Abstract
Activation of phosphatidylinositol 3-kinase (PI3K)/Akt signaling is associated with growth and progression of colorectal cancer (CRC). We have previously shown that the mammalian target of rapamycin (mTOR) kinase, a downstream effector of PI3K/Akt signaling, regulates tumorigenesis of CRC. However, the contribution of mTOR and its interaction partners towards regulating CRC progression and metastasis remains poorly understood. We found that increased expression of mTOR, Raptor and Rictor mRNA was noted with advanced stages of CRC suggesting that mTOR signaling may be associated with CRC progression and metastasis. mTOR, Raptor and Rictor protein levels were also significantly elevated in primary CRCs (stage IV) and their matched distant metastasis compared to normal colon. Inhibition of mTOR signaling, using rapamycin or stable inhibition of mTORC1 (Raptor) and mTORC2 (Rictor), attenuated migration and invasion of CRCs. Furthermore, knockdown of mTORC1 and mTORC2 induced a mesenchymal-epithelial transition and enhanced chemosensitivity of CRCs to oxaliplatin. We observed increased cell-cell contact as well as decreased actin cytoskeletal remodeling concomitant with decreased activation of the small GTPases, RhoA and Rac1, upon inhibition of both mTORC1 and mTORC2. Finally, establishment of CRC metastasis in vivo was completely abolished with targeted inhibition of mTORC1 and mTORC2 irrespective of the site of colonization. Our findings support a role for elevated mTORC1 and mTORC2 activity in regulating EMT, motility and metastasis of CRCs via RhoA and Rac1 signaling. These findings provide the rationale for including mTOR kinase inhibitors, which inhibit both mTORC1 and mTORC2, as part of the therapeutic regimen for CRC patients.
Keywords: mTOR, Rictor, Metastasis, EMT, Colorectal cancer
INTRODUCTION
Colorectal cancer (CRC) is the second leading cause of cancer death in the United States (1). The prognosis for advanced CRCs remains dismal, mainly due to the propensity for metastatic progression and resistance to chemotherapy (2). The metastatic cascade entails an orderly sequence of steps enabling tumor cells to detach from the primary tumor, migrate and invade into surrounding tissue, intravasate into systemic circulation, extravasate to distant organs and colonize at secondary sites (3). CRCs predominantly spread along the mesenteric circulation to the liver and less frequently to the lungs and bone.
It is believed that the initial step, acquisition of migratory and invasive capability, is the rate-limiting step in this cascade (4). Epithelial-mesenchymal transition (EMT) is proposed to be a crucial mechanism regulating the initial steps in metastatic progression (3). EMT is a molecular program whereby epithelial cells undergo reprogramming from a polarized, differentiated phenotype with numerous cell-cell junctions to acquire a mesenchymal phenotype including lack of polarization, decreased cell-cell junctions, increased motility and chemotherapeutic resistance (3). The plasticity of this event is highlighted by the fact that EMT is reversible – the reverse process is termed mesenchymal-epithelial transition (MET) (3). Another important regulatory mechanism in the initial steps of the metastatic cascade involves activation of small GTPases, including RhoA and Rac1, which play a crucial role in actin cytoskeletal rearrangement and cell migration (5). Specifically, RhoA induces formation of actin stress fibers and cell-cell adhesions, while Rac1 stimulates formation of lamellipodia.
Activation of PI3K/Akt signaling through activating mutations in PIK3CA (encoding the p110α catalytic subunit of PI3K) or loss of PTEN (encoding a lipid and protein phosphatase) is associated with the growth and progression of CRC (6, 7). We have previously demonstrated that PI3K and Akt2 can regulate the metastasis of CRCs (8, 9). However, identification of downstream proteins that are directly involved in regulating CRC metastasis would allow development of targeted therapies with fewer toxicities.
mTOR is a downstream serine/threonine kinase, which exists in two complexes: mTORC1 (containing mTOR, Raptor etc.) and mTORC2 (containing mTOR, Rictor etc.) (10). The bacterially derived drug, rapamycin, allosterically inhibits mTOR activity (10). mTORC1 is partially sensitive to rapamycin treatment, while mTORC2 is believed to be rapamycin-insensitive (10, 11). We have previously demonstrated that mTORC1 and mTORC2 components, Raptor and Rictor, respectively, are overexpressed in CRCs and play an important role in CRC tumorigenesis (12). In this study, we investigated whether these proteins regulate motility, EMT and metastasis of CRCs.
MATERIALS AND METHODS
Reagents, Cell Lines and Lentiviral Transduction
The human colon cancer cell lines HCT116, SW480 and KM20 were utilized as described previously (12). Identity of all cells was authenticated at the Johns Hopkins Genetic Resources Core Facility in October 2010 with short tandem repeat analysis using the Identifiler kit from Applied Biosystems (Foster City, CA). HCT116 and SW480 cells were obtained from American Type Culture Collection (Manassas, VA); KM20 cells were kindly provided by Dr. Isaiah J. Fidler (MD Anderson Cancer Center, Houston, TX). As described previously, stable knockdown HCT116, SW480 and KM20 cells were generated using shRNAs directed against human Raptor and Rictor genes constructed in pLKO.1-puro vector obtained from Addgene (Cambridge, MA) (12). A plasmid carrying non-targeting control (NTC) sequence was used to create control cells. Rapamycin and oxaliplatin were obtained from Sigma-Aldrich (St. Louis, MO), TGFβ was obtained from PeproTech (Rocky Hill, NJ), Y27632 was obtained from Cayman Chemicals (Ann Arbor, MI) and NSC23766 was obtained from Tocris Biosciences (Ellisville, MO).
Immunohistochemistry
Paraffin-embedded tissue array sections of normal colon, primary CRC and matched liver metastasis were obtained from Accurate Chemical & Scientific Corporation (Westbury, NY). Each array category consisted of tissue derived from 18 patients: 18 normal cores, 36 primary tumor cores and 36 liver metastasis cores. Staining was performed as described previously with antibodies against mTOR (1:100), Raptor (1:100) and Rictor (1:100) obtained from Bethyl Laboratory (Montgomery, TX) (12). Scoring was performed blindly by a pathologist according to a semi-quantitative seven-tier system, as described previously (12).
Quantitative RT–PCR
qRT-PCR analysis was performed as described previously (9). A panel of cDNAs derived from total RNA covering four disease stages and normal tissues from 48 patients was purchased from OriGene Technologies, Inc. (Rockville, MD). Data from the array was normalized to 18S rRNA.
Wound healing assay
A wound healing assay was used to compare the migratory ability of HCT116 and SW480 cells as described previously (13). All experiments were performed in triplicate.
Transwell Migration assay
A Boyden chamber migration assay with collagen-coated transwells was performed with HCT116, KM20 and SW480 cells over 5 h, 8 h and 18 h respectively, as described previously (14). 10% FBS was used as the chemoattractant. Cells were counted in 4 different fields with an inverted microscope. All experiments were performed in triplicate.
Transwell Invasion assay
A modified Boyden chamber invasion assay with matrigel-coated transwell chambers was performed with HCT116, KM20 and SW480 cells over 48 h as described previously (13). 10% FBS was used as the chemoattractant. Cells were counted in 4 different fields with an inverted microscope. All experiments were performed in triplicate.
Apoptosis assay
Equal numbers of cells were serum starved overnight and then treated with 10 µM oxaliplatin or DMSO (control) for 24 h in serum starved conditions. Apoptosis was quantitated using the Cell Death Detection ELISAplus (Roche, Indianapolis, IN) as detailed in the manufacturer’s instructions.
Western Blotting and Antibodies
Western blotting was performed as described previously (12). The following antibodies were from Cell Signaling (Danvers, MA): pAktSer473, total Akt, Snail, Twist, E-Cadherin, Vimentin and β-Actin. The following antibodies were from Bethyl Laboratory (Montgomery, TX): mTOR, Raptor and Rictor. Smooth muscle actin antibody was obtained from AbCam (Cambridge, MA).
Gelatin Zymography
Equal numbers of cells were seeded and treated with 5 ng/mL TGFβ for 12 h in serum free conditions. Supernatants were collected, normalized for total protein concentration, mixed with sample buffer (Invitrogen) and analyzed by electrophoresis with a 10% zymogram gel (Invitrogen) for 90 min. The gel was developed according to manufacturer instructions and stained with Coomassie Blue (Invitrogen).
Immunofluorescence
Immunofluorescence staining was performed as described previously (13). Actin, Vimentin and E-Cadherin antibodies were obtained from BD Biosciences (San Jose, CA), while Smooth muscle actin and Fibronectin antibodies were obtained from Abcam (Cambridge, MA). Images were acquired using either Nikon TE2000 inverted microscope or Nikon Total Internal Reflection Fluorescence microscope (200× objective) and NIS Elements AR3.10 software.
RhoA / Rac1 Activity Assays
RhoA and Rac1 activity was assessed using GST-tagged Rho-binding domain of Rhotekin (TRBD) and GST-tagged p21 binding domain of PAK1 (GST-PBD) pull-down assays, respectively, as described previously (14, 15). Briefly, cells were grown to ~70% confluency in regular growth medium, serum starved overnight and stimulated with 10% FBS for 5 min. For rapamycin treatment, cells were pre-incubated in 50 nM rapamycin for 24 h. RhoA monoclonal antibody was from Santa Cruz Biotechnology (Santa Cruz, CA) and Rac1 monoclonal antibody was from Millipore (Billerica, MA).
In Vivo Experimental Metastasis
1.5 × 106 KM20 sh NTC, sh Raptor or sh Rictor CRC cells were collected in 200 µL of PBS and inoculated intravenously into tail veins of six-week old male athymic Crl:NU-Foxn1nu nude mice (Charles River Labs, Wilmington, MA) (n=5 per group). All procedures were performed in an animal facility using protocols approved by the Institutional Animal Care and Use Committee. Six weeks after inoculation, mice were sacrificed by cervical dislocation and metastatic tissues were examined in paraffin-embedded sections stained with either hematoxylin and eosin or Cytokeratin 7 and Cytokeratin 20 antibodies from Abcam (Cambridge, MA).
Statistical Analysis
Analysis of variance was used to compare qRT-PCR and immunoreactivity scores across CRC stages. Contrasts were generated from the model to perform specific hypothesis tests, such as pair-wise comparisons between cancer stages and linear trend tests across increasing stage. Paired t-test or nonparametric analog were employed for comparisons of immunoreactivity scores between matched normal, primary and metastatic tumor samples. General linear mixed model was used to analyze migration, invasion and apoptosis data. Cell line, treatment, dose and location (lower, middle, upper) were included as independent variables in the model and contrasts for pair-wise comparisons between treatment versus control as well as linear trend test for increasing treatment doses were performed. Comparisons between control versus knockdown Raptor and Rictor groups of proportion of mice exhibiting metastases were performed using Fisher’s exact test. Tests for normality were performed to assess validity of parametric models.
RESULTS
Expression of mTORC1 and mTORC2 components is elevated in CRC
To delineate the extent of mTOR pathway alterations in CRC, we analyzed mTOR, Raptor and Rictor expression by quantitative RT-PCR in cDNA from 48 patients representing all stages of disease and normal colon. Comparison of mTOR, Raptor and Rictor expression profiles showed elevated mRNA expression of all three genes in stage I–IV CRC compared to normal colon (Fig. 1A). Interestingly, increasing levels of mRNA for all genes were observed with increasing stage of disease as confirmed by a test for linear trend (p<0.0001).
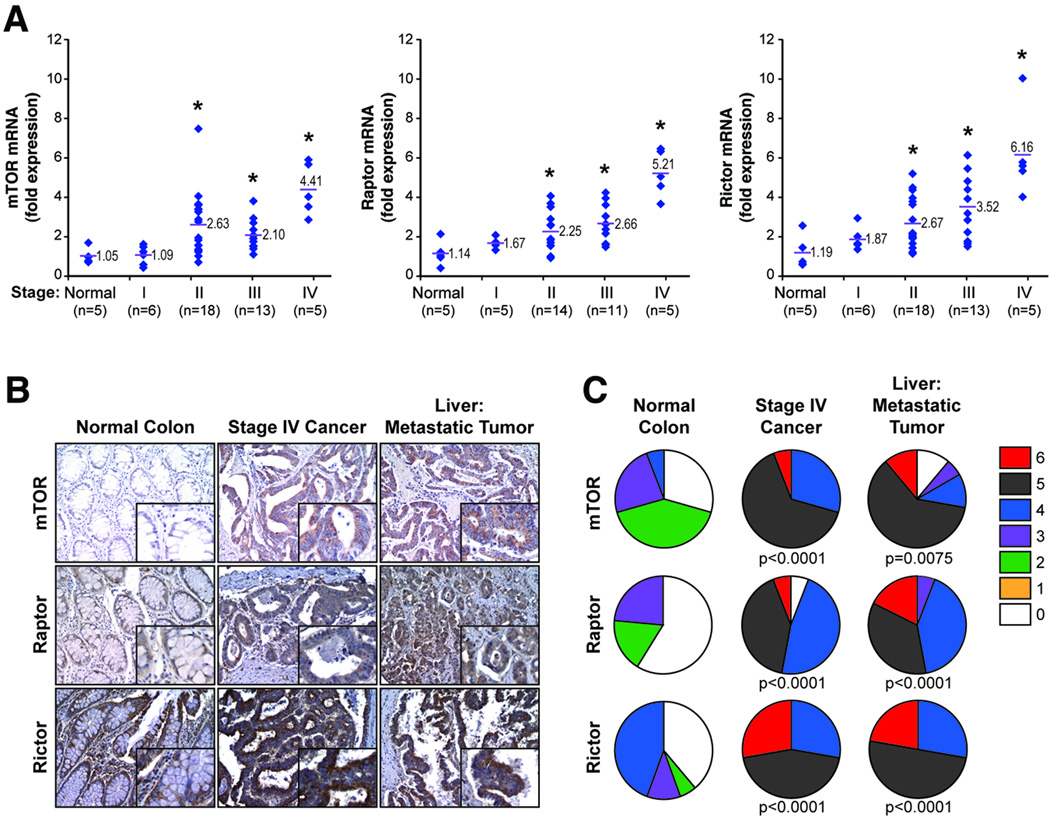
Figure 1. Expression of mTOR, Raptor and Rictor mRNA and protein levels in CRC patient specimens.
A. Relative levels of mTOR, Raptor and Rictor mRNA expression in normal colon or stage I–IV CRC derived from 48 patients. Red bars represent mean of each group. (* p<0.05 versus normal tissue) B. Expression of mTOR, Raptor and Rictor proteins in normal colon, primary CRC and matched metastatic liver tumors from stage IV patients (100×; n = 18 cases; 72 tumor cores, 18 non-neoplastic cores) C. Distribution of immunoreactivity scores in normal colon, primary CRC and matched liver metastases.
To further determine specific expression changes associated with CRC progression, we analyzed tissues from 18 stage IV patients for mTOR, Raptor and Rictor expression in normal colon, primary tumor and matched liver metastasis. Each sample was assigned an immunoreactivity score ranging from 0–6. Representative samples for each protein are shown (Fig. 1B) along with analysis (Fig. 1C). Normal colon demonstrated either negative or focal mild cytoplasmic staining for mTOR, Raptor, and Rictor. Meanwhile, primary tumor and matched liver metastasis exhibited diffuse cytoplasmic staining for all three proteins. Paired comparisons of immunoreactivity scores between normal vs. primary tumors and normal vs. metastatic tumors were significant (p<0.01) for all three proteins. No significant differences were observed between primary tumors vs. liver metastases. Taken together, these findings demonstrate that mTOR, Raptor and Rictor are overexpressed at both mRNA and protein levels in primary CRCs and this increased expression is maintained in the associated distant metastases.
mTORC1 and mTORC2 regulate CRC migration and invasion
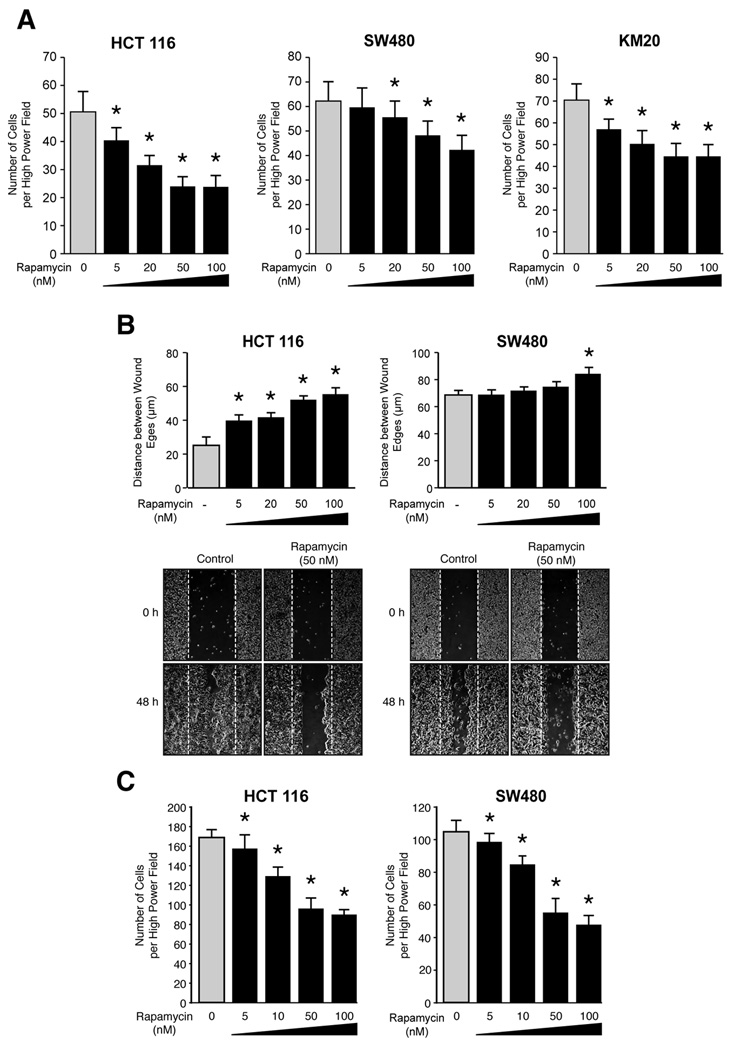
Migration and invasion are critical steps in initial progression of cancer that facilitates metastasis. We utilized HCT116, SW480 and KM20 cells to determine the effects of rapamycin upon CRC migration and invasion. First, cells were treated with increasing doses of rapamycin and migration was assessed using the short-term transwell migration assay. Treatment with rapamcyin significantly decreased migration of HCT116, SW480 and KM20 cells in a dose dependent manner (Fig. 2A). We further confirmed these findings using a wound healing assay. Consistent with results obtained from transwell migration assay, rapamycin significantly decreased migration of HCT116 and SW480 cells (Fig. 2B). Finally, the effect of rapamycin upon invasion was assessed using a modified Boyden chamber assay. Rapamycin potently decreased invasion of both HCT116 and SW480 CRC cells in a dose-dependent manner (Fig. 2C). In summary, pharmacological blockade of mTOR signaling using rapamycin significantly inhibits migration and invasion of CRC cells in a dose-dependent manner.
Figure 2. Rapamycin attenuates migration and invasion of CRCs.
A. Transwell migration assay performed with HCT116, SW480 and KM20. Cells in the control group were treated with DMSO (* p<0.05 vs. control). B. Wound healing assay performed with HCT116 and SW480 cells over 48 h. Cells in the control group were treated with DMSO (* p<0.05 vs. control). C. Transwell invasion assay was performed with HCT116 and SW480 cells over 48 h. Cells in the control group were treated with DMSO (* p<0.05 vs. control).
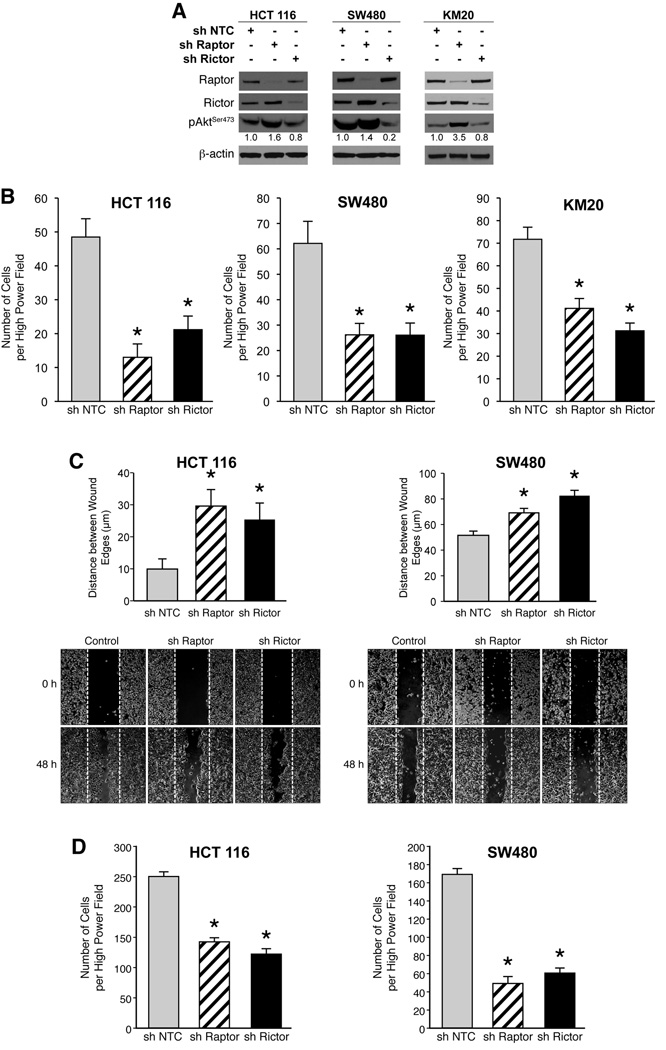
In order to decipher the contribution of mTORC1 and mTORC2 in regulating migration and invasion, we selectively silenced Raptor and Rictor, essential components of mTORC1 and mTORC2, respectively. HCT116, SW480 and KM20 cells with stable shRNA-mediated knockdown of Raptor or Rictor were generated and demonstrated >90% reduction of the targeted proteins (Fig. 3A). All cells expressing Rictor shRNA had significantly reduced levels of pAktSer473, while pAktSer473 levels in cells expressing Raptor shRNA were increased compared to control cells.
Figure 3. mTORC1 and mTORC2 regulate the migration and invasion of CRCs.
A. HCT116, SW480 and KM20 sh NTC, sh Raptor and sh Rictor cells were generated. B. Transwell migration assay was performed with HCT116, SW480 and KM20 sh Raptor, sh Rictor and sh NTC cells (* p<0.05 vs. NTC shRNA). C. Wound healing assay performed with HCT116 and SW480 sh Raptor, sh Rictor and sh NTC cells over 48 h (* p<0.05 vs. NTC shRNA). D. Transwell invasion assay was performed with HCT116 and SW480 sh Raptor, sh Rictor and sh NTC cells over 48 h (* p<0.05 vs. NTC shRNA).
Migration of stably silenced cells was assessed using the short-term transwell migration assay. Stable knockdown of both Raptor and Rictor significantly decreased migration of HCT116, SW480 and KM20 cells (Fig. 3B). Consistent with results obtained from transwell migration assay, knockdown of both Raptor and Rictor significantly decreased migration of HCT116 and SW480 cells using wound healing assay (Fig. 3C). Finally, invasion of stably silenced cells was assessed using a modified Boyden chamber assay. Targeted inhibition of both Raptor and Rictor significantly decreased invasion of HCT116 and SW480 cells (Fig. 3D). Taken together, our findings suggest that both mTORC1 and mTORC2 regulate migration and invasion, critical steps in the initial progression of CRC.
mTORC1 and mTORC2 regulate EMT in CRC
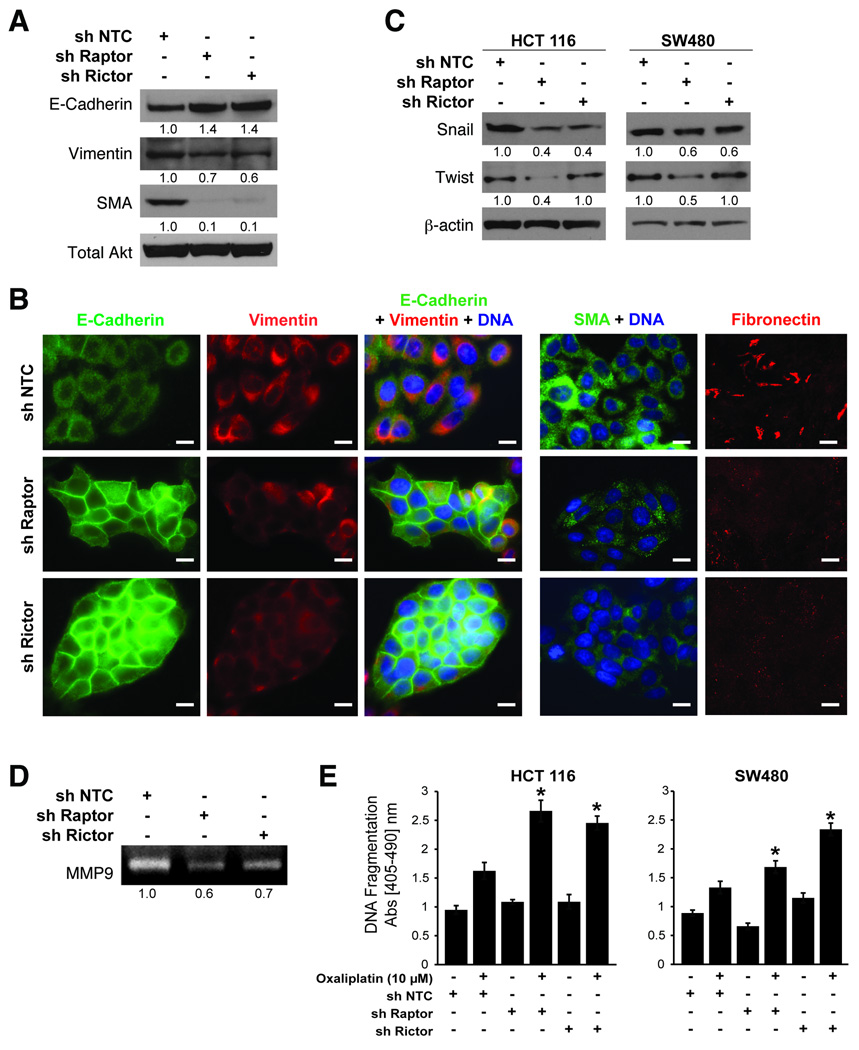
Several biochemical markers are used to characterize EMT: epithelial cells express E-cadherin predominantly, while mesenchymal cells express vimentin, smooth muscle actin (SMA) and fibronectin (3). Since targeted inhibition of mTORC1 and mTORC2 attenuated migration and invasion, we determined whether their inhibition is sufficient to induce MET by examining expression of the aforementioned markers. Stable knockdown of Raptor and Rictor in SW480 cells increased E-cadherin levels, while decreasing vimentin and SMA levels (Fig. 4A). Immunofluorescence microscopy confirmed increased levels of E-cadherin and decreased levels of vimentin, SMA and fibronectin in Raptor and Rictor knockdown cells compared to control cells (Fig. 4B).
Figure 4. mTORC1 and mTORC2 regulate epithelial-mesenchymal transition and actin cytoskeletal rearrangement of CRCs.
A. Immunoblot analysis for EMT markers in SW480 cells with sh NTC, sh Raptor and sh Rictor. B. Immunofluorescence staining in SW480 sh NTC, sh Raptor and sh Rictor cells grown in regular growth medium. Bar=50µm. C. Immunoblot analysis in HCT116 and SW480 cells. D. Gelatin zymography of supernatants collected from SW480 sh NTC, sh Raptor and sh Rictor cells stimulated with 5 ng/mL TGFβ for 12 h. E. Apoptosis assessment in HCT116 and SW480 sh NTC, sh Raptor and sh Rictor cells treated with DMSO (control) or Oxaliplatin (10µM) for 24 h (* p<0.05 vs. NTC shRNA).
The transcription factors, Snail and Twist, are implicated in transcriptional repression of E-cadherin expression and orchestrating the molecular EMT program (4). Since knockdown of Raptor and Rictor increased E-cadherin expression and induced MET, we determined whether these proteins regulate expression of Snail and Twist (Fig. 4C). Consistent with the increase in E-cadherin levels, we noted that Snail expression was decreased with knockdown of both Raptor and Rictor, while Twist expression decreased predominantly with knockdown of Raptor in HCT116 and SW480 cells.
EMT is further associated with increased production of matrix metalloproteases (MMPs), which serve to degrade extracellular matrix proteins and facilitate cell invasion. Since mTORC1 and mTORC2 can regulate the invasion of CRCs, we determined whether silencing mTORC1 or mTORC2 could decrease MMP production. As shown in Fig. 4D, knockdown of Raptor and Rictor significantly decreased levels of MMP9 secreted by SW480 cells.
Finally, EMT is associated with acquisition of chemotherapeutic resistance (3). In order to address this issue, we determined whether silencing mTORC1 or mTORC2 could enhance sensitivity to oxaliplatin-induced apoptosis. As shown in Fig. 4E, HCT116 and SW480 cells expressing shRNA targeting Raptor or Rictor were sensitized to oxaliplatin-induced apoptosis compared to control cells. Taken together, these results suggest that inhibition of mTORC1 and mTORC2 induces morphological, biochemical and functional changes reminiscent of MET.
mTORC1 and mTORC2 regulate actin cytoskeleton rearrangements involved in CRC migration via RhoA and Rac1 signaling
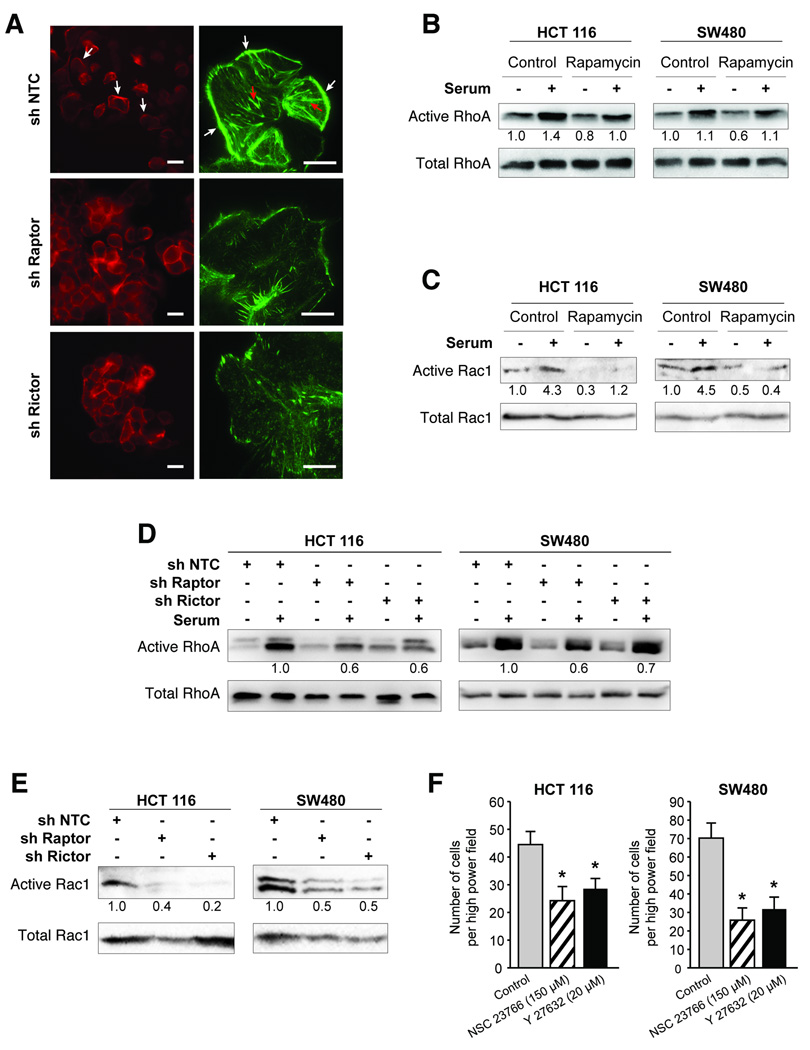
Polymerization and depolymerization of filamentous (F-) actin controls cytoskeletal reorganization leading to morphologic changes (such as lamellipodia formation) associated with motility. We investigated whether mTORC1 and mTORC2 regulate changes in the actin cytoskeleton by staining for F-actin in SW480 cells with stable Raptor or Rictor knockdown (Fig. 5A). Control cells displayed a mesenchymal morphology, spreading out evenly over the substratum with reduced cell-cell contacts (left panel) and displayed abundant lamellipodia and stress fibers (right panel). Conversely, both Raptor and Rictor knockdown cells displayed epithelial morphology, packed together closely with numerous cell-cell contacts (left panel) and displayed a dramatic decrease in lamellipodia and stress fibers (right panel).
Figure 5. mTORC1 and mTORC2 regulate activation of RhoA and Rac1.
A. Immunofluorescence staining for F-actin (using rhodamine phalloidin) in SW480 sh NTC, sh Raptor and sh Rictor cells grown in regular growth medium. White arrow=lamellipodia, red arrow=stress fibers. Left Bar=50µm Right Bar=10µm. B–E. HCT116 and SW480 cells serum starved overnight and pre-treated with DMSO or Rapamycin (50nM) for 24 h were stimulated with 10% FBS and assessed for B. RhoA-GTP and total cellular RhoA and C. Rac1-GTP and total cellular Rac1. HCT116 and SW480 sh NTC, sh Raptor and sh Rictor cells were serum starved overnight and assessed for D. RhoA-GTP and total cellular RhoA and E. Rac1-GTP and total cellular Rac1. F. Transwell migration assay was performed with HCT116 and SW480 cells treated with either NSC23766 (150 µM) or Y27632 (20 µM). Cells in control group were treated with DMSO (* p<0.05 vs. control).
RhoA and Rac1 belong to a family of GTPases that regulate F-actin assembly and disassembly while controlling cell migration. Since Raptor and Rictor knockdown cells demonstrated decreased lamellipodia, stress fibers and migration compared to control cells, we determined whether inhibiting mTOR signaling pharmacologically using rapamycin and shRNA-mediated inhibition of Raptor and Rictor can alter activation of RhoA and Rac1 using GST-TRBD and GST-PBD pull-down assays, respectively. As shown in Fig. 5B and 5C, RhoA and Rac1 activity decreased basally following rapamycin treatment in both HCT116 and SW480 cells. Serum stimulation increased activation of RhoA and Rac1, while treatment with rapamycin attenuated this activation. Next, we evaluated activation of RhoA and Rac1 in Raptor or Rictor knockdown HCT116 and SW480 cells. A 30–40% decrease in activated RhoA and 50–85% decrease in Rac1 activity was observed upon Raptor or Rictor knockdown compared to control cells (Fig. 5D and 5E). Finally, to implicate these downstream pathways in regulating migration of CRC cells, we treated HCT116 and SW480 cells with small molecule inhibitors of Rho-associated kinase (Y27632) and Rac1 (NSC23766) and assessed their effect on migration using transwell migration assay. We found that both compounds significantly reduced migration of HCT116 and SW480 cells (Fig. 5F). Taken together, these findings suggest that mTORC1 and mTORC2 are required for activation of RhoA and Rac1. The decrease in RhoA and Rac1 activity may prevent rearrangement of the actin cytoskeleton, thus accounting for the attenuated migratory capability noted upon mTORC1 and mTORC2 inhibition.
mTORC1 and mTORC2 are critical for establishment of CRC metastases
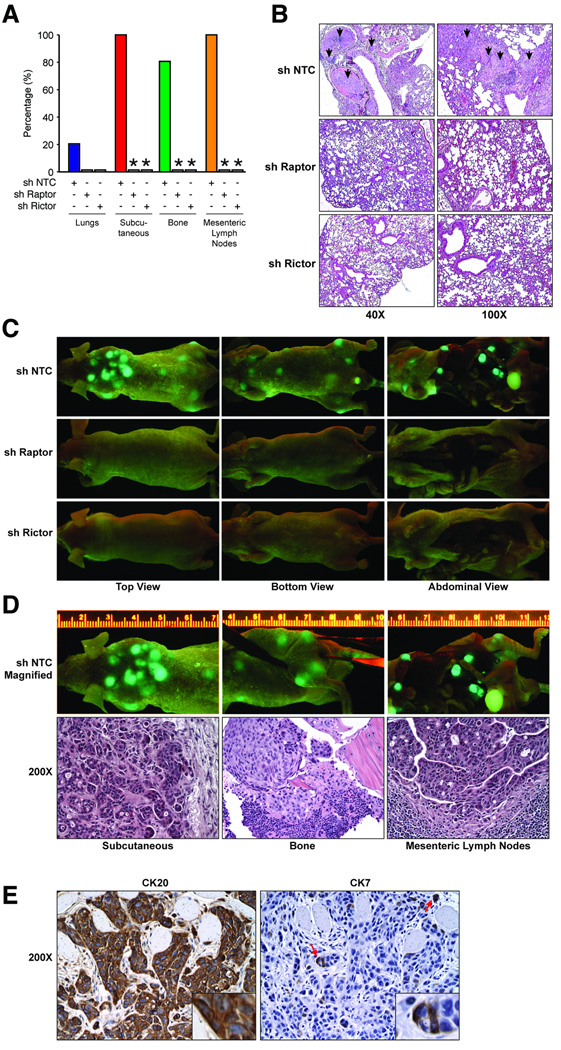
To further extend our findings in vivo, we examined the effect of mTORC1 and mTORC2 inhibition in an experimental metastasis model. GFP-labeled KM20 cells with stable knockdown of Raptor or Rictor (Fig. 3A) were injected intravenously into athymic nude mice and formation of systemic metastases was assessed; findings are summarized in Fig. 6A. Macroscopic lung nodules were not observed upon gross examination of lungs from animals inoculated with either control, Raptor or Rictor knockdown cells. However, histological examination demonstrated the presence of micrometastases in 20% of mice from the control group (Fig. 6B). In contrast, knockdown of Raptor or Rictor completely abolished formation of pulmonary micrometastases in all mice. Surprisingly, we found that 100% of mice inoculated with control cells exhibited metastatic nodules on the upper back, bone (knee joint and ribcage) and mesenteric lymph nodes (Fig. 6C and 6D). In contrast, knockdown of Raptor and Rictor completely abolished establishment of these metastases. To further confirm that the cellular origin of metastases was indeed an epithelial CRC cell line (KM20), metastatic tissues from the aforementioned locations were stained with cytokeratin 7 (CK7) and 20 (CK20). As expected for metastatic CRC, the tumor showed diffusely strong immunoreaction for CK20, while immunoreaction for CK7 was only focally noted in this poorly differentiated (high grade) CRC (Fig. 6E). (16). Taken together, our findings suggest that mTORC1 and mTORC2 are critical for establishment of CRC metastases irrespective of their site of colonization.
Figure 6. Inhibition of mTORC1 and mTORC2 abolishes establishment of metastases of CRCs in vivo.
GFP-labeled KM20 sh NTC, sh Raptor and sh Rictor cells inoculated intravenously into athymic Crl:NU-Foxn1nu nude mice and assessed after 6 weeks. A. Summary of metastases in each location (* p<0.05 vs. NTC shRNA). B. Representative images from histological assessment of lungs. C. Representative images from assessment for systemic metastases using GFP illumatool. D. Representative subcutaneous, bone and lymph node metastases along with histological analysis (200×). E. Expression of cytokeratin 20 and cytokeratin 7 in metastatic tissues (200×).
DISCUSSION
In this study, we determined the role of mTORC1 and mTORC2 in regulating motility, EMT and metastasis of CRC. First, we found increased expression of mTOR, Raptor and Rictor mRNA with more advanced stages of CRC. Additionally, mTOR, Raptor and Rictor protein levels were significantly elevated in stage IV CRCs and this overexpression profile was maintained in their matched distant metastasis. Second, we show that inhibition of mTORC1 and mTORC2 attenuated migration and invasion of CRCs concomitant with altered cytoskeletal rearrangement and decreased activation of RhoA and Rac1. Third, we demonstrate that inhibition of mTORC1 and mTORC2 induces changes reminiscent of MET. Finally, we show that establishment of metastasis in vivo was completely abolished upon targeted inhibition of mTORC1 and mTORC2. We propose that mTORC1 and mTORC2 regulate motility of CRCs via RhoA and Rac1 signaling.
While intensive studies have focused on the role of mTOR signaling in regulating growth and survival, its role in EMT, motility and metastasis of cancers is not well understood. We found that pharmacological (using rapamycin) and genetic (using RNAi) inhibition of mTORC1 (Raptor) and mTORC2 (Rictor) significantly decreased migration, invasion and establishment of metastasis in vivo. Our findings are consistent with several studies demonstrating inhibition of migration and invasion by rapamycin in various types of cancers (17). A recent study utilized knockdown of Raptor and Rictor to demonstrate that both mTORC1 and mTORC2 regulate IGF-1 stimulated migration of various cancer cells by inhibiting 4E-BP1 and p70S6K signaling (18). Finally, another recent study showed that Rictor knockdown reduces cellular chemotactic capacity and ablates pulmonary metastasis via PKCζ in breast cancer (19). These findings highlight the fact that these functions of mTOR and its interaction partners are not cell-type specific and are noted in various types of cancers.
One of the earliest detectable morphological changes observed during cell migration involves rearrangement of the actin cytoskeleton leading to formation of lamellipodia (5). Our findings indicate that inhibition of mTORC1 and mTORC2 decreases formation of lamellipodia. Consistent with our findings, a recent study showed that both mTORC1 and mTORC2 are involved in regulating F-actin reorganization and formation of lamellipodia in a panel of cancer cells (18). Moreover, previous studies have also demonstrated that silencing of mTORC2 prevents cell spreading and F-actin polymerization via PKCα in fibroblasts (20).
The Rho family of GTPases, including RhoA and Rac1, participate in regulating actin cytoskeleton reorganization and cell migration. Our findings in this study implicate signaling through RhoA and Rac1 pathways as a critical downstream mechanism by which mTORC1 and mTORC2 may regulate changes in the actin cytoskeleton and cell migration. Consistent with these findings, Jacinto et al. (21) observed that mTORC2 signals through Rac1 and found that mTOR also signals through RhoA. Similarly, Moss et al. (22) demonstrated that rapamycin increases levels of the cyclin-dependent kinase inhibitor, p27Kip1, and in turn inhibits RhoA activation, thereby blocking cytoskeletal reorganization and cell migration. Finally, a recent study showed that disruption of mTORC1 and mTORC2 by inhibiting Raptor and Rictor, respectively, inhibits the activity of RhoA and Rac1 (23). Additionally, mTORC1 was also shown to control the translation of RhoA and Rac1 mRNA’s via 4E-BP1 and p70S6K.
EMT is a key reversible step that facilitates tumor migration, invasion and metastasis. Owing to the clinical importance of this process, inhibition of EMT is an attractive therapeutic approach that can significantly alter disease outcome. However, it remains unknown which pathways should be inhibited in order to reverse EMT. Our findings implicate mTORC1 and mTORC2 as key regulators of EMT in CRCs. This conclusion is based on the observation that silencing mTORC1 and mTORC2 induces a repertoire of biochemical (increased E-cadherin and decreased vimentin, SMA and fibronectin), morphological (increased cell-cell contact, decreased formation of lamellipodia) and functional (decreased MMP9 production and enhanced sensitivity to oxaliplatin-induced apoptosis) changes characteristic of MET. Consistent with our findings, a recent study found that the cytokine TGFβ, which is known to play a major role in promoting EMT, induces activation of mTOR signaling and phosphorylation of p70S6K and 4E-BP1, which subsequently increased protein synthesis and cell size (24). Inhibition of mTOR signaling using rapamycin inhibited the increase in protein synthesis and cell size, while inhibiting cell migration and invasion associated with TGFβ-induced EMT. These studies suggest that mesenchymal-epithelial reprogramming may be an important mechanism underlying the attenuated metastasis of CRC noted upon inhibition of mTORC1 and mTORC2.
Several findings support the targeting of mTOR signaling as an anti-metastatic therapy. First, we demonstrate that mTOR, Raptor and Rictor are overexpressed at both the mRNA and protein level in primary CRCs and this increased expression is maintained in the associated distant metastases. The positive correlation noted between expression and stage suggests that mTOR signaling may contribute towards CRC progression. Second, multiple upstream pathways ultimately converge upon mTORC1 and mTORC2 to facilitate cancer cell migration and invasion. Recent studies have shown that IGF-1, EGF and TGFβ, all critical mediators of migration, EMT and metastasis, activate mTOR signaling; inhibition of mTOR, using rapamycin, potently inhibited cell motility induced by IGF-1, EGF and TGFβ (17, 19, 24). These findings suggest that mTOR signaling may be a critical node in regulating progression and metastasis of cancers. Targeting mTORC1 and mTORC2 could be a more beneficial strategy than targeting individual, upstream redundant signaling pathways. Third, advanced cancers are difficult to treat since they are chemoresistant. We demonstrate that knockdown of mTORC1 and mTORC2 sensitizes CRCs to undergo apoptosis upon treatment with oxaliplatin.
Despite the incontrovertible rationale for treating cancers addicted to PI3K/Akt/mTOR signaling, such as CRC, with rapamycin, clinical results have been disappointing. One proposed mechanism of resistance to rapamycin involves its inability to inhibit mTORC2 (10, 12). Our findings support a role for elevated mTORC1 and mTORC2 activity in regulating EMT and metastasis of CRC. Taken together with our previous results demonstrating that both mTORC1 and mTORC2 contribute to CRC tumorigenesis, we hypothesize that the inherent redundancy in functions of both complexes may allow mTORC2 to compensate for loss of mTORC1 activity upon rapamycin treatment, thereby leading to rapamycin resistance. Our findings provide the rationale for including mTOR kinase inhibitors targeting the ATP binding pocket, which inhibit both mTORC1 and mTORC2 more completely, as part of the therapeutic regimen for treating CRC patients.
ACKNOWLEDGEMENTS
Authors thank Nathan Vanderford for editing and Donna Gilbreath for graphic assistance.
This work was supported by grants P20CA1530343 (GI SPORE; BME), R01CA133429 (TG) and R01CA10913601 (KLOC) from NIH.
REFERENCES
- 1.Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA: A Cancer Journal for Clinicians. 2011 doi: 10.3322/caac.20107. [DOI] [PubMed] [Google Scholar]
- 2.Griffin MR, Bergstralh EJ, Coffey RJ, Beart RW, Jr, Melton LJ., 3rd Predictors of survival after curative resection of carcinoma of the colon and rectum. Cancer. 1987;60:2318–2324. doi: 10.1002/1097-0142(19871101)60:9<2318::aid-cncr2820600934>3.0.co;2-b. [DOI] [PubMed] [Google Scholar]
- 3.Thiery JP. Epithelial-mesenchymal transitions in tumour progression. Nat Rev Cancer. 2002;2:442–454. doi: 10.1038/nrc822. [DOI] [PubMed] [Google Scholar]
- 4.Wu Y, Zhou BP. New insights of epithelial-mesenchymal transition in cancer metastasis. Acta Biochim Biophys Sin (Shanghai) 2008;40:643–650. doi: 10.1111/j.1745-7270.2008.00443.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Hall A. Rho GTPases and the actin cytoskeleton. Science. 1998;279:509–514. doi: 10.1126/science.279.5350.509. [DOI] [PubMed] [Google Scholar]
- 6.Philp AJ, Campbell IG, Leet C, Vincan E, Rockman SP, Whitehead RH, et al. The phosphatidylinositol 3'-kinase p85alpha gene is an oncogene in human ovarian and colon tumors. Cancer Res. 2001;61:7426–7429. [PubMed] [Google Scholar]
- 7.Roy HK, Olusola BF, Clemens DL, Karolski WJ, Ratashak A, Lynch HT, et al. AKT proto-oncogene overexpression is an early event during sporadic colon carcinogenesis. Carcinogenesis. 2002;23:201–205. doi: 10.1093/carcin/23.1.201. [DOI] [PubMed] [Google Scholar]
- 8.Rychahou PG, Jackson LN, Silva SR, Rajaraman S, Evers BM. Targeted molecular therapy of the PI3K pathway: therapeutic significance of PI3K subunit targeting in colorectal carcinoma. Ann Surg. 2006;243:833–842. doi: 10.1097/01.sla.0000220040.66012.a9. discussion 43-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Rychahou PG, Kang J, Gulhati P, Doan HQ, Chen LA, Xiao SY, et al. Akt2 overexpression plays a critical role in the establishment of colorectal cancer metastasis. Proc Natl Acad Sci U S A. 2008;105:20315–20320. doi: 10.1073/pnas.0810715105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Guertin DA, Sabatini DM. Defining the role of mTOR in cancer. Cancer Cell. 2007;12:9–22. doi: 10.1016/j.ccr.2007.05.008. [DOI] [PubMed] [Google Scholar]
- 11.Sarbassov DD, Ali SM, Sengupta S, Sheen JH, Hsu PP, Bagley AF, et al. Prolonged rapamycin treatment inhibits mTORC2 assembly and Akt/PKB. Mol Cell. 2006;22:159–168. doi: 10.1016/j.molcel.2006.03.029. [DOI] [PubMed] [Google Scholar]
- 12.Gulhati P, Cai Q, Li J, Liu J, Rychahou PG, Qiu S, et al. Targeted inhibition of mammalian target of rapamycin signaling inhibits tumorigenesis of colorectal cancer. Clin Cancer Res. 2009;15:7207–7216. doi: 10.1158/1078-0432.CCR-09-1249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Bowen KA, Doan HQ, Zhou BP, Wang Q, Zhou Y, Rychahou PG, et al. PTEN loss induces epithelial--mesenchymal transition in human colon cancer cells. Anticancer Res. 2009;29:4439–4449. [PMC free article] [PubMed] [Google Scholar]
- 14.Larson Y, Liu J, Stevens PD, Li X, Li J, Evers BM, et al. Tuberous sclerosis complex 2 (TSC2) regulates cell migration and polarity through activation of CDC42 and RAC1. J Biol Chem. 2010;285:24987–24998. doi: 10.1074/jbc.M109.096917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.O'Connor KL, Nguyen BK, Mercurio AM. RhoA function in lamellae formation and migration is regulated by the alpha6beta4 integrin and cAMP metabolism. J Cell Biol. 2000;148:253–258. doi: 10.1083/jcb.148.2.253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kende AI, Carr NJ, Sobin LH. Expression of cytokeratins 7 and 20 in carcinomas of the gastrointestinal tract. Histopathology. 2003;42:137–140. doi: 10.1046/j.1365-2559.2003.01545.x. [DOI] [PubMed] [Google Scholar]
- 17.Liu L, Li F, Cardelli JA, Martin KA, Blenis J, Huang S. Rapamycin inhibits cell motility by suppression of mTOR-mediated S6K1 and 4E-BP1 pathways. Oncogene. 2006;25:7029–7040. doi: 10.1038/sj.onc.1209691. [DOI] [PubMed] [Google Scholar]
- 18.Liu L, Chen L, Chung J, Huang S. Rapamycin inhibits F-actin reorganization and phosphorylation of focal adhesion proteins. Oncogene. 2008;27:4998–5010. doi: 10.1038/onc.2008.137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhang F, Zhang X, Li M, Chen P, Zhang B, Guo H, et al. mTOR Complex Component Rictor Interacts with PKC{zeta} and Regulates Cancer Cell Metastasis. Cancer Res. 2010 doi: 10.1158/0008-5472.CAN-10-0207. [DOI] [PubMed] [Google Scholar]
- 20.Sarbassov DD, Ali SM, Kim DH, Guertin DA, Latek RR, Erdjument-Bromage H, et al. Rictor, a novel binding partner of mTOR, defines a rapamycin-insensitive and raptor-independent pathway that regulates the cytoskeleton. Curr Biol. 2004;14:1296–1302. doi: 10.1016/j.cub.2004.06.054. [DOI] [PubMed] [Google Scholar]
- 21.Jacinto E, Loewith R, Schmidt A, Lin S, Ruegg MA, Hall A, et al. Mammalian TOR complex 2 controls the actin cytoskeleton and is rapamycin insensitive. Nat Cell Biol. 2004;6:1122–1128. doi: 10.1038/ncb1183. [DOI] [PubMed] [Google Scholar]
- 22.Moss SC, Lightell DJ, Jr, Marx SO, Marks AR, Woods TC. Rapamycin regulates endothelial cell migration through regulation of the cyclin-dependent kinase inhibitor p27Kip1. J Biol Chem. 2010;285:11991–11997. doi: 10.1074/jbc.M109.066621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Liu L, Luo Y, Chen L, Shen T, Xu B, Chen W, et al. Rapamycin inhibits cytoskeleton reorganization and cell motility by suppressing RhoA expression and activity. J Biol Chem. 2010 doi: 10.1074/jbc.M110.141168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lamouille S, Derynck R. Cell size and invasion in TGF-beta-induced epithelial to mesenchymal transition is regulated by activation of the mTOR pathway. J Cell Biol. 2007;178:437–451. doi: 10.1083/jcb.200611146. [DOI] [PMC free article] [PubMed] [Google Scholar]