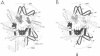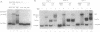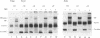Abstract
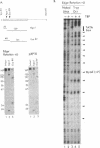
We establish that the TATA binding protein (TBP) in the presence of TFIIA recognizes the TATA box in nucleosomal DNA dependent on the dissociation of the amino-terminal tails of the core histones from the nucleosome and the position of the TATA box within the nucleosome. We examine TBP/TFIIA access to the TATA box with this sequence placed in four distinct rotational frames with reference to the histone surface and at three distinct translational positions at the edge, side and dyad axis of the nucleosome. Under our experimental conditions, we find that the preferential translational position at which TBP/TFIIA can bind the TATA box is within linker DNA at the edge of the nucleosome and that binding is facilitated if contacts made by the amino-terminal tails of the histones with nucleosomal DNA are eliminated. TBP/TFIIA binding to DNA at the edge of the nucleosome occurs with the TATA box in all four rotational positions. This is indicative of TBP/TFIIA association directing the dissociation of the TATA box from the surface of the histone octamer.
Full text
PDF



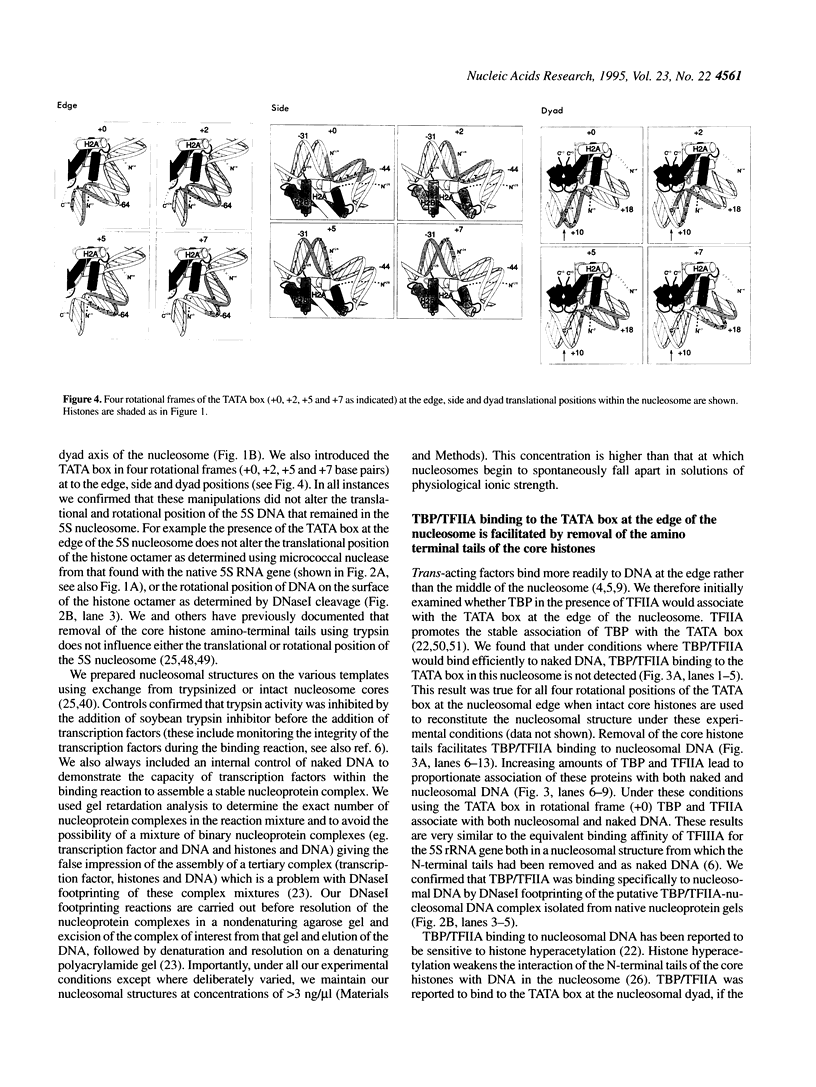



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Archer T. K., Cordingley M. G., Wolford R. G., Hager G. L. Transcription factor access is mediated by accurately positioned nucleosomes on the mouse mammary tumor virus promoter. Mol Cell Biol. 1991 Feb;11(2):688–698. doi: 10.1128/mcb.11.2.688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ausio J., Dong F., van Holde K. E. Use of selectively trypsinized nucleosome core particles to analyze the role of the histone "tails" in the stabilization of the nucleosome. J Mol Biol. 1989 Apr 5;206(3):451–463. doi: 10.1016/0022-2836(89)90493-2. [DOI] [PubMed] [Google Scholar]
- Ausio J., Seger D., Eisenberg H. Nucleosome core particle stability and conformational change. Effect of temperature, particle and NaCl concentrations, and crosslinking of histone H3 sulfhydryl groups. J Mol Biol. 1984 Jun 15;176(1):77–104. doi: 10.1016/0022-2836(84)90383-8. [DOI] [PubMed] [Google Scholar]
- Bashkin J., Hayes J. J., Tullius T. D., Wolffe A. P. Structure of DNA in a nucleosome core at high salt concentration and at high temperature. Biochemistry. 1993 Mar 2;32(8):1895–1898. doi: 10.1021/bi00059a002. [DOI] [PubMed] [Google Scholar]
- Buratowski S., Hahn S., Guarente L., Sharp P. A. Five intermediate complexes in transcription initiation by RNA polymerase II. Cell. 1989 Feb 24;56(4):549–561. doi: 10.1016/0092-8674(89)90578-3. [DOI] [PubMed] [Google Scholar]
- Cary P. D., Crane-Robinson C., Bradbury E. M., Dixon G. H. Effect of acetylation on the binding of N-terminal peptides of histone H4 to DNA. Eur J Biochem. 1982 Sep;127(1):137–143. doi: 10.1111/j.1432-1033.1982.tb06847.x. [DOI] [PubMed] [Google Scholar]
- Chen H., Li B., Workman J. L. A histone-binding protein, nucleoplasmin, stimulates transcription factor binding to nucleosomes and factor-induced nucleosome disassembly. EMBO J. 1994 Jan 15;13(2):380–390. doi: 10.1002/j.1460-2075.1994.tb06272.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cormack B. P., Struhl K. The TATA-binding protein is required for transcription by all three nuclear RNA polymerases in yeast cells. Cell. 1992 May 15;69(4):685–696. doi: 10.1016/0092-8674(92)90232-2. [DOI] [PubMed] [Google Scholar]
- Dong F., Hansen J. C., van Holde K. E. DNA and protein determinants of nucleosome positioning on sea urchin 5S rRNA gene sequences in vitro. Proc Natl Acad Sci U S A. 1990 Aug;87(15):5724–5728. doi: 10.1073/pnas.87.15.5724. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han M., Kim U. J., Kayne P., Grunstein M. Depletion of histone H4 and nucleosomes activates the PHO5 gene in Saccharomyces cerevisiae. EMBO J. 1988 Jul;7(7):2221–2228. doi: 10.1002/j.1460-2075.1988.tb03061.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayes J. J., Bashkin J., Tullius T. D., Wolffe A. P. The histone core exerts a dominant constraint on the structure of DNA in a nucleosome. Biochemistry. 1991 Aug 27;30(34):8434–8440. doi: 10.1021/bi00098a022. [DOI] [PubMed] [Google Scholar]
- Hayes J. J., Clark D. J., Wolffe A. P. Histone contributions to the structure of DNA in the nucleosome. Proc Natl Acad Sci U S A. 1991 Aug 1;88(15):6829–6833. doi: 10.1073/pnas.88.15.6829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayes J. J., Pruss D., Wolffe A. P. Contacts of the globular domain of histone H5 and core histones with DNA in a "chromatosome". Proc Natl Acad Sci U S A. 1994 Aug 2;91(16):7817–7821. doi: 10.1073/pnas.91.16.7817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayes J. J., Tullius T. D., Wolffe A. P. The structure of DNA in a nucleosome. Proc Natl Acad Sci U S A. 1990 Oct;87(19):7405–7409. doi: 10.1073/pnas.87.19.7405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayes J. J., Wolffe A. P. Histones H2A/H2B inhibit the interaction of transcription factor IIIA with the Xenopus borealis somatic 5S RNA gene in a nucleosome. Proc Natl Acad Sci U S A. 1992 Feb 15;89(4):1229–1233. doi: 10.1073/pnas.89.4.1229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayes J. J., Wolffe A. P. Preferential and asymmetric interaction of linker histones with 5S DNA in the nucleosome. Proc Natl Acad Sci U S A. 1993 Jul 15;90(14):6415–6419. doi: 10.1073/pnas.90.14.6415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffmann A., Roeder R. G. Purification of his-tagged proteins in non-denaturing conditions suggests a convenient method for protein interaction studies. Nucleic Acids Res. 1991 Nov 25;19(22):6337–6338. doi: 10.1093/nar/19.22.6337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horikoshi M., Bertuccioli C., Takada R., Wang J., Yamamoto T., Roeder R. G. Transcription factor TFIID induces DNA bending upon binding to the TATA element. Proc Natl Acad Sci U S A. 1992 Feb 1;89(3):1060–1064. doi: 10.1073/pnas.89.3.1060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horikoshi M., Wang C. K., Fujii H., Cromlish J. A., Weil P. A., Roeder R. G. Cloning and structure of a yeast gene encoding a general transcription initiation factor TFIID that binds to the TATA box. Nature. 1989 Sep 28;341(6240):299–303. doi: 10.1038/341299a0. [DOI] [PubMed] [Google Scholar]
- Imbalzano A. N., Kwon H., Green M. R., Kingston R. E. Facilitated binding of TATA-binding protein to nucleosomal DNA. Nature. 1994 Aug 11;370(6489):481–485. doi: 10.1038/370481a0. [DOI] [PubMed] [Google Scholar]
- Kahn J. D., Crothers D. M. DNA bending in transcription initiation. Cold Spring Harb Symp Quant Biol. 1993;58:115–122. doi: 10.1101/sqb.1993.058.01.015. [DOI] [PubMed] [Google Scholar]
- Kim J. L., Nikolov D. B., Burley S. K. Co-crystal structure of TBP recognizing the minor groove of a TATA element. Nature. 1993 Oct 7;365(6446):520–527. doi: 10.1038/365520a0. [DOI] [PubMed] [Google Scholar]
- Kim T. K., Roeder R. G. Involvement of the basic repeat domain of TATA-binding protein (TBP) in transcription by RNA polymerases I, II, and III. J Biol Chem. 1994 Feb 18;269(7):4891–4894. [PubMed] [Google Scholar]
- Kim Y., Geiger J. H., Hahn S., Sigler P. B. Crystal structure of a yeast TBP/TATA-box complex. Nature. 1993 Oct 7;365(6446):512–520. doi: 10.1038/365512a0. [DOI] [PubMed] [Google Scholar]
- Kladde M. P., Simpson R. T. Positioned nucleosomes inhibit Dam methylation in vivo. Proc Natl Acad Sci U S A. 1994 Feb 15;91(4):1361–1365. doi: 10.1073/pnas.91.4.1361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knezetic J. A., Luse D. S. The presence of nucleosomes on a DNA template prevents initiation by RNA polymerase II in vitro. Cell. 1986 Apr 11;45(1):95–104. doi: 10.1016/0092-8674(86)90541-6. [DOI] [PubMed] [Google Scholar]
- Lee D. Y., Hayes J. J., Pruss D., Wolffe A. P. A positive role for histone acetylation in transcription factor access to nucleosomal DNA. Cell. 1993 Jan 15;72(1):73–84. doi: 10.1016/0092-8674(93)90051-q. [DOI] [PubMed] [Google Scholar]
- Li Q., Wrange O. Translational positioning of a nucleosomal glucocorticoid response element modulates glucocorticoid receptor affinity. Genes Dev. 1993 Dec;7(12A):2471–2482. doi: 10.1101/gad.7.12a.2471. [DOI] [PubMed] [Google Scholar]
- Lorch Y., LaPointe J. W., Kornberg R. D. Nucleosomes inhibit the initiation of transcription but allow chain elongation with the displacement of histones. Cell. 1987 Apr 24;49(2):203–210. doi: 10.1016/0092-8674(87)90561-7. [DOI] [PubMed] [Google Scholar]
- Maldonado E., Ha I., Cortes P., Weis L., Reinberg D. Factors involved in specific transcription by mammalian RNA polymerase II: role of transcription factors IIA, IID, and IIB during formation of a transcription-competent complex. Mol Cell Biol. 1990 Dec;10(12):6335–6347. doi: 10.1128/mcb.10.12.6335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsui T. Transcription of adenovirus 2 major late and peptide IX genes under conditions of in vitro nucleosome assembly. Mol Cell Biol. 1987 Apr;7(4):1401–1408. doi: 10.1128/mcb.7.4.1401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meisterernst M., Horikoshi M., Roeder R. G. Recombinant yeast TFIID, a general transcription factor, mediates activation by the gene-specific factor USF in a chromatin assembly assay. Proc Natl Acad Sci U S A. 1990 Dec;87(23):9153–9157. doi: 10.1073/pnas.87.23.9153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nightingale K., Wolffe A. P. Methylation at CpG sequences does not influence histone H1 binding to a nucleosome including a Xenopus borealis 5 S rRNA gene. J Biol Chem. 1995 Mar 3;270(9):4197–4200. doi: 10.1074/jbc.270.9.4197. [DOI] [PubMed] [Google Scholar]
- Nikolov D. B., Hu S. H., Lin J., Gasch A., Hoffmann A., Horikoshi M., Chua N. H., Roeder R. G., Burley S. K. Crystal structure of TFIID TATA-box binding protein. Nature. 1992 Nov 5;360(6399):40–46. doi: 10.1038/360040a0. [DOI] [PubMed] [Google Scholar]
- Patterton H. G., Simpson R. T. Nucleosomal location of the STE6 TATA box and Mat alpha 2p-mediated repression. Mol Cell Biol. 1994 Jun;14(6):4002–4010. doi: 10.1128/mcb.14.6.4002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pennings S., Meersseman G., Bradbury E. M. Mobility of positioned nucleosomes on 5 S rDNA. J Mol Biol. 1991 Jul 5;220(1):101–110. doi: 10.1016/0022-2836(91)90384-i. [DOI] [PubMed] [Google Scholar]
- Perlmann T., Wrange O. Specific glucocorticoid receptor binding to DNA reconstituted in a nucleosome. EMBO J. 1988 Oct;7(10):3073–3079. doi: 10.1002/j.1460-2075.1988.tb03172.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piña B., Brüggemeier U., Beato M. Nucleosome positioning modulates accessibility of regulatory proteins to the mouse mammary tumor virus promoter. Cell. 1990 Mar 9;60(5):719–731. doi: 10.1016/0092-8674(90)90087-u. [DOI] [PubMed] [Google Scholar]
- Pruss D., Hayes J. J., Wolffe A. P. Nucleosomal anatomy--where are the histones? Bioessays. 1995 Feb;17(2):161–170. doi: 10.1002/bies.950170211. [DOI] [PubMed] [Google Scholar]
- Pruss D., Wolffe A. P. Histone-DNA contacts in a nucleosome core containing a Xenopus 5S rRNA gene. Biochemistry. 1993 Jul 13;32(27):6810–6814. doi: 10.1021/bi00078a002. [DOI] [PubMed] [Google Scholar]
- Ranish J. A., Lane W. S., Hahn S. Isolation of two genes that encode subunits of the yeast transcription factor IIA. Science. 1992 Feb 28;255(5048):1127–1129. doi: 10.1126/science.1546313. [DOI] [PubMed] [Google Scholar]
- Rhodes D. Structural analysis of a triple complex between the histone octamer, a Xenopus gene for 5S RNA and transcription factor IIIA. EMBO J. 1985 Dec 16;4(13A):3473–3482. doi: 10.1002/j.1460-2075.1985.tb04106.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richmond T. J., Rechsteiner T., Luger K. Studies of nucleosome structure. Cold Spring Harb Symp Quant Biol. 1993;58:265–272. doi: 10.1101/sqb.1993.058.01.031. [DOI] [PubMed] [Google Scholar]
- Ura K., Hayes J. J., Wolffe A. P. A positive role for nucleosome mobility in the transcriptional activity of chromatin templates: restriction by linker histones. EMBO J. 1995 Aug 1;14(15):3752–3765. doi: 10.1002/j.1460-2075.1995.tb00045.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vettese-Dadey M., Walter P., Chen H., Juan L. J., Workman J. L. Role of the histone amino termini in facilitated binding of a transcription factor, GAL4-AH, to nucleosome cores. Mol Cell Biol. 1994 Feb;14(2):970–981. doi: 10.1128/mcb.14.2.970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolffe A. P., Drew H. R. Initiation of transcription on nucleosomal templates. Proc Natl Acad Sci U S A. 1989 Dec;86(24):9817–9821. doi: 10.1073/pnas.86.24.9817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolffe A. P., Jordan E., Brown D. D. A bacteriophage RNA polymerase transcribes through a Xenopus 5S RNA gene transcription complex without disrupting it. Cell. 1986 Feb 14;44(3):381–389. doi: 10.1016/0092-8674(86)90459-9. [DOI] [PubMed] [Google Scholar]
- Workman J. L., Roeder R. G. Binding of transcription factor TFIID to the major late promoter during in vitro nucleosome assembly potentiates subsequent initiation by RNA polymerase II. Cell. 1987 Nov 20;51(4):613–622. doi: 10.1016/0092-8674(87)90130-9. [DOI] [PubMed] [Google Scholar]
- Yamashita S., Hisatake K., Kokubo T., Doi K., Roeder R. G., Horikoshi M., Nakatani Y. Transcription factor TFIIB sites important for interaction with promoter-bound TFIID. Science. 1993 Jul 23;261(5120):463–466. doi: 10.1126/science.8332911. [DOI] [PubMed] [Google Scholar]
- Yamashita S., Wada K., Horikoshi M., Gong D. W., Kokubo T., Hisatake K., Yokotani N., Malik S., Roeder R. G., Nakatani Y. Isolation and characterization of a cDNA encoding Drosophila transcription factor TFIIB. Proc Natl Acad Sci U S A. 1992 Apr 1;89(7):2839–2843. doi: 10.1073/pnas.89.7.2839. [DOI] [PMC free article] [PubMed] [Google Scholar]