Abstract
The cap at the growing end of the bacterial flagellum is essential for its growth, remaining stably attached while permitting the insertion of flagellin transported from the cytoplasm through the narrow central channel. We analyzed the structure of the isolated cap in its frozen hydrated state by electron cryomicroscopy. The 3D density map now shows detailed features of domains and their connections, giving reliable volumes and masses, making assignment of the domains to the amino acid sequence possible. A model of the cap–filament complex built with an atomic model of the filament allows a quantitative analysis of the cap domain movements on cap binding and rotation that promotes the efficient self assembly of flagellin during the filament growth process.
Bacteria swim by rotating flagellar filaments (1–3), each of which works as a propeller by its helical shape. In Escherichia coli and Salmonella, the motor structure called the flagellar basal body crosses both cytoplasmic and outer membrane and continues to the extracellular structure called the hook and the filament. Assembly of the flagellum begins from basal body construction, and a protein export apparatus homologous to the type III protein secretion system (4) is attached on its cytoplasmic face (5). The export apparatus selectively transfers flagellar axial proteins into the long narrow central channel of the flagellum by the energy of ATP hydrolysis by FliI, a subunit of the export apparatus (6, 7). These flagellar axial proteins travel through the channel to the distal end, where assembly occurs (8, 9). A protein, called hook-associated protein 2 (HAP2, also called FliD), forms a cap structure at the distal end of the flagellum (10), which is essential for filament growth. The cap remains stably attached while permitting flagellin insertion and binding between the filament end and itself, thereby preventing flagellin monomers from simply diffusing away. Isolated HAP2 forms a bipolar pair of pentamers in solution, but the native capping function is performed by a pentamer (11–14).
The flagellar filament is a tubular structure constructed with a single protein, flagellin. The filament is made of 11 protofilaments, whose lateral binding is axially staggered by about half a subunit, producing five indentations at the ends of the filament with a double indentation in one of these five, wider and deeper than the other four. HAP2 cap binding is therefore achieved over a symmetry mismatch, between the 5-fold circular symmetry of the cap and ≈5.5-fold helical symmetry of the filament. Vonderviszt et al. (13) proposed a model of the capping interaction, and Yonekura et al. (14) visualized the actual capping interactions by electron microscopy and 3D image reconstruction, although at a limited resolution. Based on the cap structure, with a pentagonal plate and five leg domains and the cap–filament structure showing just one binding site open for flagellin, we proposed an active role of the cap for the in vivo filament growth, namely a rotary cap mechanism that promotes efficient flagellin assembly (14). However, the boundary between the cap and the filament end could not be well resolved.
To visualize the cap–filament interaction more clearly, we have analyzed the structure of the cap at higher resolution by electron cryomicroscopy and single-particle image analysis of the frozen hydrated cap decamer. This map has more reliable volume recovery, allowing docking of this map onto an atomic model of the filament (K. Imada, K. Hasegawa, S.M.-Y., K.Y., F. A. Samatey, I. Yamashita, and K.N., unpublished results). The interactions between distal flagellin subunits and the five leg domains of the cap can now be analyzed more quantitatively, including their conformational changes. Implications for the flexibility in the domain arrangements of flagellar axial proteins will be discussed below.
Materials and Methods
Sample Preparation. HAP2 was overproduced in E. coli cells, strain BL21 (DE3), carrying pLysS and pKOT134 plasmids (15), and was prepared as described (12). Chemical crosslinking was used to stabilize the bipolar pair of caps. The crosslinking was carried out by using a zero-length crosslinker 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride (EDC), as described (11, 12), with some improvements. Briefly, 1.0 mg/ml HAP2 was incubated with 40 mM EDC and 40 mM sulfo N-hydroxysuccinimide in 20 mM sodium phosphate buffer (pH 7.0) at room temperature. After 2 h, the reaction was terminated by adding 500 mM Tris·HCl (pH 8.0) and 20 mM β-mercaptoethanol to a final concentration of 167 and 6.7 mM, respectively. The crosslinked products were dialyzed against 20 mM sodium phosphate buffer (pH 7.0) and 200 mM NaCl.
Electron Cryomicroscopy. The crosslinked products of HAP2 were suspended in the dialysis buffer containing 0.9% N-octyl-β-d-glucoside at a protein concentration of 0.5 mg/ml just before the preparation of frozen hydrated specimens. A holey carbon film on a copper grid (QUANTIFOIL R1.2/1.3 Quantifoil Micro Tools, Jena, Germany) was pretreated by glow discharge with N-amylamine, and the sample solution was applied to the grid. The grid was blotted and plunged into liquid ethane. Then, the grids were examined by using an electron microscope, JEM-3000SFF (JEOL), with a field emission gun operated at 100 kV and at a specimen temperature of 4 K. Images were recorded on SO-163 film (Eastman Kodak) at a nominal magnification of ×60,000. The electron dose was set to 13 ≈ 18 e/Å2.
Image Analysis. Electron micrographs were scanned and digitized with a LeafScan 45 linear charge-coupled device densitometer (Scitex, Tel Aviv) at a step size of 10 μm, and the image data were reduced by a factor of 2. The magnification was calibrated with the layer line spacing measured in the computed Fourier transform of the images of the straight flagellar filaments from the SJW1660 strain of Salmonella recorded on the same electron micrographs.
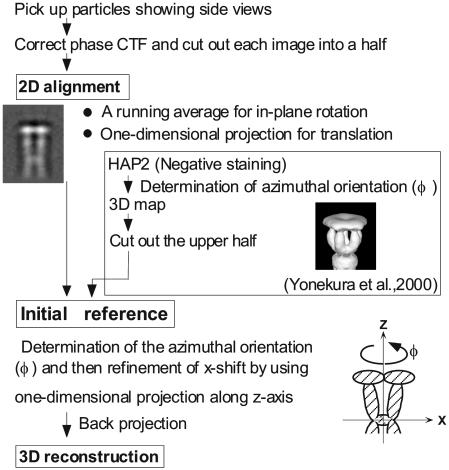
Overall procedure of image analysis and 3D image reconstruction are described in Fig. 1. First, the phase reversal caused by the contrast transfer function was corrected. Images recorded at defocus levels of 15,000–20,000 Å were used for analysis. Because HAP2 decamer images showed a tendency to bend at the connection between the two pentamers (14), we treated each pentamer separately. The end-on view images (Fig. 2A) were not used, because it was not possible to exclude images of the decamer. Only side-view images of the decamer (Fig. 2B) were selected in electron cryomicrographs to make sure that only pentamer images as halves of the HAP2 decamer were used for image alignment. In total, 422 images of the decamers were picked up by visual inspection. To treat each pentamer separately, three points were manually marked on each decamer image to indicate the center and both ends. Then, 844 pentamer images were aligned roughly based on these marked points.
Fig. 1.
Summary of the image analysis and 3D image reconstruction procedure.
Fig. 2.
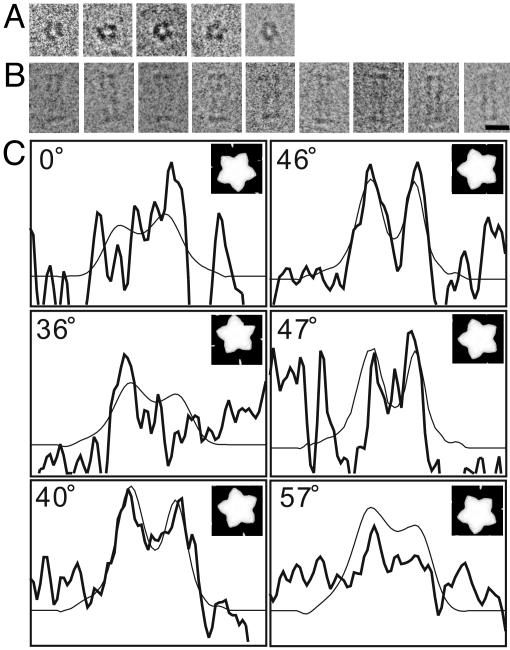
Electron cryomicrographs of the frozen hydrated HAP2 decamer complex and determination of its azimuthal orientation. (A) End-on views with a pentameric feature. (B) Side views with an elongated rectangular shape and thin plates at both ends. (Bar = 100 Å.) (C) Projection x profile for the determination of azimuthal orientation and 5-fold axis position of the cap. Six examples are shown. Thick lines represent those obtained from individual images, and thin lines represent those from the reference image. The determined azimuthal angles are 0°, 36°, 40°, 46°, 47°, and 57°. The two peaks in the density profile correspond to the leg domains on two sides of the particle and the trough to the central cavity. The interval of tick marks is 34.5 Å in the horizontal axis and an arbitrary unit in the vertical axis. These particular angles were chosen to demonstrate how this profile changes sensitively by a small change of angle.
When we first tried conventional methods such as eman (16) and spider (17), which carry out 2D alignment by crosscorrelation, they showed a tendency to cause misalignment of images. We thought that this tendency was due to the high noise level and therefore used our own 1D projection method described below. However, we later found out that the misalignment was due to off-center placement of pentamer images in individual images of the decamer being placed at the center. When we cut off pentamer images and placed them at the center of the image box, conventional alignment methods produced density maps similar to that presented here (reconstruction by eman in Fig. 6, which is published as supporting information on the PNAS web site). Although there are significant differences between the two reconstructions, particularly in the thin kinked rod subdomain regions, the Fourier shell correlation (FSC) curve between the two falls off to 0.5 at 26-Å resolution, which is comparable to those of individual reconstructions. Therefore, the differences indicate weak signal levels in those regions, probably because of some structural flexibility. We present in this report the density map obtained by the 1D method, which shows slightly better-resolved features.
2D image alignment was carried out in a stepwise manner (18). First, a running average along an ≈5-fold axis was calculated for in-plane rotational alignment. Then, two 1D density profiles obtained by projection in two orthogonal directions were used for translational alignment. We call the profile obtained by projection along the 5-fold axis of the cap structure “projection x” (x axis profile) and the profile by projection perpendicular to the 5-fold axis, “projection z” (z axis profile). From the 1D crosscorrelation between the images, the amount of x shift (shift of the axis position) was roughly determined and further refined as below. The amount of z shift (shift along the axis) was obtained from the position of maximum slope in projection z, which corresponds to the outer edge of the cap plate (14, 18). After the alignment of the images, an averaged image was obtained. By using the average as the new reference image, this 2D alignment procedure was iterated 10 times. In each step, images that showed relatively large translation or angle of rotation were excluded from the average to produce the reference for the next step. After the 10th cycle, the number of images included in the average was 664.
The azimuthal orientation of the particles in individual images was determined by projection x. The 3D density distribution of the negatively stained HAP2 pentamer (14) was first projected to produce 2D images at every 1° step of the azimuthal rotation about the 5-fold axis, and then projection x was obtained from each image and used as the initial reference. From 1D crosscorrelation between projection x of the reference and that of each image, the azimuthal orientation was determined. The position of the 5-fold axis (x shift) was also refined in finer steps at the same time. Projection x of the reference (thin solid lines in Fig. 2C) changes its profile significantly depending on the azimuthal orientation of the cap, and that of the observed images (thick solid lines in Fig. 2C), although still noisy, showed structural features significant enough to give good crosscorrelation peaks, allowing us to determine the azimuthal orientation reliably and accurately. Then, a 3D density distribution was calculated by the back-projection method with a low-pass filter. We used the same low-pass filter that we used for the reconstruction of the cap–filament complex (18) to recover correct volume. We used this reconstruction as the new reference after enforcement of the 5-fold symmetry. This procedure was iterated four times. In each step, images with a crosscorrelation coefficient <0.1 were excluded. The number of HAP2 molecules included in the final image reconstruction was 531 × 5 = 2,655. After the enforcement of the 5-fold symmetry, the resolution was 26 Å by using the 0.5 criterion in the FSC between two reconstructions calculated from randomly chosen halves of the data set. We also calculated the FSC for a density map reconstructed without the symmetry enforcement (Fig. 7, which is published as supporting information on the PNAS web site); four FSC curves between this density map and those rotated about the 5-fold axis by 72°, 144°, 216°, and 288° all crossed an FSC of 0.5 at a resolution of 29 Å (Fig. 3). (See variance map in Fig. 8, which is published as supporting information on the PNAS web site.)
Fig. 3.
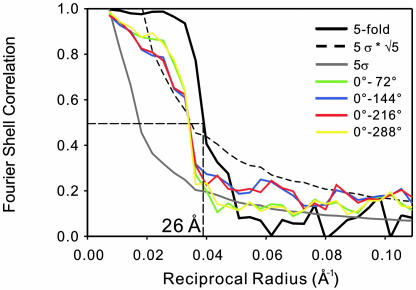
FSC profile indicating the resolution of ≈26 Å. Solid line in black is the FSC profile between two reconstructions calculated from two halves of the entire data set with 5-fold symmetry enforcement. Colored lines represent four FSC profiles between pairs of different azimuthal orientation at every 72° rotation for the reconstruction calculated without symmetry enforcement. These are also from one-half of the entire data set to be comparable with that of 5-fold averaging. The FSC curves fall off to 0.5 at 29 Å. Dotted and gray solid lines indicate the statistically expected error level in the FSC at 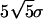 and 5 σ, with and without 5-fold symmetry taken into account, respectively.
and 5 σ, with and without 5-fold symmetry taken into account, respectively.
Model Building. An atomic model of the R type straight filament was constructed (K. Imada, K. Hasegawa, S.M.–Y., K. Yonekura, F. A. Samatey, I. Yamashita, and K.N., unpublished results) with the atomic model of the flagellar protofilament made of F41, a core fragment of flagellin (19), by using the helical symmetry of the filament (20) and the two heavy atom-binding positions as the fiducial marks. Details will be described elsewhere. The density map of the cap obtained in the present study was incorporated onto the atomic model of the filament by using the graphics program o (21). The five leg domains of the cap were cut off the plate domain by using xdispmsk (22) and rearranged without disconnection from the plate to fit into the five indentations formed at the distal end of the filament.
Results
Images of Frozen Hydrated HAP2 Decamer. The image contrast of the frozen hydrated HAP2 decamer obtained at an accelerating voltage of 300 kV was very poor because of its relatively small size, with a molecular mass of ≈250 kDa and a diameter slightly larger than 100 Å. We therefore operated the same microscope at an accelerating voltage of 100 kV to gain a higher contrast. Because the amplitude contrast is significantly higher at lower operation voltage (23), the image contrast was boosted up, and the molecular shape became much clearer. A representative set of images in end-on and side view is shown in Fig. 2 A and B, respectively.
Pentagonal particle images, called star-cap (15), corresponding to the end-on view of the HAP2 cap (Fig. 2 A) were observed, but the side views were rarely seen in the sample prepared in a solution containing 200 mM NaCl and 20 mM sodium phosphate buffer (pH 7.0). This observation indicated that the outer surface of the cap plate is relatively hydrophobic and prefers to be exposed to the air at the air–water interface. By adding a detergent, n-octyl-β-d-glucoside, to the sample solution to 0.9%, its critical micelle concentration, many side-view images appeared as shown in Fig. 2B. The bipolar structure is clearly visible in its elongated shape accentuated by the thin-plate feature at both ends. The decamer particles had a tendency to bend at the connection between the two caps, in a manner similar to those observed in negatively stained samples (11), indicating that the bending flexibility is an intrinsic property and not an artifact by negative staining.
3D Density Map of the Cap. Image analysis and 3D image reconstruction were carried out as described in Materials and Methods. The 3D density map of the HAP2 cap is shown in Fig. 4A. The side view of the whole decamer is also shown in Fig. 4B, where the density of the bottom half does not show up in many parts due to misalignment resulting from the flexible linker between the two pentamers, where the nature of disorder appears to be twisting around the 5-fold axis rather than bending. The basic architecture, consisting of a pentagonal plate that looks like a flower with five petals and five leg-like domains underneath the cap plate, is the same as the one in our previous map obtained from images of negatively stained particles (14). In the new map, however, each domain shows much more detailed features because of the significantly higher resolution. The leg domain is divided into three parts: a large subdomain attached to the plate through a narrow connection; a thin kinked rod subdomain that extends toward the linker portion adjoining the two caps; and a linker plate subdomain that adjoins the two sets of five leg domains of the two caps. The narrow connection between the large leg subdomain and the cap plate is much narrower than that in the previous map, strongly indicating that this connection is flexible enough to allow the adjustment of leg domain orientation to fit into the indentations of the distal end of the filament over the symmetry mismatch. The thin kinked rod subdomains also appear to have flexibility, which is probably responsible for the bending flexibility around the connection between the two pentamers. The volume of these kinked rod subdomains is likely to be underrepresented artificially by the contour level chosen to represent appropriate volumes of the cap plate and large leg subdomains, which occupy most of the volume of the cap dimer. The flexible nature of this portion would be responsible for variable conformations in individual decamer images, smearing out the density distribution of the bottom half of the decamer after averaging in the image reconstruction.
Fig. 4.
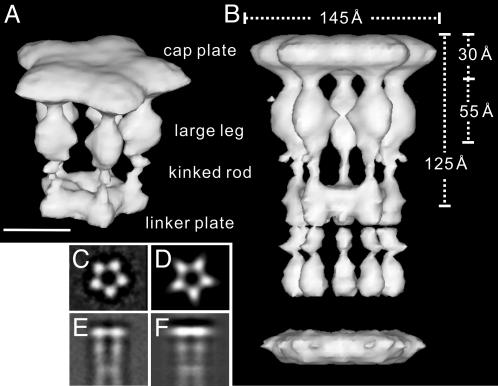
Reconstructed 3D image of the HAP2 pentamer and decamer. Oblique view of the pentamer (A) and side view of the decamer (B), in a solid surface representation with 100% volume recovery are shown. The 5-fold symmetry is enforced. (Bar = 50 Å.) (C) Averaged end-on view. (D) Projection of the 3D reconstruction along the 5-fold axis. (E) Averaged side-view. (F) Pojection of the 3D reconstruction perpendicular to the 5-fold axis.
The cap plate is 30 Å thick, and its outer diameter is 145 Å, just as observed for the cap plate in the cap–filament complex (14). The axial length of the cap is 125 Å, as measured from the outer surface of the cap plate to the center of the cap dimmer (Fig. 4B). The large leg subdomain is ≈55 Å long and 35 and 30 Å wide in its radial and azimuthal dimensions, respectively. The diameter of the empty inner space between the five legs underneath the cap plate is ≈40 Å at its narrowest position, which is also comparable to the diameter of the cavity underneath the cap plate in the cap–filament complex (14). This agreement indicates that, when the leg domains fit into the indentations at the end of the filament, radial movements are not necessary, and azimuthal movements are sufficient to allow the binding over the symmetry mismatch.
To check the validity of the method developed and used for this study, the reconstructed 3D volume was projected along and perpendicular to the 5-fold axis (Fig. 4 D and F). Those projections were compared with the 2D averages of the end-on and side-view images (Fig. 4 C and E), respectively. The close similarity between them proves the validity of the method.
We estimated the molecular mass of each domain from the volume (Table 1). The cap plate domain and the large leg, kinked rod, and linker plate subdomains occupy ≈63%, 21%, 0.3%, and 15%, respectively, and the corresponding molecular masses are ≈32, 11, 0.2, and 7.6 kDa, respectively. The value for the kinked rod domain is underrepresented, as mentioned above. Possible assignment of each domain on the amino acid sequence will be discussed later.
Table 1. Estimated volume and corresponding molecular mass of each domain.
| Domain | Volume, Å3 | Fraction, % | Molecular mass, kDa |
|---|---|---|---|
| Cap plate | 38,970 | 63 | 31.5 |
| Large leg | 12,940 | 21 | 10.5 |
| Thin rod | 210 | 0.3 | 0.2 |
| Linker plate | 9,350 | 15 | 7.6 |
Volumes and fractions were obtained from the reconstructed 3D map. The molecular mass was calculated from the volume fraction and the molecular mass of HAP2, 49.7 kDa. Note that the value for “Thin rod” is underrepresented.
Model of the Cap–Filament Complex. We docked the 3D density map of the cap onto an atomic model of the R type straight filament. The atomic model of the filament was constructed by using the atomic model of the flagellar protofilament made of F41, a core fragment of flagellin (19), as briefly described in Materials and Methods (K. Imada, K. Hasegawa, S.M.-Y., K. Yonekura, F. A. Samatey, I. Yamashita, and K.N., unpublished results). The atomic model was nicely fitted into the density map at 7-Å resolution obtained by electron cryomicroscopy (S.M.-Y., K. Yonekura, and K.N., unpublished results), further confirming the correctness of the model. Although the filament model does not contain domain D0 of flagellin, which forms the inner core of the filament, the large leg subdomains appear to interact with domains D1, which form the outer core of the filament, and therefore important interactions can be inferred in this model to identify the structural changes of the cap necessary for binding and the rotary mechanism for filament growth.
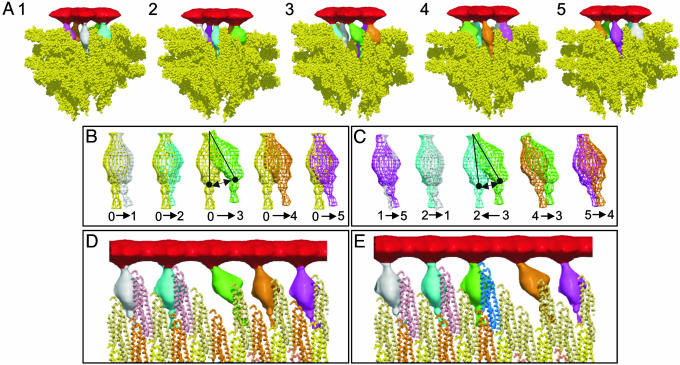
First, the cap plate was placed on the filament end according to the density map of the cap–filament complex (14), which clearly shows the relative position and azimuthal orientation of the cap plate to the helical array of the outer domains of flagellin in the filament. The filament model shows a complicated surface at its distal end but clearly demonstrates the shape, size, and position of the five indentations. The five leg domains of the cap did not fit into these indentations without rearrangement of their orientation and position. We assumed flexibility in the connection between the cap plate and leg domain for the leg domain movement. We also assumed flexibility in the connection between the large leg and the thin rod subdomains. The domains that form the linker plate were removed, because they are supposed to assume a different conformation on interaction with the inner core of the filament. We tried to fit the leg domains into the indentations so that all of the leg domains have interactions as uniform as possible with domains D1 of flagellin (Fig. 5). To produce the inverted-L-shaped opening at one site and relatively small gaps in the other four sites, as observed (14), one of the leg domains (green in Fig. 5 A–D) had to be moved and reoriented to a much larger extent than the other four, whereas the remaining four fitted nicely into the indentations with relatively small conformational changes (Fig. 5B).
Fig. 5.
A model of the cap–filament complex built with the density map of the cap and an atomic model of the flagellar filament. (A) A model of the cap–filament complex built in five different side views, showing each of the five leg conformations and the gaps between the plate and the filament end. A front portion of the model is trimmed off to show the leg domains and gaps clearly. The cap plate is red, and the five leg domains are white, blue, green, brown, and purple. For the filament portion, the flagellin F41 subunits are represented as a space-filling model in yellow. (B) Pairwise comparison of the leg domain orientation in the HAP2 decamer (yellow) with those in the cap–filament model (other colors) to show the changes in their orientation on cap binding. (C) Pairwise comparison of leg domain orientation between neighboring ones in the cap–filament model to show changes in their orientation by one-step cap rotation. Number labels in B and C correspond to those in different views in A, whereas label 0 indicates the leg domain in the HAP2 decamer. Arrows in B and C indicate the direction of leg domain movement. (D and E) The movement of the cap on binding of a flagellin subunit (blue) before and after its binding, respectively. A thick cylindrical section covering a radial range containing domains D1 of flagellin and the leg domains of HAP2 is displayed in D and E. The changes in the leg domain orientation correspond to those in C. Domain colors are consistent in B–E. The figure was made with molscript (24), raster3d (25), volcube in situs (26), and our own in-house programs.
The leg domain colored green in Fig. 5 A–D interacts with domain D1 of only one flagellin subunit, whereas the other four leg domains interact with two or three flagellin subunits (Fig. 5D). The changes in the position and orientation of the leg domains on binding of the cap to the filament end are shown by superimposition of modified leg domains on that of the isolated cap (Fig. 5B). The azimuthal displacement and the change in orientation are listed in Table 2. The largest displacement is 26 Å (the change in orientation, 33°), but the other displacements are relatively small, all within a range of 8–16 Å (6–18°). Displacements in the radial direction were not necessary.
Table 2. Displacement of large leg subdomains on cap binding and rotation.
| Leg domain
|
|||||
|---|---|---|---|---|---|
| 1, Å | 2, Å | 3, Å | 4, Å | 5, Å | |
| For binding | 8 (8°) | 8 (6°) | 26 (33°) | 16 (18°) | 10 (10°) |
| For rotation | 4 (2°) | 4 (5°) | 21 (27°) | 13 (15°) | 7 (8°) |
The displacements for the cap binding were measured at the distal (cell-proximal) end of the large leg subdomain, as marked by solid circle in Fig. 5B. The numbering also corresponds. The displacements for the cap rotation were obtained by comparing neighboring pairs, where data for leg domain 1, for instance, are from the comparison between 1 and 5. The angles are the change in domain orientation.
The changes in the position and orientation of the leg domains for each step rotation of the cap on binding of a flagellin subunit are also shown by superimposing neighboring pairs of leg domains (Fig. 5C). The displacements and the change in orientation are summarized in Table 2. The largest displacement is ≈21 Å (27°), but the other displacements are again relatively small, all within a range of 4–13 Å (2–15°).
Fig. 5 D and E show cylindrical sections at domain D1. When a new flagellin molecule (blue in Fig. 5E) that is exported from the central channel binds to the filament, one of the leg domains (green) is reoriented, making a new binding site in the next position (Fig. 5E).
Discussion
By electron cryomicroscopy of frozen hydrated HAP2 cap dimers and 3D image analysis, we recovered the shape and volume of each domain of the cap reliably. From a geometrical consideration of these data, together with the surface structure of the atomic model of the filament at its distal end (K. Imada, K. Hasegawa, S.M.-Y., K. Yonekura, F. A. Samatey, I. Yamashita, and K.N., unpublished results) and the structural features observed in the 3D density map of the cap–filament complex (14), we built a model of the cap–filament complex and identified possible ways interacting between domains of HAP2 and flagellin. Conformational changes of the cap on binding to the filament end and rotation to promote the self assembly of flagellin were preliminarily analyzed (27), but in the present study, we actually measured domain movements involved in these functions.
Possible Domain Movements. The leg domains of the cap were reoriented and fitted nicely into the indentations at the distal end of the filament, where these pivot motions required changes in the domain orientation up to ≈33° (Fig. 5B). The changes in the leg domain orientation required for a step rotation of the cap (Fig. 5C) were even smaller. The narrow linker density connecting the leg domains and the cap plate strongly suggests sufficient flexibility for all these changes in the domain orientation.
Domain-Sequence Assignment. There is evidence indicating that the leg domains are made of both terminal chains of HAP2. Approximately 40 NH2-terminal and 50 COOH-terminal residues of HAP2, which are unfolded in its monomeric or pentameric state in solution, are folded up to stabilize the cap–filament complex or the cap dimer structure (13). In the density map of the cap dimer, these terminal 90 residues would correspond to two distal leg domains: the linker plate subdomain adjoining the two caps and the thin kinked rod subdomain connecting to the large leg subdomain. From their volume, the molecular mass of these two domains together was estimated to be 8 kDa (Table 1), which roughly agrees with that of the terminal 90 residues, ≈10 kDa. Considering the possible underrepresentation of the volume of the thin kinked rod portion, this domain-sequence assignment seems reasonable. The molecular mass of the cap plate domain was estimated to be 31.5 kDa (Table 1), which is close to the apparent molecular mass (29 kDa) of a proteolytic fragment of HAP2 (13). This further supports the domain-sequence assignment for the terminal regions.
Folding of HAP2 and Implications for Export and Assembly. These results suggest that the peptide chain of HAP2 is folded in a similar manner to flagellin, having linearly connected domains with both termini in one end. The chain starts from one end, goes through all of the domains to the other end, and comes back. The peptide chain of flagellin starts from domain D0, which forms the inner core of the filament; goes out to D1, D2, and D3, which is the outermost domain; and comes back to D0 through D2 and D1 (19, 28, 29). The domains are connected by two chains forming a short antiparallel β-strand that appears to confer some flexibility in the domain arrangement on flagellin. The assembly process would be quicker if the domains are more or less folded during the export process through the central channel. It has recently been shown in the complete atomic model of the filament, however, that the diameter of the channel is smaller than the smallest dimension of the domains, suggesting that even individual domains are unfolded during transport (30). Even in that case, a flexibility in the domain arrangement would be important for each flagellin domain to properly position, orient, and bind to the indentation at the distal end of the filament just after the domains get folded in the small cavity between the cap plate and the distal end of the filament.
The peptide chain of HAP2 starts from the linker plate subdomain, because removal of unfolded terminal chains results in the loss of cap binding ability and decamer formation (13). The chain then goes through the kinked rod and the large leg subdomain and goes into the cap plate. Then, it comes back in the opposite direction. A feature distinct from flagellin is that the central portion of the chain forms the cap plate located near the filament axis, instead of the outermost domain. But the domain linkers are probably made of two antiparallel chains as well, as suggested by the narrow linker densities. These structural features are essential not only for assembly of HAP2 but also for its specific functions, namely, the stable binding over the symmetry mismatch and the stepwise cap rotation to promote the efficient self assembly of flagellin. The unfolded terminal regions of HAP2 in its monomeric state would form α-helical coiled coil on binding to the filament end, just as flagellin, but would go through repeated binding (folding) and unbinding (unfolding) for the cap rotation. This is another distinct feature from flagellin, whose terminal chains stay folded once the molecule is built into the filament structure.
New Animation for the Rotary Cap Mechanism for the Filament Growth. We have updated a graphic animation of the cap rotation and its leg domain movements promoting the efficient growth of the flagellar filament (14) to visualize more realistic motion of this molecular complex at work (see Movie 1, which is published as supporting information on the PNAS web site; also see www.npn.jst.go.jp).
Supplementary Material
Acknowledgments
We thank K. Imada for the HAP2 sample; K. Imada (in our laboratory), K. Hasegawa (Japan Synchrotron Radiation Research Institute, Hyogo, Japan), F. A. Samatey (in our laboratory) and I. Yamashita (Matsushita Electric Industrial Company, Kyoto) for making available the unpublished atomic coordinates of a straight flagellar filament; and F. Oosawa for support and encouragement.
This paper was submitted directly (Track II) to the PNAS office.
Abbreviation: FSC, Fourier shell correlation.
References
- 1.Berg, H. C. & Anderson, R. A. (1973) Nature 245, 380–382. [DOI] [PubMed] [Google Scholar]
- 2.Silverman, M. & Simon, M. (1974) Nature 249, 73–74. [DOI] [PubMed] [Google Scholar]
- 3.Larsen, S. H., Reader, R. W., Kort, E. N., Tso, W. W. & Adler, J. (1974) Nature 249, 74–77. [DOI] [PubMed] [Google Scholar]
- 4.Macnab, M. M. (1999) J. Bacteriol. 181, 7149–7153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Katayama, E., Shiraishi, T., Oosawa, K. & Aizawa, S.-I. (1996) J. Mol. Biol. 255, 458–475. [DOI] [PubMed] [Google Scholar]
- 6.Fan, F., Ohnishi, K, Francis, N. R. & Macnab, R. M. (1997) Mol. Microbiol. 26, 1035–1046. [DOI] [PubMed] [Google Scholar]
- 7.Minamino, T. & Macnab, R. M. (1999) J. Bacteriol. 181, 1388–1394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Iino, T. (1969) J. Gen. Microbiol. 56, 227–239. [DOI] [PubMed] [Google Scholar]
- 9.Emerson, S. U., Tokuyasu, K. & Simon, M. I. (1970) Science 169, 190–192. [DOI] [PubMed] [Google Scholar]
- 10.Ikeda, T., Homma, M., Iino, T., Asakura, S. & Kamiya, R. (1987) J. Bacteriol. 169, 1168–1173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Maki, S., Imada, K., Furukawa, Y., Vonderviszt, F. & Namba, K. (1998) J. Mol. Biol. 277, 771–777. [DOI] [PubMed] [Google Scholar]
- 12.Imada, K., Vonderviszt, F., Furukawa, Y., Oosawa, K. & Namba, K. (1998) J. Mol. Biol. 277, 883–891. [DOI] [PubMed] [Google Scholar]
- 13.Vonderviszt, F., Imada, K., Furukawa, Y., Uedaira, H., Taniguchi, H. & Namba, K. (1998) J. Mol. Biol. 284, 1399–1416. [DOI] [PubMed] [Google Scholar]
- 14.Yonekura, K., Maki, S., Morgan, D. G., DeRosier, D. J., Vonderviszt, F., Imada, K. & Namba, K. (2000) Science 290, 2148–2152. [DOI] [PubMed] [Google Scholar]
- 15.Ikeda, T., Oosawa, K. & Hotani, H. (1996) J. Mol. Biol. 259, 679–686. [DOI] [PubMed] [Google Scholar]
- 16.Ludtke, S. J., Baldwin, P. R. & Chiu, W. (1999) J. Struct. Biol. 128, 82–97. [DOI] [PubMed] [Google Scholar]
- 17.Frank, J., Radermacher, M., Penczek, P., Zhu, J., Li, Y., Ladjadj, M. & Leith, A. (1996) J. Struct. Biol. 116, 190–199. [DOI] [PubMed] [Google Scholar]
- 18.Yonekura, K., Maki-Yonekura, S. & Namba, K. (2001) J. Struct. Biol. 133, 246–253. [DOI] [PubMed] [Google Scholar]
- 19.Samatey, F. A., Imada, K., Nagashima, S., Vonderviszt, F., Kumasaka, T., Yamamoto, M. & Namba, K. (2001) Nature 410, 331–337. [DOI] [PubMed] [Google Scholar]
- 20.Yamashita, I., Hasegawa, K., Suzuki, H., Vonderviszt, F., Mimori-Kiyosue, Y. & Namba, K. (1998) Nat. Struct. Biol. 5, 125–132. [DOI] [PubMed] [Google Scholar]
- 21.Jones, T. A., Zhou, J. Y., Cowan, S. W. & Kjeldgaard, M. (1991) Acta Crystallogr. A 47, 110–119. [DOI] [PubMed] [Google Scholar]
- 22.Yonekura, K. & Toyoshima, C. (2000) Ultramicroscopy 84(1–2), 15–28. [DOI] [PubMed] [Google Scholar]
- 23.Toyoshima, C., Yonekura, K. & Sasabe, H. (1993) Ultramicroscopy 48, 165–176. [Google Scholar]
- 24.Merritt, E. A. & Bacon, D. J. (1997) Methods Enzymol. 277, 505–524. [DOI] [PubMed] [Google Scholar]
- 25.Sayle, R. A. & Milner-White, E. J. (1995) Trends Biochem. Sci. 20, 374–376. [DOI] [PubMed] [Google Scholar]
- 26.Wriggers, W. & Birmanns, S. (2001) J. Struct. Biol. 133, 193–202. [DOI] [PubMed] [Google Scholar]
- 27.Yonekura, K., Maki-Yonekura, S. & Namba, K. (2002) Res. Microbiol. 153, 191–197. [DOI] [PubMed] [Google Scholar]
- 28.Mimori-Kiyosue, Y., Vonderviszt, F. Yamashita, I., Fujiyoshi, Y. & Namba, K. (1996) Proc. Natl. Acad. Sci. USA 93, 15108–15113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Mimori-Kiyosue, Y., Vonderviszt, F. & Namba, K. (1997) J. Mol. Biol. 270, 222–237. [DOI] [PubMed] [Google Scholar]
- 30.Yonekura, K., Maki-Yonekura, S & Namba, K. (2003) Nature 424, 643–650. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.