Abstract
Background:
The Tissue Microarray Data Exchange Specification (TMA DES) is an eXtensible Markup Language (XML) specification for encoding TMA experiment data in a machine-readable format that is also human readable. TMA DES defines Common Data Elements (CDEs) that form a basic vocabulary for describing TMA data. TMA data are routinely subjected to univariate and multivariate statistical analysis to determine differences or similarities between pathologically distinct groups of tumors for one or more markers or between markers for different groups. Such statistical analysis tests include the t-test, ANOVA, Chi-square, Mann-Whitney U, and Kruskal-Wallis tests. All these generate output that needs to be recorded and stored with TMA data.
Materials and Methods:
We propose extending the TMA DES to include syntactic and semantic definitions of CDEs for describing the results of statistical analyses performed upon TMA DES data. These CDEs are described in this paper and it is illustrated how they can be added to the TMA DES. We created a Document Type Definition (DTD) file defining the syntax for these CDEs, and a set of ISO 11179 entries providing semantic definitions for them. We describe how we wrote a program in R that read TMA DES data from an XML file, performed statistical analyses on that data, and created a new XML file containing both the original XML data and CDEs representing the results of our analyses. This XML file was submitted to XML parsers in order to confirm that they conformed to the syntax defined in our extended DTD file. TMA DES XML files with deliberately introduced errors were also parsed in order to verify that our new DTD file could perform error checking. Finally, we also validated an existing TMA DES XML file against our DTD file in order to demonstrate the backward compatibility of our DTD.
Results:
Our experiments demonstrated the encoding of analysis results using our proposed CDEs. We used XML parsers to confirm that these XML data were syntactically correct and conformed to the rules specified in our extended TMA DES DTD. We also demonstrated that this extended DTD was capable of being used to successfully perform error checking, and was backward compatible with pre-existing TMA DES data which did not use our new CDEs.
Conclusions:
The TMA DES allows Tissue Microarray data to be shared. A variety of statistical tests are used to analyze such data. We have proposed a set of CDEs as an extension to the TMA DES which can be used to annotate TMA DES data with the results of statistical analyses performed on that data. We performed experiments which demonstrated the usage of TMA DES data containing our proposed CDEs.
Keywords: CDEs, DTD, statistical analysis, tissue microarray, TMA Data Exchange Specification, XML
BACKGROUND
Tissue microarray (TMA) is a cost-effective and high-throughput technology that allows hundreds of tissue samples to be represented and analyzed in a single paraffin histology block.[1] It has been developed into a very effective tool for rapid molecular analysis of tissue to provide new diagnostic and prognostic biomarkers, as well as potential therapeutic targets in disease. Critically, TMA allows tiny representative cores to be taken from a tissue sample, meaning whole samples are not exhausted by a single research study. TMA blocks can be cut to provide thin tissue sections that are subsequently mounted onto glass slides and, most commonly, stained by immunohistochemical techniques using antibodies specific for a protein of interest. Use of TMAs has had particular success in cancer research, providing new diagnostic biomarkers for different tumor types and shedding new light on tumor biology.[2]
Like other array-based techniques, TMAs are typically associated with a range of data. Such data include identifiers for cores, slides, blocks, the creator of the block, data generated by performing experiments on cell samples contained in the block, etc. These data must be stored electronically in order for it to be archived and analyzed efficiently. If researchers in different laboratories wish to share their data, or if heterogeneous software applications which analyze that data need to interoperate, there is a requirement for a uniform means of describing the data. The TMA Data Exchange Specification (TMA DES)[3] allows description of TMAs and their associated data. It was created by the Technical Standards Committee of the Association for Pathology Informatics. The “Common Data Elements” (CDEs) defined in this specification have universally agreed upon semantics, and represent TMA blocks, slides and cores, and data associated with these objects. TMA DES uses eXtensible Markup Language (XML)[4] to encode documents describing TMA blocks and data associated with those blocks. TMA DES documents are composed of 4 types of data, contained in header, block, slide, and core CDEs. Although a TMA DES document contains only one header CDE, multiple block, slide, and core CDEs can be present in these files. CDEs contained within the header CDE describe the actual document itself, e.g., who created it, when it was created, etc. The block CDE describes a TMA block, e.g., how many cores the block has, how they are arranged, etc. The slide CDE describes slides that are generated by sectioning a block, and the core CDE describes cores in a TMA block. A Document Type Definition (DTD) file specifies the valid syntax and structure for TMA DES data. Examples of applications which allow TMA data to be exported in the TMA DES format include Xperanto-TMA[5] and Tissue Array Management and Evaluation Environment (TAMEE).[6]
Data acquired from TMA experiments are routinely subjected to both univariate and multivariate statistical analysis tests. Table 1 illustrates a number of such tests and their purpose.
Table 1.
Statistical tests and their purposes
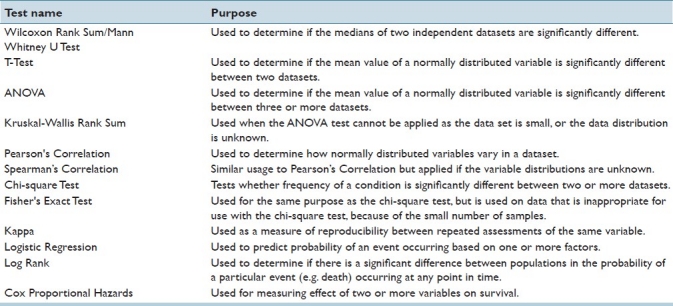

The following examples demonstrate the diversity of statistical tests applied to TMA data and the varying domains of application. Pathologists seeking novel biomarkers for prostate cancer diagnosis use Spearman's Correlation to assess the relationship between this diagnosis and expression of the GOLPH2 protein in tissue.[7] A study involving immunohistochemical staining on TMA slides containing tissue taken from a cohort of breast cancer patients[8] used the Cox Proportional Hazards model to assess the prognostic power of a panel of biomarkers. Often, multiple statistical tests are applied to data in a study and a typical example involved analysis of data generated from immunohistochemistry stained TMA slides containing pathologic white matter taken from Alzheimer's disease (AD) patients.[9] In this study, the Spearman, Kruskal-Wallis, and Mann-Whitney statistical tests were used to determine if there were group differences and a correlation between vessel quantities and the neuropathological severity of AD. More complex analyses involve application of Cox Proportional Hazards for survival analysis. An example of this type of analysis is given by Rubin et al., where models were generated to predict time to prostate-specific antigen recurrence after radical prostatectomy for clinically localized prostate cancer using Ki-67 immunohistochemical data from TMA slides.[10]
There are a number of existing XML specifications which describe statistical data, for example, the Predictive Model Markup Language (PMML)[11] and StatDataML.[12] However, these specifications do not define CDEs describing the results of widely used statistical tests which are applied to TMA data. For example, PMML contains CDEs for describing the results of two statistical tests (logistic regression and ANOVA), but does not contain CDEs for describing the results of some previously mentioned tests (e.g., Cox Proportional Hazards, Spearman's Correlation, Kruskal-Wallis). To facilitate storage of data analysis results and sharing of results within the cancer research domain, there is a pressing need for such CDEs to be included in the TMA DES. These CDEs can be used to create XML representations of the many forms of statistical analysis applied to TMA data. In light of this, we propose extending the TMA DES to include such CDEs. We will illustrate the use of these CDEs to create XML descriptions of a number of statistical tests. TMA DES was designed so that it could be supplemented by internal DTD extensions for locally defined TMA data elements (LDEs). Our proposed CDEs can serve as “building blocks” used by these LDEs to describe the results of further statistical tests that have not been mentioned in this paper. However, DTDs do not describe the types of data that CDEs must contain, or the semantic definition of CDEs. A document containing such definitions[13] was included with an earlier paper describing the TMA DES. This document conforms to the ISO 11179 specification,[14] a standard for specifying attributes of CDEs such as data types, semantic definitions, language, whether or not the CDE is mandatory, or what the maximum allowed occurrence of the CDE is. We have prepared ISO 11179 semantic definitions for the statistical analysis CDEs described in this paper (Supplementary File 3, CDEDefinitions.rtf).
In the “Methods” segment of this paper, we detail our proposed extension of the TMA DES to include the results of statistical analysis to TMA data. In the “Results” segment, we describe validation of these TMA DES extensions and show application to an example dataset.
MATERIALS AND METHODS
The TMA DES, block, slide, and core CDEs can be extended with new LDEs. However, statistical tests could be carried out on multiple blocks, or slides taken from multiple blocks, etc. In such cases, the test results are independent of any particular block/slide/core. Therefore, this paper proposes an optional segment for TMA DES to contain the results of statistical analysis performed on data associated with multiple blocks, slides, or cores.
The particular blocks, slides, or cores associated with the data to which statistical analysis has been applied are specified using a CDE called “datasetForAnalysis.” The datasetForAnalysis CDE can contain any of the following CDEs:
Block_identifier
This CDE identifies a block associated with data on which analysis was performed. Unless specific slide/core identifiers are also included in the datasetForAnalysis CDE, all nested slides and cores associated with this block are included in the analysis data. This may lead to cases where the block_identifier can be incorrectly used (e.g., it is falsely assumed that all slides from that block are stained identically, and it is critical for the statistical analysis being performed that this is the case). If someone wanted to unambiguously identify one of several slides/stains associated with a block, they should do so by using the slide_identifier CDE.
Slide_identifier
This CDE identifies a slide associated with data on which analysis was performed. Only core data generated from cores on this slide are included in the analysis dataset. Unless specific core_identifiers are also included in the datasetForAnalysis CDE, all cores explicitly associated with this slide are included in the analysis data.
Core_array-identifier
This CDE identifies a core associated with data on which analysis was performed. Core sections are present in every slide sectioned from a TMA block (assuming that they have not fallen off, and the sectioning has been performed correctly). The core_array-identifier CDE must be a child element of a slide_identifier CDE.
An “analysis dataset” is thus defined with a datasetForAnalysis CDE. Some of the tests listed in Table 1 are applied to a single analysis dataset; others must be applied to two, others to three or more. The definition of the CDE representing each test specifies the valid number of analysis datasets that the test can be applied to. For example, the two sample t-tests are applied to two analysis datasets, to determine if their means are significantly different. See the examples in Figures 1 and 2.
Figure 1.
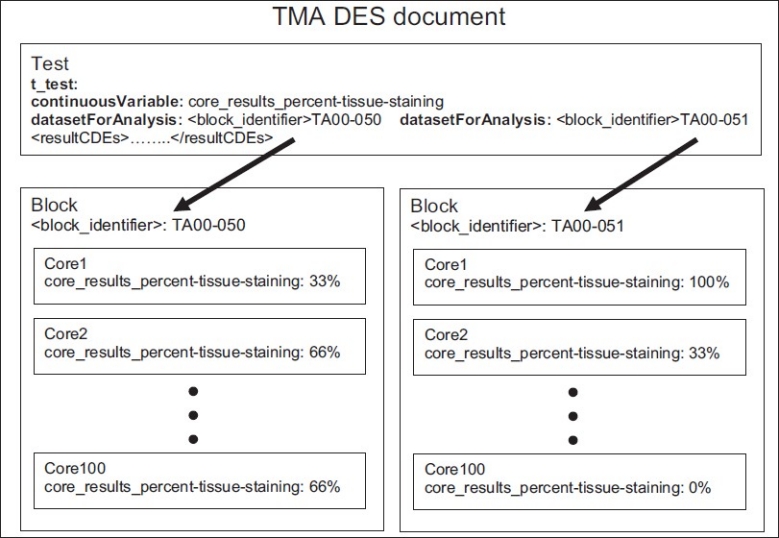
Simplified representation of an example TMA DES document that can define inputs to a t-test. Two datasetForAnalysis CDEs define two analysis datasets, each specifying the identifier for a block that is analyzed. Each core has a core_results_percent-tissue-staining CDE with a value from 0 to 100%. The mean core_results_percent-tissue-staining (because it is identified as the continuousVariable) for each block are inputs to the t-test
Figure 2.
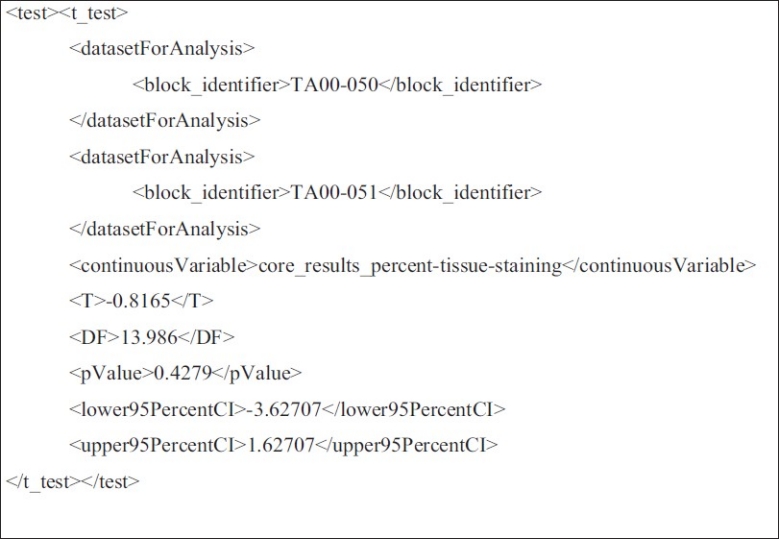

CDEs for describing results of a t-test
Multiple values (e.g., staining and intensity) from a single analysis dataset may be evaluated using one of several types of tests. Figure 3 shows an example of such a situation involving the Pearson's Correlation test.
Figure 3.
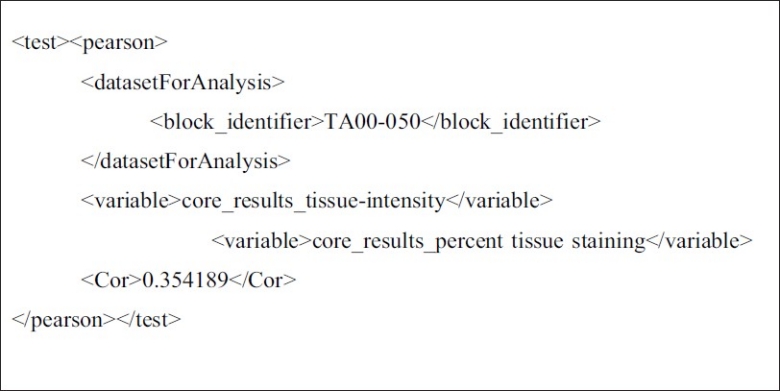

XML representation of the inputs to a Pearson's Correlation test applied to one analysis dataset. This test measures correlation between two results. Two variable CDEs identify core_results_tissue-intensity (the intensity of a stained tissue sample) and core_results_percent-tissue-staining (the percent of that same sample that is a certain color) as being tested for correlation. Correlation is determined for each core associated with “TA00-050” block
The results of correlation tests can also be described using two datasetForAnalysis CDEs. For example, a Spearman's Correlation could be performed to determine if there was a correlation between the values of an intensity scoring variable between two blocks, as shown in Figure 4. We have defined CDEs for all the statistical tests previously listed in Table 1. Examples of XML descriptions of the results of these tests can be found in Supplementary File 1 (tmaStatsAnalysis.xml). The results of further statistical tests can be represented using our CDEs as “building blocks.”
Figure 4.
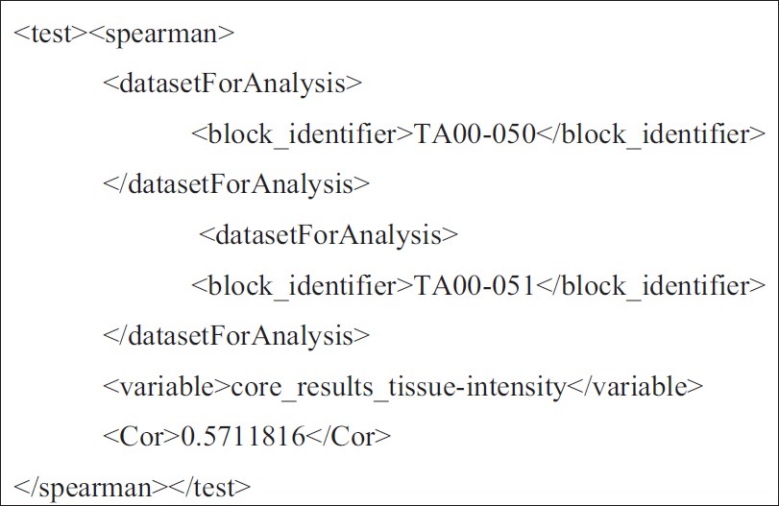

CDEs for describing results of Spearman's Correlation between a single variable in two analysis datasets
An XML document may be “well formed” and/or “valid.” A well-formed document conforms to the syntax rules defined by the XML standard. A valid XML document is composed using only CDEs specified in a DTD schema document. A TMA DES DTD[13] has been designed by Nohle et al., which outlines the structure of a TMA DES file, and specifies what CDEs a TMA DES file can contain. We propose a set of additions to the TMA DES DTD, which specify new CDEs representing the results of statistical analysis performed on TMA data. These extensions are in Supplementary File 2 (extendedTMADES.dtd). As the DTD extract below shows, we have added a child element to the existing TMA DES CDE “tma” called “analysis.”
<!ELEMENT tma (header+, block+, analysis*)>
This line indicates that the tma CDE can contain 3 child CDEs, “header,” “block,” and “analysis.” The symbols beside these CDEs indicate their cardinality. The “+” symbol indicates that there can be any number of this CDE in the range 1..N. The “*” symbol indicates that there can be any number of this CDE in the range 0..N. There must be at least one of the header and block CDEs in a tma CDE, while the analysis CDE is optional (as there may exist TMA data upon which no statistical analysis has been performed). The analysis CDE is defined within our extended TMA DES DTD as follows:
<!ELEMENT analysis (test+, analysisID, date)>
The test CDE contains further CDEs, representing the statistical method used to perform that test, as shown below:
<!ELEMENT test (kappa | t_test | wilcoxon | one_way_anova | kruskal_wallis | pearson | spearman | fisher | chiSquare | log_rank | logisticRegression | coxProportionalHazard)>
Each of these CDEs contains further CDEs containing the results of the analysis applied to data in that tma object. Figure 5 illustrates a complete example of the “pearson” CDE, containing the results returned when Pearson's Correlation is applied to a set of data.
Figure 5.
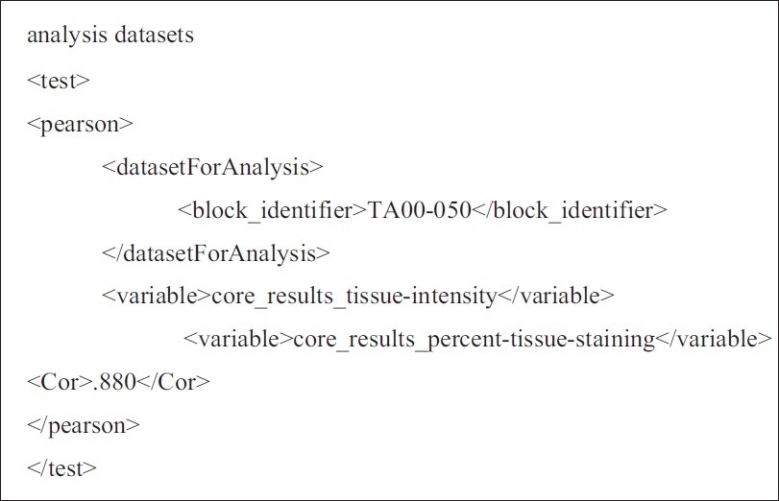

Example of XML describing inputs to a Pearson's Correlation test and the test results
An example of our ISO 11179 semantic definition of the pearson CDE in this document is shown in Figure 6.
Figure 6.

ISO 11179 compliant semantic definition of Pearson CDE
We wrote a program using the R Statistical Software package (http://www.r-project.org/) which demonstrated how existing applications could read in TMA DES data, perform statistical tests on the data, and annotate that data with the results of those tests. R has XML data structures that can store the contents of XML files. The script extracted the following data from a TMA DES XML file containing details of samples from colorectal carcinomas, low-grade dysplasia, high-grade dysplasia, and normal tissue and corresponding marker staining:
Stain: Two markers, p16 and p53.
Diagnosis: Either “Normal,” “Low grade dysplasia,” “High grade dysplasia,” or “Carcinoma.”
Score: scores ranged from 0 to 3, depending on the extent to which the core sample was stained.
We used the Mann Whitney U Test to determine if the difference between the medians of p53 scores for the “Normal” and “Carcinoma” cores was significant. Another statistical test we performed was a Spearman's Correlation between p53 and p16 scores in "Carcinoma" cores. The R script constructed XML CDEs that represented the results of these statistical tests, and inserted these CDEs into the original XML which was then exported to a file (Supplementary File 4, R_Output.xml).
RESULTS
We used a web-based XML parser, www.xmlvalidation.com, which provides an interface to the Simple API for XML (SAX) parsing software to verify that TMA DES XML data containing our proposed CDEs were both well-formed (i.e., conformed to the XML syntax), and was valid (i.e., conformed to the structure outlined in our extended TMA DES DTD). The output from our R script which applied statistical tests to TMA DES data (R_Output.xml) was successfully validated using this parser. We extended the TA00-050.XML file that was included as an additional file with the paper describing the TMA DES DTD[15] to include CDEs representing the results of the statistical tests listed in Table 1. When it was submitted to the SAX parser, no errors or warnings were reported. Another XML parser, Richard Tobin's XML well-formedness checker and validator (RXP),[16] also parsed our extended TMA DES XML data and validated it against our extended TMA DES DTD without reporting any errors. To verify that our DTD correctly detected syntax errors in TMA DES data, deliberate errors were submitted to the SAX parser; see Table 2 which describes these errors and the output they produced from the parser.
Table 2.
Messages produced by the SAX parser when errors were introduced into TMA DES XML

We checked that the DTD correctly defined the structure for statistical data by swapping CDEs between test results, and submitting the altered XML to the SAX parser. For example, the contents of a log_rank CDE were swapped with the contents of a chiSquare CDE. The parser returned the following errors:
The content of element type “chiSquare” must match “(datasetForAnalysis,datasetForAnalysis+,condition,pearsonChiSquare,likelihoodRatio,numberOfValidCases).”
The content of element type “log_rank” must match “(datasetForAnalysis,datasetForAnalysis+,condition,chiSquareValue,DF,Sig).”
Finally, to demonstrate that the CDEs in our extended TMA DES DTD are optional, we validated the original TA00-050.XML file, which contained none of the statistical CDEs defined in our DTD, and observed the results. No errors were reported when this XML file was validated against our extended TMA DES DTD using the SAX parser.
CONCLUSIONS
We have extended the TMA DES DTD to include definitions of CDEs representing the results of statistical analysis on TMA DES data. An ISO 11179 compliant file containing a list of semantic definitions for these CDEs was also written. We have used these CDEs to construct XML descriptions of the results of a number of statistical tests routinely applied to TMA datasets. These CDEs could also be used as “building blocks” to create descriptions of other statistical tests not mentioned in this paper. Thus, the extended TMA DES we present is itself further extensible.
We used XML parsers to validate both TMA DES files using the CDEs defined in our TMA DES extensions. We confirmed that our DTD successfully detected a range of syntactic and structural errors in TMA DES files. We also successfully validated existing TMA DES data which contained no statistical data against our extended TMA DES DTD, demonstrating that our DTD is backward compatible with existing TMA DES data. The CDEs we propose will permit developers to incorporate these into software to allow standardization of storage as well as sharing of statistical analysis results from TMA experiments.
Although the TMA DES DTD is not itself an XML document, the syntax and structure of TMA DES data can be defined in an XML document, such as the tissue microarray OWL schema.[17] XML Schema,[18] an XML-based method for describing the structure of XML documents, can also be used to specify the syntax and structure of an XML document, along with the data types of CDEs (e.g., Boolean, string, decimal, float), valid ranges of values for these elements and more precise values for numbers of occurrences of elements (e.g., 1-5, instead of 1-Infinity) then can be specified using DTDs. At present, this additional information is specified in an ISO 11179 file. Future work could involve redefining the TMA DES specification as an XML Schema document.
Existing markup languages for describing statistical data do not contain CDEs for describing the results of many of the statistical tests described in this paper. We suggest that many of the CDEs proposed in this paper could also serve as a template for describing the results of statistical analyses in other dedicated biomedical markup languages.
Acknowledgments
PL is supported by the Welsh Assembly Government. PQ is supported by Yorkshire Cancer Research and the Experimental Cancer Medicine Centre and NCRI informatics initiative for infrastructural support. PQ and DT are supported by the NCRI informatics initiative for infrastructure support.
Footnotes
Available FREE in open access from: http://www.jpathinformatics.org/text.asp? 2011/2/1/17/78263
REFERENCES
- 1.Kononen J, Bubendorf L, Kallioniemi A, Barlund M, Schraml P, Leighton S, et al. Tissue microarrays for high-throughput molecular profiling of tumor specimens. Nat Med. 1998;4:844–7. doi: 10.1038/nm0798-844. [DOI] [PubMed] [Google Scholar]
- 2.Voduc D, Kenney C, Nielsen T. Tissue Microarrays in Clinical Oncology. Semin Radiat Oncol. 2008;18:89–97. doi: 10.1016/j.semradonc.2007.10.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Berman JJ, Edgerton ME, Friedman BA. The tissue microarray data exchange specification: A community-based, open source tool for sharing tissue microarray data. BMC Med Inform Decis Mak. 2003;3:5. doi: 10.1186/1472-6947-3-5. Available from: http://www.biomedcentral.com/1472-6947/3/5 . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bray T, Paoli J, Sperberg-McQueen CM, Maler E, Yergeau F. Extensible Markup Language (XML) 1.0 (5th edition) W3C Recommendation. 2008. Nov 08, Available from: http://www.w3.org/TR/2008/REC-xml-20081126/ W3C Recommendation.
- 5.Xperanto-TMA. [Last accessed on 2010 Sep 01]. Available from: http://xperanto.snubi.org/TMA/
- 6.Thallinger GG, Baumgartner K, Pirklbauer M, Uray M, Pauritsch E, Mehes G, et al. TAMEE: data management and analysis for tissue microarrays. BMC Bioinformatics. 2007;8:81. doi: 10.1186/1471-2105-8-81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kristiansen G, Fritzsche FR, Wassermann K, Jäger C, Tölls A, Lein M, et al. GOLPH2 protein expression as a novel tissue biomarker for prostate cancer: implications for tissue-based diagnostics. Br J Cancer. 2008;99:939–48. doi: 10.1038/sj.bjc.6604614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Crabb SJ, Bajdik CD, Leung S, Speers CH, Kennecke H, Huntsman DG, et al. Can clinically relevant prognostic subsets of breast cancer patients with four or more involved axillary lymph nodes be identified through immune histochemical biomarkers?. A tissue microarray feasibility study. Breast Cancer Res. 2008;10:R6. doi: 10.1186/bcr1847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Sjöbeck M, Haglund M, Persson A, Sturesson K, Englund E. Brain tissue microarrays in dementia research: White matter microvascular pathology in Alzheimer's disease. Neuropathol. 2003;23:290–5. doi: 10.1046/j.1440-1789.2003.00515.x. [DOI] [PubMed] [Google Scholar]
- 10.Rubin MA, Dunn R, Strawderman M, Pienta KJ. Tissue microarray sampling strategy for prostate cancer biomarker analysis. Am J Surg Pathol. 2002;26:312–9. doi: 10.1097/00000478-200203000-00004. [DOI] [PubMed] [Google Scholar]
- 11.Predictive Model Markup Language. [Last accessed on 2010 Sep 01]. Available from: http://www.dmg.org/v4-0/GeneralStructure.html .
- 12.The StatDataML package. [Last accessed on 2010 Sep 01]. Available from: http://www.omegahat.org/StatDataML/
- 13.Edgerton M. Assoc Pathol Inform 1/27/03 Tissue MicroArray Common Data Elements. [Last accessed on 2010 Sep 01]. Available from: http://www.biomedcentral.com/content/supplementary/1472-6947-3-5-s1.htm .
- 14.Solbrig HR. Metadata and the reintegration of clinical information: ISO 11179. MD Comput. 2000;3:25–8. [PubMed] [Google Scholar]
- 15.Nohle D, Ayers L. The tissue microarray data exchange specification: A document type definition to validate and enhance XML data. BMC Med Inform Decis Mak. 2005;5:12. doi: 10.1186/1472-6947-5-12. Available from: http://www.biomedcentral.com/1472-6947/5/12 . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Richard Tobin's XML well-formedness checker and validator at. [Last accessed on 2010 Sep 01]. Available from: http://www.cogsci.ed.ac.uk/%7Erichard/xml-check.html .
- 17.Kang HP, Borromeo CD, Berman JJ, Becich MJ. The tissue microarray OWL schema: An open-source tool for sharing tissue microarray data. J Pathol Inform. 2010;1:9. doi: 10.4103/2153-3539.65347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Thompson H, Beech D, Maloney M, Mendelsohn N. XML Schema Part 1: Structures 2nd edition Recommendation. [Last accessed on 28 Nov 2004]. Available from: http://www.w3.org/TR/xmlschema-1/ W3C Recommendation.


