Integrated transcriptomic and genomic data for AIMAH provides supporting evidence that larger adrenal nodules accumulate an increased number of genetic, and consequently, transcript abnormalities.
Abstract
Context:
Massive macronodular adrenocortical disease or ACTH-independent macronodular adrenal hyperplasia (AIMAH) is a clinically and genetically heterogeneous disorder.
Objective and Design:
Whole-genome expression profiling and oligonucleotide array comparative genomic hybridization changes were analyzed in samples of different nodules from the same patients with AIMAH. Quantitative RT-PCR and staining were employed to validate the mRNA array data.
Results:
Chromosomal gains were more frequent in larger nodules when compared with smaller nodules from the same patients. Among the 50 most overexpressed genes, 50% had a chromosomal locus that was amplified in the comparative genomic hybridization data. Although the list of most over- and underexpressed genes was similar between the nodules of different size, the gene set enrichment analysis identified different pathways associated with AIMAH that corresponded to the size; the smaller nodules were mainly enriched for metabolic pathways, whereas p53 signaling and cancer genes were enriched in larger nodules. Confirmatory studies demonstrated that BCL2, E2F1, EGF, c-KIT, MYB, PRKCA, and CTNNB1 were overexpressed in the larger nodules at messenger and/or protein levels. Chromosomal enrichment analysis showed that chromosomes 20q13 and 14q23 might be involved in progression of AIMAH from smaller to larger tumors.
Conclusion:
Integrated transcriptomic and genomic data for AIMAH provides supporting evidence to the hypothesis that larger adrenal lesions, in the context of this chronic, polyclonal hyperplasia, accumulate an increased number of genomic and, subsequently, transcript abnormalities. The latter shows that the disease appears to start with mainly tissue metabolic derangements, as suggested by the study of the smaller nodules, but larger lesions showed aberrant expression of oncogenic pathways.
ACTH-independent macronodular adrenal hyperplasia (AIMAH), also known as massive macronodular adrenal disease, is known as a rare cause of Cushing's syndrome (CS), accounting for less than 1% of all cases of ACTH-independent CS (1–3). However, the diagnosis of AIMAH associated with subclinical hypercortisolism has lately increased. This is not surprising, because approximately 10–15% of the presumably clinically silent adrenal masses that are detected after various imaging studies are indeed bilateral (4).
AIMAH is a sporadic disease in the majority of cases, but several families with AIMAH, in whom the disease was inherited as an autosomal dominant trait, have been reported (5–7). Cortisol production in AIMAH can be regulated by the aberrant expression of G protein-coupled receptors other than ACTH, such as those for glucose-dependent insulinotropic peptide (GIPR), β-adrenergic receptors (β-AR), vasopressin (V2-V3-vasopressin receptor), serotonin (5-HT7 receptor), angiotensin II (AT1R), glucagon (GCGR), and LH/human chorionic gonadotropin (LH/hCGR) (8, 9).
A number of genetic abnormalities have been detected in adrenocortical adenomas and carcinomas (10–16). Bourdeau et al. (17) showed somatic losses of the 17q22-24 region and protein kinase A (PKA) subunit and enzymatic activity changes in AIMAH, demonstrating that cAMP/PKA signaling is altered in this disorder, not unlike the case in primary pigmented nodular adrenocortical disease caused by PRKAR1A-inactivating mutations or sporadic adrenal tumors that harbor 17q (the PRKAR1A locus) losses (16). Whole-genome expression studies in AIMAH and primary pigmented nodular adrenocortical disease, the main forms of bilateral adrenal hyperplasia, confirmed the involvement of the cAMP signaling pathway (18, 19) but also pointed to the overexpression of genes that regulate or are part of the Wnt signaling pathway such as WISP2, GSK3B, and CTNNB1 (20, 21).
Previous cytogenetic analysis of adrenal tumors identified no gains or losses in adenomas less than 5 cm by comparative genomic hybridization (CGH) (10). Adenomas more than 5 cm size had one chromosomal gain or loss or both, whereas adrenal carcinomas (range, 7–20 cm) frequently had losses involving chromosomes 2, 11q, and 17p and gains at chromosomes 4 and 5 (10). Sidhu et al. (11) also found a significantly high frequency of CGH changes in adrenal carcinomas when compared with adenomas. Although a strong correlation between the size of adrenal tumor and the number of cytogenetic changes has been demonstrated by several reports (10–14), the hypothesis that genetic changes in adrenal hyperplasias correlates with the nodule size in the same patients (22) remains to be explored.
In the present study, we tested this hypothesis after microdissection of different nodules from each of two unrelated patients with AIMAH and analysis of the whole genome at the DNA and RNA levels. The data point to the significance of size of adrenocortical lesions as a major determining factor of their tumor potential. It is not only the number of changes that increased with size; the data also show a qualitative difference in the pathways altered in smaller vs. larger nodules within AIMAH, suggesting a progression from a relatively simple metabolic derangement at the beginning of the process to the gradual involvement of oncogenic pathways in larger nodules.
Subjects and Methods
Subjects
Two patients with AIMAH were admitted to the National Institutes of Health (NIH) Warren Magnuson Clinical Center from 2000–2006 for the work-up and treatment of adrenocortical tumors under protocol 00-CH-160. The Eunice Kennedy Shriver National Institute of Child Health and Human Development Institutional Review Board approved this study, and informed consents were obtained from the two subjects.
Patient 1
A 42-yr-old male patient with a history of mild hypertension and weight gain (60 lbs over 10 yr), plethoric face, and violaceous striae was admitted to the NIH Clinical Center to investigate CS. The hormonal evaluation revealed absence of plasma cortisol suppression after 1 mg overnight dexamethasone (DEX) test and, after the DEX-ovine corticotropin-releasing hormone test, high 24 h urinary free cortisol levels (159.1 μg; normal, <90 μg/24 h), abnormal midnight serum cortisol levels (11.6 μg/dl), and suppressed ACTH levels (<5 pg/ml). Bilateral enlargement of the adrenal glands with macronodules was seen on the computer tomography scan. The patient was diagnosed with ACTH-independent CS and underwent bilateral adrenalectomy (Fig. 1A).
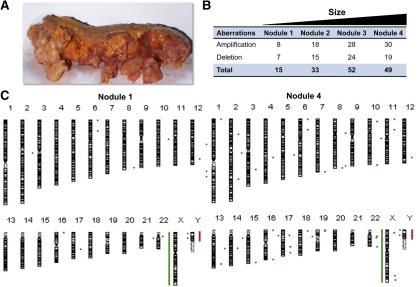
Fig. 1.
AIMAH patient 1. A, Macroscopic appearance of the left adrenal gland and a summary of chromosomal aberrations; B, all chromosomes with identified gains and losses; C, from two nodules of different size: red bars and points (*) to the right of the chromosome ideogram indicate a gain, whereas green bars and points (*) to the left indicate loss of genetic material.
Patient 2
A 42-yr-old female patient presented with a 4-yr history of secondary amenorrhea, easy bruising, hirsutism, weight gain (25 lbs), muscle weakness, and behavioral changes. Biochemical evaluation showed absence of plasma cortisol suppression after 1 mg overnight DEX test and, after the DEX-ovine corticotropin-releasing hormone test, high 24-h urinary free cortisol levels (270 μg; normal, <90 μg/24 h) and suppressed ACTH levels (<5 pg/ml). Computer tomography scan revealed bilateral adrenal macronodules. She was diagnosed with ACTH-independent CS and underwent laparoscopic bilateral adrenalectomy (Fig. 2A).
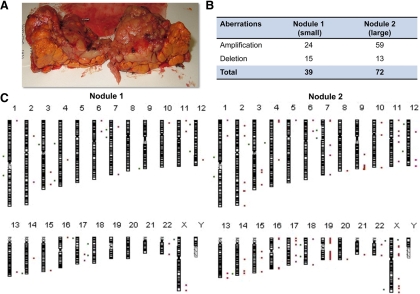
Fig. 2.
AIMAH patient 2. A, Macroscopic appearance of the right adrenal in patient 2, showing macronodules and a summary of chromosomal aberrations; B, all chromosomes with indentified gains and losses; C, from two nodules of different size: red bars and points (*) to the right of the chromosome ideogram indicate a gain, whereas green bars and points (*) to the left indicate loss of genetic material.
Aberrant G protein-coupled receptor expression (GIPR, AGTR1, ADRB1, -2, and -3, AVPR1, -2, and -3, HTR7, GCGR, and LHCGR) was not detected in tissue samples from both patients; however, these patients were not tested in vivo for their illegitimate receptor responses.
Hormone assays
Plasma cortisol and ACTH levels were measured as described elsewhere (23). Urinary free cortisol excretion was measured by direct RIA (24). The intraassay coefficient of variation was 5%, and the interassay coefficient of variation was 10%.
Comparative genome hybridization
Adrenal tissue was collected from both patients, and the nodules were macroscopically dissected from the surrounding tissue and snap frozen in liquid nitrogen. DNA extraction was performed in the four nodules (microdissected tissue) from patient 1 (sizes were 1, 3, 5, and 7 cm, respectively) and in two available from patient 2 (sizes were 1 and 3.5 cm, respectively) using the QIAGEN DNeasy blood and tissue kit (QIAGEN, Valencia, CA). Oligonucleotide array CGH (oligo-aCGH) by GeneDX was used to identify regions of gains or losses throughout the chromosomes. A set of 105,000 oligonucleotide probes was used to cover 37 kb of the human genome sequence (22 autosomes and X and Y chromosomes) per probe (GenomeDx version 2.0). Agilent CGH Analytics version 3.5 software was used to analyze the oligo-aCGH data.
Microarray analysis
Total RNA extraction was performed from seven different-sized nodules using the Trizol reagent method (Invitrogen, Carlsbad, CA). The RNA samples were further purified using the RNeasy columns (QIAGEN), and the quality of RNA was assessed using the Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA). Three commercially available pools of human adrenal total RNA (Clontech, Mountain View, CA; BioChain, Hayward, CA; and Ambion, Austin, TX) were employed as reference samples. Preparation of cRNA from total RNA, hybridization in Sentrix HumanRef-8 Expression BeadChips, scanning, and image analysis was done as previously described (25). Analysis of Illumina data was performed using Illumina BeadStudio software (Illumina, San Diego, CA), which returns the trimmed mean average intensity for each single gene probe type (nonnormalized). Any gene consistently with a P detection value above 0.01 for all samples was eliminated from further analysis. Z-transformation for normalization was performed for each Illumina sample/array (25, 26). First, the raw intensity data for each sample was log10-transformed, and Z scores were calculated by subtracting the overall average gene intensity (within a single experiment) from the raw intensity data for each gene, then, dividing that result by the sd of all of the measured intensities. Changes in gene expression (ratio) between different Z-transformed datasets (nodules compared with the average of the normal adrenal pools) were calculated as differences between the corresponding Z scores and then divided by the sd of each Z difference dataset (25). A 2-fold change was employed as the cutoff value to identify over- and underexpressed genes in adrenal nodules in relation to normal adrenal pools. Microarray data are in compliance with the minimal information about a microarray experiment (MIAME) format. The raw and normalized microarray data reported in this paper have been deposited in the Gene Expression Omnibus (GEO) database (accession no. GSE25031).
Analysis of the mRNA profiling
The functional analysis of the whole-genome transcriptome profiling was performed using the Database for Annotation, Visualization and Integrated Discovery (DAVID) Bioinformatic Resources 2008 (National Institute of Allergy and Infectious Diseases, NIH, http://david.abcc.ncifcrf.gov/home.jsp) (27, 28). The lists of genes (induced or repressed) were submitted to the DAVID database (http://david.abcc.ncifcrf.gov); genes are there clustered according to a series of common keywords. The proportion of each keyword in the list is compared with the one in the whole genome, making it possible to compute P values and enrichment scores (geometric mean of the inverse log of each P value). The detailed information of gene alterations was systematically reported on KEGG pathways.
Gene set enrichment analysis (GSEA) was performed by GSEA Software version 2.0 (www.broad.mit.edu) in pairwise comparisons (29). Gene expression results derived from microarray experiments were correlated with chromosome gene sets. GSEA was performed using gene set permutation type as default, and the number of permutations was set to 1000. Statistical significance levels were defined as nominal P value <0.05.
Real-time quantitative RT-PCR
Quantitative real-time PCR was performed in the ABI Prism 7700 sequence detector using using the Oncogene and Tumor Suppressor Genes PCR Array Plate (Supplemental Table 1, published on The Endocrine Society's Journals Online web site at http://jcem.endojournals.org; SABiosciences, Frederick, MD). An average of three commercially available pools of human adrenal total RNA (Clontech, BioChain, and Ambion) was used as control. Relative quantification was performed using the 2−ΔΔCT method (30).
Immunohistochemistry
All immunohistochemistry was performed in collaboration with Histoserv, Inc. (Germantown, MD) using standard procedures. The following primary antibodies were used: CTNNB1 (ab6302; Abcam), BCL-2 (C-2, sc-7382; Santa Cruz Biotechnology, Santa Cruz, CA), and c-KIT (E1, sc-17806), Cα PKA subunit (C-20, sc-903), Cβ PKA subunit (C-20, sc-904), and PRKX (gift from Dr. Robert M. Kotin, National Heart, Lung, and Blood Institute, NIH, Bethesda, MD). The following grading system was used for staining evaluation: negative (absence of expression), weak staining (from 1–25% of immunoreactive cells), and strong staining (>25% of immunoreactive cells). Immunoreactivity for BCL-2, c-KIT, and PKA subunits was evaluated in the cytoplasm, whereas CTNNB1 staining was assessed in the nucleus.
PKA activity
PKA enzymatic activity was measured in tissue extracts as previously described (31).
Statistical analysis
All statistical analyses were performed with the SPSS version 16.0 (SPSS Inc., Chicago, IL). Continuous data are expressed as mean ± sd. All the experiments were performed in triplicate. A two-sample t test was used for paired samples. A P value <0.05 was considered significant.
Results
Oligo-aCGH analysis
Oligo-aCGH was used to map DNA copy number aberrations that occur in nodules of different sizes. A summary of all the changes is presented for patients 1 and 2 (Figs. 1 and 2, respectively). Chromosomal gains were more frequent in larger nodules when compared with smaller nodules. In patient 1, 30 amplifications were detected in nodule 4 (larger), whereas only eight amplifications were found in nodule 1 (smaller) (Fig. 1, B and C). Similarly, a positive correlation between nodule size and number of gains of genetic material was observed for patient 2 (Fig. 2, B and C). Copy number increases (amplifications) were found for 59 chromosomal regions in the large nodule from patient 2 but only for 24 chromosomal regions in the smaller nodule. Chromosomal losses were not as frequent as gains in both cases (35.8 vs. 64.2%). The number of chromosomal deletions was also associated with the nodule size in patient 1.
Whole transcriptome profiling
Microarray analysis was performed in three normal adrenal pools and in seven adrenal nodules from the same subject with AIMAH (patient 2) using the Illumina Beadarrays system. All genes were displayed in the heat maps constructed by processing the data using unsupervised hierarchical clustering (Fig. 3A). The most differentially expressed genes in AIMAH were also compared with normal adrenal cortex (Fig. 3A and Supplemental Table 2). Among the 50 most overexpressed genes, 50% of them were found to be amplified in the CGH data. However, only nine of the 50 most underexpressed genes (18%) were located in regions of chromosomal losses.
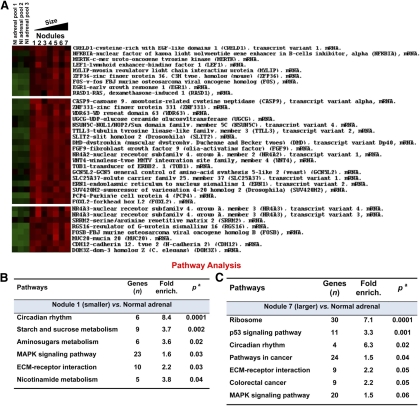
Fig. 3.
A, Heatmap visualization of gene expression data displaying differentially expressed genes in normal adrenals and AIMAH nodules from patient 2; B and C, functional analysis of whole-genome transcriptome profiling of nodule 1 (small) and nodule 7 (large) compared with normal adrenal tissue. The array functional analysis was performed using DAVID Bioinformatics Resources 2008, National Institute of Allergy and Infectious Diseases, NIH (http://david.abcc.ncifcrf.gov/home.jsp).
The overexpressed genes associated with chromosomal gains included TFPI2 [7q22; ratio (log10) 6.45], TNFRSF12A (16p13.3; ratio 6.14), CDH12 [5p14.3; ratio 4.9], CASP9 (1p36.21; ratio 4.6), FOS (14q24.3; ratio 4.4), and FOSB (19q13.32; ratio 4.5), among others (Supplemental Table 2). On the other hand, some of the top overexpressed genes not associated with gene amplification were TSPAN8 (12q14.1-q21.1; ratio 9.0), NR4A2 (2q22-q23; ratio 7.17), IL7R (5p13; ratio 6.6), RGS1 (1q31; ratio 4.8), and FGF9 (3q11-q12, ratio 4.3).
Wnt signaling activation in AIMAH was demonstrated in a previous gene array study from our group (21). In the current analysis, WNT4 was overexpressed similarly in all AIMAH nodules [nodule 1, ratio (log10) 4.0; nodule 2 = 4.5; nodule 3 = 3.8; nodule 4 = 3.5; nodule 5 = 3.8; nodule 6 = 3.3; and nodule 7 = 4.2]. Additionally, WISP1 was also overexpressed in AIMAH, regardless of the nodule size (nodule 1 = 2.2; nodule 2 = 1.7; nodule 3 = 1.8; nodule 4 = 1.8; nodule 5 = 2.5; nodule 6 = 1.5; and nodule 7 = 1.7).
Although the list of most over- and underexpressed genes was similar between nodules, the GSEA identified different pathways associated with AIMAH depending on the size of the examined nodules (Fig. 3B and Supplemental Table 3). The functional analysis of the whole-genome transcriptome profiling was performed using the DAVID Bioinformatic Resources 2008 (National Institute of Allergy and Infectious Diseases, NIH) (28). Circadian rhythm and metabolic pathways (sucrose and aminosugars metabolism) were the most significantly enriched pathways in nodule 1 (smaller) in relation to normal adrenal (P < 0.05) (Fig. 3B). MAPK signaling and extracellular matrix (ECM)-interaction pathways were also overexpressed in nodule 1 (P = 0.03). Interestingly, the functional analysis of nodule 7 (the largest one) revealed different pathways associated with AIMAH-tumor signature. An enrichment for p53 signaling (P = 0.001), other cancer pathways (P = 0.04), and, especially, genes involved in colorectal cancer (P = 0.05) were found in nodule 7 (Fig. 3B).
Correlation of chromosomal enrichment by gene set enrichment analysis (GSEA) and cytogenetic data
GSEA was performed and correlated with CGH data. GSEA revealed chromosome 20q13 and 14q23 enrichment in nodule 7 (large) when compared with nodule 1 (small) gene signature (P = 0.01) (Table 1). Both chromosome 20q13 and 14q23 regions had a higher amplification in nodule 7 in relation to nodule 1. The enriched gene set in chromosome 20q13 included LAMA5, BIRC7, PKIG, PRIC285, and RGS19 (Table 1). In chromosome 14q23, RHOJ, TMEM30B, and PRKCH mainly accounted for the chromosomal enrichment of this amplified region.
Table 1.
Chromosomal enrichment by GSEA in a larger nodule when compared with a smaller nodule from AIMAH
| Enrichment (%) | |
|---|---|
| Enriched genes at chromosomal 20q13 | |
| PFDN4: prefoldin subunit 4 | 47.3 |
| ZNF334, zinc finger protein 334 (ZNF334) | 73.8 |
| CSE1 liter, CSE1 chromosome segregation 1-like (yeast) | 10.6 |
| SS18L1, synovial sarcoma translocation gene on chromosome 18-like 1 | 10.8 |
| SLC2A4RG, SLC2A4 regulator | 14.5 |
| TCEA2, transcription elongation factor A (SII) | 92.9 |
| GTPBP5, GTP binding protein 5 | 16.9 |
| PPP1R3D, protein phosphatase 1, regulatory subunit 3D | 15.5 |
| PRIC285, peroxisomal proliferator-activated receptor A interacting complex 285 | 24 |
| RGS19, regulator of G-protein signaling 19 | 18 |
| LAMA5, laminin, α 5 | 100 |
| PKIG, protein kinase (cAMP-dependent, catalytic) inhibitor γ | 40 |
| UBE2V1, ubiquitin-conjugating enzyme E2 variant 1 | 16.6 |
| BIRC7, baculoviral IAP repeat-containing 7 (livin) | 69.1 |
| TFAP2C, transcription factor AP-2 γ | 13.5 |
| Enriched genes at chromosomal 14q23 | |
| TMEM30B, transmembrane protein 30B | 44.6 |
| RHOJ, ras homolog gene family, member J | 45.7 |
| NMNAT1, nicotinamide nucleotide adenylyltransferase 1 | 10 |
| VTI1B, vesicle transport through interaction with t-SNAREs homolog 1B (yeast) | 26.2 |
| PRKCH, protein kinase C, η | 11 |
Expression of oncogenes and PKA catalytic subunits in AIMAH nodules of different size
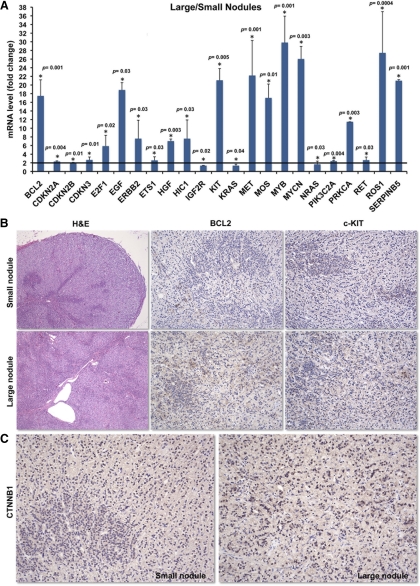
A quantitative RT-PCR array carrying 84 genes of oncogenic pathways was performed to confirm the microarray data and evaluate which genes might be involved in tumor progression in AIMAH (Supplemental Table 1). Gene expression was compared between two groups, the smaller (nodules 1 and 2) and the larger (nodules 6 and 7) tumors (Fig. 4A). Overexpressed oncogenes in larger AIMAH nodules (and, thus, possibly associated with tumor progression) were BCL2, E2F1, EGF, c-KIT, MYB, PRKCA, and SERPINB5, among others. At protein level, BCL2 and c-KIT expression was strong in large nodules and weak in small nodules (Fig. 4B). Similarly, nuclear CTNNB1 staining was strong in a large nodule and weak in a small nodule (Fig. 4C).
Fig. 4.
A, A quantitative RT-PCR array including 84 genes involved in oncogenesis demonstrated that key oncogenes were overexpressed in the larger nodules when compared with smaller AIMAH nodules; B, a small and large nodule from patient 2, stained by hematoxylin and eosin (magnification, ×5): BCL-2 and c-KIT were highly expressed in the large nodule but not as much in the smaller nodule (magnification, ×10); C, strong CTNNB1 staining in a large nodule and weak staining in the small nodule (magnification, ×10).
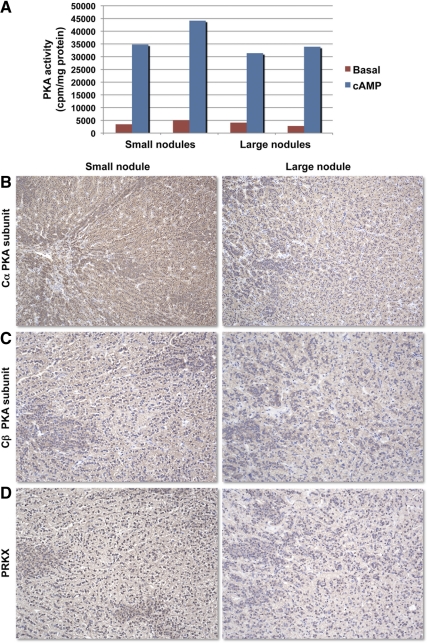
PKA activity declined in larger nodules
PKA activity was measured in four AIMAH nodules (Fig. 5A). After cAMP exposure, smaller nodules had a tendency to have higher total kinase activity compared with larger ones (39.453 ± 4.682 vs. 32.637 ± 1261 cpm/mg protein). Expression of PKA catalytic subunits (Cα, Cβ, and PRKX) decreased in macronodules when compared with micronodules (Fig. 5, B–D); this decrease in expression of the main catalytic subunits, may account for the tendency for lower total PKA activity in larger nodules.
Fig. 5.
A, Basal and cAMP-stimulated PKA activity in AIMAH nodules of different size from the same subject; C–D, Cα (B) and Cβ (C) PKA catalytic subunits and PRKX (D) staining was weaker in a large nodule compared with the small nodule from the same patient (magnification, ×5).
Discussion
AIMAH is a clinically and genetically heterogeneous disorder that can be associated with aberrant hormone receptors (8, 9, 18). It is frequently associated with subclinical hypercortisolism or atypical CS (1, 2). Given the genetic heterogeneity of AIMAH, genomic integrated approaches can reveal novel aspects involved in tumor progression and offer the possibility of more informed clinical decision making and may yield novel therapeutic targets (15, 32–34). In this study, our data set represents, to our knowledge, the first fully integrated analysis of AIMAH with whole-genome profiling and oligo-aCGH in nodules of different size from the same patient.
Previous cytogenetic studies in adrenal tumors showed that the number of chromosomal aberrations is strongly correlated with the size of the tumor (10, 11, 13). Here, we first demonstrated a positive association between chromosomal gains and the size of the nodules within the same adrenal gland from AIMAH patients. With the direct comparison of gene profiling and CGH results performed on the same samples, an overlap between significantly overexpressed genes and chromosomal amplification was established in about half of the genes and respective genetic loci.
The tetraspanin 8 (TSPAN8) gene was the most overexpressed transcript in all AIMAH nodules. Interestingly, TSPAN8 overexpression was not associated with gene amplification in the CGH data. Tetraspanin 8 is a cell surface glycoprotein that is known to complex with integrins and mediates signal transduction events that play a role in the regulation of cell development, activation, growth, and motility (35, 36). Tetraspanin 8 is an important angiogenesis inducer within tumors and also in tumor-free tissues (37).
Although the most over- and underexpressed genes were similar in all nodules, the functional analysis revealed different gene set enrichment that corresponded to the nodule size. The smallest nodules were mostly enriched for metabolic pathways, ECM interaction, and MAPK signaling pathways, whereas a high statistical association with p53 signaling and cancer pathways was found in the largest nodules.
It should be noted that, although we have identified activation of oncogenic pathways in AIMAH, there is no clinical evidence, so far, supporting a high risk of malignancy among patients with AIMAH. Confirmatory studies demonstrated that BCL2, E2F1, EGF, c-KIT, MYB, PRKCA, and CTNNB1 were overexpressed in larger nodules at the messenger and/or protein levels but, AIMAH is a polyclonal and genetically heterogeneous disorder (18, 38). Chromosomal gains and enrichment of oncogenic pathways apparently provide functional advantage in longstanding AIMAH but may not necessarily predispose to cancer.
Chromosomal enrichment analysis by GSEA showed that chromosomes 20q13 and 14q23 were overexpressed in larger nodules when compared with smaller ones. Duplication of 14q23 has been previously associated with congenital abnormalities, and 14q is frequently amplified in various sarcomas (39, 40). On the other hand, chromosomal 20q13 amplification has been associated with tumor progression and poor prognosis in ovarian and breast cancer (41–43). In our study, an enriched gene set in chromosome 20q13 revealed important genes that can be associated with AIMAH progression, such as LAMA5, BIRC7, PKIG, PRIC285, and RGS19. Laminin α5 protein encoded by LAMA5 gene belongs to the α-subfamily of laminin chains and is a major component of basement membranes. Recently, Paquet-Fifield et al. (44) showed that laminin α5 promotes epithelial cell proliferation and skin regeneration by modifying the ECM microenvironment.
Wnt signaling pathway is a master regulator of tumorigenesis driven by PKA dysregulation (21, 26, 45–47). We demonstrated here that WISP1 and WNT4 were significantly overexpressed in AIMAH nodules, as previously reported (21). Interestingly, nuclear CTNNB1 staining was found to be higher in macronodules when compared with micronodules. Furthermore, PKA catalytic subunits were strongly expressed in both micro- and macronodules, but their expression was lower in macronodules at the protein level. PKA signaling involvement in AIMAH was initially shown by our group (17); AIMAH displayed somatic 17q22-24 allelic losses and an important increase in cAMP responsiveness.
In conclusion, our data provide a unique public resource for the molecular features of AIMAH; in this study, we showed for the first time that larger nodules in this polyclonal disorder represented a progression to a more tumor-like profile with an increased number of chromosomal aberrations and an expression signature that was enriched with cancer-involved pathways and oncogenic transformation. This is supportive of the original hypothesis (22) that proposed that adrenal tissue was not different from other tissues in the process of carcinogenesis; just like polyps in colonic tissues, smaller polyclonal nodules in adrenal cortex, when they grow, have the capacity of activating oncogenic pathways and, thus, develop the potential, at least in theory, of monoclonal neoplastic transformation.
Acknowledgments
This work was supported by U.S. National Institutes of Health, Eunice Kennedy Shriver National Institute of Child Health and Human Development intramural project Z01-HD-000642-04 (to C.A.S.).
Current address for H.-P.H.: Department of Pediatrics, Kaohsiung Municipal HsiaoKang Hospital, Kaohsiung Medical University, Taiwan.
Disclosure Summary: The authors have nothing to disclose.
Footnotes
- AIMAH
- ACTH-independent macronodular adrenal hyperplasia
- CGH
- comparative genomic hybridization
- CS
- Cushing's syndrome
- DEX
- dexamethasone
- ECM
- extracellular matrix
- GSEA
- gene set enrichment analysis
- oligo-aCGH
- oligonucleotide array CGH
- PKA
- protein kinase A.
References
- 1. Lacroix A. 2009. ACTH-independent macronodular adrenal hyperplasia. Best Pract Res Clin Endocrinol Metab 23:245–259 [DOI] [PubMed] [Google Scholar]
- 2. Lacroix A, Bourdeau I, Lampron A, Mazzuco TL, Tremblay J, Hamet P. 2010. Aberrant G-protein coupled receptor expression in relation to adrenocortical overfunction. Clin Endocrinol (Oxf) 73:1–15 [DOI] [PubMed] [Google Scholar]
- 3. Stratakis CA. 2009. New genes and/or molecular pathways associated with adrenal hyperplasias and related adrenocortical tumors. Mol Cell Endocrinol 300:152–157 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Barzon L, Sonino N, Fallo F, Palu G, Boscaro M. 2003. Prevalence and natural history of adrenal incidentalomas. Eur J Endocrinol 149:273–285 [DOI] [PubMed] [Google Scholar]
- 5. Findlay JC, Sheeler LR, Engeland WC, Aron DC. 1993. Familial adrenocorticotropin-independent Cushing's syndrome with bilateral macronodular adrenal hyperplasia. J Clin Endocrinol Metab 76:189–191 [DOI] [PubMed] [Google Scholar]
- 6. Minami S, Sugihara H, Sato J, Tatsukuchi A, Sugisaki Y, Sasano H, Wakabayashi I. 1996. ACTH independent Cushing's syndrome occurring in siblings. Clin Endocrinol (Oxf) 44:483–488 [DOI] [PubMed] [Google Scholar]
- 7. Gagliardi L, Hotu C, Casey G, Braund WJ, Ling KH, Dodd T, Manavis J, Devitt PG, Cutfield R, Rudzki Z, Scott HS, Torpy DJ. 2009. Familial vasopressin-sensitive ACTH-independent macronodular adrenal hyperplasia (VPs-AIMAH): clinical studies of three kindreds. Clin Endocrinol (Oxf) 70:883–891 [DOI] [PubMed] [Google Scholar]
- 8. Lacroix A, Baldacchino V, Bourdeau I, Hamet P, Tremblay J. 2004. Cushing's syndrome variants secondary to aberrant hormone receptors. Trends Endocrinol Metab 15:375–382 [DOI] [PubMed] [Google Scholar]
- 9. Lacroix A, Ndiaye N, Tremblay J, Hamet P. 2001. Ectopic and abnormal hormone receptors in adrenal Cushing's syndrome. Endocr Rev 22:75–110 [DOI] [PubMed] [Google Scholar]
- 10. Kjellman M, Kallioniemi OP, Karhu R, Höög A, Farnebo LO, Auer G, Larsson C, Bäckdahl M. 1996. Genetic aberrations in adrenocortical tumors detected using comparative genomic hybridization correlate with tumor size and malignancy. Cancer Res 56:4219–4223 [PubMed] [Google Scholar]
- 11. Sidhu S, Marsh DJ, Theodosopoulos G, Philips J, Bambach CP, Campbell P, Magarey CJ, Russell CF, Schulte KM, Röher HD, Delbridge L, Robinson BG. 2002. Comparative genomic hybridization analysis of adrenocortical tumors. J Clin Endocrinol Metab 87:3467–3474 [DOI] [PubMed] [Google Scholar]
- 12. Figueiredo BC, Stratakis CA, Sandrini R, DeLacerda L, Pianovsky MA, Giatzakis C, Young HM, Haddad BR. 1999. Comparative genomic hybridization analysis of adrenocortical tumors of childhood. J Clin Endocrinol Metab 84:1116–1121 [DOI] [PubMed] [Google Scholar]
- 13. Zhao J, Speel EJ, Muletta-Feurer S, Rütimann K, Saremaslani P, Roth J, Heitz PU, Komminoth P. 1999. Analysis of genomic alterations in sporadic adrenocortical lesions. Gain of chromosome 17 is an early event in adrenocortical tumorigenesis. Am J Pathol 155:1039–1045 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Dohna M, Reincke M, Mincheva A, Allolio B, Solinas-Toldo S, Lichter P. 2000. Adrenocortical carcinoma is characterized by a high frequency of chromosomal gains and high-level amplifications. Genes Chromosomes Cancer 28:145–152 [PubMed] [Google Scholar]
- 15. Szabó PM, Tamási V, Molnár V, Andrásfalvy M, Tömböl Z, Farkas R, Kövesdi K, Patócs A, Tóth M, Szalai C, Falus A, Rácz K, Igaz P. 2010. Meta-analysis of adrenocortical tumour genomics data: novel pathogenic pathways revealed. Oncogene 29:3163–3172 [DOI] [PubMed] [Google Scholar]
- 16. Bertherat J, Groussin L, Sandrini F, Matyakhina L, Bei T, Stergiopoulos S, Papageorgiou T, Bourdeau I, Kirschner LS, Vincent-Dejean C, Perlemoine K, Gicquel C, Bertagna X, Stratakis CA. 2003. Molecular and functional analysis of PRKAR1A and its locus (17q22-24) in sporadic adrenocortical tumors: 17q losses, somatic mutations, and protein kinase A expression and activity. Cancer Res 63:5308–5319 [PubMed] [Google Scholar]
- 17. Bourdeau I, Matyakhina L, Stergiopoulos SG, Sandrini F, Boikos S, Stratakis CA. 2006. 17q22-24 chromosomal losses and alterations of protein kinase a subunit expression and activity in adrenocorticotropin-independent macronodular adrenal hyperplasia. J Clin Endocrinol Metab 91:3626–3632 [DOI] [PubMed] [Google Scholar]
- 18. Hsiao HP, Kirschner LS, Bourdeau I, Keil MF, Boikos SA, Verma S, Robinson-White AJ, Nesterova M, Lacroix A, Stratakis CA. 2009. Clinical and genetic heterogeneity, overlap with other tumor syndromes, and atypical glucocorticoid hormone secretion in adrenocorticotropin-independent macronodular adrenal hyperplasia compared with other adrenocortical tumors. J Clin Endocrinol Metab 94:2930–2937 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Kirschner LS, Carney JA, Pack SD, Taymans SE, Giatzakis C, Cho YS, Cho-Chung YS, Stratakis CA. 2000. Mutations of the gene encoding the protein kinase A type I-α regulatory subunit in patients with the Carney complex. Nat Genet 26:89–92 [DOI] [PubMed] [Google Scholar]
- 20. Horvath A, Mathyakina L, Vong Q, Baxendale V, Pang AL, Chan WY, Stratakis CA. 2006. Serial analysis of gene expression in adrenocortical hyperplasia caused by a germline PRKAR1A mutation. J Clin Endocrinol Metab 91:584–596 [DOI] [PubMed] [Google Scholar]
- 21. Bourdeau I, Antonini SR, Lacroix A, Kirschner LS, Matyakhina L, Lorang D, Libutti SK, Stratakis CA. 2004. Gene array analysis of macronodular adrenal hyperplasia confirms clinical heterogeneity and identifies several candidate genes as molecular mediators. Oncogene 23:1575–1585 [DOI] [PubMed] [Google Scholar]
- 22. Stratakis CA. 2003. Genetics of adrenocortical tumors: gatekeepers, landscapers and conductors in symphony. Trends Endocrinol Metab 14:404–410 [DOI] [PubMed] [Google Scholar]
- 23. Papanicolaou DA, Yanovski JA, Cutler GB, Jr, Chrousos GP, Nieman LK. 1998. A single midnight serum cortisol measurement distinguishes Cushing's syndrome from pseudo-Cushing states. J Clin Endocrinol Metab 83:1163–1167 [DOI] [PubMed] [Google Scholar]
- 24. Gomez MT, Malozowski S, Winterer J, Vamvakopoulos NC, Chrousos GP. 1991. Urinary free cortisol values in normal children and adolescents. J Pediatr 118:256–258 [DOI] [PubMed] [Google Scholar]
- 25. Cheadle C, Vawter MP, Freed WJ, Becker KG. 2003. Analysis of microarray data using Z score transformation. J Mol Diagn 5:73–81 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Almeida MQ, Muchow M, Boikos S, Bauer AJ, Griffin KJ, Tsang KM, Cheadle C, Watkins T, Wen F, Starost MF, Bossis I, Nesterova M, Stratakis CA. 2010. Mouse Prkar1a haploinsufficiency leads to an increase in tumors in the Trp53+/− or Rb1+/− backgrounds and chemically induced skin papillomas by dysregulation of the cell cycle and Wnt signaling. Hum Mol Genet 19:1387–1398 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Huang da W, Sherman BT, Tan Q, Kir J, Liu D, Bryant D, Guo Y, Stephens R, Baseler MW, Lane HC, Lempicki RA. 2007. DAVID bioinformatics resources: expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res 35(Web Server issue):W169–W75 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Huang da W, Sherman BT, Lempicki RA. 2009. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57 [DOI] [PubMed] [Google Scholar]
- 29. Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP. 2005. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Livak KJ, Schmittgen TD. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408 [DOI] [PubMed] [Google Scholar]
- 31. Rohlff C, Clair T, Cho-Chung YS. 1993. 8-Cl-cAMP induces truncation and down-regulation of the RI α-subunit and up-regulation of the RII β-subunit of cAMP-dependent protein kinase leading to type II holoenzyme-dependent growth inhibition and differentiation of HL-60 leukemia cells. J Biol Chem 268:5774–5782 [PubMed] [Google Scholar]
- 32. Taylor BS, Schultz N, Hieronymus H, Gopalan A, Xiao Y, Carver BS, Arora VK, Kaushik P, Cerami E, Reva B, Antipin Y, Mitsiades N, Landers T, Dolgalev I, Major JE, Wilson M, Socci ND, Lash AE, Heguy A, Eastham JA, Scher HI, Reuter VE, Scardino PT, Sander C, Sawyers CL, Gerald WL. 2010. Integrative genomic profiling of human prostate cancer. Cancer Cell 18:11–22 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Chitale D, Gong Y, Taylor BS, Broderick S, Brennan C, Somwar R, Golas B, Wang L, Motoi N, Szoke J, Reinersman JM, Major J, Sander C, Seshan VE, Zakowski MF, Rusch V, Pao W, Gerald W, Ladanyi M. 2009. An integrated genomic analysis of lung cancer reveals loss of DUSP4 in EGFR-mutant tumors. Oncogene 28:2773–2783 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Ding L, Getz G, Wheeler DA, Mardis ER, McLellan MD, Cibulskis K, Sougnez C, Greulich H, Muzny DM, Morgan MB, Fulton L, Fulton RS, Zhang Q, Wendl MC, Lawrence MS, Larson DE, Chen K, Dooling DJ, Sabo A, Hawes AC, Shen H, Jhangiani SN, Lewis LR, Hall O, Zhu Y, Mathew T, Ren Y, Yao J, Scherer SE, Clerc K, Metcalf GA, Ng B, Milosavljevic A, Gonzalez-Garay ML, Osborne JR, Meyer R, Shi X, Tang Y, Koboldt DC, Lin L, Abbott R, Miner TL, Pohl C, Fewell G, Haipek C, Schmidt H, Dunford-Shore BH, Kraja A, Crosby SD, Sawyer CS, Vickery T, Sander S, Robinson J, Winckler W, Baldwin J, Chirieac LR, Dutt A, Fennell T, Hanna M, Johnson BE, Onofrio RC, Thomas RK, Tonon G, Weir BA, Zhao X, Ziaugra L, Zody MC, Giordano T, Orringer MB, Roth JA, Spitz MR, Wistuba II, Ozenberger B, Good PJ, Chang AC, Beer DG, Watson MA, Ladanyi M, Broderick S, Yoshizawa A, Travis WD, Pao W, Province MA, Weinstock GM, Varmus HE, Gabriel SB, Lander ES, Gibbs RA, Meyerson M, Wilson RK. 2008. Somatic mutations affect key pathways in lung adenocarcinoma. Nature 455:1069–1075 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Roskoski R., Jr 2007. Vascular endothelial growth factor (VEGF) signaling in tumor progression. Crit Rev Oncol Hematol 62:179–213 [DOI] [PubMed] [Google Scholar]
- 36. Gesierich S, Berezovskiy I, Ryschich E, Zöller M. 2006. Systemic induction of the angiogenesis switch by the tetraspanin D6.1A/CO-029. Cancer Res 66:7083–7094 [DOI] [PubMed] [Google Scholar]
- 37. Nazarenko I, Rana S, Baumann A, McAlear J, Hellwig A, Trendelenburg M, Lochnit G, Preissner KT, Zöller M. 2010. Cell surface tetraspanin Tspan8 contributes to molecular pathways of exosome-induced endothelial cell activation. Cancer Res 70:1668–1678 [DOI] [PubMed] [Google Scholar]
- 38. Beuschlein F, Reincke M, Karl M, Travis WD, Jaursch-Hancke C, Abdelhamid S, Chrousos GP, Allolio B. 1994. Clonal composition of human adrenocortical neoplasms. Cancer Res 54:4927–4932 [PubMed] [Google Scholar]
- 39. Cohen MM, Charrow J, Balkin NE, Harris CJ. 1983. Partial trisomy 14 (q23 leads to qter) via segregation of a 14/X translocation. Am J Hum Genet 35:635–644 [PMC free article] [PubMed] [Google Scholar]
- 40. Ou Z, Martin DM, Bedoyan JK, Cooper ML, Chinault AC, Stankiewicz P, Cheung SW. 2008. Branchiootorenal syndrome and oculoauriculovertebral spectrum features associated with duplication of SIX1, SIX6, and OTX2 resulting from a complex chromosomal rearrangement. Am J Med Genet A 146A:2480– 2489 [DOI] [PubMed] [Google Scholar]
- 41. Tominaga E, Tsuda H, Arao T, Nishimura S, Takano M, Kataoka F, Nomura H, Hirasawa A, Aoki D, Nishio K. 2010. Amplification of GNAS may be an independent, qualitative, and reproducible biomarker to predict progression-free survival in epithelial ovarian cancer. Gynecol Oncol 118:160–166 [DOI] [PubMed] [Google Scholar]
- 42. Hines WC, Bazarov AV, Mukhopadhyay R, Yaswen P. 2010. BORIS (CTCFL) is not expressed in most human breast cell lines and high grade breast carcinomas. PLoS One 5:e9738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Plevova P, Cerna D, Balcar A, Foretova L, Zapletalova J, Silhanova E, Curik R, Dvorackova J. 2010. CCND1 and ZNF217 gene amplification is equally frequent in BRCA1 and BRCA2 associated and non-BRCA breast cancer. Neoplasma 57:325–332 [DOI] [PubMed] [Google Scholar]
- 44. Paquet-Fifield S, Schlüter H, Li A, Aitken T, Gangatirkar P, Blashki D, Koelmeyer R, Pouliot N, Palatsides M, Ellis S, Brouard N, Zannettino A, Saunders N, Thompson N, Li J, Kaur P. 2009. A role for pericytes as microenvironmental regulators of human skin tissue regeneration. J Clin Invest 119:2795–2806 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Gaujoux S, Tissier F, Groussin L, Libé R, Ragazzon B, Launay P, Audebourg A, Dousset B, Bertagna X, Bertherat J. 2008. Wnt/β-catenin and 3′,5′-cyclic adenosine 5′-monophosphate/protein kinase A signaling pathways alterations and somatic β-catenin gene mutations in the progression of adrenocortical tumors. J Clin Endocrinol Metab 93:4135–4140 [DOI] [PubMed] [Google Scholar]
- 46. Tadjine M, Lampron A, Ouadi L, Horvath A, Stratakis CA, Bourdeau I. 2008. Detection of somatic β-catenin mutations in primary pigmented nodular adrenocortical disease (PPNAD). Clin Endocrinol (Oxf) 69:367–373 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Iliopoulos D, Bimpaki EI, Nesterova M, Stratakis CA. 2009. MicroRNA signature of primary pigmented nodular adrenocortical disease: clinical correlations and regulation of Wnt signaling. Cancer Res 69:3278–3282 [DOI] [PMC free article] [PubMed] [Google Scholar]