| M062R forward | ATCGTATTGGAACCGGAGTG |
| M062R reverse | GCCGAATAGATGCGTTTGAT |
| M071L forward | CCTCGGAATCCAGAAACAGT |
| M071L reverse | TCAAGCAACGTCGTATCGTC |
| O.Cuniculus18S forward | TGACTCAACACGGGAAACCT |
| O.Cuniculus18S reverse | GACAAATCGCTCCACCAACT |
Acknowledgments
Our laboratory is funded by NIH grants AI080607 and CA13854 and the Bankhead Coley Foundation.
We thank Yan Xiang for sharing plasmids encoding V5-tagged homologs of C7L family, Jason Liem for the technique support in the electronic formatting and software usage, and Westley Reeves for sharing his real-time PCR equipment for this study.
Footnotes
Published ahead of print on 19 January 2011.
Supplemental material for this article may be found at http://jvi.asm.org/.
REFERENCES
- 1.Adams, M. M., B. H. van Leeuwen, G. McFadden, and P. J. Kerr. 2008. Construction and testing of a novel host-range defective myxoma virus vaccine with the M063 gene inactivated that is non-permissive for replication in rabbit cells. Vet. Res. 39:60. [DOI] [PubMed] [Google Scholar]
- 2.Backes, S., et al. 2010. Viral host-range factor C7 or K1 is essential for modified vaccinia virus Ankara late gene expression in human and murine cells, irrespective of their capacity to inhibit protein kinase R-mediated phosphorylation of eukaryotic translation initiation factor 2α. J. Gen. Virol. 91:470-482. [DOI] [PubMed] [Google Scholar]
- 3.Barrett, J. W., et al. 2007. Identification of host range mutants of myxoma virus with altered oncolytic potential in human glioma cells. J. Neurovirol. 13:549-560. [DOI] [PubMed] [Google Scholar]
- 4.Barrett, J. W., et al. 2007. Myxoma virus M063R is a host range gene essential for virus replication in rabbit cells. Virology 361:123-132. [DOI] [PubMed] [Google Scholar]
- 5.Barry, M., S. F. Lee, L. Boshkov, and G. McFadden. 1995. Myxoma virus induces extensive CD4 downregulation and dissociation of p56lck in infected rabbit CD4+ T lymphocytes. J. Virol. 69:5243-5251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Chefetz, I., et al. 2008. Normophosphatemic familial tumoral calcinosis is caused by deleterious mutations in SAMD9, encoding a TNF-alpha responsive protein. J. Invest. Dermatol. 128:1423-1429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hershkovitz, D., et al. 2011. Functional characterization of SAMD9, a protein deficient in normophosphatemic familial tumoral calcinosis. J. Invest. Dermatol., 131:662-669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Johnston, J. B., and G. McFadden. 2004. Technical knockout: understanding poxvirus pathogenesis by selectively deleting viral immunomodulatory genes. Cell Microbiol. 6:695-705. [DOI] [PubMed] [Google Scholar]
- 9.Lee, P. Y., et al. 2008. TLR7-dependent and FcγR-independent production of type I interferon in experimental mouse lupus. J. Exp. Med. 205:2995-3006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Li, C. F., et al. 2007. Human sterile alpha motif domain 9, a novel gene identified as down-regulated in aggressive fibromatosis, is absent in the mouse. BMC Genomics 8:92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Liu, J., et al. 2008. Reduction in severity of a herpes simplex virus type 1 murine infection by treatment with a ribozyme targeting the UL20 gene RNA. J. Virol. 82:7467-7474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Liu, J., S. Wennier, and G. McFadden. 2010. The immunoregulatory properties of oncolytic myxoma virus and their implications in therapeutics. Microbes Infect. 12:1144-1152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Liu, J., et al. 2009. Myxoma virus expressing interleukin-15 fails to cause lethal myxomatosis in European rabbits. J. Virol. 83:5933-5938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lun, X., et al. 2005. Myxoma virus is a novel oncolytic virus with significant antitumor activity against experimental human gliomas. Cancer Res. 65:9982-9990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Magnani, E., L. Bartling, and S. Hake. 2006. From Gateway to MultiSite Gateway in one recombination event. BMC Mol. Biol. 7:46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Meng, X., J. Chao, and Y. Xiang. 2008. Identification from diverse mammalian poxviruses of host-range regulatory genes functioning equivalently to vaccinia virus C7L. Virology 372:372-383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Meng, X., et al. 2009. Vaccinia virus K1L and C7L inhibit antiviral activities induced by type I interferons. J. Virol. 83:10627-10636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Metzker, A., B. Eisenstein, J. Oren, and R. Samuel. 1988. Tumoral calcinosis revisited—common and uncommon features: report of ten cases and review. Eur. J. Pediatr. 147:128-132. [DOI] [PubMed] [Google Scholar]
- 19.Nash, P., et al. 2000. Post-translational modification of the myxoma-virus anti-inflammatory serpin SERP-1 by a virally encoded sialyltransferase. Biochem. J. 347:375-382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ramsey-Ewing, A. L., and B. Moss. 1996. Complementation of a vaccinia virus host-range K1L gene deletion by the nonhomologous CP77 gene. Virology 222:75-86. [DOI] [PubMed] [Google Scholar]
- 21.Smallwood, S. E., M. M. Rahman, D. W. Smith, and G. McFadden. 2010. Myxoma virus: propagation, purification, quantification, and storage. Curr. Protoc. Microbiol. Chapter 14:Unit 14A1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Tanaka, M., T. Shimbo, Y. Kikuchi, M. Matsuda, and Y. Kaneda. 2010. Sterile alpha motif containing domain 9 is involved in death signaling of malignant glioma treated with inactivated Sendai virus particle (HVJ-E) or type I interferon. Int. J. Cancer 126:1982-1991. [DOI] [PubMed] [Google Scholar]
- 23.Topaz, O., et al. 2006. A deleterious mutation in SAMD9 causes normophosphatemic familial tumoral calcinosis. Am. J. Hum. Genet. 79:759-764. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.