Abstract
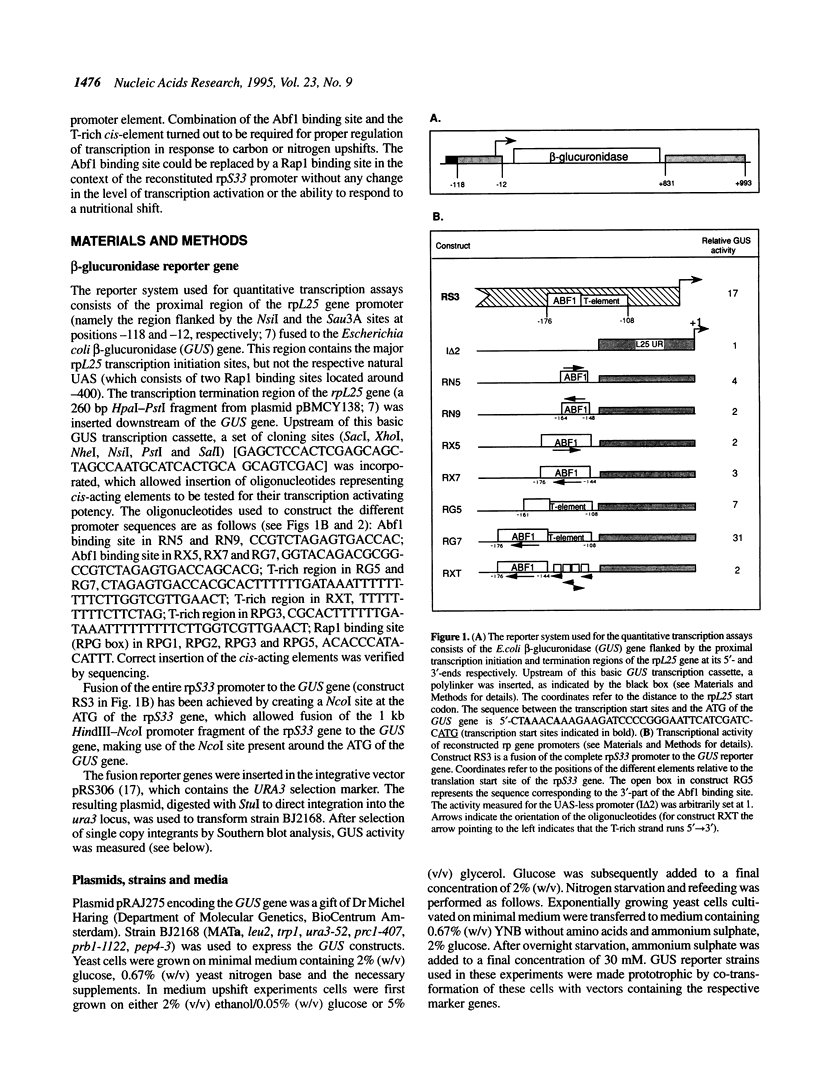
All ribosomal protein (rp) gene promoters from Saccharomyces cerevisiae studied so far contain either (usually two) binding sites for the global gene regulator Rap1p or one binding site for another global factor, Abf1p. Previous analysis of the rpS33 and rpL45 gene promoters suggested that apart from the Abf1 binding site, additional cis-acting elements play a part in transcription activation of these genes. We designed a promoter reconstruction system based on the beta-glucuronidase reporter gene to examine the role of the Abf1 binding site and other putative cis-acting elements in promoting transcription. An isolated Abf1 binding site turned out to be a weak activating element. A T-rich sequence derived from the rpS33 proximal promoter was found to be stronger, but full transcription activation was only achieved by a combination of these elements. Both in the natural rpL45 promoter and in the reconstituted promoter, a Rap1 binding site could functionally replace the Abf1 binding site. Characteristic rp gene nutritional control of transcription, evoked by a carbon source upshift or by nitrogen re-feeding to nitrogen starved cells, could only be mediated by the combined Abf1 (or Rap1) binding site and T-rich element and not by the individual elements. These results demonstrate that Abf1p and Rap1p do not activate rp genes in a prototypical fashion, but rather may serve to potentiate transcription activation through the T-rich element.
Full text
PDF



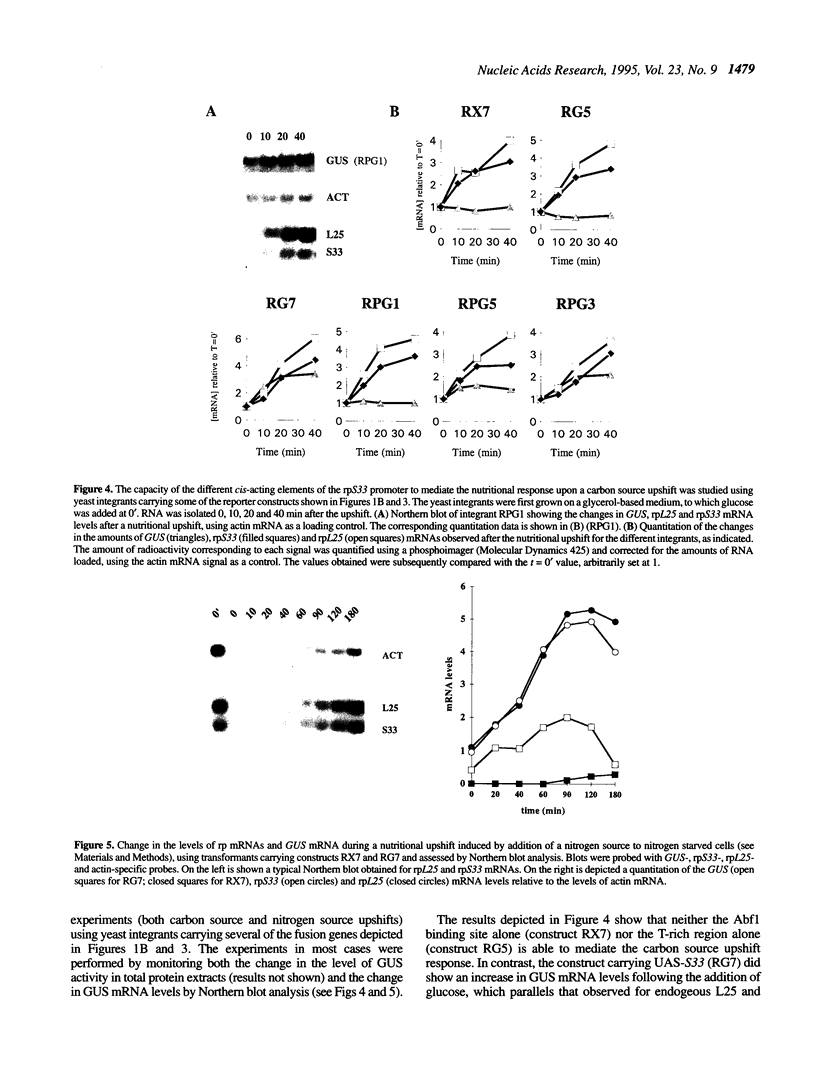

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Brand A. H., Micklem G., Nasmyth K. A yeast silencer contains sequences that can promote autonomous plasmid replication and transcriptional activation. Cell. 1987 Dec 4;51(5):709–719. doi: 10.1016/0092-8674(87)90094-8. [DOI] [PubMed] [Google Scholar]
- Buchman A. R., Kornberg R. D. A yeast ARS-binding protein activates transcription synergistically in combination with other weak activating factors. Mol Cell Biol. 1990 Mar;10(3):887–897. doi: 10.1128/mcb.10.3.887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Winde J. H., Van Leeuwen H. C., Grivell L. A. The multifunctional regulatory proteins ABF1 and CPF1 are involved in the formation of a nuclease-hypersensitive region in the promoter of the QCR8 gene. Yeast. 1993 Aug;9(8):847–857. doi: 10.1002/yea.320090805. [DOI] [PubMed] [Google Scholar]
- Della Seta F., Ciafré S. A., Marck C., Santoro B., Presutti C., Sentenac A., Bozzoni I. The ABF1 factor is the transcriptional activator of the L2 ribosomal protein genes in Saccharomyces cerevisiae. Mol Cell Biol. 1990 May;10(5):2437–2441. doi: 10.1128/mcb.10.5.2437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devlin C., Tice-Baldwin K., Shore D., Arndt K. T. RAP1 is required for BAS1/BAS2- and GCN4-dependent transcription of the yeast HIS4 gene. Mol Cell Biol. 1991 Jul;11(7):3642–3651. doi: 10.1128/mcb.11.7.3642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diffley J. F., Stillman B. Similarity between the transcriptional silencer binding proteins ABF1 and RAP1. Science. 1989 Nov 24;246(4933):1034–1038. doi: 10.1126/science.2511628. [DOI] [PubMed] [Google Scholar]
- Dorsman J. C., Doorenbosch M. M., Maurer C. T., de Winde J. H., Mager W. H., Planta R. J., Grivell L. A. An ARS/silencer binding factor also activates two ribosomal protein genes in yeast. Nucleic Acids Res. 1989 Jul 11;17(13):4917–4923. doi: 10.1093/nar/17.13.4917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dorsman J. C., van Heeswijk W. C., Grivell L. A. Yeast general transcription factor GFI: sequence requirements for binding to DNA and evolutionary conservation. Nucleic Acids Res. 1990 May 11;18(9):2769–2776. doi: 10.1093/nar/18.9.2769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilson E., Roberge M., Giraldo R., Rhodes D., Gasser S. M. Distortion of the DNA double helix by RAP1 at silencers and multiple telomeric binding sites. J Mol Biol. 1993 May 20;231(2):293–310. doi: 10.1006/jmbi.1993.1283. [DOI] [PubMed] [Google Scholar]
- Griffioen G., Mager W. H., Planta R. J. Nutritional upshift response of ribosomal protein gene transcription in Saccharomyces cerevisiae. FEMS Microbiol Lett. 1994 Oct 15;123(1-2):137–144. doi: 10.1111/j.1574-6968.1994.tb07213.x. [DOI] [PubMed] [Google Scholar]
- Hamil K. G., Nam H. G., Fried H. M. Constitutive transcription of yeast ribosomal protein gene TCM1 is promoted by uncommon cis- and trans-acting elements. Mol Cell Biol. 1988 Oct;8(10):4328–4341. doi: 10.1128/mcb.8.10.4328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herruer M. H., Mager W. H., Woudt L. P., Nieuwint R. T., Wassenaar G. M., Groeneveld P., Planta R. J. Transcriptional control of yeast ribosomal protein synthesis during carbon-source upshift. Nucleic Acids Res. 1987 Dec 23;15(24):10133–10144. doi: 10.1093/nar/15.24.10133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kraakman L. S., Griffioen G., Zerp S., Groeneveld P., Thevelein J. M., Mager W. H., Planta R. J. Growth-related expression of ribosomal protein genes in Saccharomyces cerevisiae. Mol Gen Genet. 1993 May;239(1-2):196–204. doi: 10.1007/BF00281618. [DOI] [PubMed] [Google Scholar]
- Kraakman L. S., Mager W. H., Grootjans J. J., Planta R. J. Functional analysis of the promoter of the gene encoding the acidic ribosomal protein L45 in yeast. Biochim Biophys Acta. 1991 Oct 8;1090(2):204–210. doi: 10.1016/0167-4781(91)90102-r. [DOI] [PubMed] [Google Scholar]
- Leer R. J., van Raamsdonk-Duin M. M., Hagendoorn M. J., Mager W. H., Planta R. J. Structural comparison of yeast ribosomal protein genes. Nucleic Acids Res. 1984 Sep 11;12(17):6685–6700. doi: 10.1093/nar/12.17.6685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mager W. H., Planta R. J. Coordinate expression of ribosomal protein genes in yeast as a function of cellular growth rate. 1991 May 29-Jun 12Mol Cell Biochem. 104(1-2):181–187. doi: 10.1007/BF00229818. [DOI] [PubMed] [Google Scholar]
- Mager W. H., Planta R. J. Multifunctional DNA-binding proteins mediate concerted transcription activation of yeast ribosomal protein genes. Biochim Biophys Acta. 1990 Aug 27;1050(1-3):351–355. doi: 10.1016/0167-4781(90)90193-6. [DOI] [PubMed] [Google Scholar]
- McBroom L. D., Sadowski P. D. DNA bending by Saccharomyces cerevisiae ABF1 and its proteolytic fragments. J Biol Chem. 1994 Jun 10;269(23):16461–16468. [PubMed] [Google Scholar]
- Planta R. J., Raué H. A. Control of ribosome biogenesis in yeast. Trends Genet. 1988 Mar;4(3):64–68. doi: 10.1016/0168-9525(88)90042-x. [DOI] [PubMed] [Google Scholar]
- Rotenberg M. O., Woolford J. L., Jr Tripartite upstream promoter element essential for expression of Saccharomyces cerevisiae ribosomal protein genes. Mol Cell Biol. 1986 Feb;6(2):674–687. doi: 10.1128/mcb.6.2.674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmitz U. K., Lonsdale D. M., Jefferson R. A. Application of the beta-glucuronidase gene fusion system to Saccharomyces cerevisiae. Curr Genet. 1990 Mar;17(3):261–264. doi: 10.1007/BF00312618. [DOI] [PubMed] [Google Scholar]
- Sikorski R. S., Hieter P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics. 1989 May;122(1):19–27. doi: 10.1093/genetics/122.1.19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Struhl K. Naturally occurring poly(dA-dT) sequences are upstream promoter elements for constitutive transcription in yeast. Proc Natl Acad Sci U S A. 1985 Dec;82(24):8419–8423. doi: 10.1073/pnas.82.24.8419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teem J. L., Abovich N., Kaufer N. F., Schwindinger W. F., Warner J. R., Levy A., Woolford J., Leer R. J., van Raamsdonk-Duin M. M., Mager W. H. A comparison of yeast ribosomal protein gene DNA sequences. Nucleic Acids Res. 1984 Nov 26;12(22):8295–8312. doi: 10.1093/nar/12.22.8295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tornow J., Zeng X., Gao W., Santangelo G. M. GCR1, a transcriptional activator in Saccharomyces cerevisiae, complexes with RAP1 and can function without its DNA binding domain. EMBO J. 1993 Jun;12(6):2431–2437. doi: 10.1002/j.1460-2075.1993.tb05897.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woudt L. P., Mager W. H., Nieuwint R. T., Wassenaar G. M., van der Kuyl A. C., Murre J. J., Hoekman M. F., Brockhoff P. G., Planta R. J. Analysis of upstream activation sites of yeast ribosomal protein genes. Nucleic Acids Res. 1987 Aug 11;15(15):6037–6048. doi: 10.1093/nar/15.15.6037. [DOI] [PMC free article] [PubMed] [Google Scholar]






