SUMMARY
The repressor element 1-silencing transcription factor (REST) functions as a master regulator to maintain neural stem/progenitor cells (NPCs). REST undergoes proteasomal degradation through β-TrCP-mediated ubiquitination during neuronal differentiation. However, reciprocal mechanisms that stabilize REST in NPCs are undefined. Here we show that deubiquitinase HAUSP counterbalances REST ubiquitination and prevents NPC differentiation. HAUSP expression declines concordantly with REST upon neuronal differentiation and reciprocally with β-TrCP levels. HAUSP knockdown in NPCs decreases REST and induces differentiation. In contrast, HAUSP overexpression up-regulates REST by overriding β-TrCP-mediated ubiquitination. A consensus site (310-PYSS-313) of human REST is required for HAUSP-mediated REST deubiquitination. Furthermore, REST overexpression in NPCs rescues the differentiation phenotype induced by HAUSP knockdown. These data demonstrate that HAUSP stabilizes REST through deubiquitination and antagonizes β-TrCP in regulating REST at post-translational level. Thus, the HAUSP-mediated deubiquitination represents a critical regulatory mechanism involved in the maintenance of NPCs.
Transcriptional regulators of stem cell maintenance and differentiation require exquisite control to direct cell fate determination. Uncontrolled activation of core stem cell pathways drives transformation while loss of function in these cellular mechanisms leads to degenerative conditions. As regenerative medicine advances, understanding the regulation of self-renewal and lineage commitment becomes imperative. The brain has been the focus of numerous investigations into molecular mechanisms informing the maintenance and differentiation of neural stem cells due to the devastating nature of brain cancers and neurodegenerative diseases. REST (also known as neuron restrictive silencer factor, NRSF) is a critical transcription factor in regulating NPC self-renewal and lineage specification1–4, but REST itself is also regulated at both transcriptional and post-transcriptional levels. During neuronal differentiation, REST protein is targeted for proteosomal degradation by the E3 ubiquitin ligase SCF-β-TrCP (Skp1-Cul1-F-box protein)5, 6. As REST functions as a central transcriptional repressor of neuronal differentiation-associated genes to promote NPC maintenance2, 7, 8, aberrant REST function is associated with neurodegenerative diseases (e.g. Huntington’s disease) and other pathological states9–12. Neural tumors, specifically medulloblastomas and neuroblastomas, express high levels of REST and forced expression of REST can promote malignant transformation of neural progenitors13–15, although REST has been also reported as a tumor suppressor of colon cancer16. The effects of inappropriate REST expression would suggest that unidirectional negative regulation of REST protein by the β-TrCP-mediated ubiquitination during neuronal differentiation are balanced by a reciprocal mechanism that promotes REST stabilization via deubiquitination for NPC maintenance. Here, we demonstrated that deubiquitinase HAUSP (Herpesvirus-Associated Ubiquitin-Specific Protease, also known as Ubiquitin-Specific Protease 7, USP7)17, 18 prevents REST degradation through deubiquitination and promotes NPC maintenance.
RESULTS
HAUSP positively regulates REST protein levels in neural progenitor cells
Post-translational modifications are capable of rapidly regulating protein function and stability in response to cell state or external stimuli creating optimal points of network control for systems requiring precise regulation, including stem cell pathways. As irreversible commitment to a neuronal fate is controlled by the loss of REST protein by ubiquitination-mediated proteosomal degradation, we reasoned that this mechanism requires counterbalancing deubiquitination to prevent an instability of the control of NPC maintenance. We therefore screened for deubiquitinases that are nuclear in location and decline in expression during NPC differentiation. Using these criteria, we identified HAUSP as a deubiquitinase that gradually decreased in expression coordinated with decreased REST levels as fetal NPCs were induced to differentiate with all-trans retinoic acid (RA) treatment (Fig. 1a). HAUSP levels inversely correlated with lineage commitment as measured by acquisition of the neuronal marker TUJ1 (type III β-tubulin, a REST target gene) and the E3 ubiquitin ligase β-TrCP that targets REST for degradation (Fig. 1a). These results were confirmed by immunofluorescent staining of NPCs undergoing RA-induced differentiation (Fig. 1b, c). HAUSP and REST were highly expressed in nuclei of NPCs and their protein levels declined coordinately during the process of differentiation (Fig. 1b), while the expression of the neuronal lineage marker TUJ1 significantly increased upon differentiation (Fig. 1c). Thus, HAUSP deubiquitinase is positively associated with REST protein levels in NPCs.
Figure 1.
HAUSP and REST protein levels decline coordinately during neuronal differentiation. (a) Immunoblotting of HAUSP, β-TrCP, REST and TUJ1 (a neuronal differentiation marker) during differentiation. 15167 NPCs (neural stem/progenitor cells derived from a fetal brain by Lonza) were induced by all-trans retinoic acid (RA) to undergo cellular differentiation for the indicated times. HAUSP and REST protein levels gradually decreased, while the β-TrCP E3 ubiquitin ligase and the TUJ1 (type III β-tubulin, a REST target gene) levels increased during NPC differentiation. (b) Immunofluorescent staining confirmed that both HAUSP (red) and REST (green) protein levels declined during NPC differentiation. 15167 NPCs were induced by RA to undergo differentiation for the indicated times, then fixed and immunostained with anti-HAUSP and anti-REST specific antibodies. Nuclei were counterstained with DAPI (blue). (c) Immunofluorescent staining showed that HAUSP (red) decreased but the neuronal differentiation marker TUJ1 (green) increased during differentiation. ENStemA NPCs (derived from a human ES cell line by Chemicon/Millipore) were differentiated by RA treatment for the indicated times, fixed and stained with anti-HAUSP and anti-TUJ1 specific antibodies. Nuclei were counterstained with DAPI (blue). Uncropped images of blots are shown in Supplementary Information, Fig. S8.
To interrogate a mechanistic link between HAUSP activity and REST stability, we examined the effects of targeting HAUSP with shRNA (short hairpin RNA) on REST protein levels in NPCs. NPCs were transduced with lentiviruses expressing either a non-targeting (NT) control shRNA or one of two non-overlapping shRNA sequences (designated B2 and B5) against human HAUSP resulting in attenuation of HAUSP levels (70–85% reduction compared to the non-targeting control). HAUSP knockdown reduced REST protein levels but not the co-repressor CoREST in two separate NPC lines (Fig. 2 a, b), despite the presence of CoREST in a repressor complex with REST3, 19 supporting a specificity of HAUSP on REST rather than a general effect on the repressor complex. Immunofluorescent staining also showed that HAUSP knockdown resulted in reduced REST protein levels in NPC nuclei (Fig. 2c). This result was confirmed in an additional NPC line by puromycin selection of cells expressing the HAUSP-targeting shRNA reflected in nearly universal reduction of REST (Fig. 2d). In contrast, ectopic expression of HAUSP (HA-tagged HAUSP) in NPCs increased REST protein levels (Supplementary Information, Fig. S1a). Immunofluorescent staining validated that single cells overexpressing HA-HAUSP showed elevated REST protein levels (Supplementary Information, Fig. S1b). Collectively, these data demonstrate that HAUSP deubiquitinase positively regulates REST protein levels.
Figure 2.
HAUSP knockdown reduces REST protein levels in NPCs. (a, b) Immunoblotting showed that HAUSP knockdown by two distinct shRNAs (B2 and B5) decreased protein levels of REST but not CoREST in ENStemA (a) and 15167 (b) NPCs. NPCs were infected with lentiviruses expressing shHAUSP or non-targeting (NT) control shRNA for 48 hours, whole cell lysates were harvested for immunoblotting with the specific antibodies as indicated. (c, d) Immunofluorescent staining confirmed that HAUSP knockdown reduced REST levels in ENStemA (c) and 17231 (d) NPCs. Cells were cultured and attached on cover glasses coated with BD Matrigel hESC-qualified matrix, infected with lentiviruses expressing shHAUSP or control NT shRNA, treated without (c) or with (d) puromycin to select for infected cells, fixed and immunostained with anti-HAUSP and anti-REST specific antibodies. HAUSP was labeled in green, and REST was labeled in red. Nuclei were counterstained with DAPI (blue). Nuclei with reduced HAUSP and REST proteins are indicated by arrows in c. All Puromycin-selected cells infected with lentiviruses expressing HAUSP-targeting shRNA showed reduced HAUSP and REST protein levels in d. Lentiviral infection efficiency in NPCs with GFP-expressing lentiviruses is shown in Supplementary Information, Fig. S7. Uncropped images of blots are shown in Supplementary Information, Fig. S8.
HAUSP knockdown in neural progenitor cells induces neuronal differentiation and disrupts self-renewal
As REST prevents neuronal differentiation1–3, 5, 7 and we have found that HAUSP controls REST, we investigated the impact of targeting HAUSP on NPC neuronal differentiation. To interrogate the effect of HAUSP knockdown on cell fate determination in NPCs, we assayed Nestin as an NPC marker and TUJ1 as a neuronal marker by immunofluorescent staining. NPCs were transduced with either HAUSP-targeting shRNA (shHAUSP) or control NT shRNA. The overwhelming majority of NPCs expressing NT shRNA were Nestin-positive and rarely TUJ1-positive (Fig. 3a, top panel), indicating maintenance of an undifferentiated NPC state. In contrast, most cells targeted with shHAUSP became TUJ1-positive and only rarely remained Nestin-positive (Fig. 3 a, bottom panel), suggesting that targeting HAUSP induced neuronal differentiation. Indeed, quantified data indicate that HAUSP knockdown significantly increased the proportion of differentiated cells (Fig. 3b), suggesting that HAUSP deubiquitinase prevents NPC differentiation. To further causally link REST regulation to the effect of HAUSP on preventing NPC differentiation, we attempted to rescue the differentiation phenotype induced by HAUSP knockdown by overexpressing REST. Immunofluorescent staining showed that HAUSP knockdown in NPCs transfected with a vector control retained a differentiation phenotype but this effect was attenuated by overexpression of Flag-REST (Fig. 3c, d). This result was further validated by immunoblotting analysis demonstrating that the TUJ1 expression induced by HAUSP knockdown was reduced by the ectopic expression of Flag-REST in NPCs (Fig. 3e). These data indicate that ectopic expression of REST largely overrode the cell differentiation induced by HAUSP knockdown. In addition, knockdown of REST itself to the levels caused by HAUSP knockdown in NPCs induced similar differentiation (Supplementary Information, Fig. S2a–c), suggesting that REST knockdown phenocopied HAUSP knockdown in the induction of neuronal differentiation. Collectively, these data demonstrate that HAUSP functions largely through REST to prevent NPC differentiation.
Figure 3.
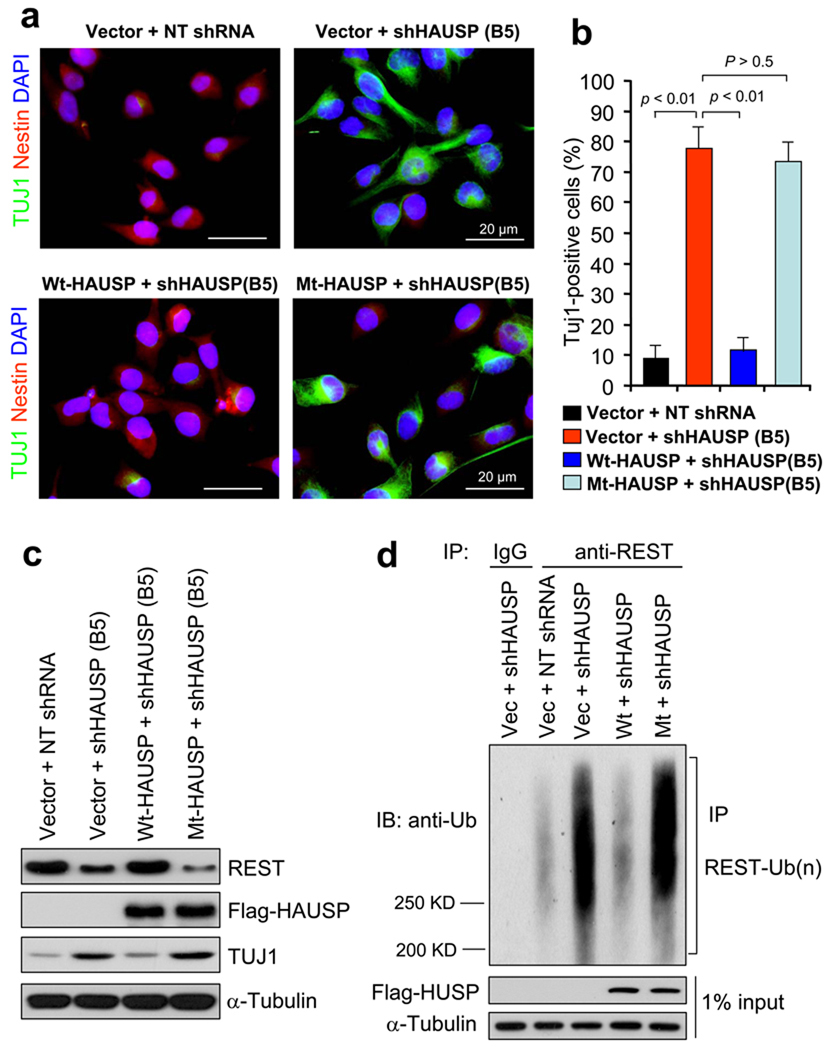
HAUSP knockdown promotes neural differentiation and decreases NPC self-renewal, and REST overexpression rescues the differentiation phenotype induced by HAUSP knockdown. (a) Targeting HAUSP with shRNA promotes neuronal differentiation. 15167 NPCs were infected with lentiviruses expressing shHAUSP (B5 clone) or non-targeting (NT) shRNA for 126 hours and immuno-stained for Nestin (an NPC maker, in red) and TUJ1 (a neuronal differentiation marker, in green). (b) Quantified data from a confirmed that HAUSP knockdown increased neuronal lineage specification. The fraction of cells expressing TUJ1 (green) significantly (p < 0.001) increased and the fraction of cells expressing Nestin (red) decreased after HAUSP knockdown in the NPCs. Data are presented as means ± s.d. [n = 3 (200 cells/experiment)]. (c) Immunofluorescent staining showed that ectopic REST expression rescued the differentiation phenotype induced by HAUSP knockdown. 17231 NPCs were transfected with Flag-REST or vector control, and infected with shHAUSP (B5 clone) or NT shRNA lentiviruses for 126 hours and immuno-stained for Nestin (red) and TUJ1 (green). (d) Quantified data from c indicated that ectopic expression of REST significantly (p < 0.001) attenuated the increased fraction of cells expressing TUJ1 induced by HAUSP knockdown. Data are presented as means ± s.d. [n = 3 (200 cells/experiment)]. (e) Immunoblotting confirmed that ectopic expression of REST repressed the TUJ1 expression induced by HAUSP knockdown. 17231 NPCs were transfected with Flag-REST or vector, and infected with shHAUSP (B5 clone) or NT shRNA lentiviruses for 96 hours, and immunoblotted with specific antibodies against HAUSP, Flag, TUJ1 and α-Tubulin. (f) Neurosphere formation assay showed that HAUSP knockdown reduced NPC self-renewal potential. 15167 NPCs were infected with shHAUSP or NT shRNA lentiviruses and allowed to form neurospheres in serum-free suspension culture. HAUSP knockdown reduced the neurosphere size and induced the attachment of neurosphere on the uncoated dishes. (g) Quantified data from f confirmed that HAUSP knockdown with two specific shRNAs (B2 and B5) significantly (p < 0.001) decreased the number of neurosphere formation of 15167 NPCs. Data are means ± s.d. (n = 3). Uncropped images of blots are shown in Supplementary Information, Fig. S8.
Self-renewal is a defining characteristic of stem cells so we examined the role of HAUSP in regulating NPC self-renewal potential. Targeting HAUSP via shRNA reduced the self-renewal of two different NPC preparations as demonstrated by the neurosphere formation assay (Fig. 3f, g; Supplementary Information, Fig. S3a, b). shHAUSP not only attenuated neurosphere formation frequency (a measure of self-renewal) and the neurosphere size (a measure of proliferation) but also induced cell attachment under neurosphere culture conditions (Fig. 3f; Supplementary Information, Fig. S3a). Moreover, the reduced neurosphere formation induced by HAUSP knockdown was significantly rescued by ectopic expression of Flag-REST (Supplementary Information, Fig. S3c, d). Taken together, these data demonstrate that HAUSP deubiquitinase maintains NPCs mainly by controlling REST levels.
HAUSP stabilizes REST protein through deubiquitination
To determine the mechanism through which HAUSP regulates REST expression, we interrogated effects of HAUSP on REST at different regulating levels. Theoretically, HAUSP might indirectly modulate REST through transcriptional control but we confirmed that HAUSP knockdown did not alter REST mRNA expression on RT-PCR analysis (Fig. 4). HAUSP functions as a deubiquitinase making post-translational regulation, most likely deubiquitination, as a potential link to REST control. We determined that HAUSP physically interacts with REST as demonstrated in co-immunoprecipitation assays (Fig. 5a, b; Supplementary Information, Fig. S4a, b), suggesting that HAUSP may mediate REST deubiquitination to prevent REST proteosomal degradation. We therefore examined effects of HAUSP knockdown on REST ubiquitination in NPCs. NPCs were transduced with NT shRNA or shHAUSP and treated with the MG132 proteasome inhibitor to assay the ubiquitinated REST. HAUSP knockdown increased REST poly-ubiquitination as demonstrated by both anti-REST immunoprecipitation (IP) and anti-Ubiquitin reciprocal IP (Fig. 5c; Supplementary Information, Fig. S4c). To confirm this result, we examined the effect of HAUSP overexpression on REST ubiquitination. NPCs were transduced with lentiviruses expressing Flag-tagged wild-type (Wt) HAUSP or catalytically dead mutant (Mt) HAUSP (C223S)20 and then assessed for REST ubiquitination. Ectopic expression of Wt-HAUSP but not the Mt-HAUSP decreased REST ubiquitination and increased REST protein levels (Fig. 5d; Supplementary Information, Fig. S4d). Collectively, these data demonstrate that HAUSP negatively regulates REST ubiquitination and promotes REST stabilization.
Figure 4.
Real-time PCR (RT-PCR) analysis indicated that reduced HAUSP expression by shRNA did not significantly alter REST mRNA expression but increased TUJ1 (a REST target gene) mRNA levels. ENStemA NPCs were targeted with shHAUSP (B5 clone) or control non-targeting (NT) shRNA for 72 hours through lentiviral infection. RNA samples were prepared for RT-PCR analysis with specific primers for HAUSP, REST, Co-REST and TUJ1. HAUSP mRNA was significantly down-regulated but REST mRNA levels were not significantly affected. Data are presented as means ± s.d. (n = 3).
Figure 5.
HAUSP mediates REST deubiquitination to regulate REST protein levels. (a, b) Immunoprecipitation (IP) showed that HAUSP and REST interact in NPCs. Cell lysates of ENStemA NPCs were immunoprecipitated with anti-REST (mAb) or anti-HAUSP antibody or IgG control and then immunoblotted with anti-HAUSP and anti-REST (rabbit polyclonal) antibodies. (c) Ubiquitination assays showed that HAUSP knockdown increased REST ubiquitination in NPCs. 17231 NPCs were infected with lentiviruses expressing shHAUSP or non-targeting (NT) shRNA for 48 hours and then treated with the proteasome inhibitor MG132 for 6 hours before harvest for IP. Cell lysates were immunoprecipitated with an anti-REST or anti-Ubiquitin specific antibody or control IgG, and then immunobloted with anti-Ubiquitin or anti-REST specific antibody. Both IP with anti-REST antibody and the reciprocal IP with anti-Ubiquitin antibody confirmed that HAUSP knockdown increased REST poly-ubiquitination. (d) Ectopic expression of wild-type HAUSP (Wt-HAUSP) but not the catalytic dead mutant (223C→S) HAUSP (Mt-HAUSP) reduced REST ubiquitination. 17231 NPCs were transfected with the Flag-tagged Wt-HAUSP (Wt), Mt-HAUSP (Mt) or vector (V) control through lentiviral infection for 36 hours, then treated with the proteasome inhibitor MG132 for 6 hours, and subjected for analysis of REST ubiquitination. Ectopic expression of Wt-HAUSP but not Mt-HAUSP reduced REST ubiquitination in the NPCs. (e) In vitro deubquitination assay showed that the β-TrCP-mediated REST ubiquitination was specifically inhibited by Wt-HAUSP (lane 3) but not by the Mt-HAUSP (a catalytic dead mutant, lane 4) or control deubiquitinase USP1 (lane 5). Flag-Wt-HAUSP, Flag-Mt-HAUSP, Myc-REST, HA-β-TrCP and Flag-USP1 were individually overexpressed in 293T cells, and then purified with the specific antibody or the corresponding tag antibody for this assay. (f) In vivo deubiquitination assay confirmed that the β-TrCP-mediated REST ubiquitination was specifically attenuated by the Wt-HAUSP but not the catalytic dead Mt-HAUSP. 293T cells were transfected with the indicated sets of plasmids for 48 hours, treated with the proteasome inhibitor MG132 for 6 hours and then subjected for analysis of REST ubiquitination in the samples expressing different set of proteins as indicated. Uncropped images of blots are shown in Supplementary Information, Fig. S8.
To further address whether REST deubiquitination is directly and specifically mediated by HAUSP, we performed an in vitro deubiquitination assay20 with purified wild-type HAUSP (Flag-Wt-HAUSP), catalytic dead mutant HAUSP (Flag-Mt-HAUSP), or a control deubiquitinase USP1 (Flag-USP1). Wt-HAUSP specifically counteracted β-TrCP-mediated REST ubiquitination, while the Mt-HAUSP or the control deubiquitinase USP1 did not alter the REST ubiquitination (Fig. 5e), suggesting that HAUSP directly and specifically deubiquitinated REST and suppressed REST ubiquitination. This result was validated by the in vivo deubiquitination assay in 293T cells expressing Wt-HAUSP or the catalytic dead Mt-HAUSP in combination with REST, Ubiquitin and β-TrCP (Fig. 5f). Consistently, expression of β-TrCP increased REST poly-ubiquitination (Fig. 5f, lane 2, 3), which was attenuated by expression of Wt-HAUSP (Fig. 5f, lane 2–4). In contrast, expression of the catalytic dead Mt-HAUSP did not modulate the REST poly-ubiquitination induced by β-TrCP expression (Fig. 5f, lane 3–5). These data demonstrated that HAUSP counteracts β-TrCP-mediated REST ubiquitination through specific deubiquitination.
As HAUSP knockdown reduced REST protein levels and promoted NPC differentiation, we further examined whether ectopic expression of Wt-HAUSP overrides shHAUSP-induced differentiation. Because the B5 shHAUSP targets the 3’-end non-coding region of endogenous HAUSP mRNA while the Wt-HAUSP and Mt-HAUSP constructs do not contain the 3’-end non-coding sequence, mRNA from Wt-HAUSP or Mt-HAUSP is not targeted by B5 shHAUSP. Thus, we were able to simultaneously knock down endogenous HAUSP and overexpress Wt-HAUSP or Mt-HAUSP in NPCs. Immunofluorescent staining showed that ectopic expression of Wt-HAUSP but not the Mt-HAUSP almost fully rescued the differentiation phenotype induced by knockdown of endogenous HAUSP with the B5 shHAUSP (Fig. 6a, b). Immunoblotting confirmed that ectopic expression of Wt-HAUSP but not the Mt-HAUSP attenuated TUJ1 expression induced by knockdown of endogenous HAUSP (Fig. 6c). Moreover, deubiquitination assay demonstrated that expression of Wt-HAUSP but not the Mt-HAUSP attenuated the REST ubiquitination induced by B5 shHAUSP (Fig. 6d), showing that Wt-HAUSP functioned through deubiquitination to stabilize REST and rescued the differentiation phenotype induced by knockdown of endogenous HAUSP. These data demonstrate that expression of functional HAUSP is critical for preventing REST ubiquitination and suppressing NPC differentiation.
Figure 6.
Neuronal differentiation induced by knockdown of endogenous HAUSP was rescued by ectopic expression of wild-type HAUSP but not the catalytic dead HAUSP mutant. As the B5 shHAUSP clone targets 3’-end non-coding region of endogenous HAUSP mRNA, ectopic expression of the wild-type and mutant HAUSP that lack the 3’-end non-coding region was not affected by the B5 HAUSP shRNA. (a) Immunofluorescent staining showed that ectopic expression of wild-type HAUSP (Wt-HAUSP) but not the catalytic dead mutant HAUSP (Mt-HAUSP) rescued the neuronal differentiation phenotype induced by knockdown of endogenous HAUSP. 17231 NPCs were transfected with Flag-tagged Wt-HAUSP or Mt-HAUSP, or vector control through lentiviral infection, and infected with lentiviruses expressing shHAUSP (B5 clone) or control non-targeting (NT) shRNA for 96 hours and immuno-stained with specific antibodies against Nestin (red) and TUJ1 (green). Ectopic expression of Wt-HAUSP but not the Mt-HAUSP suppressed the expression of neuronal marker TUJ1 that was induced by knockdown of endogenous HAUSP. (b) Quantified data from a indicated that ectopic expression of the Wt-HAUSP but not the Mt-HAUSP in NPCs almost fully rescued the differentiation phenotype induced by knockdown of the endogenous HAUSP. Data are means ± s.d. [n = 4 (200 cells/experiment)]. (c) Immunoblotting validated that ectopic expression of Wt-HAUSP but not the Mt-HAUSP resulted in suppression of TUJ1 expression. 17231 NPCs were transfected with Flag-tagged Wt-HAUSP or Mt-HAUSP, or vector control through lentiviral infection, and infected with lentiviruses expressing shHAUSP (B5 clone) or NT shRNA for 72 hours and immuno-stained with specific antibodies against REST, Flag, TUJ1 and α-Tubulin (loading control). (d) Deubiquitination assay showed that ectopic expression of Wt-HAUSP but not the catalytic dead Mt-HAUSP attenuated the REST ubiquitination induced by knockdown of endogenous HAUSP. 15167 NPCs were transfected with Flag-tagged Wt-HAUSP or Mt-HAUSP, or vector control through lentiviral infection, and infected with lentiviruses expressing shHAUSP (B5 clone) or NT shRNA for 48 hours, treated with the proteasome inhibitor MG132 for 6 hours, immunoprecipitated (IP) with anti-REST specific antibody or the IgG control, and immunobloted with anti-Ubiquitin specific antibody. Uncropped images of blots are shown in Supplementary Information, Fig. S8.
A consensus binding site (310-PYSS-313) of human REST is required for the HAUSP-mediated REST deubiquitination
Our data demonstrate that HAUSP directly deubiquitinates REST. As the known HAUSP substrates usually contain a consensus sequence (P/AXXS) for the HAUSP-mediated specific deubiquitination21, 22, we sought to identify the HAUSP consensus binding sequences on the human REST protein. Human REST contains five P/AXXS sequences (Supplementary Information, Fig. S5). To determine which sequence is critical for the HAUSP-mediated REST deubiquitination, we sequentially mutated each of the critical amino acid Ser to Ala (S→A) of these potential sites and expressed mutant REST constructs in NPCs. We found that only the S313A mutation of 310-PYSS-313 of human REST disrupted the HAUSP-mediated REST deubiquitination (Fig. 7a, b). In contrast, a similar mutation (S1042A) on other potential sites such as 1039-PQES-1042 did not prevent the HAUSP-mediated REST deubiquitination, and S313A/S1042A double mutations (Flag-REST-AA) showed similar effect to the S313A single mutation on suppressing the HAUSP-mediated REST deubiquitination (Fig. 7b). Thus, we have identified the consensus HAUSP binding site (310-PYSS-313) on human REST that is required for the HAUSP-mediated REST deubiquitination. These data further support that REST deubiquitination is specifically mediated by HAUSP.
Figure 7.
A consensus site of human REST (310-PYSS-313) is required for the HAUSP-mediated REST deubquitination and HAUSP counteracts β-TrCP-mediated REST ubiquitination. (a) Diagram of human REST protein. A consensus sequence (310-PYSS-313) of human REST required for the HAUSP-mediated REST deubiquitination and a critical mutation (S313A) on this site that disrupts the deubiquitination are shown. (b) In vivo deubiquitination assay showed that a critical mutation (S313A) on the consensus sequence (310-PYSS-313) of human REST attenuated the HAUSP-mediated REST deubiquitination (see lanes 2–4), while a similar mutation (S1042A) on another potential site (1039-PQES-1042) did not alter the HAUSP-mediated REST deubiquitination (see lane 3–5). S313A/S1042A double mutations (Flag-REST-AA) and S313A single mutation showed similar effect on the HAUSP-mediated REST deubiquitination (see lane 4–6). 293T cells were transfected with the indicated set of expression plasmids, and treated with the proteasome inhibitor MG132 for 6 hours, and then subjected for analysis of REST ubiquitination. (c) REST ubiquitination increased during neuronal differentiation. 17231 NPCs differentiation was induced by all-trans retinoic acid (RA) for four days, treated with MG132 for 6 hours, and harvested for a ubiquitination assay. Cell lysates from NPCs or differentiated cells were subjected to immunoprecipitation with anti-REST antibody (Rabbit) and immunoblotted with anti-Ubiquitin and the anti-REST antibodies (mAb). (d) Double knockdown analysis confirmed that REST protein is controlled by both β-TrCP-mediated ubiquitination and the HAUSP-mediated deubiquitination in NPCs. 15167 NPCs were treated with RA for only 24 hours to initiate differentiation, and transduced with shHAUSP, shβ-TrCP, both shHAUSP and shβ-TrCP, or non-targeting (NT) shRNA for 36 hours through lentiviral infection, treated with MG132 for 6 hours, and harvested for ubiquitination assessment. HAUSP knockdown alone increased REST ubiquitination (lanes 1, 2), while β-TrCP knockdown alone reduced REST ubiqutination (lanes 1, 3). However, HAUSP and β-TrCP double knockdown restored REST ubiquitination that was reduced by β-TrCP knockdown and abolishes REST ubiquitination induced by HAUSP knockdown (lanes 1–4). Uncropped images of blots are shown in Supplementary Information, Fig. S8.
HAUSP and β-TrCP function as oppositional counterparts to control REST protein stability at post-translational level
As both HAUSP and REST protein levels gradually decline and the E3 ubiquitin ligase β-TrCP increases during NPC differentiation (Fig. 1), we next examined the roles of reduced HAUSP and increased β-TrCP in regulating REST ubiquitination during neural differentiation. Under RA-induced differentiation of NPCs, REST poly-ubiquitination increased with reduction of total REST protein levels (Fig. 7c; Supplementary Information, Fig. S6a). Because HAUSP knockdown increased REST ubiquitination (Fig. 5c) and β-TrCP promoted REST ubiquitination (Fig. 5e, f), it is likely that both decreased HAUSP and increased β-TrCP contribute to increased REST ubiquitination during neuronal differentiation. We hypothesized that HAUSP functions as a critical counterbalance to β-TrCP to inhibit REST ubiquitination and maintain NPCs. Thus, we examined the effects of knocking down both HAUSP and β-TrCP to determine REST ubiquitination during NPC differentiation. To initiate neuronal differentiation, NPCs were treated with RA for 48 hours and transduced with NT shRNA, shHAUSP, β-TrCP – targeting shRNA (shβ-TrCP), or both shHAUSP and shβ-TrCP. Cells were treated with RA long enough to initiate differentiation but assessed before terminal differentiation so both HAUSP and β-TrCP were expressed. As expected, shHAUSP increased REST ubiquitination (Fig. 7d, lane 1, 2), while shβ-TrCP decreased REST ubiquitination (Fig. 7d, lane 1, 3). Simultaneous targeting of HAUSP and β-TrCP caused intermediate levels of REST ubiquitination (Fig. 7d, lane 2–4), suggesting that both HAUSP and β-TrCP regulate REST ubiquitination during NPC differentiation. Collectively, these data demonstrated HAUSP deubiquitinase and β-TrCP ubiquitin E3 ligase function as oppositional counterparts (“Ying-Yang”) to control REST protein levels (Fig. 8). To further confirm this important point, we reconstituted a regulatory system by overexpressing Myc-REST and HA-ubiquitin in combination with HAUSP, Flag-β-TrCP or both HAUSP and Flag-β-TrCP in 293T cells that have minimal expression of endogenous REST and HAUSP. Thus, we were able to directly assess REST regulation by the β-TrCP-mediated ubiquitination and the HAUSP-mediated deubiquitination in a defined cellular system. Consistently, expression of HAUSP reduced REST poly-ubiquitination (Supplementary Information, Fig. S6b, lanes 2, 1), and overexpression of β-TrCP increased REST ubiquitination (Supplementary Information, Fig. S6b, lanes 3, 2). Importantly, overexpression of HAUSP with β-TrCP together abolished the increased REST ubiquitination induced by β-TrCP overexpression (Supplementary Information, Fig. S6b, lanes 4, 3). Taken together, these data further validated that both HAUSP-mediated deubiquitination and the β-TrCP-mediated ubiquitination serve opposite roles to control REST protein stability at the post-translational level. HAUSP stabilizes REST protein through deubiquitination and promotes NPC maintenance, while β-TrCP mediates REST degradation through ubiquitination and stimulates NPC differentiation. The balance between ubiquitination and deubiquitination may determine the net REST protein levels and cellular fate. This post-translational regulatory mechanism of REST is crucial for determining NPC maintenance or differentiation.
Figure 8.
“Ying-Yang” control model of REST protein at the post-translational level. Both HAUSP-mediated deubiquitination (“Ying”) and β-TrCP-mediated ubiquitination (“Yang”) regulate REST protein levels in NPCs. HAUSP deubiquitinase stabilizes REST protein to promote NPC maintenance. In contrast, the β-TrCP E3 ubiquitin ligase mediates REST ubiquitination and degradation to promote neuronal differentiation. The net balance between the β-TrCP-mediated ubiquitination and the HAUSP-mediated deubiquitination controls REST protein levels and determine cellular fate.
DISCUSSION
We now demonstrate that HAUSP stabilizes REST through deubiquitination and promotes maintenance of NPCs. HAUSP was originally identified to be associated with viral proteins such as ICP0 (Herpesvirus) and EBNA1 (EB virus) during viral infection17, 18, 22. A series of elegant studies have demonstrated that HAUSP also regulates the stability and functions of several important proteins under normal and stress conditions20, 21, 23–25. Our study expands the roles of the HAUSP deubiquitinase in the maintenance of NPCs by stabilizing REST to repress neuronal differentiation. Although HAUSP has several molecular targets, the ability to rescue the effects of HAUSP modulation on NPC maintenance and differentiation by REST expression indicates that HAUSP functions largely through REST to prevent NPC differentiation. However, it is possible that HAUSP may also control other stem cell transcription factors for the maintenance of other stem cells.
REST is a key transcriptional repressor of neuronal differentiation genes1–3, 7, 8 to prevent NPC differentiation1–8. In accordance with its role in silencing both neuronal and non-neuronal genes, REST is also essential for embryonic development and for a number of cellular responses8, 26, 27. REST has also been implicated in the regulation of mitotic fidelity in non-neural cells6 and was proposed as a tumor suppressor16. Although there is controversy over the requirement of REST for maintaining ESC pluripotency28–30, the important role of REST in repressing neuronal differentiation and promoting NPC maintenance has been extensively demonstrated1–8. Our study suggested that post-translational control of REST by ubiquitination and deubiquitination is critical for regulating REST protein levels to determine cellular fate of NPCs.
We have demonstrated that HAUSP specifically stabilizes REST protein through deubiquitination and a critical consensus site (310-PYSS-313) on human REST is required for the HAUSP-mediated REST deubiquitination. It has been reported that the association of TRF2 (telomere repeat factor 2) with REST also prevents the ubiquitin proteasome-mediated degradation of REST31. However, as TRF2 does not have deubiquitinase activity, TRF2 is unlikely to deubiquitinate REST. Whether TRF2 modulates HAUSP or β-TrCP function to indirectly affect REST stability is under investigation. Our study establishes the ubiquitination-deubiquitination system as a critical post-translational control mechanism to regulate the key stem cell transcriptional factors and thus determine stem cell maintenance or differentiation. Based on our results and previous studies5, 6, we propose that deubiquitination and ubiquitination system works as “Ying-Yang” control system to regulate REST protein levels (Fig. 8) with the net balance between the HAUSP-mediated deubiquitination and β-TrCP-mediated ubiquitination controlling REST protein levels to direct cell fate. The relative activity of deubiquitination and ubiquitination for REST may define the maintenance of “stemness” or initiation of neuronal differentiation. When the HAUSP-mediated deubiquitination overrides β-TrCP-mediated ubiquitination, REST is stabilized to suppress differentiation and promote NPC maintenance. In contrast, when the β-TrCP-mediated ubiquitination exceeds HAUSP-mediated deubiquitination, REST is targeted for degradation, which promotes cell differentiation by releasing repression of differentiation-associated genes. This paradigm of reciprocal yoked post-translational control may be present in other stem cell regulatory networks to permit cells to efficiently and finely tune the levels of critical factors that have potent cellular effects in excess or in shortage.
In conclusion, our study uncovered a crucial post-translational control mechanism to regulate the key stem cell transcription factor for cell fate determination of NPCs. NPCs have been proposed as regenerative resources for some CNS diseases. As the use of these cells requires sufficient proliferation to supply requisite cell numbers while permitting appropriate differentiation for the stem cell-based therapies, understanding the molecular mechanisms controlling NPC maintenance and differentiation will have significant impacts on improving the treatment of neural degenerative diseases.
METHODS
Cells and cell culture
15167 and 17231 NPCs (derived from human fetal brains, Lonza) and the ENStemA NPCs (derived from a human embryonic stem cell line, Chemicon/Millipore) were cultured in neurobasal medium supplemented with B27 and epidermal growth factor (EGF)/basic fibroblast growth factor (bFGF) as described in our previous reports32, 33. These cells were cultured either in neurosphere suspension or in attached monolayer on the tissue culture dishes coated with the BD Matrigel hESC-qualified matrix (BD Bioscience). To induce differentiation, cells were treated with 1 µM of all-trans retinoic acid (RA, Sigma-Aldrich) for the indicated time.
Immunoblotting and immunofluorescent staining
These basic methods were performed as described in our previous reports32–34. Anti-human REST monoclonal antibody (murine Clone 7D1.3, Millipore), rabbit polyclonal anti-REST antibodies (Abcam, Santa Cruz), rabbit polyclonal anti-HAUSP antibody (Abcam), anti-human HAUSP monoclonal antibody (Bethyl laboratories), anti-β-TrCP antibodies (Santa Cruz), anti-Flag (Sigma-Aldrich), anti-HA (Sigma-Aldrich), anti-TUJ1 (Covance), anti-Nestin (Millipore), anti-Myc tag (Sigma-Aldrich), anti-Ubiquitin (P4D1, Santa Cruz) and anti-α-Tubulin (Sigma-Aldrich) antibodies were used for immunoblotting, immunoprecipitation or immunofluorescent staining. Briefly, for immunofluorescent staining NPCs cultured on the BD Matrigel-coated cover glasses were induced by 1 µM of RA for differentiation or infected with lentiviruses expressing HAUSP-targeting shRNA (shHAUSP) or non-targeting (NT) control shRNA, fixed in 4% PFA (paraformaldehyde) for 15 minutes, permeablized, blocked by blocking buffer containing 0.1% BSA (bovine serum albumin) and 0.3% triton X-100 for 30 minutes, incubated with the primary antibodies over night at 4 °C, incubated with the fluorescence-conjugated secondary antibodies (Invitrogen) at room temperature for 1 hour. Nuclei were counterstained with DAPI.
Knockdown by lentiviral vector-mediated shRNA
shRNA clones for knocking down human HAUSP, REST and β-TrCP were obtained by screening lentivirus shRNA sets (Mission shRNA, Sigma-Aldrich). The production of lentivirus and viral infection of NPCs were performed as previously described33, 34. Lentiviral infection efficiency in NPCs was determined with the lentiviruses expressing GFP (green-fluorescent protein). When the MOI (Multiplicity of Infection) of 3 was used for the infection, the majority (>93%) of NPCs were infected by the viruses and expressed GFP (Supplementary Information, Fig. S7).
Generations of plasmid constructs
Human cDNA clones for REST (BC132859), β-TrCP (MGC: 40028) and ubiquitin (MGC: 8385) were obtained from Open Biosystems. pCI-neo HAUSP was acquired from B. Vogelstein through Addgene. Open reading frames of these cDNAs were PCR-amplified using Platinum High Fidelity PCR Supermix (Invitrogen) according to manufacturer’s protocol and then subcloned into XbaI and BamHI sites of lentivector pLCMV-Neo (a kind gift of Dr. Peter Chumakov) with either Flag, HA, His or Myc tag coding sequences in frame and verified by sequencing. All specific mutations were generated by using QuikChange® II XL Site-Directed Mutagenesis Kit (Stratagene) according to manufacturer’s instruction. The catalytic dead mutant (223C→S) HAUSP (Mt-HAUSP) was generated with PCR primers 5’-GAA TCA GGG AGC GAC TTC TTA CAT GAA CAG CCT GC-3’ and 5’-GCA GGC TGT TCA TGT AAG AAG TCG CTC CCT GAT TC-3’. The REST-S313A mutant (HAUSP binding site mutation) were generated with PCR primers 5’-CTT TGT CCT TAC TCA GCT TCT CAG AAG ACT CATC-3’ and 5’-GAT GAG TCT TCT GAG AAG CTG AGT AAG GAC AAA G-3’.
In vivo ubiquitination and deubiquitination assay in NPCs
Human NPCs (before and after HAUSP or β-TrCP knockdown), RA-induced differentiated cells, or the cells overexpressing HAUSP, β-TrCP, REST or targeting shRNAs subjected for the ubiquitination assays were treated with the proteasome inhibitor MG132 (20 µM, Sigma-Aldrich) for 6 hours and then harvested for immunoprecipitation (IP) with anti-REST, anti-Flag, anti-HA, or anti-Ubiquitin antibody followed immunobloting with an anti-Ubiquitin, anti-REST, or anti-HA antibody. Briefly, cell lysates (500 µg of total protein) were incubated with 1 µg of anti-REST antibody (H-290, Santa Cruz), anti-Ubiquitin antibody (P4D1, Santa Cruz), or normal rabbit IgG with constant rotation overnight at +4°C. Immunocomplexes were captured by 20 µl of Protein A/G Plus Agarose beads (Santa Cruz) for 1 hour at +4°C, washed 3 times with ice-cold lysis buffer and eluted in 2× loading buffer by boiling for 10 minutes, and then analyzed by immunoblotting. Proteins were resolved on Tris-acetate gels (Invitrogen), blotted onto PVDF (polyvinylidene) membranes and probed by antibodies specific to REST (Millipore) and Ubiquitin (P4D1, Santa Cruz).
In vitro deubiquitination assay
The in vitro deubiquitination assay was performed as described6. Myc-REST-WT was immunoprecipitated with anti-REST antibody from 293T cells transfected with Myc-REST-WT plasmid. Flag-β-TrCP, Flag-USP1, Flag-WT-HAUSP and Flag-Mt-HAUSP were purified by the same way. In vitro ubiquitination assay was performed in a 25 µl mixture including the indicated purified proteins and 50 mM Tris-HCl pH 7.6, 5 mM MgCl2, 0.6 mM dithiothreitol, 2 mM ATP, 1.5 ng/µl His-E1, 10 ng/µl His-Ubc3, 10 ng/µl His-Ubc5, 2.5 µg/µl Ubiquitin, and 1 µM ubiquitin aldehyde (all from Boston Biochem). Then the reactions were incubated at 30°C for 1 hour and analyzed by anti-REST Western blot.
REST-HAUSP interaction
REST Immunoprecipitation: 293T cells were transfected with plasmids expressing Myc-REST and Flag-HAUSP using Lipofectamine and Plus reagent (Invitrogen) according to manufacturer’s recommendations. Cells were harvested in lysis buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 5 mM EDTA, 1% NP-40 supplemented with protease inhibitors) and lysates were pre-cleared by centrifugation at 15k g for 15 minutes. 500 µg of total protein lysate were subjected to immunoprecipitation with 2 µg of α-REST (NRSF) antibody (H-290, Santa Cruz) or normal rabbit IgG for 3 hours at 4°C. Immunocomplexes were captured by 20 µl of protein A/G Plus agarose beads (Santa Cruz) for 1 hour at 4°C, washed 5 times with lysis buffer, eluted in 2× Laemmli buffer by boiling for 5 minutes and resolved by SDS-PAGE. HA (HAUSP) Immuoprecipitation: 293T cells were transfected with plasmids expressing HA-HAUSP (or empty vector) and Myc-REST using Lipofectamine and Plus reagent (Invitrogen) according to manufacturer’s instructions. Cells were harvested in lysis buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 5 mM EDTA, 1% NP-40 with protease inhibitors) and lysates were precleared by centrifugation at 15k g for 15 minutes. 500 µg of total protein lysate were incubated with 25 µl of EZview Red Anti-HA Affinity Gel (Sigma-Aldrich) for 3 hours at 4°C and washed 5 times with lysis buffer. Captured proteins were eluted in 2× Laemmli buffer by boiling for 5 minutes and resolved by SDS-PAGE.
REST ubiquitination and deubiquitination assays in 293T
293T cells were transfected using Lipofectamine and Plus reagent (Invitrogen) to express HA-Ub, Myc-REST, Flag-β-TrCP, HAUSP or corresponding empty vectors as indicated. Prior to harvesting cells were treated with 20 µM MG-132 proteasome inhibitor for 4 hours. Cells were harvested in lysis buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 5 mM EDTA, 1% NP-40 supplemented with protease inhibitors) and lysates were precleared by centrifugation at 15k g for 15 minutes. 500 µg of total protein lysate were incubated with 25 µl of EZview Red Anti-HA Affinity Gel (Sigma-Aldrich) for 3 hours at 4°C with constant rotation, beads were washed 5 times in lysis buffer and captured proteins were eluted in 2× Laemmli buffer by boiling for 5 minutes.
Real Time PCR
mRNA samples were isolated from 15167 or ENStemA NPCs infected with lentiviruses expressing HAUSP-targeting shRNA or non-targeting control shRNA for 72 hours, and then subjected for real-time PCR (RT-PCR) analysis using the following primer pair: HAUSP (Forward Primer: 5’-ACT TTG AGC CAC AGC CCG GTA ATA-3’, Reverse Primer: 5’-GCC TTG AAC ACA CCA GCT TGG AAA-3’); REST (Forward Primer: 5’-CGC CAT GCA AGA CAG GTT CAC AAT-3’, Reverse Primer: 5’-AGC TGC ATA GTC ACA TAC AGG GCA-3’); Co-REST (Forward Primer: 5’-AAC GGG ACA ATC TTG GCA TGT TGG-3’, Reverse Primer: 5’-AGA GCC TGT TCC ATG TTG TAC CCA-3’) and TUJ1 (Forward Primer: 5’-ATC AGC AAG GTG CGT GAG GAG TAT-3’, Reverse Primer: 5’-TCG TTG TCG ATG CAG TAG GTC TCA-3’) in the 7900HT Fast Real-Time PCR System (AB Applied Biosystems).
Statistical analysis
All grouped data are presented as mean ± standard deviation (s.d.). Difference between groups was assessed by one-way ANOVA or one-way ANOVA on ranks tests. SigmaStat Software (Version 3.5) was used for all statistical analyses.
Supplementary Material
ACKNOWLEDGEMENTS
We thank members in Dr. Rich’s laboratory for helpful discussions and suggestions. We are also grateful to the Flow Cytometry Core, Imaging Core and Central Cell Services at Cleveland Clinic Lerner Research Institute for their help and services. This work was supported by a research fund from Cleveland Clinic Foundation and a NIH grant (NS070315) to S.B.
Footnotes
AUTHOR CONTRIBUTIONS
Z.H., Q.W., O.G. and L.C. performed and planed all experiments. S.B. developed the hypothesis, coordinated the study, oversaw the research and results and wrote the manuscript. J.R. helped to write the manuscript and provided input into design and interpretation. W.S. provided reagents and helpful suggestions. The work was carried out in the laboratory of S.B.
COMPETING FINANCIAL INTERESTS
The authors declare that they have no competing financial interests.
References
- 1.Chong JA, et al. REST: a mammalian silencer protein that restricts sodium channel gene expression to neurons. Cell. 1995;80:949–957. doi: 10.1016/0092-8674(95)90298-8. [DOI] [PubMed] [Google Scholar]
- 2.Schoenherr CJ, Anderson DJ. The neuron-restrictive silencer factor (NRSF): a coordinate repressor of multiple neuron-specific genes. Science. 1995;267:1360–1363. doi: 10.1126/science.7871435. [DOI] [PubMed] [Google Scholar]
- 3.Ballas N, Grunseich C, Lu DD, Speh JC, Mandel G. REST and its corepressors mediate plasticity of neuronal gene chromatin throughout neurogenesis. Cell. 2005;121:645–657. doi: 10.1016/j.cell.2005.03.013. [DOI] [PubMed] [Google Scholar]
- 4.Abrajano JJ, et al. Differential deployment of REST and CoREST promotes glial subtypes specificcatio and oligodendrocyte lineage maturation. PLoS One. 2010;4:e7665. doi: 10.1371/journal.pone.0007665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Westbrook TF, et al. SCFβ-TRCP controls oncogenic transformation and neural differentiation through REST degradation. Nature. 2008;452:370–374. doi: 10.1038/nature06780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Guardavaccaro D, et al. Control of chromosome stability by the β-TrCP-REST-Mad2 axis. Nature. 2008;452:365–369. doi: 10.1038/nature06641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ballas N, Mandel G. The many faces of REST oversee epigenetic programming of neuronal genes. Curr. Opin. Neurobiol. 2005;15:500–506. doi: 10.1016/j.conb.2005.08.015. [DOI] [PubMed] [Google Scholar]
- 8.Johnson R, et al. REST regulates distinct transcriptional networks in embryonic and neural stem cells. PLoS Biol. 2008;6:2205–2219. doi: 10.1371/journal.pbio.0060256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zuccato C, et al. Huntingtin interacts with REST/NRSF to modulate the transcription of NRSE-controlled neuronal genes. Nature Genet. 2003;35:76–83. doi: 10.1038/ng1219. [DOI] [PubMed] [Google Scholar]
- 10.Lepagnol-Bestel AM, et al. Huntingtin interacts with REST/NRSF to modulate the transcription of NRSE-controlled neuronal genes. Hum. Mol. Genet. 2009;18:1405–1414. [Google Scholar]
- 11.Tahiliani M, et al. The histone H3K4 demethylase SMCX links REST target genes to X-linked mental retardation. Nature. 2007;447:601–605. doi: 10.1038/nature05823. [DOI] [PubMed] [Google Scholar]
- 12.Canzonetta C, et al. DYRK1A-dosage imbalance perturbs NRSF/REST levels, deregulating pluripotency and embryonic stem cell fate in Down syndrome. Am. J. Hum. Genet. 2008;83:388–400. doi: 10.1016/j.ajhg.2008.08.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lawinger P, et al. The neuronal repressor REST/NRSF is an essential regulator in medulloblastoma cells. Nature Med. 2000;6:826–831. doi: 10.1038/77565. [DOI] [PubMed] [Google Scholar]
- 14.Su X, et al. Abnormal expression of REST/NRSF and Myc in neural stem/progenitor cells causes cerebellar tumors by blocking neuronal differentiation. Mol. Cell Biol. 2006;26:1666–1678. doi: 10.1128/MCB.26.5.1666-1678.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Fuller GN, et al. Many human medulloblastoma tumors overexpress repressor element-1 silencing transcription (REST)/neuron-restrictive silencer factor, which can be functionally countered by REST-VP16. Mol. Cancer Ther. 2005;4:343–349. doi: 10.1158/1535-7163.MCT-04-0228. [DOI] [PubMed] [Google Scholar]
- 16.Westbrook T, et al. A genetic screen for candidate tumor suppressors identifies REST. Cell. 2005;121:837–848. doi: 10.1016/j.cell.2005.03.033. [DOI] [PubMed] [Google Scholar]
- 17.Holowaty MN, et al. Protein profiling with Epstein-Barr nuclear antigen-1 reveals an interaction with the herpesvirus-associated ubiquitin-specific protease HAUSP/USP7. J. Biol. Chem. 2003;278:29987–29994. doi: 10.1074/jbc.M303977200. [DOI] [PubMed] [Google Scholar]
- 18.Boutell C, Canning M, Orr A, Everett RD. Reciprocal activities between herpes simplex virus type 1 regulatory protein ICP0, a ubiquitin E3 ligase, and ubiquitin-specific protease USP7. J. Virol. 2005;79:12342–12354. doi: 10.1128/JVI.79.19.12342-12354.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Andres ME, et al. CoREST: a functional corepressor required for regulation of neural-specific gene expression. Proc. Natl. Acad. Sci. USA. 1999;96:9873–9878. doi: 10.1073/pnas.96.17.9873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Van der Horst A, et al. FOXO4 transcriptional activity is regulated by monoubiquitination and USP7/HAUSP. Nature Cell Biol. 2006;8:1064–1073. doi: 10.1038/ncb1469. [DOI] [PubMed] [Google Scholar]
- 21.Sheng Y, et al. Molecular recognition of p53 and MDM2 by USP7/HAUSP. Nature Struct. Mol. Biol. 2006;13:285–291. doi: 10.1038/nsmb1067. [DOI] [PubMed] [Google Scholar]
- 22.Saridakis V, et al. Structure of the p53 binding domain of HAUSP/USP7 bound to Epstein-Barr nuclear antigen 1 implications for EBV-mediated immortalization. Mol. Cell. 2005;18:25–36. doi: 10.1016/j.molcel.2005.02.029. [DOI] [PubMed] [Google Scholar]
- 23.Li M, Brooks CL, Kon N, Gu W. A dynamic role of HAUSP in the p53-Mdm2 pathway. Mol. Cell. 2004;13:879–886. doi: 10.1016/s1097-2765(04)00157-1. [DOI] [PubMed] [Google Scholar]
- 24.Cummins JM, et al. Tumour suppression: disruption of HAUSP gene stabilizes p53. Nature. 2004;428:6982. doi: 10.1038/nature02501. [DOI] [PubMed] [Google Scholar]
- 25.Song MS, et al. The deubiquitinylation and localization of PTEN are regulated by a HAUSP-PML network. Nature. 2008;455:813–817. doi: 10.1038/nature07290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Jorgensen HF, et al. REST selectively represses a subset of RE1-containing neuronal genes in mouse embryonic stem cells. Development. 2009;136:715–721. doi: 10.1242/dev.028548. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Chen ZF, Paquette AJ, Anderson DJ. NRSF/REST is required in vivo for repression of multiple neuronal target genes during embryogenesis. Nature Genet. 1998;20:136–142. doi: 10.1038/2431. [DOI] [PubMed] [Google Scholar]
- 28.Singh SK, Kagalwala MN, Parker-Thornburg J, Adams H, Majumder S. REST maintains self-renewal and pluripotency of embryonic stem cells. Nature. 2008;453:223–227. doi: 10.1038/nature06863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Buckley NJ, Johnson R, Sun YM, Stanton LW. Is REST a regulator of pluripotency? Nature. 2009;457:E5–E6. doi: 10.1038/nature07784. [DOI] [PubMed] [Google Scholar]
- 30.Yamada Y, Aoki H, Kunisada T, Hara A. Rest promotes the early differentiation of mouse ESCs but is not required for their maintenance. Cell Stem Cell. 2010;6:10–15. doi: 10.1016/j.stem.2009.12.003. [DOI] [PubMed] [Google Scholar]
- 31.Zhang P, et al. Nontelomeric TRF2-REST interaction modulates neuronal gene silencing and fate of tumor and stem cells. Curr. Biol. 2008;18:1489–1494. doi: 10.1016/j.cub.2008.08.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Bao S, et al. Cancer stem cells promote radioresistance of malignant glioma through preferential activation of DNA damage checkpoint and repair. Nature. 2006;444:756–760. doi: 10.1038/nature05236. [DOI] [PubMed] [Google Scholar]
- 33.Bao S, et al. Targeting cancer stem cells through L1CAM suppresses glioma growth. Cancer Res. 2008;68:6043–6048. doi: 10.1158/0008-5472.CAN-08-1079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Li Z, et al. Hypoxia-inducible factors regulate tumorigenic capacity of glioma stem cells. Cancer Cell. 2009;15:501–513. doi: 10.1016/j.ccr.2009.03.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.