Abstract
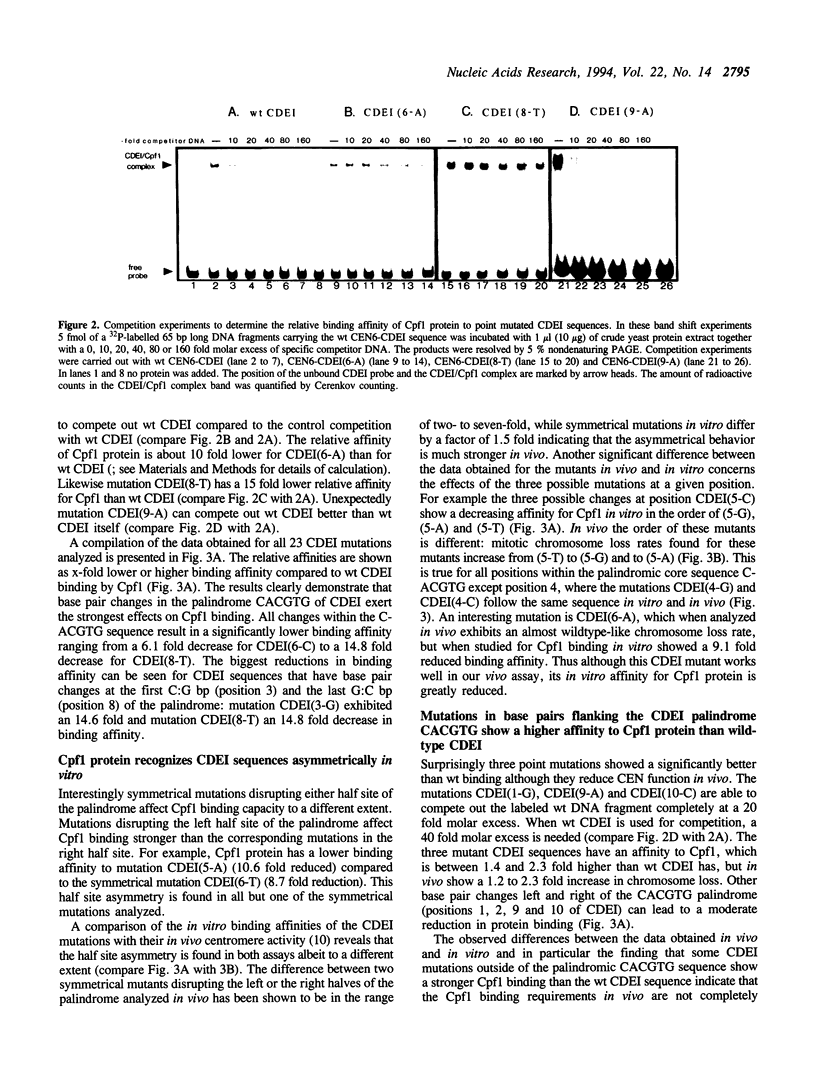
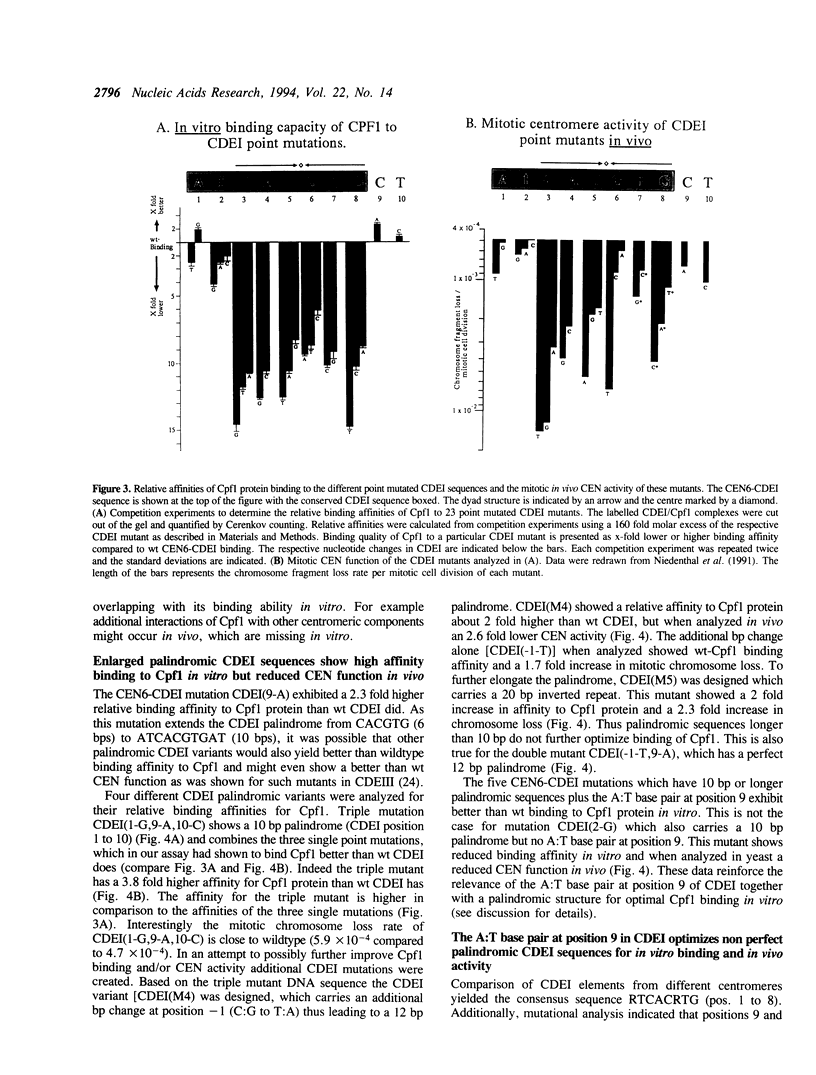
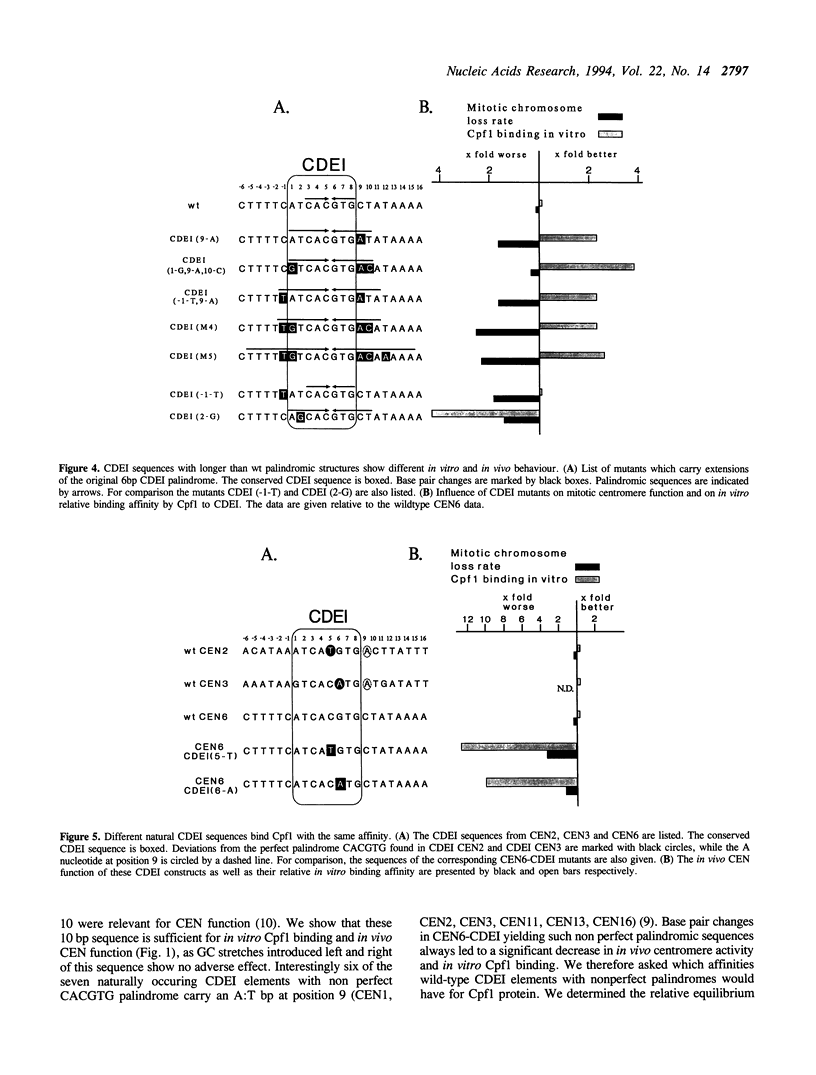
The centromere and promoter factor Cpf1 binds centromere DNA element I found in all centromere DNAs from the yeast Saccharomyces cerevisiae. We analyzed thirty different point mutations in or around CEN6-CDEI (ATCACGTG) for their relative binding affinity to Cpf1 and these data were compared with the in vivo centromere function of these mutants. We show that the minimal length of the Cpf1 binding site needed for full in vitro binding and in vivo activity is 10 base pairs long comprised of CDEI plus the two base pairs 3' of this sequence. The palindromic core sequence CACGTG is most important for in vivo CEN function and in vitro Cpf1 binding. Symmetrical mutations in either halfsite of the core sequence affect in vitro Cpf1 binding and in vivo mitotic centromere function asymmetrically albeit to a different extent. Enlarging the CDEI palindrome to 12 or 20 bps increases in vitro Cpf1 binding but results in increased chromosome loss rates suggesting a need for asymmetrical Cpf1 binding sequences. Additionally, the ability of Cpf1 protein to bind a mutant CDEI element in vitro does not parallel the ability of that mutant to confer in vivo CEN activity. Our data indicate that the in vitro binding characteristics of Cpf1 to CDEI only partly overlap with their corresponding activity within the centromere complex, thus suggesting that in the in vivo situation the CDEI/Cpf1 complex might undergo interactions with other centromere DNA/protein complexes.
Full text
PDF






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Amati B., Brooks M. W., Levy N., Littlewood T. D., Evan G. I., Land H. Oncogenic activity of the c-Myc protein requires dimerization with Max. Cell. 1993 Jan 29;72(2):233–245. doi: 10.1016/0092-8674(93)90663-b. [DOI] [PubMed] [Google Scholar]
- Baker R. E., Fitzgerald-Hayes M., O'Brien T. C. Purification of the yeast centromere binding protein CP1 and a mutational analysis of its binding site. J Biol Chem. 1989 Jun 25;264(18):10843–10850. [PubMed] [Google Scholar]
- Baker R. E., Masison D. C. Isolation of the gene encoding the Saccharomyces cerevisiae centromere-binding protein CP1. Mol Cell Biol. 1990 Jun;10(6):2458–2467. doi: 10.1128/mcb.10.6.2458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benson N., Adams C., Youderian P. Mutant lambda repressors with increased operator affinities reveal new, specific protein-DNA contacts. Genetics. 1992 Jan;130(1):17–26. doi: 10.1093/genetics/130.1.17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blackwood E. M., Eisenman R. N. Max: a helix-loop-helix zipper protein that forms a sequence-specific DNA-binding complex with Myc. Science. 1991 Mar 8;251(4998):1211–1217. doi: 10.1126/science.2006410. [DOI] [PubMed] [Google Scholar]
- Bram R. J., Kornberg R. D. Isolation of a Saccharomyces cerevisiae centromere DNA-binding protein, its human homolog, and its possible role as a transcription factor. Mol Cell Biol. 1987 Jan;7(1):403–409. doi: 10.1128/mcb.7.1.403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai M., Davis R. W. Yeast centromere binding protein CBF1, of the helix-loop-helix protein family, is required for chromosome stability and methionine prototrophy. Cell. 1990 May 4;61(3):437–446. doi: 10.1016/0092-8674(90)90525-j. [DOI] [PubMed] [Google Scholar]
- Cherest H., Thomas D., Surdin-Kerjan Y. Nucleotide sequence of the MET8 gene of Saccharomyces cerevisiae. Nucleic Acids Res. 1990 Feb 11;18(3):659–659. doi: 10.1093/nar/18.3.659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cottarel G., Shero J. H., Hieter P., Hegemann J. H. A 125-base-pair CEN6 DNA fragment is sufficient for complete meiotic and mitotic centromere functions in Saccharomyces cerevisiae. Mol Cell Biol. 1989 Aug;9(8):3342–3349. doi: 10.1128/mcb.9.8.3342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dang C. V., Dolde C., Gillison M. L., Kato G. J. Discrimination between related DNA sites by a single amino acid residue of Myc-related basic-helix-loop-helix proteins. Proc Natl Acad Sci U S A. 1992 Jan 15;89(2):599–602. doi: 10.1073/pnas.89.2.599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Winde J. H., Van Leeuwen H. C., Grivell L. A. The multifunctional regulatory proteins ABF1 and CPF1 are involved in the formation of a nuclease-hypersensitive region in the promoter of the QCR8 gene. Yeast. 1993 Aug;9(8):847–857. doi: 10.1002/yea.320090805. [DOI] [PubMed] [Google Scholar]
- Densmore L., Payne W. E., Fitzgerald-Hayes M. In vivo genomic footprint of a yeast centromere. Mol Cell Biol. 1991 Jan;11(1):154–165. doi: 10.1128/mcb.11.1.154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feldmann T., Alex R., Suckow J., Dildrop R., Kisters-Woike B., Müller-Hill B. Single exchanges of amino acids in the basic region change the specificity of N-Myc. Nucleic Acids Res. 1993 Nov 11;21(22):5050–5058. doi: 10.1093/nar/21.22.5050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher D. E., Parent L. A., Sharp P. A. High affinity DNA-binding Myc analogs: recognition by an alpha helix. Cell. 1993 Feb 12;72(3):467–476. doi: 10.1016/0092-8674(93)90122-7. [DOI] [PubMed] [Google Scholar]
- Fisher F., Goding C. R. Single amino acid substitutions alter helix-loop-helix protein specificity for bases flanking the core CANNTG motif. EMBO J. 1992 Nov;11(11):4103–4109. doi: 10.1002/j.1460-2075.1992.tb05503.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halazonetis T. D., Kandil A. N. Determination of the c-MYC DNA-binding site. Proc Natl Acad Sci U S A. 1991 Jul 15;88(14):6162–6166. doi: 10.1073/pnas.88.14.6162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hegemann J. H., Fleig U. N. The centromere of budding yeast. Bioessays. 1993 Jul;15(7):451–460. doi: 10.1002/bies.950150704. [DOI] [PubMed] [Google Scholar]
- Hegemann J. H., Shero J. H., Cottarel G., Philippsen P., Hieter P. Mutational analysis of centromere DNA from chromosome VI of Saccharomyces cerevisiae. Mol Cell Biol. 1988 Jun;8(6):2523–2535. doi: 10.1128/mcb.8.6.2523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jehn B., Niedenthal R., Hegemann J. H. In vivo analysis of the Saccharomyces cerevisiae centromere CDEIII sequence: requirements for mitotic chromosome segregation. Mol Cell Biol. 1991 Oct;11(10):5212–5221. doi: 10.1128/mcb.11.10.5212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang W. D., Philippsen P. Purification of a protein binding to the CDEI subregion of Saccharomyces cerevisiae centromere DNA. Mol Cell Biol. 1989 Dec;9(12):5585–5593. doi: 10.1128/mcb.9.12.5585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones N. Transcriptional regulation by dimerization: two sides to an incestuous relationship. Cell. 1990 Apr 6;61(1):9–11. doi: 10.1016/0092-8674(90)90207-u. [DOI] [PubMed] [Google Scholar]
- Keleher C. A., Goutte C., Johnson A. D. The yeast cell-type-specific repressor alpha 2 acts cooperatively with a non-cell-type-specific protein. Cell. 1988 Jun 17;53(6):927–936. doi: 10.1016/s0092-8674(88)90449-7. [DOI] [PubMed] [Google Scholar]
- Korch C., Mountain H. A., Byström A. S. Cloning, nucleotide sequence, and regulation of MET14, the gene encoding the APS kinase of Saccharomyces cerevisiae. Mol Gen Genet. 1991 Sep;229(1):96–108. doi: 10.1007/BF00264218. [DOI] [PubMed] [Google Scholar]
- Lechner J., Carbon J. A 240 kd multisubunit protein complex, CBF3, is a major component of the budding yeast centromere. Cell. 1991 Feb 22;64(4):717–725. doi: 10.1016/0092-8674(91)90501-o. [DOI] [PubMed] [Google Scholar]
- Lehming N., Sartorius J., Kisters-Woike B., von Wilcken-Bergmann B., Müller-Hill B. Mutant lac repressors with new specificities hint at rules for protein--DNA recognition. EMBO J. 1990 Mar;9(3):615–621. doi: 10.1002/j.1460-2075.1990.tb08153.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Masison D. C., O'Connell K. F., Baker R. E. Mutational analysis of the Saccharomyces cerevisiae general regulatory factor CP1. Nucleic Acids Res. 1993 Aug 25;21(17):4133–4141. doi: 10.1093/nar/21.17.4133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKenzie E. A., Kent N. A., Dowell S. J., Moreno F., Bird L. E., Mellor J. The centromere and promoter factor, 1, CPF1, of Saccharomyces cerevisiae modulates gene activity through a family of factors including SPT21, RPD1 (SIN3), RPD3 and CCR4. Mol Gen Genet. 1993 Sep;240(3):374–386. doi: 10.1007/BF00280389. [DOI] [PubMed] [Google Scholar]
- Mellor J., Jiang W., Funk M., Rathjen J., Barnes C. A., Hinz T., Hegemann J. H., Philippsen P. CPF1, a yeast protein which functions in centromeres and promoters. EMBO J. 1990 Dec;9(12):4017–4026. doi: 10.1002/j.1460-2075.1990.tb07623.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mellor J., Rathjen J., Jiang W., Barnes C. A., Dowell S. J. DNA binding of CPF1 is required for optimal centromere function but not for maintaining methionine prototrophy in yeast. Nucleic Acids Res. 1991 Jun 11;19(11):2961–2969. doi: 10.1093/nar/19.11.2961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murre C., McCaw P. S., Baltimore D. A new DNA binding and dimerization motif in immunoglobulin enhancer binding, daughterless, MyoD, and myc proteins. Cell. 1989 Mar 10;56(5):777–783. doi: 10.1016/0092-8674(89)90682-x. [DOI] [PubMed] [Google Scholar]
- Nelson H. C., Sauer R. T. Lambda repressor mutations that increase the affinity and specificity of operator binding. Cell. 1985 Sep;42(2):549–558. doi: 10.1016/0092-8674(85)90112-6. [DOI] [PubMed] [Google Scholar]
- Niedenthal R. K., Sen-Gupta M., Wilmen A., Hegemann J. H. Cpf1 protein induced bending of yeast centromere DNA element I. Nucleic Acids Res. 1993 Oct 11;21(20):4726–4733. doi: 10.1093/nar/21.20.4726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niedenthal R., Stoll R., Hegemann J. H. In vivo characterization of the Saccharomyces cerevisiae centromere DNA element I, a binding site for the helix-loop-helix protein CPF1. Mol Cell Biol. 1991 Jul;11(7):3545–3553. doi: 10.1128/mcb.11.7.3545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ogawa N., Oshima Y. Functional domains of a positive regulatory protein, PHO4, for transcriptional control of the phosphatase regulon in Saccharomyces cerevisiae. Mol Cell Biol. 1990 May;10(5):2224–2236. doi: 10.1128/mcb.10.5.2224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sasmor H. M., Betz J. L. Symmetric lac operator derivatives: effects of half-operator sequence and spacing on repressor affinity. Gene. 1990 Apr 30;89(1):1–6. doi: 10.1016/0378-1119(90)90198-z. [DOI] [PubMed] [Google Scholar]
- Schüller H. J., Hahn A., Tröster F., Schütz A., Schweizer E. Coordinate genetic control of yeast fatty acid synthase genes FAS1 and FAS2 by an upstream activation site common to genes involved in membrane lipid biosynthesis. EMBO J. 1992 Jan;11(1):107–114. doi: 10.1002/j.1460-2075.1992.tb05033.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas D., Barbey R., Surdin-Kerjan Y. Gene-enzyme relationship in the sulfate assimilation pathway of Saccharomyces cerevisiae. Study of the 3'-phosphoadenylylsulfate reductase structural gene. J Biol Chem. 1990 Sep 15;265(26):15518–15524. [PubMed] [Google Scholar]
- Thomas D., Cherest H., Surdin-Kerjan Y. Elements involved in S-adenosylmethionine-mediated regulation of the Saccharomyces cerevisiae MET25 gene. Mol Cell Biol. 1989 Aug;9(8):3292–3298. doi: 10.1128/mcb.9.8.3292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas D., Jacquemin I., Surdin-Kerjan Y. MET4, a leucine zipper protein, and centromere-binding factor 1 are both required for transcriptional activation of sulfur metabolism in Saccharomyces cerevisiae. Mol Cell Biol. 1992 Apr;12(4):1719–1727. doi: 10.1128/mcb.12.4.1719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wechsler D. S., Dang C. V. Opposite orientations of DNA bending by c-Myc and Max. Proc Natl Acad Sci U S A. 1992 Aug 15;89(16):7635–7639. doi: 10.1073/pnas.89.16.7635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Vliet P. C., Verrijzer C. P. Bending of DNA by transcription factors. Bioessays. 1993 Jan;15(1):25–32. doi: 10.1002/bies.950150105. [DOI] [PubMed] [Google Scholar]




