Abstract
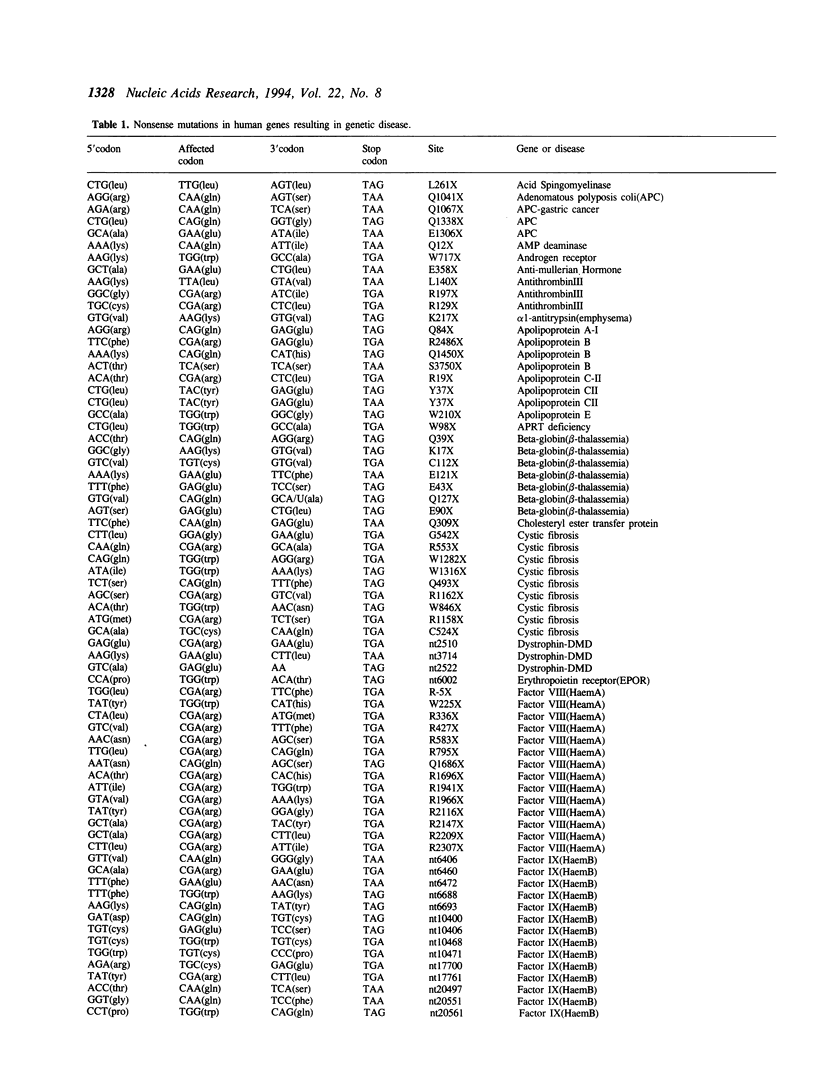
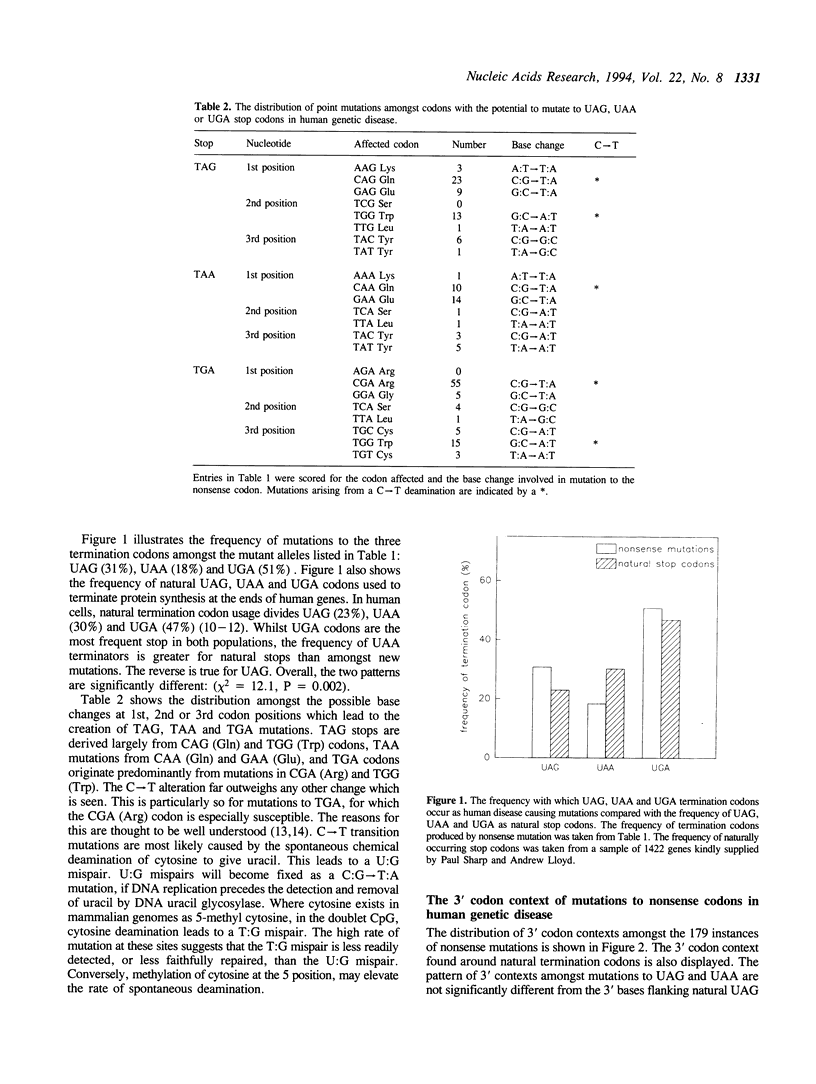
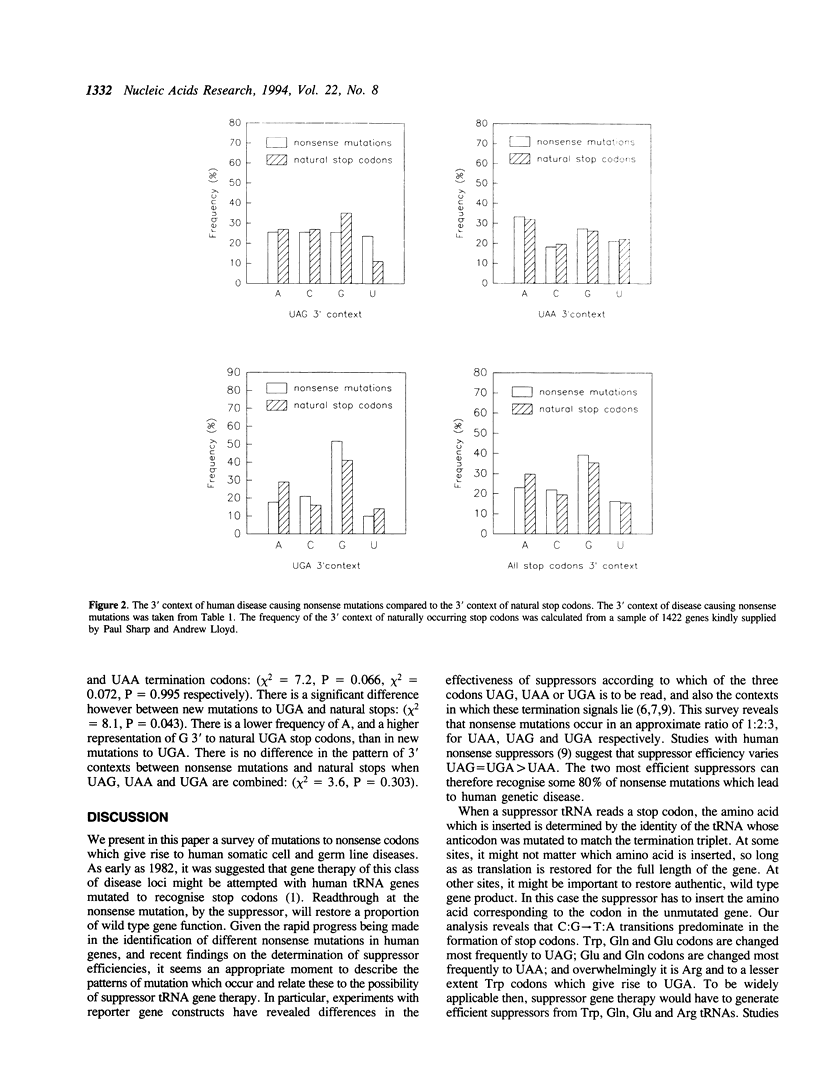
Nonsense suppressor tRNAs have been suggested as potential agents for human somatic gene therapy. Recent work from this laboratory has described significant effects of 3' codon context on the efficiency of human nonsense suppressors. A rapid increase in the number of reports of human diseases caused by nonsense codons, prompted us to determine how the spectrum of mutation to either UAG, UAA or UGA codons and their respective 3' contexts, might effect the efficiency of human suppressor tRNAs employed for purposes of gene therapy. This paper presents a survey of 179 events of mutations to nonsense codons which cause human germline or somatic disease. The analysis revealed a ratio of approximately 1:2:3 for mutation to UAA, UAG and UGA respectively. This pattern is similar, but not identical, to that of naturally occurring stop codons. The 3' contexts of new mutations to stop were also analysed. Once again, the pattern was similar to the contexts surrounding natural termination signals. These results imply there will be little difference in the sensitivity of nonsense mutations and natural stop codons to suppression by nonsense suppressor tRNAs. Analysis of the codons altered by nonsense mutations suggests that efforts to design human UAG suppressor tRNAs charged with Trp, Gln, and Glu; UAA suppressors charged with Gln and Glu, and UGA suppressors which insert Arg, would be an essential step in the development of suppressor tRNAs as agents of human somatic gene therapy.
Full text
PDF




Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arkov A. L., Korolev S. V., Kisselev L. L. Termination of translation in bacteria may be modulated via specific interaction between peptide chain release factor 2 and the last peptidyl-tRNA(Ser/Phe). Nucleic Acids Res. 1993 Jun 25;21(12):2891–2897. doi: 10.1093/nar/21.12.2891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernardi G. The vertebrate genome: isochores and evolution. Mol Biol Evol. 1993 Jan;10(1):186–204. doi: 10.1093/oxfordjournals.molbev.a039994. [DOI] [PubMed] [Google Scholar]
- Bienz M., Kubli E., Kohli J., deHenau S., Huez G., Marbaix G., Grosjean H. Usage of the three termination codons in a single eukaryotic cell, the Xenopus laevis oocyte. Nucleic Acids Res. 1981 Aug 11;9(15):3835–3850. doi: 10.1093/nar/9.15.3835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown C. M., Dalphin M. E., Stockwell P. A., Tate W. P. The translational termination signal database. Nucleic Acids Res. 1993 Jul 1;21(13):3119–3123. doi: 10.1093/nar/21.13.3119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown C. M., Stockwell P. A., Trotman C. N., Tate W. P. Sequence analysis suggests that tetra-nucleotides signal the termination of protein synthesis in eukaryotes. Nucleic Acids Res. 1990 Nov 11;18(21):6339–6345. doi: 10.1093/nar/18.21.6339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown C. M., Stockwell P. A., Trotman C. N., Tate W. P. The signal for the termination of protein synthesis in procaryotes. Nucleic Acids Res. 1990 Apr 25;18(8):2079–2086. doi: 10.1093/nar/18.8.2079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Capone J. P. Modulation of the phenotypic expression of a human serine tRNA gene by 5'-flanking sequences. DNA. 1988 Sep;7(7):459–468. doi: 10.1089/dna.1.1988.7.459. [DOI] [PubMed] [Google Scholar]
- Capone J. P., Sedivy J. M., Sharp P. A., RajBhandary U. L. Introduction of UAG, UAA, and UGA nonsense mutations at a specific site in the Escherichia coli chloramphenicol acetyltransferase gene: use in measurement of amber, ochre, and opal suppression in mammalian cells. Mol Cell Biol. 1986 Sep;6(9):3059–3067. doi: 10.1128/mcb.6.9.3059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cavener D. R., Ray S. C. Eukaryotic start and stop translation sites. Nucleic Acids Res. 1991 Jun 25;19(12):3185–3192. doi: 10.1093/nar/19.12.3185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooper D. N. Human gene mutations affecting RNA processing and translation. Ann Med. 1993 Feb;25(1):11–17. doi: 10.3109/07853899309147851. [DOI] [PubMed] [Google Scholar]
- Dietz H. C., Valle D., Francomano C. A., Kendzior R. J., Jr, Pyeritz R. E., Cutting G. R. The skipping of constitutive exons in vivo induced by nonsense mutations. Science. 1993 Jan 29;259(5095):680–683. doi: 10.1126/science.8430317. [DOI] [PubMed] [Google Scholar]
- Eyre-Walker A. C. An analysis of codon usage in mammals: selection or mutation bias? J Mol Evol. 1991 Nov;33(5):442–449. doi: 10.1007/BF02103136. [DOI] [PubMed] [Google Scholar]
- Geiduschek E. P., Tocchini-Valentini G. P. Transcription by RNA polymerase III. Annu Rev Biochem. 1988;57:873–914. doi: 10.1146/annurev.bi.57.070188.004301. [DOI] [PubMed] [Google Scholar]
- Gutman G. A., Hatfield G. W. Nonrandom utilization of codon pairs in Escherichia coli. Proc Natl Acad Sci U S A. 1989 May;86(10):3699–3703. doi: 10.1073/pnas.86.10.3699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kleina L. G., Masson J. M., Normanly J., Abelson J., Miller J. H. Construction of Escherichia coli amber suppressor tRNA genes. II. Synthesis of additional tRNA genes and improvement of suppressor efficiency. J Mol Biol. 1990 Jun 20;213(4):705–717. doi: 10.1016/S0022-2836(05)80257-8. [DOI] [PubMed] [Google Scholar]
- Kohli J., Grosjean H. Usage of the three termination codons: compilation and analysis of the known eukaryotic and prokaryotic translation termination sequences. Mol Gen Genet. 1981;182(3):430–439. doi: 10.1007/BF00293932. [DOI] [PubMed] [Google Scholar]
- Li G., Rice C. M. The signal for translational readthrough of a UGA codon in Sindbis virus RNA involves a single cytidine residue immediately downstream of the termination codon. J Virol. 1993 Aug;67(8):5062–5067. doi: 10.1128/jvi.67.8.5062-5067.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin R. On the relationship between preferred termination codon contexts and nonsense suppression in human cells. Nucleic Acids Res. 1994 Jan 11;22(1):15–19. doi: 10.1093/nar/22.1.15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin R., Phillips-Jones M. K., Watson F. J., Hill L. S. Codon context effects on nonsense suppression in human cells. Biochem Soc Trans. 1993 Nov;21(4):846–851. doi: 10.1042/bst0210846. [DOI] [PubMed] [Google Scholar]
- Miller J. H., Albertini A. M. Effects of surrounding sequence on the suppression of nonsense codons. J Mol Biol. 1983 Feb 15;164(1):59–71. doi: 10.1016/0022-2836(83)90087-6. [DOI] [PubMed] [Google Scholar]
- Mottagui-Tabar S., Björnsson A., Isaksson L. A. The second to last amino acid in the nascent peptide as a codon context determinant. EMBO J. 1994 Jan 1;13(1):249–257. doi: 10.1002/j.1460-2075.1994.tb06255.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Normanly J., Kleina L. G., Masson J. M., Abelson J., Miller J. H. Construction of Escherichia coli amber suppressor tRNA genes. III. Determination of tRNA specificity. J Mol Biol. 1990 Jun 20;213(4):719–726. doi: 10.1016/S0022-2836(05)80258-X. [DOI] [PubMed] [Google Scholar]
- Phillips-Jones M. K., Watson F. J., Martin R. The 3' codon context effect on UAG suppressor tRNA is different in Escherichia coli and human cells. J Mol Biol. 1993 Sep 5;233(1):1–6. doi: 10.1006/jmbi.1993.1479. [DOI] [PubMed] [Google Scholar]
- Raftery L. A., Egan J. B., Cline S. W., Yarus M. Defined set of cloned termination suppressors: in vivo activity of isogenetic UAG, UAA, and UGA suppressor tRNAs. J Bacteriol. 1984 Jun;158(3):849–859. doi: 10.1128/jb.158.3.849-859.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp P. M., Bulmer M. Selective differences among translation termination codons. Gene. 1988;63(1):141–145. doi: 10.1016/0378-1119(88)90553-7. [DOI] [PubMed] [Google Scholar]
- Temple G. F., Dozy A. M., Roy K. L., Kan Y. W. Construction of a functional human suppressor tRNA gene: an approach to gene therapy for beta-thalassaemia. Nature. 1982 Apr 8;296(5857):537–540. doi: 10.1038/296537a0. [DOI] [PubMed] [Google Scholar]
- Yarus M. Translational efficiency of transfer RNA's: uses of an extended anticodon. Science. 1982 Nov 12;218(4573):646–652. doi: 10.1126/science.6753149. [DOI] [PubMed] [Google Scholar]
- Youssoufian H., Kazazian H. H., Jr, Phillips D. G., Aronis S., Tsiftis G., Brown V. A., Antonarakis S. E. Recurrent mutations in haemophilia A give evidence for CpG mutation hotspots. 1986 Nov 27-Dec 3Nature. 324(6095):380–382. doi: 10.1038/324380a0. [DOI] [PubMed] [Google Scholar]



