Abstract
In this paper we describe the advantages of collecting infrared microspectral data in imaging mode opposed to point mode. Imaging data are processed using the PapMap algorithm, which co-adds pixel spectra that have been scrutinized for R-Mie scattering effects as well as other constraints. The signal-to-noise quality of PapMap spectra will be compared to point spectra for oral mucosa cells deposited onto low-e slides. Also the effects of software atmospheric correction will be discussed. Combined with the PapMap algorithm, data collection in imaging mode proves to be a superior method for spectral cytopathology.
Keywords: FT-IR, IR-MSP, Point Mode, Imaging Mode, Spectral Cytopathology, Oral Mucosa, Principal Component Analysis
1. Introduction
Monitoring the onset of disease by detecting premalignant and malignant cells using the Papanicolaou test [1] has greatly reduced the mortality rate of cervical cancer. Nevertheless, the process of screening Pap smears is labor intensive and has changed little since it was first described by Papanicolaou about 80 years ago [2]. The Pap test is performed by exfoliating endo- and ectocervical cells from a patient's cervix using a spatula or brush. Cervical disease often originates from the cervical transformation zone, i.e., the border between the endocervix (covered by glandular or columnar cells) and the ectocervix (covered by stratified squamous cells); therefore, cells from this zone are harvested by the exfoliation procedure. The exfoliated cells are subsequently smeared, or otherwise deposited, onto a microscope slide which is then “Pap stained” with hematoxylin/eosin (H&E) and several other counter stains, and examined microscopically.
The microscopic examination is a tedious process, and requires a cytologist or cytotechnologist to visually scrutinize all the fields of a slide to detect few (about 10 out of 10,000) aberrant cells in a specimen. Consequently, the detection of abnormal specimens depends on the proper sample collection by the physician, the level of a cytologist's experience, quality of the smear preparation, and the work load. As a result of these concerns and the resulting low accuracy of classical cytopathology [3], attempts have been made to both automate the Pap screening process, and develop other more objective alternatives. Recent developments in classical cytolopathology have focused on preparing better cell deposits by eliminating clumps of cells and other confounding materials such as mucus, erythrocytes, etc.
Other techniques focus on improving the actual diagnostic step, using automated image analysis systems which aid cytologists in the visual inspection of cells. These methods select cells that need further human inspection by eliminating the most normal cells from the cell population. However, these techniques are expensive, labor intensive, and do not aid in all desirable cell diagnoses. Another option which may improve the diagnosis of dysplasia is testing for the presence of HPV (human papillomavirus). In this state-of-the-art technique, the presence of the oncogenic HPV strains is detected via the viral DNA. Studies have shown that this test improves the accuracy of the Pap test results to over 99% [4]. However, co-testing is not cost effective and will not replace traditional methods.
Consequently, a need exists to improve diagnostic techniques, using a revolutionary, rather than an evolutionary approach. This revolutionary method relies on detecting changes in a cell’s overall biochemical composition, rather than morphology, during the progression from normal to cancerous states. At the Laboratory for Spectral Diagnoses (LSpD) at Northeastern University (Boston, MA USA) and a few other research laboratories worldwide, a novel technique for the diagnosis of exfoliated cells was developed recently [5, 6]. This method determines a snapshot of a cell’s biochemical composition, and variations therein, via a global spectral measurement using infrared microspectroscopy [7], and is referred to as Spectral Cytopathology (SCP).
Infrared microspectroscopy (IR-MSP) is a well-established technique of analytical chemistry, and was reviewed in detail recently [7–9]. Its application to medical and biochemical problems dates back to the last decade when instruments with sufficient sensitivity to rapidly acquire single cell data became commercially available. All early efforts to use IR-MSP for cellular diagnostics were carried out in a “point-and-shoot” mode, in which a cell was centered in the field of view of an infrared microspectrometer. An aperture was used to block areas of the field of view not occupied by the cell, and a spectrum of the cell was acquired within a few seconds. Since a microscope slide contains hundreds or thousands of individual cells, this process was automated using the instrument manufacturer’s software to select positions, aperture sizes and orientations of hundreds of cells visually, store these parameters, and after have the micro-spectrometer acquire data at each of the stored cell positions. Although this method was quite successful in preliminary studies [6] this process was tedious and time consuming, and may have omitted cells on the slide that were assumed to be debris, since at the time of data acquisition the slide is unstained, and the recognition of cells is difficult.
Here, we report a method in which the surface of the entire sample area is automatically scanned at a pixel size smaller than the diffraction limit, regardless of whether or not the surface is occupied by a cell, producing a large spectral dataset of the entire sample area. Subsequently, the spectra of individual cells are reconstituted from pixel spectra of areas identified to belong to one cell. We shall refer to this approach as the “PapMap” method, which was submitted for patent protection [10].
2. Experimental
2.1 Sample Preparation
Cells are exfoliated using a cytobrush, which is immediately submersed into fixative after sample collection. SurePath® solution [Burlington, NC USA] is the fixative of choice in our laboratory because this is the fixative the hospital utilizes in practice. SurePath contains less than 24% ethanol and approximately 1% each of methanol and isopropanol. This type of fixative renders proteins insoluble by dehydration, and also removes lipids. Nonetheless, substantial changes in the protein composition coupled with concomitant changes in the intensity of phosphate specific bands, which in the absence of lipid can be more readily attributed to RNA and DNA abundance, are observed for populations of oral and cervical diseased cells [11, 12]. This preservative is not ideal in all instances where the lipid composition is vital to characterize tissue, cell or disease type, for example in brain or skin tissue. However, recent investigations upon frozen and de-paraffinised tissue have displayed similar results in their ability to classify normal and diseased anatomical regions of the brain [13, 14].
Cells were shaken off the collection device by vortexing the brush and container, and the resulting cell suspension was purified by filtering with an 11 µm Nylon net filter [Millipore, Billerica, MA USA], and repeated centrifugation (2×, 600g, 15 minutes) and washing with Hank’s Balanced Salt Solution [ATCC, Manassas, VA USA] to eliminate very small cells such as red blood cells and other debris. The purified squamous cells were deposited onto reflectively coated glass slides [Kevley Technologies, Chesterland, OH USA] by cytocentrifugation using a Cytospin [Thermo, Waltham MA USA]. Sparse cell deposits (ca. 500–1000 cells in a 5 mm diameter sample spot) are routinely produced using this approach. Cells are not stained at this point, since the stains would interfere with the cellular spectra. A desirable cell density is shown in Figure 1, Panel A.
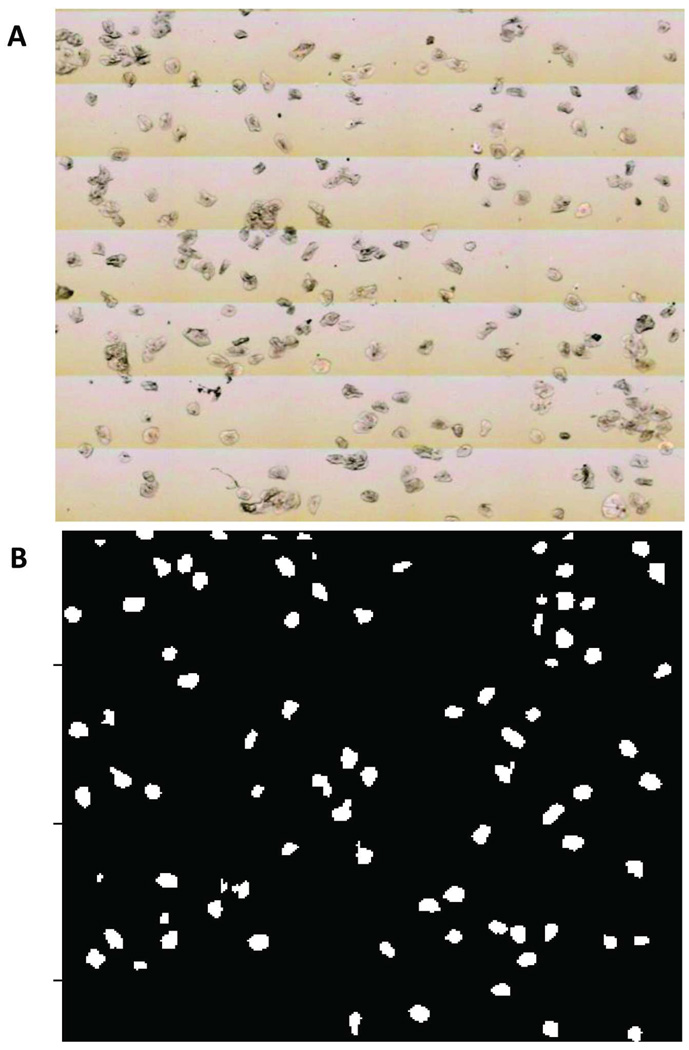
Figure 1.
(A) Visual image of a 2 mm × 2 mm area on a sample spot collected on the PE Spotlight microscope showing desirable cell density, and (B) its corresponding binary mask constructed using the PapMap algorithm.
The sample preparation method has been applied to cervical, oral and urine-borne cells, and can be used, with suitable minor modification in the purification step, for other exfoliated cells, or cells collected by any other means (lavages, thin needle aspiration, etc.).
2.2 Data Acquisition
Spectral data of the sample spot are collected by raster-scanning the slide through the focus of an infrared microscope. At the LSpD, commercial instruments, Perkin Elmer Spectrum One/Spotlight 400 imaging IR micro-spectrometers [Sheldon, CT USA], henceforth referred to as the PE 400, incorporating a 16 × 1 element photoconductive HgCdTe linear array detector, are used for data acquisition. In these instruments, the pixel size for imaging can be selected to cover 50 µm × 50 µm, 25 µm × 25 µm or 6.25 µm × 6.25 µm on edge areas. Although typical squamous cells measure between 25 and 60 µm in size, we found that only the pixel size of 6.25 µm produces datasets of sufficient spectral detail for data analysis. In addition, the PE 400 also incorporates a 100 µm single element HgCdTe detector for single spectrum acquisition. The instrument optical bench, microscope and microscope stage chamber are continuously purged with dry air (−40 °C dew point) to reduce atmospheric water vapor contribution to spectra.
The actual spatial resolution of the PE 400 infrared micro-spectrometer was established using military targets, and was found to be ca. 12 µm at 1000 cm−1. Thus, imaging the sample area using a 6.25 µm pixel size represents an approximate factor 2 of oversampling. Despite an increase in acquisition time, this methodology is vastly preferable to under-sampled (e.g. 25 µm pixel size) datasets, since spectra at this resolution often display very large scattering contributions [15–17], as shown in Figure 3 below.
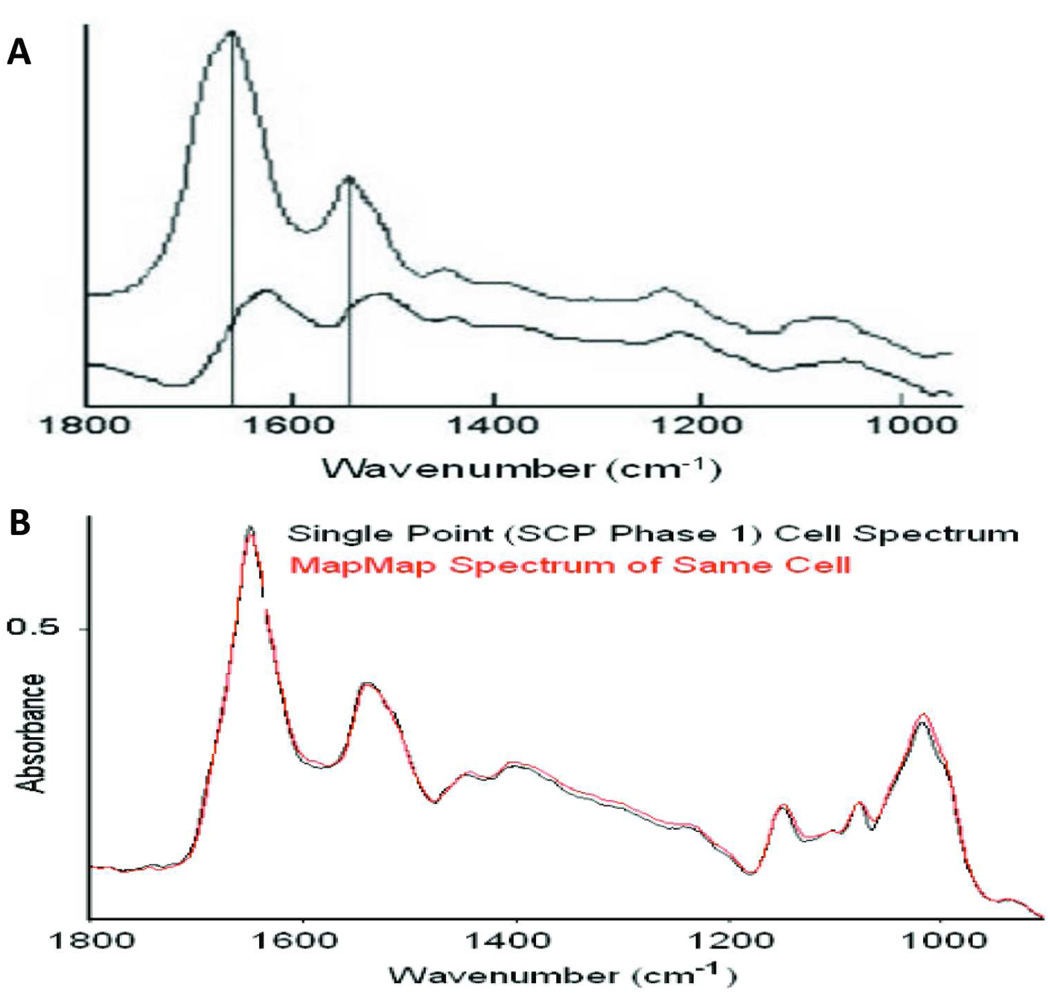
Figure 3.
(A) Distortion of the spectral band shapes and positions in the fingerprint region (1800–900 cm−1), as the result of R-Mie scattering effects. The most significant changes are in the intensity ratio of the amide I to amide II bands, but there are distortive changes throughout the spectral region. (B) R-Mie effects are minimized in imaging mode using PapMap since the algorithm scrutinizes which pixels are co-added by subjecting them to various constraints. The spectrum of the same cell collected in imaging mode and processed using PapMap is almost identical to the spectrum collected in single point mode.
A 4 mm × 4 mm area within the 5 mm diameter sample spot is scanned at 6.25 µm pixel size, resulting in a dataset of 409,600 individual spectra, and occupying about 2.5 GByte. Spectra were acquired in transflection mode at 4 cm−1 spectral resolution, co-adding 2 interferograms per pixel, between 4000 and 700 cm−1. Fourier transformation was performed using Norton-Beer apodization and one level zero-filling. The resulting single beam spectra were ratioed against a background collected for each detector element (720 interferograms), to produce spectral vectors with 1651 data points. At 8 cm−1 resolution and single interferogram collection, the maximum data acquisition rate of the PE 400 is ca. 170 pixels/s; thus, for 4 cm−1 resolution and co-addition of 2 interferograms, the acquisition rate is ca. 40 pixel/s. The background collection, image acquisition, Fourier transformation and storage of 409,600 spectra therefore requires about 4 hours.
2.3 PapMap Data Manipulation
For data analysis, the 2.5 GByte datasets in native PE 400 format (.fsm files) were transferred to workstations incorporating 8 GByte RAM, operating under Windows XP64.
The PapMap algorithm subsequently reconstructs the spectrum of each individual cell in the dataset. Briefly, pixel spectra are first converted from transmission to absorbance, and further processed in MATLAB [The Mathworks, Natick, MA USA] single precision (4 byte) format to reduce the size of the dataset. Next, all spectra are off-set corrected. The algorithm determines which pixel spectra belong to a cell by constructing a binary mask. This binary mask (shown in Figures 1B and 2B) is established by removing the pixels with an intensity value below a determined threshold for the amide I, typically 0.1 OD. An enlarged region of a similar image to the one displayed in Figure 1 is shown in Figure 2. Figure 2A depicts the visual image stored by the PE 400 before infrared spectral data acquisition, and Figure 2B shows the corresponding binary mask. Contiguous white regions in this image are assumed to belong to one or a few overlapping cells. The MATLAB function “bw label” is used to detect contiguous white areas, number them consecutively and determine the number of pixels in each of them. These contiguous regions will be henceforth referred to as “raw cells”.
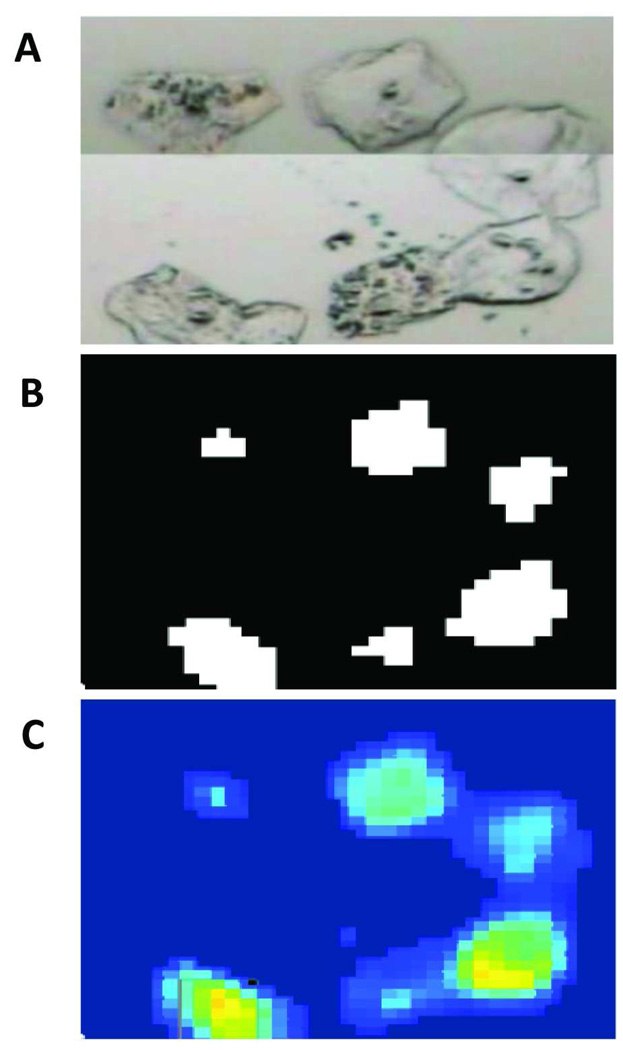
Figure 2.
Enlarged region of a small area (approximately 170 µm × 130 µm), displaying the (A) visual image collected on the PE Spotlight microscope, (B) its corresponding binary mask constructed using the PapMap algorithm, and (C) the amide I intensity in each of the contiguous “raw cell” regions where the bright yellow hues exhibit pixels with the highest amide I intensity values.
The following step refines the binary mask by removing clumps of overlapping or partially overlapping cells, and debris such as broken cells, cellular nuclei or bacteria. This is accomplished by defining lower and upper boundaries for the number of pixels associated with a cell. Since each pixel is collected from an area of 39 µm2, and a squamous cell may occupy areas between ca. 350 and 4000 µm2, the number of pixels responsible for one individual cell may range between 9 and 100. Any “raw cell” regions falling outside these ranges will be eliminated. The number of cells eliminated at this point is strictly sample dependent, but may be on average about 20% for squamous samples. The cells eliminated are predominately clumps of 2 or more squamous cells. However, spectra of cell clumps larger than 100 pixels can be examined by adjusting the number of co-added pixels. Likewise, the parameter can be adjusted for cells smaller than 9 pixels (approximately 18 µm in diameter). The parameters for cells containing between 9 and 100 pixels is sufficient for samples of normal oral mucosa cells included in this study.
Next, spectra of individual cells are calculated by averaging all pixel spectra in a given “raw cell” that are further subjected to two constraints: peak location and peak intensity. These constraints are necessary since the pixels from the edge of cells (“edge effects”) are often very weak, and therefore, exhibit a poor signal-to-noise ratio. Adding these spectra to the final spectra of the cell may degrade the signal quality of the final cellular spectrum. The second constraint is due to an optical effect that has recently attracted much interest. At the edges of spherical objects, an effect referred to as “resonance Mie (R-Mie) scattering” is frequently observed. This effect, first described by Gardner and his group [15–17], effectively adds a distorted line shape to the absorptive component, which produces an effect shown in Figure 3. The result of this R-Mie effect causes significant changes throughout the spectral region, including a shift of the amide I peak towards lower wavenumbers, and a distortion of the amide I and amide II intensity ratios. We decrease the influence of this effect in the cellular spectrum by controlling which pixels are included in the computation of the cell spectrum, as follows. In each of the contiguous “raw cell” regions, the pixel with the highest amide I intensity is determined. These pixels are shown in Figure 2C by the brightest (yellow) hues. We assume that these pixels are from the nucleus of the cell, which always has the highest amide I intensity. † We further assume that this pixel, generally from the interior of a cell, is relatively free of the “edge effects”. This spectrum must pass stringent conditions before it can be used as a “seed spectrum” to reconstruct the cellular spectrum: the intensity maximum must occur between 1630 and 1670 cm−1, and must be above an intensity threshold typical for nuclear spectra. We selected to use the amide I intensity value opposed to ratioing the amide I to amide II because it is a more sensitive and strict criterion. Starting from this “seed spectrum”, all spectra in the contiguous area of the “raw cell” are co-added to the seed spectrum if their amide I intensity is above a certain percentage (67%) of the amide I intensity of the seed spectrum, and if their amide I frequency is within a shift of 4 cm−1 of the amide I frequency of the seed spectrum. If the pixel within the contiguous area does not fulfill these criteria, then it will be excluded from the co-addition procedure.
Figure 3B displays a comparison between a cellular spectrum constructed by co-adding 15 (out of 36) individual pixel spectra from a single cell in the binary map shown in Figure 2C, and a cellular spectrum obtained using single point data acquisition. This figure illustrates that the co-addition process described above results in a spectrum that is nearly identical to the single point cellular spectrum. The loss of signal-to-noise ratio in any given pixel, caused by sampling at an aperture that is smaller than a cell, is recovered by the co-addition process.
After spectral data acquisition, cells on the slide are stained and cover-slipped and high resolution digital images are collected automatically for each individual cell on the substrate. This is accomplished by an imaging system constructed in-house around an Olympus BX50 microscope fitted with an automatic stage and a QImaging GO3 3MB digital color camera. Cell positions determined by the PapMap algorithm are used to find the cells and automatically capture their images. These images are linked with the calculated cellular spectra, and stored in a database for further processing.
3. Results and discussion
Exfoliated oral mucosa cells collected from laboratory volunteers were utilized to investigate any confounding differences between data collection in imagining mode versus single point mode, as well as effects caused by using an atmospheric correction (provided with the PE400 software) and varying the aperture size in single point mode. For data collected in imaging mode, the individual spectra were calculated using the PapMap algorithm. Data from 30 individual cells from one volunteer, ranging in size from 25 µm to 55 µm in diameter are presented in these results. For simplicity purposes, only cells from one volunteer are presented in this manuscript, since normal oral mucosa cells from multiple volunteers have been shown to lack spectral differentiation [6, 11].
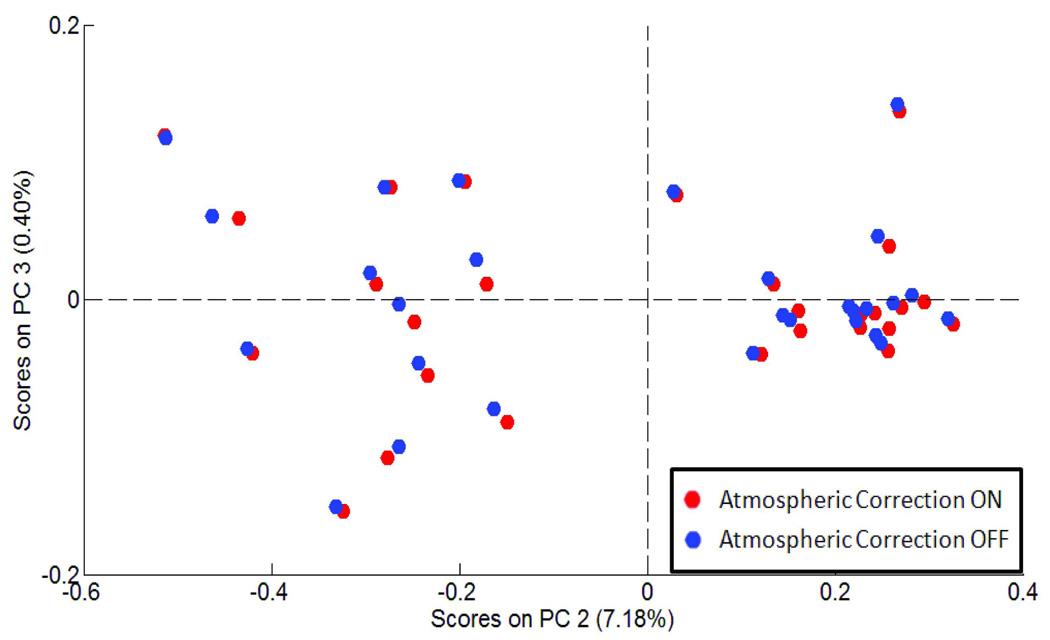
Although the instrument is purged with dry air as described above, water vapour and CO2 rot-vibrational contributions may still be present in the final cellular absorption spectra. An atmospheric correction can be applied to correct for any remaining water or CO2 contamination in the spectral data. In single point mode, this correction is performed for every scan. In imagining mode, atmospheric correction is performed after data collection is complete, and is calculated for every pixel. We first present results comparing spectral data collected with and without the software atmospheric correction. The result of this comparison are shown in Figure 4. This figure represents a PCA scores plot [18] of about 30 oral mucosa cells, collected in imaging mode and processed via PapMap as discussed above. Similar results were observed for spectral data collected in the single point mode (data not shown). The PCA scores plot shows that there are no statistical differences between the imaging data collected with and without the software atmospheric correction. In fact, the results for each individual cell appear as pairs in PCA space. From this, it can be concluded that the purge of the instrument optical bench, infrared microscope, and external microscope enclosure box is effective in eliminating atmospheric artifacts, and that the software atmospheric correction is not required.
Figure 4.
PCA scores plot of oral mucosa cells collected via imaging mode with atmospheric water correction on (red) and off (blue) where each symbol represents the spectrum of an individual cell.
Next, the effect of the aperture size in point mode will be discussed, and the spectral quality of point and imaging modes will be compared. The atmospheric correction was not applied for either point or imaging data. For single point acquisition, 128 interferograms were co-added. In imagining mode, 2 scans per pixel were co-added.
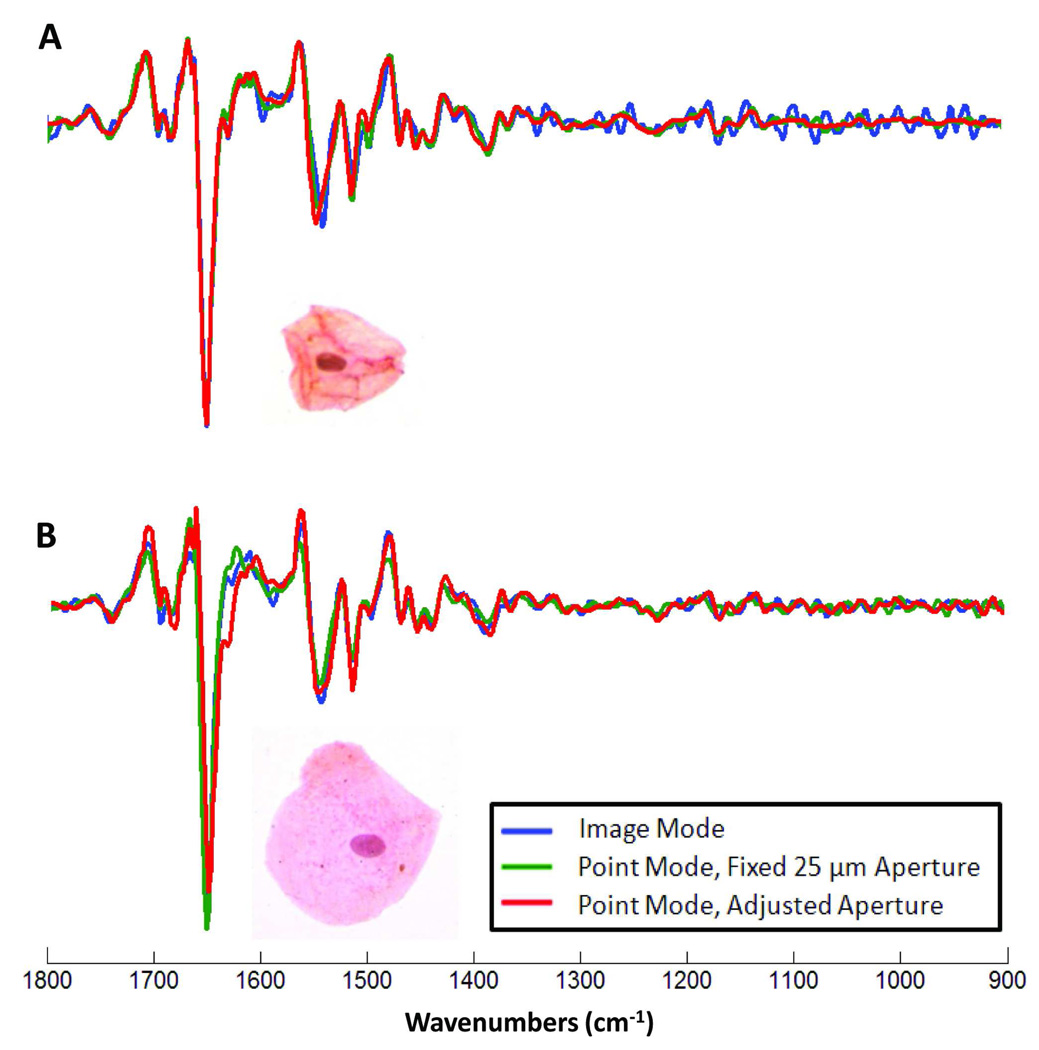
In single point mode, the size of the aperture determines the amount of light reaching the detector. This light level, in turn, affects the spectral quality. Furthermore, as discussed above, spectra from the edges of cells usually are contaminated by components with reflective line-shapes (R-Mie scattering components). In order to better visualize small spectral changes, the spectral data shown in Figure 5 were converted to second derivatives via a 9 point smooting/derivative window function [19]. In these plots, the blue trace is from the imaging datasets (PapMap), whereas the green and red traces are from single point data acquisition. The green spectrum was collected using a fixed aperture size of 25 µm, centered at the nucleus, whereas the red trace was collected using a larger aperture selected to fit the cell while avoiding the edge of the cell. The cells whose spectra are shown in Figure 5, Panels A and B, measure ca. 25 µm and 50 µm in size, respectively. For the small cell, shown in Panel A, the image spectrum has significantly more noise in the lower wavenumber region compared to the point spectra. This is due to the fact that only about 20 pixel spectra were co-added. Of all spectra, the red trace is the least noisy since it was collected at the largest aperture; however, the red and green traces are quite similar. The shape of the amide I band between 1700 and 1600 cm−1 appears nearly unchanged between the three modes of data acquisition. This indicates that the dispersion effects due to R-Mie scattering, which affect the amide I band most strongly, are negligible in all three spectra. Interestingly, the position of the amide II high frequency component at position ca. 1550 cm−1 is shifted between the three spectra, with the imaging data showing lowest wavenumber position, and the large aperture spectrum showing highest wavenumber position. Since all other band components between 1700 and 1400 cm−1 are unshifted, we do not believe that the wavenumber shift in the amide II component is due to the R-Mie effect, but due to some random sampling variance.
Figure 5.
Second derivative vector normalized spectra for the spectral range 1800–900 cm−1 where (A) represents a small cell ca. 20 pixels and (B) represents a large cell ca. 80 pixels. The blue trace is the spectrum calculated using the PapMap algorithm. The green and red traces are of spectra collected in point mode using two aperture sizes: 25 µm and an adjusted aperture to encompass the inside of the cell, respectively. High resolution images of a representative small cell and a large cell are depicted with each respective second derivative trace.
For the larger cell, shown in Panel B, the image spectrum has significantly less noise, comparable to the signal to noise (S/N) ratio in the point spectra. For this cell, about 80 pixels were co-added, resulting in an increase of the S/N ratio of a factor of ca. 2. Both the amide I and amide II peak positions remain consistent in the image and point collection modes. Some differences in band shape and peak intensity in the point spectrum using the adjusted aperture may be due to collecting data from regions of the cell, particularly at the edges, which may have been eliminated in the co-addition of pixel spectra in PapMap for the image data. For the large cell, the image spectrum more closely resembles the fixed aperture spectrum.
All in all, it can be concluded that the spectral quality is not significantly affected by the collection mode, single point versus imaging. Although the spectra are slightly more comparable for larger cells in terms of the S/N ratio, the spectral quality can be improved for small cells collected in imaging mode by co-adding more scans per pixel. Therefore, since collection in imaging mode is more reasonable in terms of experimental time and potential to automate, it has been shown to be superior to point mode data collection.
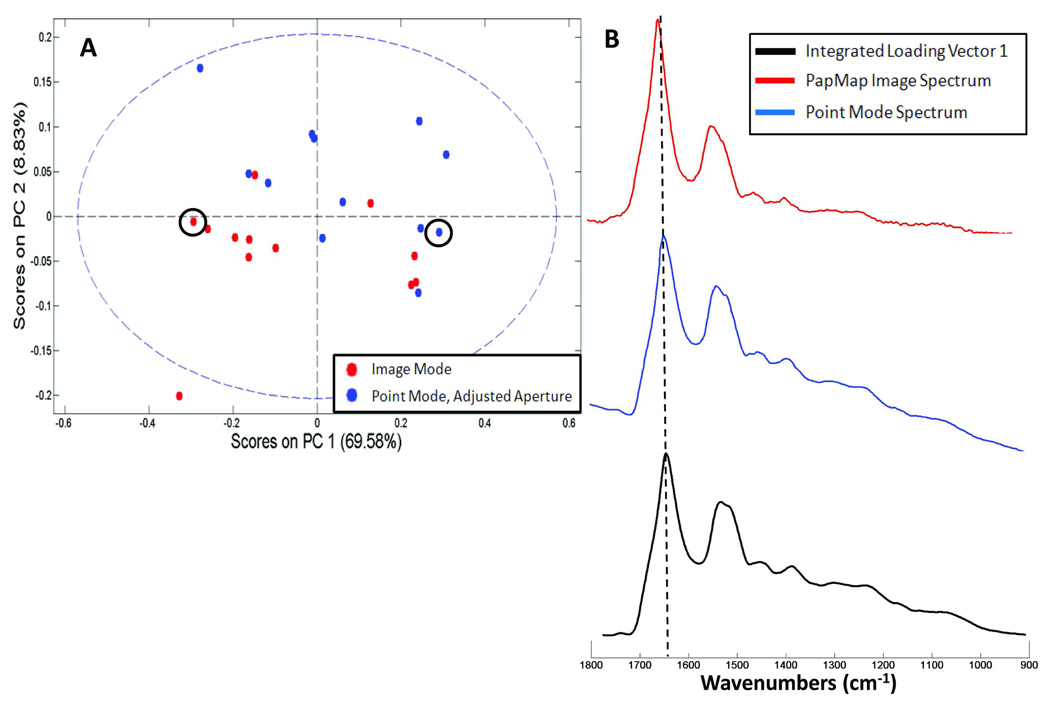
Additionally, data collected from 12 cells in both imaging and point modes were analyzed using PCA to evaluate any spectral differences between techniques. Only cells with a diameter between 35–55 µm were included in order to eliminate variations in the S/N ratio of smaller cells. Point data using the adjusted aperture were analyzed in order to evaluate the effectiveness of removing R-Mie scatter affected pixel spectra using PapMap for imaging data. The PCA scores plot in Figure 6A shows grouping by collection method across PC1. The loading vector for PC1 shown in Figure 6B, which was integrated from derivative format to an absorbance profile, describes spectral features attributable to R-Mie scattering. Both the amide I and amide II bands are shifted approximately 8 cm−1 to 1644 and 1532 cm−1, respectively. The point spectra in this analysis are mostly populated on the right hand side, or positive side of the scores plot, and suggest that these cellular spectra display larger scattering contributions. Whereas the majority of reconstructed spectra calculated by PapMap are grouped on the left, or negative side of the scores plot. However, there are still some PapMap derived spectra clustered on the postive side of the PC1 axis. These results would indicate that the PapMap method does not remove all of the pixels that display R-Mie scattering from the reconsructed spectra, but does remove the most highly scattered pixel spectra for each cell.
Figure 6.
(A) PCA scores plot of 12 oral mucosa cells from one volunteer collected via imaging mode (red) and point mode using an adjusted aperture (blue). (B) Integrated loading vector 1 (black trace) for PC1 of the scores plot shown in A. This spectrum is severely affected by R-Mie scattering: the amide I and amide II bands are located at 1642 and 1532 cm−1, respectively. The red and blue traces are from individual cells on either side of PC1 as indicated in A.
4. Conclusion
The PapMap algorithm was developed to process imaging data in order to decrease experimental time and begin to develop an automated process. In this study we have shown that data collected in imaging mode is comparable to data collected in single point mode. Although the PapMap spectra are slightly noisier for small cells, the S/N ratio can be improved by co-adding more scans per pixel. The dry air purge on the instrument, optical bench, and sample chamber is sufficient in mimimising spectral contamination from atmospheric water and CO2. By use of the PapMap methodology, spectra that desribe large scattering contributions are removed by using stringent constraints that allow cellular spectra to be constructed from pixels with little or no spectral distortion. However, despite these improvements in data acqutisiton and processing of large infrared images from cellular deposits, the reconstructed cellular spectra still desrcibe some R-Mie scattering contributions. Therefore, data sets constructed from such analyses still require a correction for these scatterting effects[15–17, 20].
Acknowledgements
Support of this research from grant CA 090346 from the National Institutes of Health is gratefully acknowledged.
Biographies

Jennifer M. Schubert received her bachelor degree in chemistry and biology from Assumption College in 2006. She is currently a senior graduate student pursuing her Ph.D. in Physical Chemistry under the direction of Dr. Max Diem at Northeastern University. Her thesis project involves applying Spectral Cytopathology to cervical samples, and developing automated screening methods for cervical and oral cytology.

Antonella I. Mazur received her bachelor degree in chemistry and biochemistry from Monmouth College and is currently a Ph.D. candidate in the Chemistry and Chemical Biology Department at Northeastern University under the advisement of Dr. Max Diem. She is currently investigating the application of Spectral Cytopathology to cells of the nasopharynx.

Benjamin Bird received his bachelor degree in chemistry from the University of Hertfordshire in 2002. He then worked under the direction of Prof. Mike Chesters and Prof. Mike George at the University of Nottingham where he received his Ph.D. in 2006. His thesis title was “FTIR imaging: A Route Toward Automated Histopathology” and documented his work that investigated the feasibility of using infrared imaging as a tool to characterize and classify both exfoliated and biopsied clinical samples of lymph nodes and the cervix. Since 2006 he has been working as a postdoctoral fellow in the research laboratory of Prof. Max Diem where he works in close collaboration with the Pathology Department of Tufts Medical Center. Currently he is involved in several projects focused toward the automation of spectral cytology and spectral histopathology.

Miloš Miljković received his bachelor degree in chemistry from the Belgrade University in 1997 before taking up position of R&D scientist in Paints and Coating Division of Duga IBL. He joined the lab of Prof. Max Diem in 2001 and defended his Ph.D. thesis in 2006 at the City University of New York. In 2006 he moved to Boston and took up position of associate research scientist at the Department of Chemistry and Chemical Biology at Northeastern University. Currently he is focused on data analysis within several projects involving automation of spectral cytology and histopathology, as well as monitoring drug uptake by cells via Raman spectroscopy.

Max Diem received his Vordiplom in Chemistry at the Universität Karlsruhe, Germany in 1970, and his Ph.D. in Physical Chemistry in 1976 at the University of Toledo, OH USA. He worked in the laboratory of Prof. Laurence Nafie at Syracuse University (Syracuse, NY USA) as a postdoc between 1976 and 1978. He then joined the faculty of the Chemistry Department of the City University of New York, Hunter College, where he was promoted to Professor of Chemistry in 1987. In 2005, he moved his research group to the Department of Chemistry and Chemical Biology at Northeastern University in Boston, MA. Prof. Diem’s research interests during the first two decades of his academic career dealt mostly with the emerging field of vibrational optical activity (infrared circular dichroism and Raman optical activity). Later he started to focus his attention on the application of vibrational spectroscopy for medical diagnosis. He is the author or co-author of about two books, ten book chapters, and over 150 papers.
Footnotes
The nuclear amide I intensities are higher than those of the cytoplasm primarily due to two reasons: the nucleus is thicker than the cytoplasm (by a factor of about 3–5) for dried cells. Secondly, the abundance of nuclear proteins is very high. Thus, pixels from the nucleus often have absorbance values of 0.2 to 0.5 OD, whereas pixels from the cytoplasm typically have absorbance values of 0.02 OD.
References
- 1.Papanicolaou GN, Traut H. The diagnostic value of vaginal smears in carcinoma of the uterus. Am J Obstet Gynecol. 1941;42:193–206. [PubMed] [Google Scholar]
- 2.Papanicolaou GN. New Cancer Diagnosis. Proceedings of the Third Race Betterment C. 1928:528. [Google Scholar]
- 3.Stoler MH, Schiffman M. Interobserver Reproducibility of Cervical Cytologic and Histologic Interpretations. JAMA. 2001;285:1500–1505. doi: 10.1001/jama.285.11.1500. [DOI] [PubMed] [Google Scholar]
- 4.Brink AATP, Zielinski GD, Steenbergen RDM, Snijders PJF, Meijer CJLM. Clinical relevance of human papillomavirus testing in cytopathology. Cytopathology. 2005;16:7–12. doi: 10.1111/j.1365-2303.2004.00227.x. [DOI] [PubMed] [Google Scholar]
- 5.Romeo MJ, Mohlenhoff B, Diem M. Infrared microspectroscopy of human cells: causes for the spectral variance of oral mucosa (buccal) cells. Vib. Spectrosc. 2006;42:9–14. doi: 10.1016/j.vibspec.2006.04.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Romeo MJ, Mohlenhoff B, Jennings M, Diem M. Infrared micro-spectroscopic studies of epithelial cells. Biochim. Biophys. Acta. 2006;1758:915–922. doi: 10.1016/j.bbamem.2006.05.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Romeo MJ, Boydston-White S, Matthaus C, Miljkovic M, Bird B, Chernenko T, Lasch P, Diem M. Infrared and Raman Microspectroscopic Studies of Individual Human Cells. In: Diem M, Griffiths PR, Chalmers JM, editors. Vibrational Spectroscopy for Medical Diagnosis. Chichester: John Wiley & Sons Ltd.; 2008. [Google Scholar]
- 8.Diem M, Romeo M, Boydston-White S, Miljkovic M, Matthaeus C. A decade of vibrational microspectroscopy of human cells and tissue (1994–2004) Analyst (Cambridge, United Kingdom) 2004;129:880–885. doi: 10.1039/b408952a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Levin IW, Bhargava R. Fourier Transform Infrared Vibrational Spectroscopic Imaging: Integrating Microscopy and Molecular Recgonition. Annu. Rev. Phys. Chem. 2005;56:429–474. doi: 10.1146/annurev.physchem.56.092503.141205. [DOI] [PubMed] [Google Scholar]
- 10.Diem M, Miljkovic M, Romeo M, Bird B, Schubert J. Method of Reconstituting Cellular Spectra from Spectral Mapping Data. USA: United States Patent and Trademark Office; 2009. [Google Scholar]
- 11.Papamarkakis K, Bird B, Schubert JM, Miljkovic M, Bedrossian K, Laver N, Diem M. Cytopathology by Optical Methods: Spectral Cytopathology of the Oral Mucosa. 2009;90:589–598. doi: 10.1038/labinvest.2010.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Schubert JM, Bird B, Papamarkakis K, Miljkovic M, Bedrossian K, Laver N, Diem M. Spectral Cytopathology of Cervical Samples: Detecting Cellular Abnormalities in Cytologically Normal Cells. Laboratory Investigation. 2010 doi: 10.1038/labinvest.2010.72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Amharref N, Beljebbar A, Dukic S, Venteo L, Schneider L, Pluot M, Vistelle R, Manfait M. Brain tissue characterisation by infrared imagining in a rat glioma model. Biochim. Biophys. Acta. 2006;1758:892–899. doi: 10.1016/j.bbamem.2006.05.003. [DOI] [PubMed] [Google Scholar]
- 14.Bambery KR, Schultke E, Wood BR, MacDonald STR, Ataelmannan K, Griebel RW, Juurlink BHJ, McNaughton D. A Fourier transform infrared microspectroscopic imaging investigation into an animal model exhibiting glioblastoma multiforme. Biochim. Biophys. Acta. 2006;1758:900–907. doi: 10.1016/j.bbamem.2006.05.004. [DOI] [PubMed] [Google Scholar]
- 15.Bassan P, Byrne HJ, Bonnier F, Lee J, Dumas P, Gardner P. Resonant Mie scattering in frared spectroscopy of biological materials - understanding the 'dispersion artefact'. Analyst. 2009;134:1586–1593. doi: 10.1039/b904808a. [DOI] [PubMed] [Google Scholar]
- 16.Bassan P, Byrne HJ, Lee J, Bonnier F, Clarke C, Dumas P, Gazi E, Brown MD, Clarke NW, Gardner P. Reflection contributions to the dispersion artefact in FTIR spectra of single biological cells. Analyst. 2009;134:1171–1175. doi: 10.1039/b821349f. [DOI] [PubMed] [Google Scholar]
- 17.Bassan P, Kohler A, Martens H, Lee J, Byrne HJ, Dumas P, Gazi E, Brown M, Clarke N, Gardner P. Resonant Mie Scattering (RMieS) correction of infrared spectra from highly scattering biological samples. Analyst. 2010 doi: 10.1039/b921056c. [DOI] [PubMed] [Google Scholar]
- 18.Adams MJ. Chemometrics in Analytical Spectroscopy. 2nd ed. Cambridge: Royal Society of Chemistry; 2004. [Google Scholar]
- 19.Savitzky A, Golay MJE. Smoothing and Differentiation of Data by Simplified Least Squares Procedures. Analytical Chemistry. 1964;36:1627–1639. [Google Scholar]
- 20.Bird B, Miljkovic M, Diem M. Two Step Resonant Mie Scattering Correction of Infrared Micro-Spectral Data: Human Lympth Node Tissue. Journal of Biophotonics. 2010 doi: 10.1002/jbio.201000024. in press. [DOI] [PubMed] [Google Scholar]