Abstract
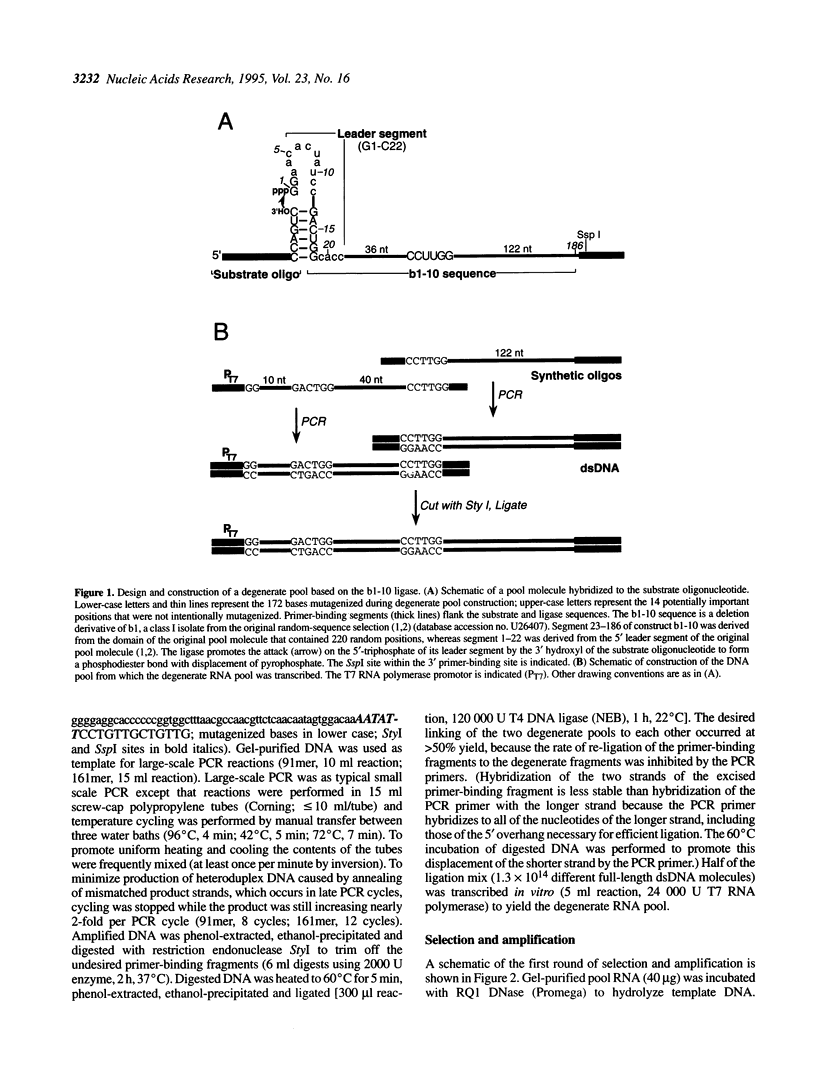
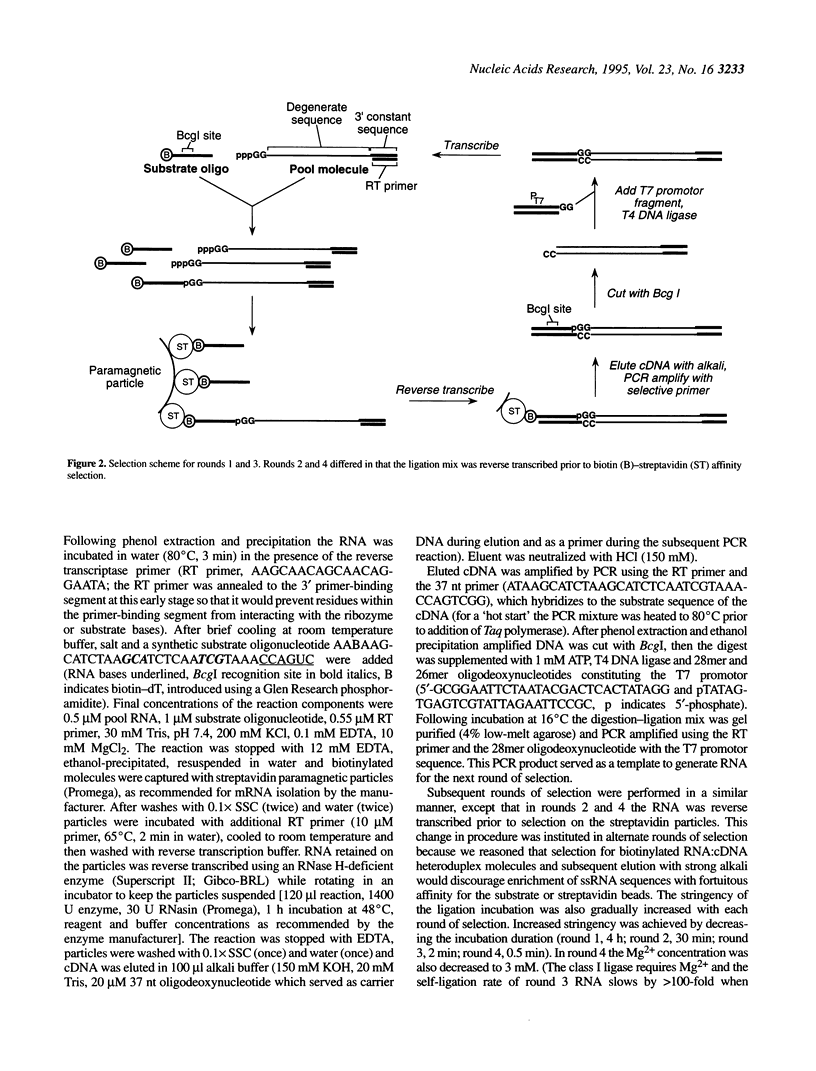
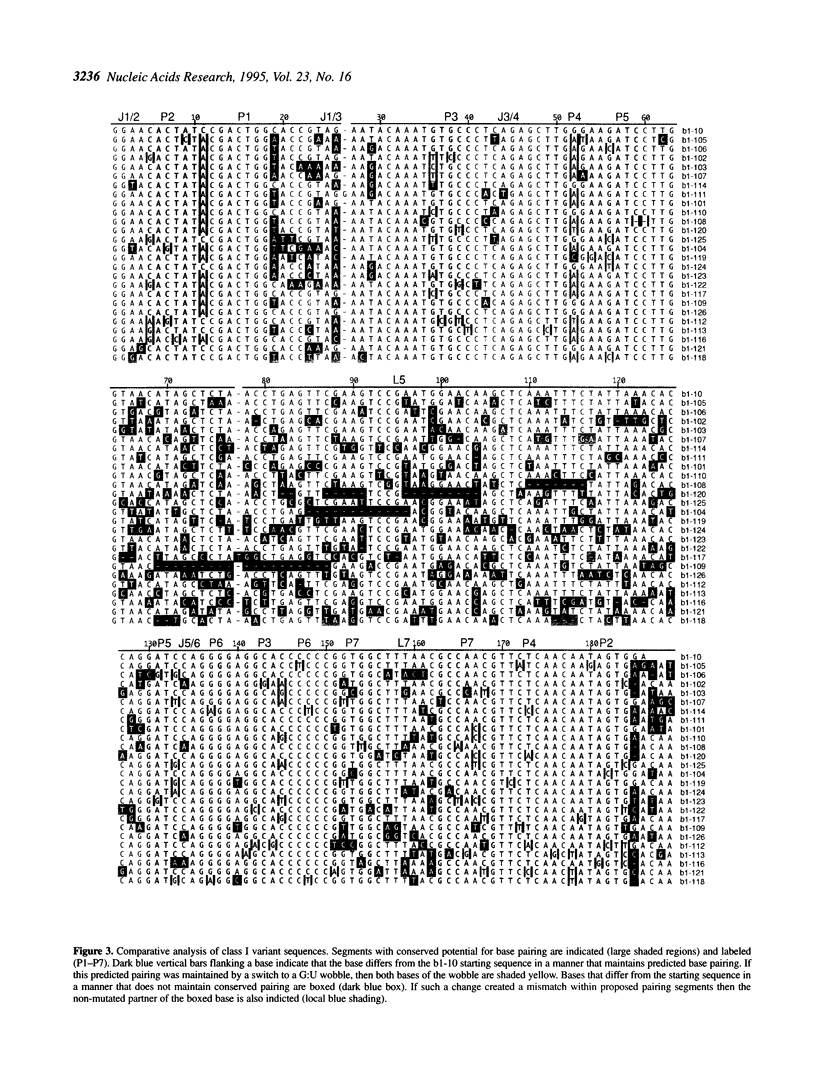
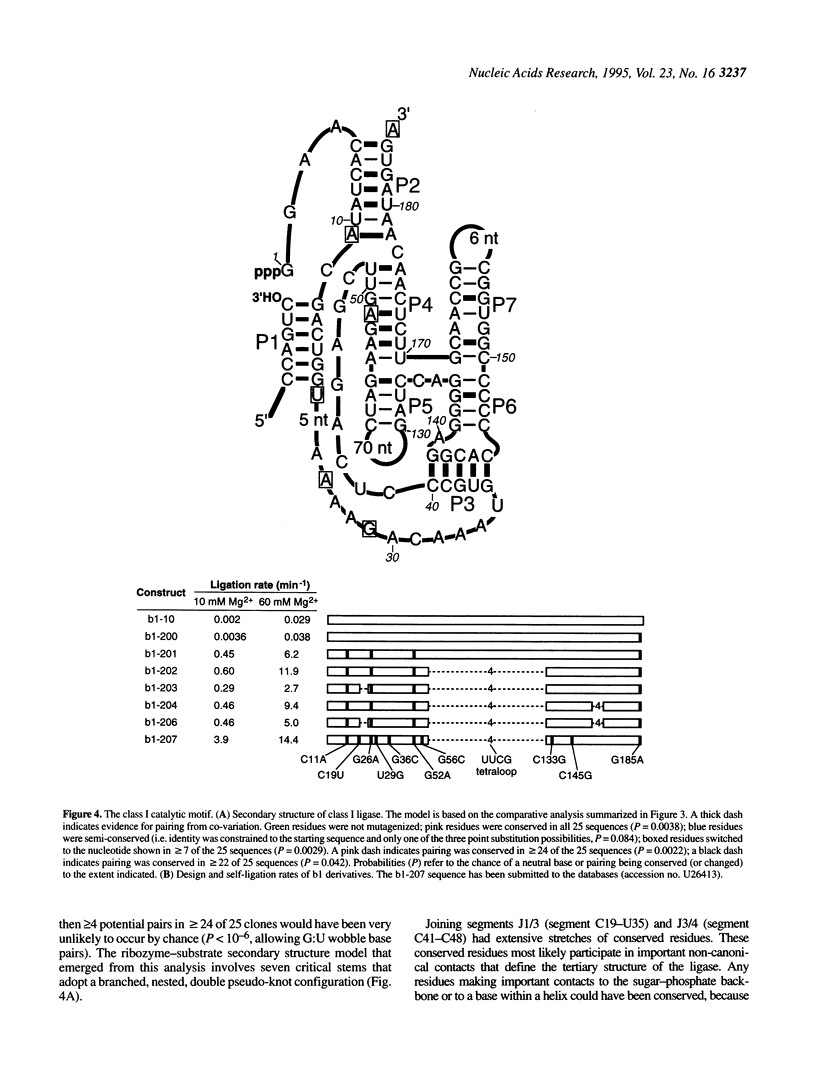
In vitro selection can generate functional sequence variants of an RNA structural motif that are useful for comparative analysis. The technique is particularly valuable in cases where natural variation is unavailable or non-existent. We report the extension of this approach to a new extreme--the identification of a 112 nt ribozyme secondary structure imbedded within a 186 nt RNA. A pool of 10(14) variants of an RNA ligase ribozyme was generated using combinatorial chemical synthesis coupled with combinatorial enzymatic ligation such that 172 of the 186 relevant positions were partially mutagenized. Active variants of this pool were enriched using an in vitro selection scheme that retains the sequence variability at positions very close to the ligation junction. Ligases isolated after four rounds of selection catalyzed self-ligation up to 700 times faster than the starting sequence. Comparative analysis of the isolates indicated that when complexed with substrate RNAs the ligase forms a nested, double pseudo-knot secondary structure with seven stems and several important joining segments. Comparative analysis also suggested the identity of mutations that account for the increased activity of the selected ligase variants; designed constructs incorporating combinations of these changes were more active than any of the individual ligase isolates.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bartel D. P., Szostak J. W. Isolation of new ribozymes from a large pool of random sequences [see comment]. Science. 1993 Sep 10;261(5127):1411–1418. doi: 10.1126/science.7690155. [DOI] [PubMed] [Google Scholar]
- Bartel D. P., Zapp M. L., Green M. R., Szostak J. W. HIV-1 Rev regulation involves recognition of non-Watson-Crick base pairs in viral RNA. Cell. 1991 Nov 1;67(3):529–536. doi: 10.1016/0092-8674(91)90527-6. [DOI] [PubMed] [Google Scholar]
- Berzal-Herranz A., Joseph S., Burke J. M. In vitro selection of active hairpin ribozymes by sequential RNA-catalyzed cleavage and ligation reactions. Genes Dev. 1992 Jan;6(1):129–134. doi: 10.1101/gad.6.1.129. [DOI] [PubMed] [Google Scholar]
- Berzal-Herranz A., Joseph S., Chowrira B. M., Butcher S. E., Burke J. M. Essential nucleotide sequences and secondary structure elements of the hairpin ribozyme. EMBO J. 1993 Jun;12(6):2567–2573. doi: 10.1002/j.1460-2075.1993.tb05912.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burke J. M., Belfort M., Cech T. R., Davies R. W., Schweyen R. J., Shub D. A., Szostak J. W., Tabak H. F. Structural conventions for group I introns. Nucleic Acids Res. 1987 Sep 25;15(18):7217–7221. doi: 10.1093/nar/15.18.7217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burke J. M., Berzal-Herranz A. In vitro selection and evolution of RNA: applications for catalytic RNA, molecular recognition, and drug discovery. FASEB J. 1993 Jan;7(1):106–112. doi: 10.1096/fasebj.7.1.8422956. [DOI] [PubMed] [Google Scholar]
- Ekland E. H., Szostak J. W., Bartel D. P. Structurally complex and highly active RNA ligases derived from random RNA sequences. Science. 1995 Jul 21;269(5222):364–370. doi: 10.1126/science.7618102. [DOI] [PubMed] [Google Scholar]
- Green R., Ellington A. D., Szostak J. W. In vitro genetic analysis of the Tetrahymena self-splicing intron. Nature. 1990 Sep 27;347(6291):406–408. doi: 10.1038/347406a0. [DOI] [PubMed] [Google Scholar]
- Green R., Szostak J. W. In vitro genetic analysis of the hinge region between helical elements P5-P4-P6 and P7-P3-P8 in the sunY group I self-splicing intron. J Mol Biol. 1994 Jan 7;235(1):140–155. doi: 10.1016/s0022-2836(05)80022-1. [DOI] [PubMed] [Google Scholar]
- Joyce G. F. In vitro evolution of nucleic acids. Curr Opin Struct Biol. 1994;4:331–336. doi: 10.1016/s0959-440x(94)90100-7. [DOI] [PubMed] [Google Scholar]
- Levitt M. Detailed molecular model for transfer ribonucleic acid. Nature. 1969 Nov 22;224(5221):759–763. doi: 10.1038/224759a0. [DOI] [PubMed] [Google Scholar]
- Michel F., Westhof E. Modelling of the three-dimensional architecture of group I catalytic introns based on comparative sequence analysis. J Mol Biol. 1990 Dec 5;216(3):585–610. doi: 10.1016/0022-2836(90)90386-Z. [DOI] [PubMed] [Google Scholar]
- Pan T., Uhlenbeck O. C. In vitro selection of RNAs that undergo autolytic cleavage with Pb2+. Biochemistry. 1992 Apr 28;31(16):3887–3895. doi: 10.1021/bi00131a001. [DOI] [PubMed] [Google Scholar]
- Tsai D. E., Harper D. S., Keene J. D. U1-snRNP-A protein selects a ten nucleotide consensus sequence from a degenerate RNA pool presented in various structural contexts. Nucleic Acids Res. 1991 Sep 25;19(18):4931–4936. doi: 10.1093/nar/19.18.4931. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tuerk C., Gauss P., Thermes C., Groebe D. R., Gayle M., Guild N., Stormo G., d'Aubenton-Carafa Y., Uhlenbeck O. C., Tinoco I., Jr CUUCGG hairpins: extraordinarily stable RNA secondary structures associated with various biochemical processes. Proc Natl Acad Sci U S A. 1988 Mar;85(5):1364–1368. doi: 10.1073/pnas.85.5.1364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tuerk C., Gold L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science. 1990 Aug 3;249(4968):505–510. doi: 10.1126/science.2200121. [DOI] [PubMed] [Google Scholar]
- Tuerk C., MacDougal S., Gold L. RNA pseudoknots that inhibit human immunodeficiency virus type 1 reverse transcriptase. Proc Natl Acad Sci U S A. 1992 Aug 1;89(15):6988–6992. doi: 10.1073/pnas.89.15.6988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woese C. R., Gutell R., Gupta R., Noller H. F. Detailed analysis of the higher-order structure of 16S-like ribosomal ribonucleic acids. Microbiol Rev. 1983 Dec;47(4):621–669. doi: 10.1128/mr.47.4.621-669.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]



