Abstract
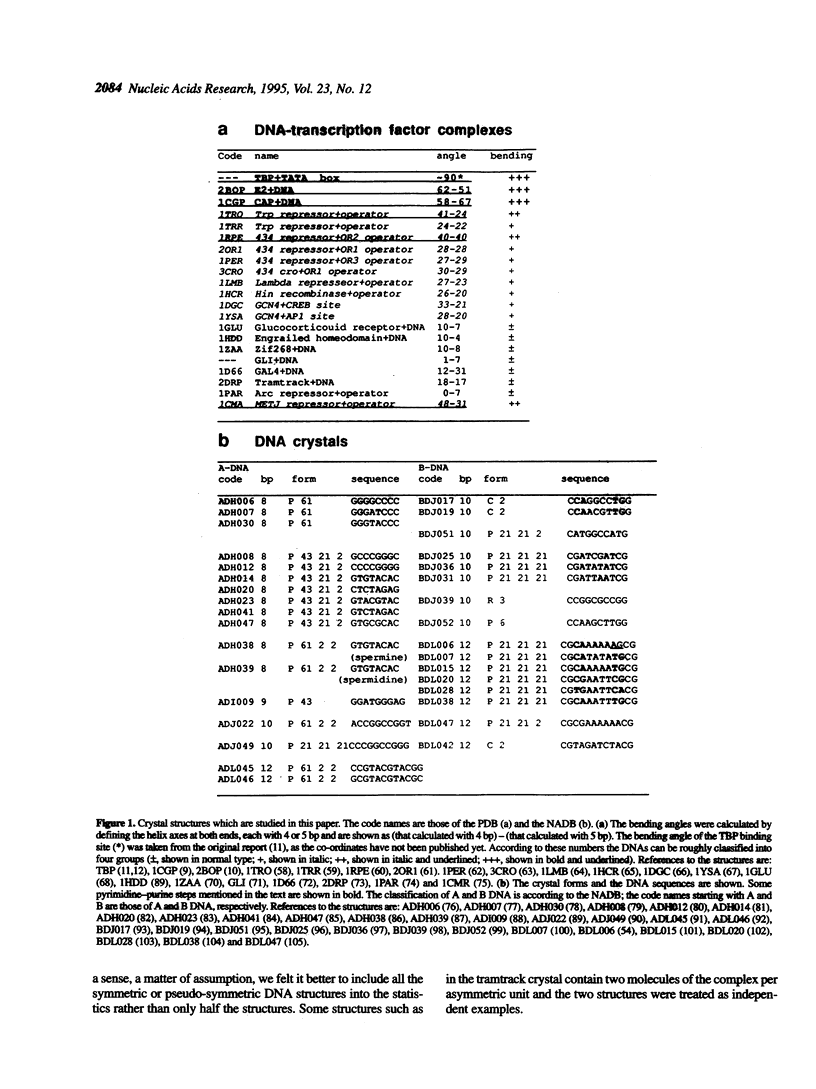
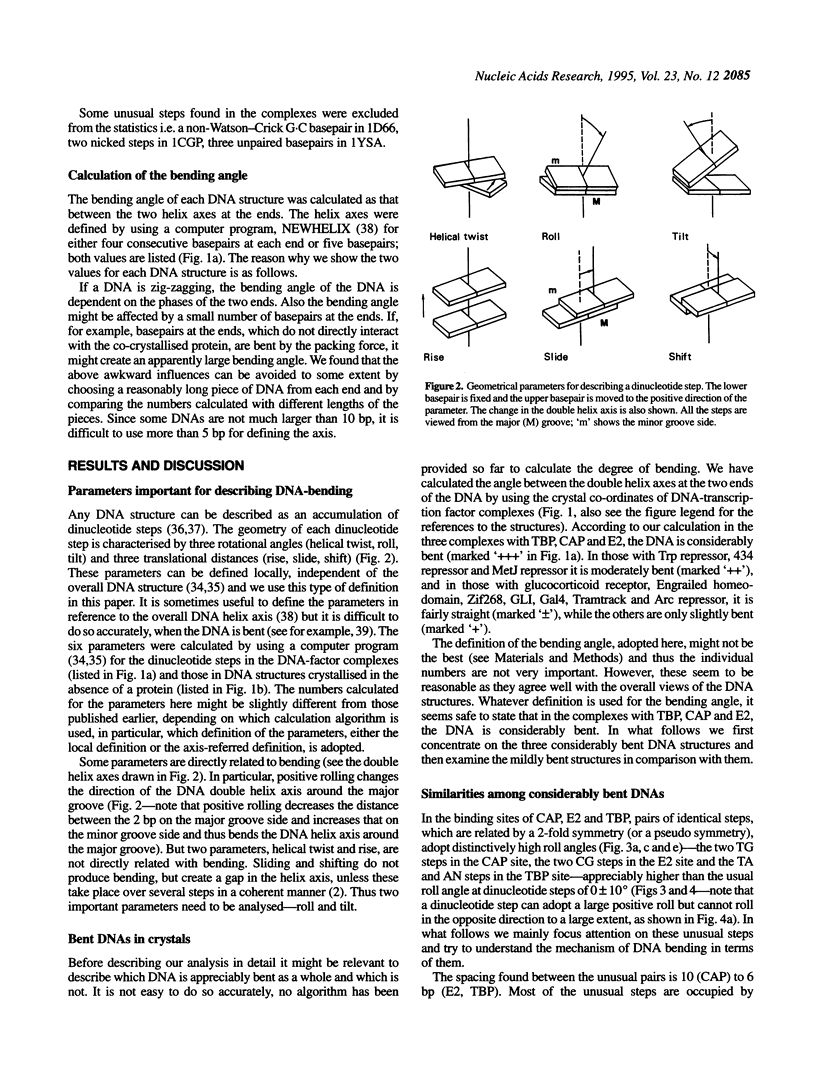
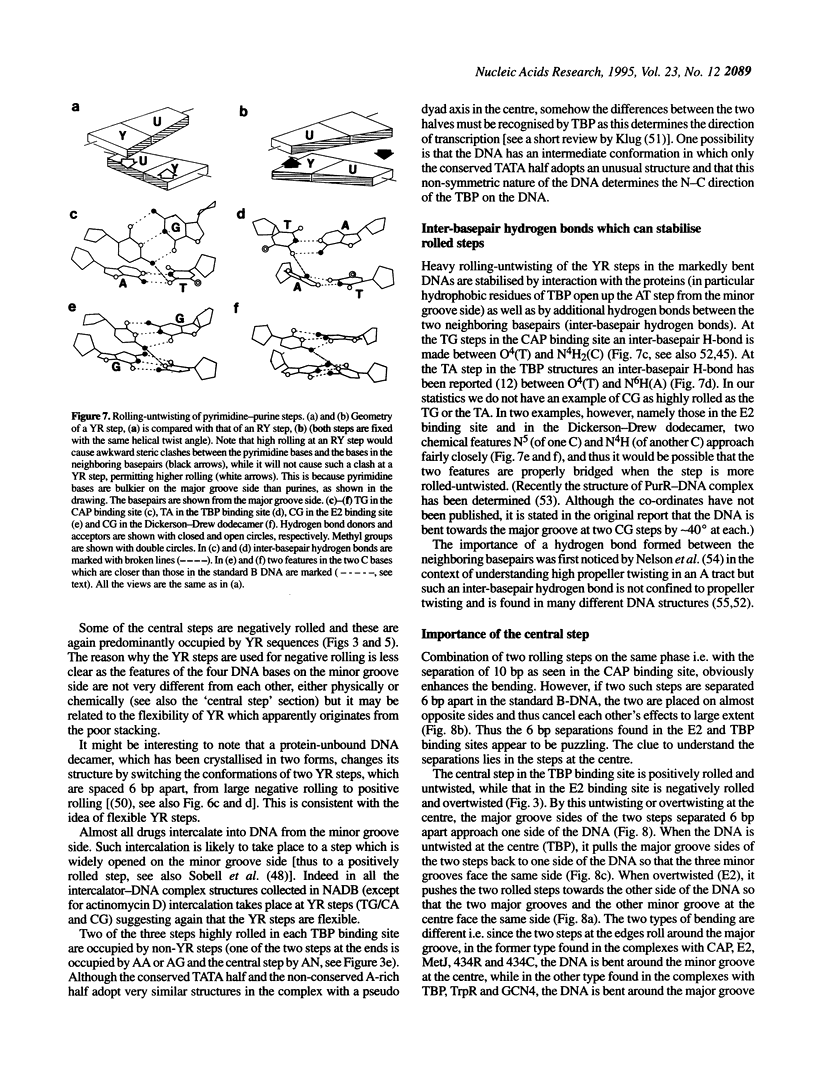
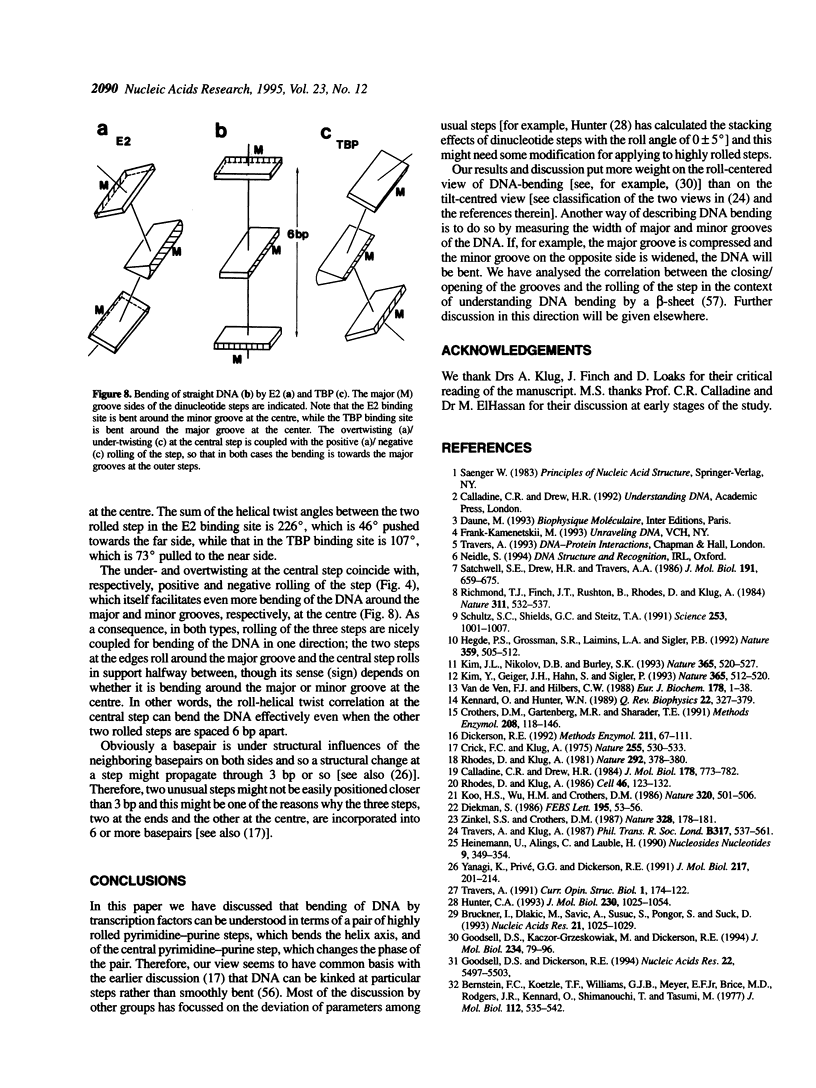
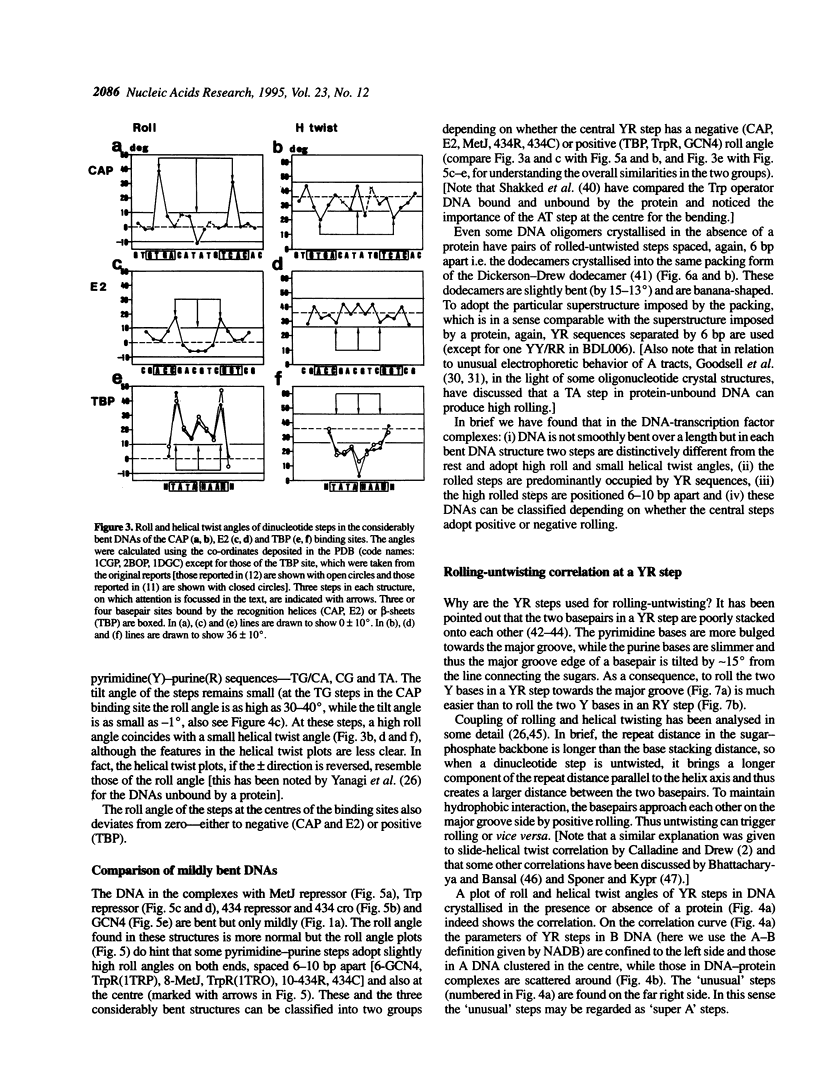
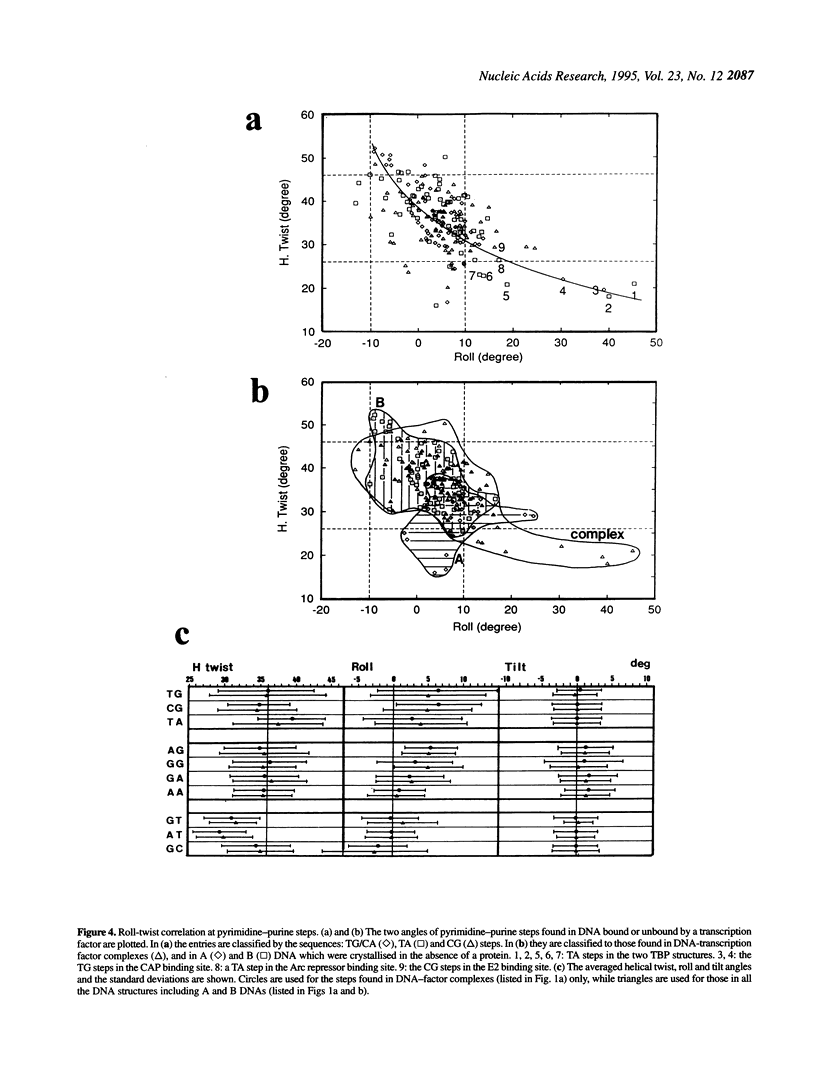
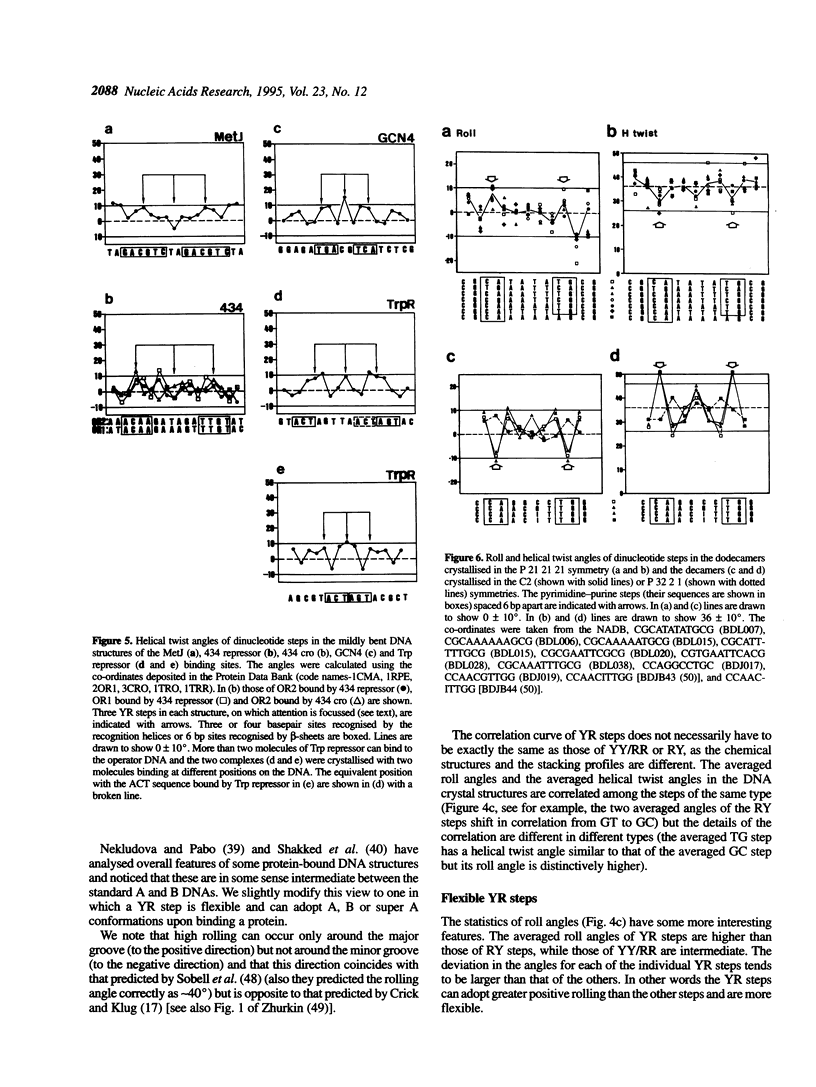
Superstructure-formation of DNA plays an important role in transcription regulation as well as in chromatin formation. To understand the stereochemical basis of DNA bending by proteins we analysed the structural characteristics of dinucleotide steps which occur at the site where DNA is bent upon binding a transcription factor. When DNA is considerably bent in a crystal structure the bending is not spread smoothly over a length, but the DNA is kinked at a pair of crucial steps which are highly rolled and untwisted. These rolled steps are spaced 6-10 bp apart and are predominantly occupied by pyrimidine-purine sequences. In association with another dinucleotide step at the centre, which combines 6 bp-spaced rolled steps towards the same side of the DNA, these produce two essentially different types of DNA bending.
Full text
PDF





Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aggarwal A. K., Rodgers D. W., Drottar M., Ptashne M., Harrison S. C. Recognition of a DNA operator by the repressor of phage 434: a view at high resolution. Science. 1988 Nov 11;242(4880):899–907. doi: 10.1126/science.3187531. [DOI] [PubMed] [Google Scholar]
- Babcock M. S., Pednault E. P., Olson W. K. Nucleic acid structure analysis. Mathematics for local Cartesian and helical structure parameters that are truly comparable between structures. J Mol Biol. 1994 Mar 18;237(1):125–156. doi: 10.1006/jmbi.1994.1213. [DOI] [PubMed] [Google Scholar]
- Babcock M. S., Pednault E. P., Olson W. K. Nucleic acid structure analysis: a users guide to a collection of new analysis programs. J Biomol Struct Dyn. 1993 Dec;11(3):597–628. doi: 10.1080/07391102.1993.10508018. [DOI] [PubMed] [Google Scholar]
- Bernstein F. C., Koetzle T. F., Williams G. J., Meyer E. F., Jr, Brice M. D., Rodgers J. R., Kennard O., Shimanouchi T., Tasumi M. The Protein Data Bank: a computer-based archival file for macromolecular structures. J Mol Biol. 1977 May 25;112(3):535–542. doi: 10.1016/s0022-2836(77)80200-3. [DOI] [PubMed] [Google Scholar]
- Bhattacharyya D., Bansal M. Groove width and depth of B-DNA structures depend on local variation in slide. J Biomol Struct Dyn. 1992 Aug;10(1):213–226. doi: 10.1080/07391102.1992.10508639. [DOI] [PubMed] [Google Scholar]
- Bingman C. A., Zon G., Sundaralingam M. Crystal and molecular structure of the A-DNA dodecamer d(CCGTACGTACGG). Choice of fragment helical axis. J Mol Biol. 1992 Oct 5;227(3):738–756. doi: 10.1016/0022-2836(92)90221-5. [DOI] [PubMed] [Google Scholar]
- Bingman C., Jain S., Zon G., Sundaralingam M. Crystal and molecular structure of the alternating dodecamer d(GCGTACGTACGC) in the A-DNA form: comparison with the isomorphous non-alternating dodecamer d(CCGTACGTACGG). Nucleic Acids Res. 1992 Dec 25;20(24):6637–6647. doi: 10.1093/nar/20.24.6637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bingman C., Li X., Zon G., Sundaralingam M. Crystal and molecular structure of d(GTGCGCAC): investigation of the effects of base sequence on the conformation of octamer duplexes. Biochemistry. 1992 Dec 29;31(51):12803–12812. doi: 10.1021/bi00166a014. [DOI] [PubMed] [Google Scholar]
- Brukner I., Dlakic M., Savic A., Susic S., Pongor S., Suck D. Evidence for opposite groove-directed curvature of GGGCCC and AAAAA sequence elements. Nucleic Acids Res. 1993 Feb 25;21(4):1025–1029. doi: 10.1093/nar/21.4.1025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calladine C. R., Drew H. R. A base-centred explanation of the B-to-A transition in DNA. J Mol Biol. 1984 Sep 25;178(3):773–782. doi: 10.1016/0022-2836(84)90251-1. [DOI] [PubMed] [Google Scholar]
- Calladine C. R. Mechanics of sequence-dependent stacking of bases in B-DNA. J Mol Biol. 1982 Oct 25;161(2):343–352. doi: 10.1016/0022-2836(82)90157-7. [DOI] [PubMed] [Google Scholar]
- Clarke N. D., Beamer L. J., Goldberg H. R., Berkower C., Pabo C. O. The DNA binding arm of lambda repressor: critical contacts from a flexible region. Science. 1991 Oct 11;254(5029):267–270. doi: 10.1126/science.254.5029.267. [DOI] [PubMed] [Google Scholar]
- Coll M., Frederick C. A., Wang A. H., Rich A. A bifurcated hydrogen-bonded conformation in the d(A.T) base pairs of the DNA dodecamer d(CGCAAATTTGCG) and its complex with distamycin. Proc Natl Acad Sci U S A. 1987 Dec;84(23):8385–8389. doi: 10.1073/pnas.84.23.8385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crick F. H., Klug A. Kinky helix. Nature. 1975 Jun 12;255(5509):530–533. doi: 10.1038/255530a0. [DOI] [PubMed] [Google Scholar]
- Crothers D. M., Gartenberg M. R., Shrader T. E. DNA bending in protein-DNA complexes. Methods Enzymol. 1991;208:118–146. doi: 10.1016/0076-6879(91)08011-6. [DOI] [PubMed] [Google Scholar]
- Definitions and nomenclature of nucleic acid structure parameters. J Mol Biol. 1989 Feb 20;205(4):787–791. doi: 10.1016/0022-2836(89)90324-0. [DOI] [PubMed] [Google Scholar]
- Definitions and nomenclature of nucleic acid structure parameters. EMBO J. 1989 Jan;8(1):1–4. doi: 10.1002/j.1460-2075.1989.tb03339.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DiGabriele A. D., Sanderson M. R., Steitz T. A. Crystal lattice packing is important in determining the bend of a DNA dodecamer containing an adenine tract. Proc Natl Acad Sci U S A. 1989 Mar;86(6):1816–1820. doi: 10.1073/pnas.86.6.1816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DiGabriele A. D., Steitz T. A. A DNA dodecamer containing an adenine tract crystallizes in a unique lattice and exhibits a new bend. J Mol Biol. 1993 Jun 20;231(4):1024–1039. doi: 10.1006/jmbi.1993.1349. [DOI] [PubMed] [Google Scholar]
- Dickerson R. E. Base sequence and helix structure variation in B and A DNA. J Mol Biol. 1983 May 25;166(3):419–441. doi: 10.1016/s0022-2836(83)80093-x. [DOI] [PubMed] [Google Scholar]
- Dickerson R. E. DNA structure from A to Z. Methods Enzymol. 1992;211:67–111. doi: 10.1016/0076-6879(92)11007-6. [DOI] [PubMed] [Google Scholar]
- Dickerson R. E., Drew H. R. Structure of a B-DNA dodecamer. II. Influence of base sequence on helix structure. J Mol Biol. 1981 Jul 15;149(4):761–786. doi: 10.1016/0022-2836(81)90357-0. [DOI] [PubMed] [Google Scholar]
- Diekmann S. Sequence specificity of curved DNA. FEBS Lett. 1986 Jan 20;195(1-2):53–56. doi: 10.1016/0014-5793(86)80128-4. [DOI] [PubMed] [Google Scholar]
- Eisenstein M., Frolow F., Shakked Z., Rabinovich D. The structure and hydration of the A-DNA fragment d(GGGTACCC) at room temperature and low temperature. Nucleic Acids Res. 1990 Jun 11;18(11):3185–3194. doi: 10.1093/nar/18.11.3185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ellenberger T. E., Brandl C. J., Struhl K., Harrison S. C. The GCN4 basic region leucine zipper binds DNA as a dimer of uninterrupted alpha helices: crystal structure of the protein-DNA complex. Cell. 1992 Dec 24;71(7):1223–1237. doi: 10.1016/s0092-8674(05)80070-4. [DOI] [PubMed] [Google Scholar]
- Fairall L., Schwabe J. W., Chapman L., Finch J. T., Rhodes D. The crystal structure of a two zinc-finger peptide reveals an extension to the rules for zinc-finger/DNA recognition. Nature. 1993 Dec 2;366(6454):483–487. doi: 10.1038/366483a0. [DOI] [PubMed] [Google Scholar]
- Feng J. A., Johnson R. C., Dickerson R. E. Hin recombinase bound to DNA: the origin of specificity in major and minor groove interactions. Science. 1994 Jan 21;263(5145):348–355. doi: 10.1126/science.8278807. [DOI] [PubMed] [Google Scholar]
- Fratini A. V., Kopka M. L., Drew H. R., Dickerson R. E. Reversible bending and helix geometry in a B-DNA dodecamer: CGCGAATTBrCGCG. J Biol Chem. 1982 Dec 25;257(24):14686–14707. [PubMed] [Google Scholar]
- Frederick C. A., Quigley G. J., Teng M. K., Coll M., Van der Marel G. A., Van Boom J. H., Rich A., Wang A. H. Molecular structure of an A-DNA decamer d(ACCGGCCGGT). Eur J Biochem. 1989 May 1;181(2):295–307. doi: 10.1111/j.1432-1033.1989.tb14724.x. [DOI] [PubMed] [Google Scholar]
- Goodsell D. S., Dickerson R. E. Bending and curvature calculations in B-DNA. Nucleic Acids Res. 1994 Dec 11;22(24):5497–5503. doi: 10.1093/nar/22.24.5497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodsell D. S., Kaczor-Grzeskowiak M., Dickerson R. E. The crystal structure of C-C-A-T-T-A-A-T-G-G. Implications for bending of B-DNA at T-A steps. J Mol Biol. 1994 May 27;239(1):79–96. doi: 10.1006/jmbi.1994.1352. [DOI] [PubMed] [Google Scholar]
- Goodsell D. S., Kopka M. L., Cascio D., Dickerson R. E. Crystal structure of CATGGCCATG and its implications for A-tract bending models. Proc Natl Acad Sci U S A. 1993 Apr 1;90(7):2930–2934. doi: 10.1073/pnas.90.7.2930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grzeskowiak K., Goodsell D. S., Kaczor-Grzeskowiak M., Cascio D., Dickerson R. E. Crystallographic analysis of C-C-A-A-G-C-T-T-G-G and its implications for bending in B-DNA. Biochemistry. 1993 Aug 31;32(34):8923–8931. doi: 10.1021/bi00085a025. [DOI] [PubMed] [Google Scholar]
- Grzeskowiak K., Yanagi K., Privé G. G., Dickerson R. E. The structure of B-helical C-G-A-T-C-G-A-T-C-G and comparison with C-C-A-A-C-G-T-T-G-G. The effect of base pair reversals. J Biol Chem. 1991 May 15;266(14):8861–8883. doi: 10.2210/pdb1d23/pdb. [DOI] [PubMed] [Google Scholar]
- Haran T. E., Shakked Z., Wang A. H., Rich A. The crystal structure of d(CCCCGGGG): a new A-form variant with an extended backbone conformation. J Biomol Struct Dyn. 1987 Oct;5(2):199–217. doi: 10.1080/07391102.1987.10506390. [DOI] [PubMed] [Google Scholar]
- Hegde R. S., Grossman S. R., Laimins L. A., Sigler P. B. Crystal structure at 1.7 A of the bovine papillomavirus-1 E2 DNA-binding domain bound to its DNA target. Nature. 1992 Oct 8;359(6395):505–512. doi: 10.1038/359505a0. [DOI] [PubMed] [Google Scholar]
- Heinemann U., Alings C., Bansal M. Double helix conformation, groove dimensions and ligand binding potential of a G/C stretch in B-DNA. EMBO J. 1992 May;11(5):1931–1939. doi: 10.1002/j.1460-2075.1992.tb05246.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heinemann U., Alings C. Crystallographic study of one turn of G/C-rich B-DNA. J Mol Biol. 1989 Nov 20;210(2):369–381. doi: 10.1016/0022-2836(89)90337-9. [DOI] [PubMed] [Google Scholar]
- Heinemann U., Lauble H., Frank R., Blöcker H. Crystal structure analysis of an A-DNA fragment at 1.8 A resolution: d(GCCCGGGC). Nucleic Acids Res. 1987 Nov 25;15(22):9531–9550. doi: 10.1093/nar/15.22.9531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunter C. A. Sequence-dependent DNA structure. The role of base stacking interactions. J Mol Biol. 1993 Apr 5;230(3):1025–1054. doi: 10.1006/jmbi.1993.1217. [DOI] [PubMed] [Google Scholar]
- Hunter W. N., D'Estaintot B. L., Kennard O. Structural variation in d(CTCTAGAG). Implications for protein-DNA interactions. Biochemistry. 1989 Mar 21;28(6):2444–2451. doi: 10.1021/bi00432a015. [DOI] [PubMed] [Google Scholar]
- Jain S., Zon G., Sundaralingam M. Base only binding of spermine in the deep groove of the A-DNA octamer d(GTGTACAC). Biochemistry. 1989 Mar 21;28(6):2360–2364. doi: 10.1021/bi00432a002. [DOI] [PubMed] [Google Scholar]
- Kennard O., Hunter W. N. Oligonucleotide structure: a decade of results from single crystal X-ray diffraction studies. Q Rev Biophys. 1989 Aug;22(3):327–379. doi: 10.1017/s0033583500002997. [DOI] [PubMed] [Google Scholar]
- Kim J. L., Nikolov D. B., Burley S. K. Co-crystal structure of TBP recognizing the minor groove of a TATA element. Nature. 1993 Oct 7;365(6446):520–527. doi: 10.1038/365520a0. [DOI] [PubMed] [Google Scholar]
- Kim Y., Geiger J. H., Hahn S., Sigler P. B. Crystal structure of a yeast TBP/TATA-box complex. Nature. 1993 Oct 7;365(6446):512–520. doi: 10.1038/365512a0. [DOI] [PubMed] [Google Scholar]
- Kissinger C. R., Liu B. S., Martin-Blanco E., Kornberg T. B., Pabo C. O. Crystal structure of an engrailed homeodomain-DNA complex at 2.8 A resolution: a framework for understanding homeodomain-DNA interactions. Cell. 1990 Nov 2;63(3):579–590. doi: 10.1016/0092-8674(90)90453-l. [DOI] [PubMed] [Google Scholar]
- Klug A., Jack A., Viswamitra M. A., Kennard O., Shakked Z., Steitz T. A. A hypothesis on a specific sequence-dependent conformation of DNA and its relation to the binding of the lac-repressor protein. J Mol Biol. 1979 Jul 15;131(4):669–680. doi: 10.1016/0022-2836(79)90196-7. [DOI] [PubMed] [Google Scholar]
- Klug A. Transcription. Opening the gateway. Nature. 1993 Oct 7;365(6446):486–487. doi: 10.1038/365486a0. [DOI] [PubMed] [Google Scholar]
- Koo H. S., Wu H. M., Crothers D. M. DNA bending at adenine . thymine tracts. Nature. 1986 Apr 10;320(6062):501–506. doi: 10.1038/320501a0. [DOI] [PubMed] [Google Scholar]
- König P., Richmond T. J. The X-ray structure of the GCN4-bZIP bound to ATF/CREB site DNA shows the complex depends on DNA flexibility. J Mol Biol. 1993 Sep 5;233(1):139–154. doi: 10.1006/jmbi.1993.1490. [DOI] [PubMed] [Google Scholar]
- Lauble H., Frank R., Blöcker H., Heinemann U. Three-dimensional structure of d(GGGATCCC) in the crystalline state. Nucleic Acids Res. 1988 Aug 25;16(16):7799–7816. doi: 10.1093/nar/16.16.7799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawson C. L., Carey J. Tandem binding in crystals of a trp repressor/operator half-site complex. Nature. 1993 Nov 11;366(6451):178–182. doi: 10.1038/366178a0. [DOI] [PubMed] [Google Scholar]
- Lipanov A., Kopka M. L., Kaczor-Grzeskowiak M., Quintana J., Dickerson R. E. Structure of the B-DNA decamer C-C-A-A-C-I-T-T-G-G in two different space groups: conformational flexibility of B-DNA. Biochemistry. 1993 Feb 9;32(5):1373–1389. doi: 10.1021/bi00056a024. [DOI] [PubMed] [Google Scholar]
- Luisi B. F., Xu W. X., Otwinowski Z., Freedman L. P., Yamamoto K. R., Sigler P. B. Crystallographic analysis of the interaction of the glucocorticoid receptor with DNA. Nature. 1991 Aug 8;352(6335):497–505. doi: 10.1038/352497a0. [DOI] [PubMed] [Google Scholar]
- Marmorstein R., Carey M., Ptashne M., Harrison S. C. DNA recognition by GAL4: structure of a protein-DNA complex. Nature. 1992 Apr 2;356(6368):408–414. doi: 10.1038/356408a0. [DOI] [PubMed] [Google Scholar]
- McCall M., Brown T., Hunter W. N., Kennard O. The crystal structure of d(GGATGGGAG): an essential part of the binding site for transcription factor IIIA. Nature. 1986 Aug 14;322(6080):661–664. doi: 10.1038/322661a0. [DOI] [PubMed] [Google Scholar]
- McCall M., Brown T., Kennard O. The crystal structure of d(G-G-G-G-C-C-C-C). A model for poly(dG).poly(dC). J Mol Biol. 1985 Jun 5;183(3):385–396. doi: 10.1016/0022-2836(85)90009-9. [DOI] [PubMed] [Google Scholar]
- Mondragón A., Harrison S. C. The phage 434 Cro/OR1 complex at 2.5 A resolution. J Mol Biol. 1991 May 20;219(2):321–334. doi: 10.1016/0022-2836(91)90568-q. [DOI] [PubMed] [Google Scholar]
- Narayana N., Ginell S. L., Russu I. M., Berman H. M. Crystal and molecular structure of a DNA fragment: d(CGTGAATTCACG). Biochemistry. 1991 May 7;30(18):4449–4455. doi: 10.1021/bi00232a011. [DOI] [PubMed] [Google Scholar]
- Nekludova L., Pabo C. O. Distinctive DNA conformation with enlarged major groove is found in Zn-finger-DNA and other protein-DNA complexes. Proc Natl Acad Sci U S A. 1994 Jul 19;91(15):6948–6952. doi: 10.1073/pnas.91.15.6948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nelson H. C., Finch J. T., Luisi B. F., Klug A. The structure of an oligo(dA).oligo(dT) tract and its biological implications. Nature. 1987 Nov 19;330(6145):221–226. doi: 10.1038/330221a0. [DOI] [PubMed] [Google Scholar]
- Otwinowski Z., Schevitz R. W., Zhang R. G., Lawson C. L., Joachimiak A., Marmorstein R. Q., Luisi B. F., Sigler P. B. Crystal structure of trp repressor/operator complex at atomic resolution. Nature. 1988 Sep 22;335(6188):321–329. doi: 10.1038/335321a0. [DOI] [PubMed] [Google Scholar]
- Pavletich N. P., Pabo C. O. Crystal structure of a five-finger GLI-DNA complex: new perspectives on zinc fingers. Science. 1993 Sep 24;261(5129):1701–1707. doi: 10.1126/science.8378770. [DOI] [PubMed] [Google Scholar]
- Pavletich N. P., Pabo C. O. Zinc finger-DNA recognition: crystal structure of a Zif268-DNA complex at 2.1 A. Science. 1991 May 10;252(5007):809–817. doi: 10.1126/science.2028256. [DOI] [PubMed] [Google Scholar]
- Privé G. G., Yanagi K., Dickerson R. E. Structure of the B-DNA decamer C-C-A-A-C-G-T-T-G-G and comparison with isomorphous decamers C-C-A-A-G-A-T-T-G-G and C-C-A-G-G-C-C-T-G-G. J Mol Biol. 1991 Jan 5;217(1):177–199. doi: 10.1016/0022-2836(91)90619-h. [DOI] [PubMed] [Google Scholar]
- Ramakrishnan B., Sundaralingam M. High resolution crystal structure of the A-DNA decamer d(CCCGGCCGGG). Novel intermolecular base-paired G*(G.C) triplets. J Mol Biol. 1993 May 20;231(2):431–444. doi: 10.1006/jmbi.1993.1292. [DOI] [PubMed] [Google Scholar]
- Raumann B. E., Rould M. A., Pabo C. O., Sauer R. T. DNA recognition by beta-sheets in the Arc repressor-operator crystal structure. Nature. 1994 Feb 24;367(6465):754–757. doi: 10.1038/367754a0. [DOI] [PubMed] [Google Scholar]
- Rhodes D., Klug A. An underlying repeat in some transcriptional control sequences corresponding to half a double helical turn of DNA. Cell. 1986 Jul 4;46(1):123–132. doi: 10.1016/0092-8674(86)90866-4. [DOI] [PubMed] [Google Scholar]
- Rhodes D., Klug A. Sequence-dependent helical periodicity of DNA. Nature. 1981 Jul 23;292(5821):378–380. doi: 10.1038/292378a0. [DOI] [PubMed] [Google Scholar]
- Richmond T. J., Finch J. T., Rushton B., Rhodes D., Klug A. Structure of the nucleosome core particle at 7 A resolution. Nature. 1984 Oct 11;311(5986):532–537. doi: 10.1038/311532a0. [DOI] [PubMed] [Google Scholar]
- Rodgers D. W., Harrison S. C. The complex between phage 434 repressor DNA-binding domain and operator site OR3: structural differences between consensus and non-consensus half-sites. Structure. 1993 Dec 15;1(4):227–240. doi: 10.1016/0969-2126(93)90012-6. [DOI] [PubMed] [Google Scholar]
- Satchwell S. C., Drew H. R., Travers A. A. Sequence periodicities in chicken nucleosome core DNA. J Mol Biol. 1986 Oct 20;191(4):659–675. doi: 10.1016/0022-2836(86)90452-3. [DOI] [PubMed] [Google Scholar]
- Schultz S. C., Shields G. C., Steitz T. A. Crystal structure of a CAP-DNA complex: the DNA is bent by 90 degrees. Science. 1991 Aug 30;253(5023):1001–1007. doi: 10.1126/science.1653449. [DOI] [PubMed] [Google Scholar]
- Schumacher M. A., Choi K. Y., Zalkin H., Brennan R. G. Crystal structure of LacI member, PurR, bound to DNA: minor groove binding by alpha helices. Science. 1994 Nov 4;266(5186):763–770. doi: 10.1126/science.7973627. [DOI] [PubMed] [Google Scholar]
- Shakked Z., Guzikevich-Guerstein G., Frolow F., Rabinovich D., Joachimiak A., Sigler P. B. Determinants of repressor/operator recognition from the structure of the trp operator binding site. Nature. 1994 Mar 31;368(6470):469–473. doi: 10.1038/368469a0. [DOI] [PubMed] [Google Scholar]
- Shimon L. J., Harrison S. C. The phage 434 OR2/R1-69 complex at 2.5 A resolution. J Mol Biol. 1993 Aug 5;232(3):826–838. doi: 10.1006/jmbi.1993.1434. [DOI] [PubMed] [Google Scholar]
- Sobell H. M., Tsai C. C., Jain S. C., Gilbert S. G. Visualization of drug-nucleic acid interactions at atomic resolution. III. Unifying structural concepts in understanding drug-DNA interactions and their broader implications in understanding protein-DNA interactions. J Mol Biol. 1977 Aug 15;114(3):333–365. doi: 10.1016/0022-2836(77)90254-6. [DOI] [PubMed] [Google Scholar]
- Somers W. S., Phillips S. E. Crystal structure of the met repressor-operator complex at 2.8 A resolution reveals DNA recognition by beta-strands. Nature. 1992 Oct 1;359(6394):387–393. doi: 10.1038/359387a0. [DOI] [PubMed] [Google Scholar]
- Timsit Y., Vilbois E., Moras D. Base-pairing shift in the major groove of (CA)n tracts by B-DNA crystal structures. Nature. 1991 Nov 14;354(6349):167–170. doi: 10.1038/354167a0. [DOI] [PubMed] [Google Scholar]
- Travers A. A., Klug A. The bending of DNA in nucleosomes and its wider implications. Philos Trans R Soc Lond B Biol Sci. 1987 Dec 15;317(1187):537–561. doi: 10.1098/rstb.1987.0080. [DOI] [PubMed] [Google Scholar]
- Trifonov E. N. Curved DNA. CRC Crit Rev Biochem. 1985;19(2):89–106. doi: 10.3109/10409238509082540. [DOI] [PubMed] [Google Scholar]
- Van de Ven F. J., Hilbers C. W. Nucleic acids and nuclear magnetic resonance. Eur J Biochem. 1988 Dec 1;178(1):1–38. doi: 10.1111/j.1432-1033.1988.tb14425.x. [DOI] [PubMed] [Google Scholar]
- Westhof E. Re-refinement of the B-dodecamer d(CGCGAATTCGCG) with a comparative analysis of the solvent in it and in the Z-hexamer d(5BrCG5BrCG5BrCG). J Biomol Struct Dyn. 1987 Dec;5(3):581–600. doi: 10.1080/07391102.1987.10506414. [DOI] [PubMed] [Google Scholar]
- Yanagi K., Privé G. G., Dickerson R. E. Analysis of local helix geometry in three B-DNA decamers and eight dodecamers. J Mol Biol. 1991 Jan 5;217(1):201–214. doi: 10.1016/0022-2836(91)90620-l. [DOI] [PubMed] [Google Scholar]
- Yoon C., Privé G. G., Goodsell D. S., Dickerson R. E. Structure of an alternating-B DNA helix and its relationship to A-tract DNA. Proc Natl Acad Sci U S A. 1988 Sep;85(17):6332–6336. doi: 10.1073/pnas.85.17.6332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuan H., Quintana J., Dickerson R. E. Alternative structures for alternating poly(dA-dT) tracts: the structure of the B-DNA decamer C-G-A-T-A-T-A-T-C-G. Biochemistry. 1992 Sep 1;31(34):8009–8021. [PubMed] [Google Scholar]
- Zhurkin V. B. Sequence-dependent bending of DNA and phasing of nucleosomes. J Biomol Struct Dyn. 1985 Feb;2(4):785–804. doi: 10.1080/07391102.1985.10506324. [DOI] [PubMed] [Google Scholar]
- Zinkel S. S., Crothers D. M. DNA bend direction by phase sensitive detection. Nature. 1987 Jul 9;328(6126):178–181. doi: 10.1038/328178a0. [DOI] [PubMed] [Google Scholar]



