Abstract
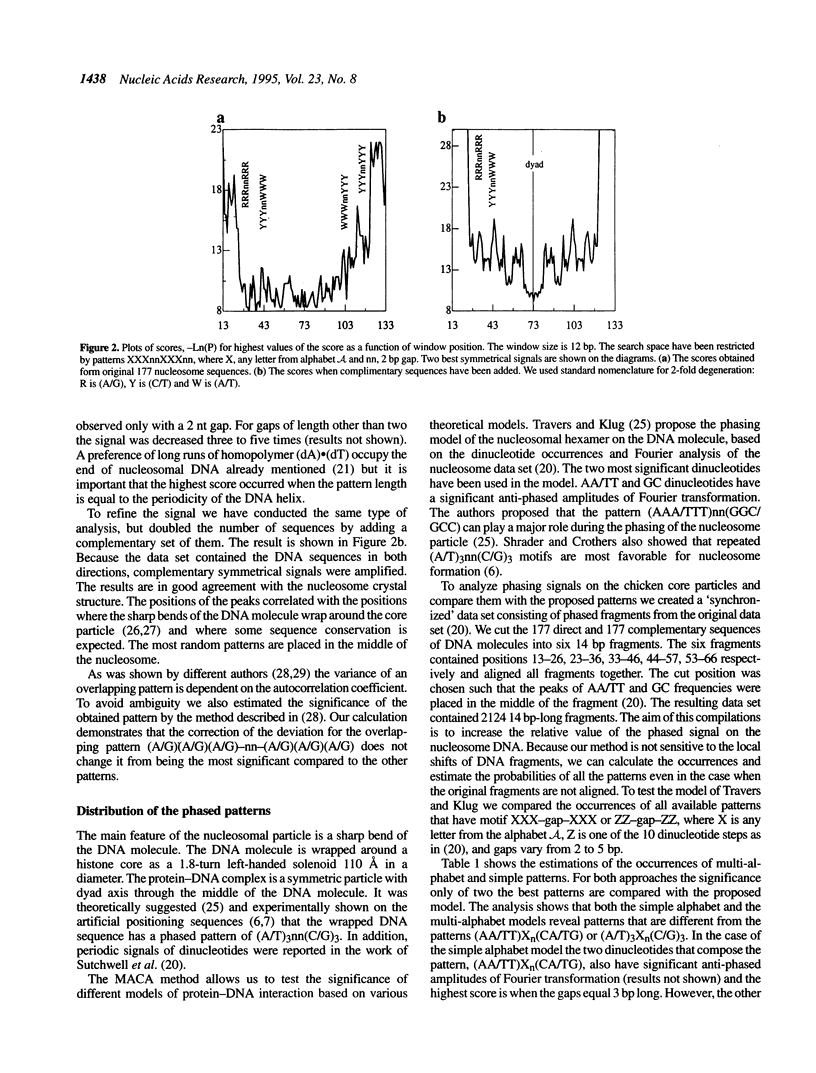
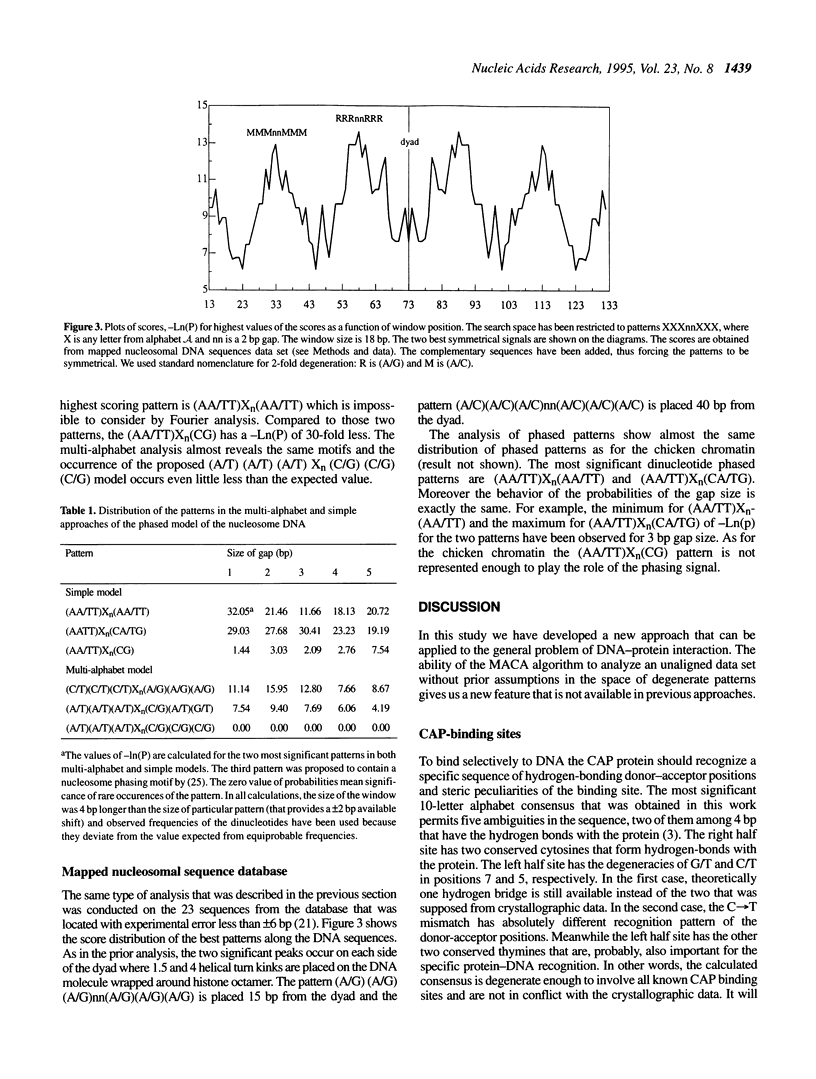
A method for the identification and characterization of protein-DNA interactions is presented. We have developed an approach for finding unknown multiple patterns that occur imperfectly in a set of several sequences. The pattern may contain letters from the nucleotide alphabet (A, C, G and T) including ambiguous characters (A/C, A/G, A/T; A/C/G, etc.). This method reveals weak DNA signals on an unaligned set of DNA fragments known to be functionally related and assumes no prior information on the sequences' alignment. It determines the locations of the signals from only the information intrinsic to the sequences themselves. We have applied this method to analyze the binding sites of cAMP receptor protein (CRP). The consensus based on these data are discussed and a comparison of the consensus with the crystal structure of CAP-DNA complex is presented. We further show that in a mixture of DNA sequences, containing binding sites for two different proteins, both classes of binding sites can be discovered simultaneously by this method. The DNA sequences of nucleosome cores from chicken erythrocyte and a set of the other known nucleosomal sequences show existence of symmetrical features in nucleosome-binding DNA sequences. We also show multi-alphabet patterns that can play a role in the phasing signal on the nucleosome DNA molecule and have compared the results with existing models of nucleosome positioning.
Full text
PDF




Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arratia R., Gordon L. Tutorial on large deviations for the binomial distribution. Bull Math Biol. 1989;51(1):125–131. doi: 10.1007/BF02458840. [DOI] [PubMed] [Google Scholar]
- Berg O. G., von Hippel P. H. Selection of DNA binding sites by regulatory proteins. II. The binding specificity of cyclic AMP receptor protein to recognition sites. J Mol Biol. 1988 Apr 20;200(4):709–723. doi: 10.1016/0022-2836(88)90482-2. [DOI] [PubMed] [Google Scholar]
- Berg O. G., von Hippel P. H. Selection of DNA binding sites by regulatory proteins. Statistical-mechanical theory and application to operators and promoters. J Mol Biol. 1987 Feb 20;193(4):723–750. doi: 10.1016/0022-2836(87)90354-8. [DOI] [PubMed] [Google Scholar]
- Drew H. R., Calladine C. R. Sequence-specific positioning of core histones on an 860 base-pair DNA. Experiment and theory. J Mol Biol. 1987 May 5;195(1):143–173. doi: 10.1016/0022-2836(87)90333-0. [DOI] [PubMed] [Google Scholar]
- Galas D. J., Eggert M., Waterman M. S. Rigorous pattern-recognition methods for DNA sequences. Analysis of promoter sequences from Escherichia coli. J Mol Biol. 1985 Nov 5;186(1):117–128. doi: 10.1016/0022-2836(85)90262-1. [DOI] [PubMed] [Google Scholar]
- Hertz G. Z., Hartzell G. W., 3rd, Stormo G. D. Identification of consensus patterns in unaligned DNA sequences known to be functionally related. Comput Appl Biosci. 1990 Apr;6(2):81–92. doi: 10.1093/bioinformatics/6.2.81. [DOI] [PubMed] [Google Scholar]
- Hogan M. E., Rooney T. F., Austin R. H. Evidence for kinks in DNA folding in the nucleosome. Nature. 1987 Aug 6;328(6130):554–557. doi: 10.1038/328554a0. [DOI] [PubMed] [Google Scholar]
- Ioshikhes I., Trifonov E. N. Nucleosomal DNA sequence database. Nucleic Acids Res. 1993 Oct 25;21(21):4857–4859. doi: 10.1093/nar/21.21.4857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kleffe J., Borodovsky M. First and second moment of counts of words in random texts generated by Markov chains. Comput Appl Biosci. 1992 Oct;8(5):433–441. doi: 10.1093/bioinformatics/8.5.433. [DOI] [PubMed] [Google Scholar]
- Lawrence C. E., Reilly A. A. An expectation maximization (EM) algorithm for the identification and characterization of common sites in unaligned biopolymer sequences. Proteins. 1990;7(1):41–51. doi: 10.1002/prot.340070105. [DOI] [PubMed] [Google Scholar]
- Mengeritsky G., Smith T. F. Recognition of characteristic patterns in sets of functionally equivalent DNA sequences. Comput Appl Biosci. 1987 Sep;3(3):223–227. doi: 10.1093/bioinformatics/3.3.223. [DOI] [PubMed] [Google Scholar]
- Pevzner P. A., Borodovsky MYu, Mironov A. A. Linguistics of nucleotide sequences. I: The significance of deviations from mean statistical characteristics and prediction of the frequencies of occurrence of words. J Biomol Struct Dyn. 1989 Apr;6(5):1013–1026. doi: 10.1080/07391102.1989.10506528. [DOI] [PubMed] [Google Scholar]
- Raibaud O., Vidal-Ingigliardi D., Richet E. A complex nucleoprotein structure involved in activation of transcription of two divergent Escherichia coli promoters. J Mol Biol. 1989 Feb 5;205(3):471–485. doi: 10.1016/0022-2836(89)90218-0. [DOI] [PubMed] [Google Scholar]
- Richmond T. J., Finch J. T., Rushton B., Rhodes D., Klug A. Structure of the nucleosome core particle at 7 A resolution. Nature. 1984 Oct 11;311(5986):532–537. doi: 10.1038/311532a0. [DOI] [PubMed] [Google Scholar]
- Satchwell S. C., Drew H. R., Travers A. A. Sequence periodicities in chicken nucleosome core DNA. J Mol Biol. 1986 Oct 20;191(4):659–675. doi: 10.1016/0022-2836(86)90452-3. [DOI] [PubMed] [Google Scholar]
- Schultz S. C., Shields G. C., Steitz T. A. Crystal structure of a CAP-DNA complex: the DNA is bent by 90 degrees. Science. 1991 Aug 30;253(5023):1001–1007. doi: 10.1126/science.1653449. [DOI] [PubMed] [Google Scholar]
- Shrader T. E., Crothers D. M. Artificial nucleosome positioning sequences. Proc Natl Acad Sci U S A. 1989 Oct;86(19):7418–7422. doi: 10.1073/pnas.86.19.7418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shrader T. E., Crothers D. M. Effects of DNA sequence and histone-histone interactions on nucleosome placement. J Mol Biol. 1990 Nov 5;216(1):69–84. doi: 10.1016/S0022-2836(05)80061-0. [DOI] [PubMed] [Google Scholar]
- Simpson R. T. Nucleosome positioning: occurrence, mechanisms, and functional consequences. Prog Nucleic Acid Res Mol Biol. 1991;40:143–184. doi: 10.1016/s0079-6603(08)60841-7. [DOI] [PubMed] [Google Scholar]
- Stormo G. D. Consensus patterns in DNA. Methods Enzymol. 1990;183:211–221. doi: 10.1016/0076-6879(90)83015-2. [DOI] [PubMed] [Google Scholar]
- Stormo G. D., Hartzell G. W., 3rd Identifying protein-binding sites from unaligned DNA fragments. Proc Natl Acad Sci U S A. 1989 Feb;86(4):1183–1187. doi: 10.1073/pnas.86.4.1183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stormo G. D., Strobl S., Yoshioka M., Lee J. S. Specificity of the Mnt protein. Independent effects of mutations at different positions in the operator. J Mol Biol. 1993 Feb 20;229(4):821–826. doi: 10.1006/jmbi.1993.1088. [DOI] [PubMed] [Google Scholar]
- Stormo G. D., Yoshioka M. Specificity of the Mnt protein determined by binding to randomized operators. Proc Natl Acad Sci U S A. 1991 Jul 1;88(13):5699–5703. doi: 10.1073/pnas.88.13.5699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thoma F. Nucleosome positioning. Biochim Biophys Acta. 1992 Feb 28;1130(1):1–19. doi: 10.1016/0167-4781(92)90455-9. [DOI] [PubMed] [Google Scholar]
- Travers A. A., Klug A. The bending of DNA in nucleosomes and its wider implications. Philos Trans R Soc Lond B Biol Sci. 1987 Dec 15;317(1187):537–561. doi: 10.1098/rstb.1987.0080. [DOI] [PubMed] [Google Scholar]
- Trifonov E. N., Sussman J. L. The pitch of chromatin DNA is reflected in its nucleotide sequence. Proc Natl Acad Sci U S A. 1980 Jul;77(7):3816–3820. doi: 10.1073/pnas.77.7.3816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waterman M. S., Arratia R., Galas D. J. Pattern recognition in several sequences: consensus and alignment. Bull Math Biol. 1984;46(4):515–527. doi: 10.1007/BF02459500. [DOI] [PubMed] [Google Scholar]
- Weber I. T., Steitz T. A. Model of specific complex between catabolite gene activator protein and B-DNA suggested by electrostatic complementarity. Proc Natl Acad Sci U S A. 1984 Jul;81(13):3973–3977. doi: 10.1073/pnas.81.13.3973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Crombrugghe B., Busby S., Buc H. Cyclic AMP receptor protein: role in transcription activation. Science. 1984 May 25;224(4651):831–838. doi: 10.1126/science.6372090. [DOI] [PubMed] [Google Scholar]



