Abstract
Numerous studies have recently focused on the identification of specific glycan biomarkers; given the important roles that protein linked glycans play, for example, during development and disease progression. The identification of protein glycobiomarkers, which are part of a very complex proteome, has involved the use of fractionation techniques such as lectin affinity chromatography. In this study, the glycoproteomic characterization of pluripotent murine embryonic stem cells (ES) and from ES cells that were differentiated into embroid bodies (EB) was performed using immobilized Concanavalin A (ConA). This procedure allowed the isolation of glycopeptides that express biantennary and hybrid N-linked structures (ConA2 fraction) as well as high mannose glycans (ConA3 fraction), that were abundant in both ES and EB stages. A total of 293 unique N-linked glycopeptide sequences (from 180 glycoproteins) were identified in the combined data sets from ES and EB cells. Of these glycopeptides, a total of 119 sequences were identified exclusively in only one of the lectin bound fractions, (24 in the ES-ConA2, 15 in the ES-ConA3, 16 in the EB-ConA2 and 64 in the EB-ConA3). Results from this study allowed the identification of individual N-glycosylation sites of proteins that express specific glycan types. The absence of some of these lectin bound glycopeptides in a cell stage suggested that they were derived from proteins that were either expressed exclusively on a defined developmental stage, or were expressed in both cell stages but carried the lectin bound oligosaccharides in only one of them. Therefore, these lectin bound glycopeptides can be considered as stage specific glycobiomarkers.
Keywords: Glycoproteomics, LC-MS/MS, Glycopeptides, N-linked glycosylation sites, Lectin affinity chromatography, embryonic stem cells
1. Introduction
It has been estimated that approximately 60% of total human proteins and virtually all of secreted or membrane bound proteins are glycosylated 1. The oligosaccharide moieties of these glycoproteins play crucial roles in various processes, such as protein folding' cell-cell recognition, signal transduction, inflammation, tumorigenesis, differentiation, as well as cell-cell recognition2-4. Efforts to characterize these glycoproteins involve the identification of specific glycan structures, the identification of the proteins that express each glycan, the identification of specific glycosylation sites in proteins and specific glycan structures expressed at these sites, constituting the emerging field of glycoproteomics 5,6.
Recently, the efforts of many research groups have focused on the use of glycoproteomic methodologies for the identification of particular protein markers that express specific glycan structures 6, 7. Some of these efforts have been successful in the identification of specific glycoprotein biomarkers that may prove to be useful for the early detection of diseases such as liver cancer and colon cancer 7, 8. These glycosylation specific protein markers have been designated as “glycobiomarkers” 6.
For the identification of potential glycobiomarkers, it is desirable to enrich for glycoproteins or glycopeptides, and several strategies have been implemented. Some of these methodologies are based on the use of recognition molecules that bind to carbohydrate moieties to capture and separate glycoproteins of interest. Lectins are proteins of non-immune origin that recognize and bind to glycan structures which can be either free in solution, or covalently linked to glycoproteins, glycolipids and proteoglycans. Given the availability of many lectins with specificities for a great variety of glycan structures, these molecules have been used in glycoproteomics studies to identify glycoproteins from serum that express a specific set of glycan structures 9-11. Moreover, lectins have been used in the identification of glycosylation sites in a set of glycoproteins 10-15, as well as being used in the identification of glycans that are expressed at specific glycosylation sites in glycoproteins of interest 16.
Lectins have also been used to investigate changes in the glycan repertoire that occur during oncogenesis. For example, an increase in the expression of N-linked glycans with the Man-β-1,6-GlcNAc branch, which is recognized by the leucocyte hemagluttinin from Phaseolus vulgaris (L-PHA), has been shown to correlate with increased malignancy and metastases in patients with breast cancer, melanoma and several other types of cancer 17, 18. Similarly, there are glycosylation epitopes that are expressed during the initial steps of embryonic development. Examples of this are the Lewis-X (SSEA-1) and Forssmann antigens that are expressed specifically during early mouse embryogenesis 19.
In mammals, pluripotent embryonic stem (ES) cells are derived from the inner cell mass (ICM) of blastocyst-stage embryos. When cultured over extensive periods of time under appropriate conditions, ES cells retain many of the characteristic associated with pluripotent cells of the ICM, including the capacity to generate the three embryonic germ lineages (ectoderm, endoderm and mesoderm) as well as the extra embryonic tissues that support development. This pluripotency of ES cells provides the basis for replicating a wide variety of somatic and extra embryonic tissues. Understanding the molecular mechanisms for stem cell differentiation and directing these mechanisms to obtain specific stem cell populations are critical areas of contemporary research because of the potential therapeutic applications in the treatment of diseases 20. A significant impediment to this research is the ability to isolate pure populations of differentiated cells of interest. Identifying cell-type specific markers that allow these types of isolation, therefore, is of paramount importance.
Differentiation of stem cells into embroid bodies or other defined cell types is reflected in the expression of specific proteins in the proteomic repertoire of the differentiated cells 21-23. As a result of these differences, protein markers such as CD9 or alkaline phosphatase (AP) have been identified as stage specific markers for the pluripotent ES stage in murine or human stem cells 20. The molecular differences between ES cells and their differentiated cell lineages are also manifested in changes in their glycan repertoire, examples of this are the Lewis-X (SSEA-1) antigens that are expressed specifically during the ES stage but disappear upon differentiation of the cells into other developmental stages, or the expression of glycoproteins with terminal α-linked GalNAc residues that bind to the lectin DBA during the ES stage, and disappears upon differentiation of the cells 19.
Preliminary studies 24 (L. Wells and M. Tiemeyer, personal communication) have shown that both murine embryonic stem (ES) cells and cells in differentiated embryoid bodies (EB) expressed large amounts of high mannose, hybrid, or complex biantennary N-linked glycans. In order to determine differences in the identities of the proteins that express these structures in both cell stages and to attempt to define stage-specific glycobiomarkers, we used Concanavalin A (ConA) lectin affinity for the isolation and glycoproteomic analysis of glycopeptides prepared from ES and EB cells. The sequences of the N-linked glycopeptides in the Con A-separated fractions containing high mannose/hybrid and biantennary glycans were identified after deglycosylation and isotopic tagging of glycosylation sites by PNGase F treatment in H218O. These glycoproteomic analyses resulted in the identification of abundant glycoproteins present in both ES and EB cell stages. However, a set of glycoproteins was found exclusively in the ConA bound glycopeptide fraction from ES cells, while a separate set of glycoproteins was identified as present only in the EB stage. The results presented in this study indicate that this glycoproteomic strategy can serve as a basis for identification of potential cell type-specific glycobiomarkers, either the glycopeptides themselves, or the glycoproteins from which they are derived.
2. Experimental Section
Cell culture
D3 and R1 mESCs 25 were cultured in the absence of feeders on tissue culture grade plastic ware precoated with 0.1% gelatin-phosphate buffered saline (PBS), as described previously 26. mESC culture medium consisted of Dulbecco's Modified Eagle Medium (DMEM, Gibco BRL) supplemented with 10% fetal calf serum (FCS), 1 mM L-glutamine, 0.1 mM 2-mercaptoethanol, and 103 U/ml recombinant human leukemia inhibitory factor (LIF, ESGRO, Chemicon International, Temecula CA) at 37°C under 10% CO2. Differentiation of mESCs into embryoid bodies (EBs) was carried out as described 27. ES cells were harvested by trypsinization converting suspensions of single ESCs into aggregates and seeded into 10 cm bacteriological dishes at a density of 1 × 105 cells/mL, in 10 mL mESC-medium lacking LIF. EBs were harvested daily, the medium was changed every 2 days, and cultures were split one into two at day 4.
Preparation of Glycopeptides from stem cell pellets
Murine stem cells and embroid bodies pellets (ca. 1× 107 cells). Cells were harvested and transferred into 15 mL conical tubes where they pelleted by centrifugation at 1000 ×g. The cells were washed three times with a phosphate buffered solution (50 mM Na2HPO4, 150 mM NaCl, pH 7.6), centrifuging at 1000 ×g after each wash. All supernatant was removed from the tube and the cell pellets were stored at − 80 °C until analysis. For oligosaccharide extraction, lipids were first extracted from the cells using a modification of the procedure by Svennerholm 28. The cell pellets (ca. 1 × 107 cells) were added with 2 mL of water. The tubes were then placed in an ice bath and the pellets were sonicated 40 seconds (in four pulses of 10 seconds each) in a probe sonicator at an intensity of 15 watts. The solubilized cells were then mixed with methanol and chloroform to a final proportion of chloroform: methanol: water of 4:8:3. The resulting mixture was incubated 2 hours at room temperature and then mixed with water to modify the chloroform: methanol: water proportion to 4:8:5.6. The mixture was then centrifuged at 5000 ×g and three phases were separated. The lower (chloroform rich) and upper (aqueous) phases were carefully removed with a Pasteur pipette and the intermediate layer (protein rich) was mixed with 1 mL of acetone and centrifuged at 5000 ×g. The acetone supernatant was removed and the delipidated protein pellet was washed once more with cold acetone, then mixed with 2 mL of water, sonicated as described above and lyophilized overnight. The lyophilized proteins (10-20 mg) were dissolved in 1 mL of 50 mM, Tris 2M Urea, pH 8.5 and dissolved by sonication. The proteins were reduced by adding dithiotreitol (DTT) to a concentration of 25 mM for 45 min at 50 °C and then carbamidomethylated by adding iodoacetamide to a concentration of 90 mM during 1 hour in the dark. Trypsin (10 μg for each milligram of protein) was then added and the proteolytic digestion was carried out overnight at 37 °C. The resulting mixture of peptides and glycopeptides was desalted through Sep-Pak C18 cartridge column. The cartridge was activated with 10 mL of methanol and then equilibrated with 10 ml volumes of 5% acetic acid. Glycopeptides were eluted stepwise, first with 3 ml of 20% isopropyl alcohol in 5% acetic acid and then with 3 ml of 40% isopropyl alcohol in 5% acetic acid. The 20 and 40% isopropyl alcohol steps were pooled and evaporated to dryness.
Lectin affinity chromatography
Lectin affinity chromatography with modifications of the technique described by Cummings et al.29. Briefly, Glycopeptide fractions from ES and EB cells were dissolved in a Tris Buffered solution (TBS; 20mM Tris HCL, 150 mM NaCl, 1mM MgCl2, 1mM CaCl2, pH 7.4) and applied to a disposable column packed with 0.5 mL of Concanavalin A (Con A) Sepharose (GE-Healthcare, Piscataway, NJ), which had been washed with 5 mL of TBS buffer prior to the addition of the peptide mixture. The Con A column was next eluted with 2.5 mL of TBS to recover the unbound glycopeptides. The column was then eluted with 2.5 ml of 10mM α-D-methylglucopyranoside in TBS to obtain the biantennary and hybrid N-linked glycopeptide fraction (ConA2). The high mannose glycopeptide fraction (ConA3) was eluted with 2.5 mL of 100 mM α-D-methyl mannoside in TBS. The eluted glycopeptides were desalted on a 60 mg Oasis MCX cartridge (Waters). The cartridge was washed with methanol and then equilibrated with 5% acetic acid. After application of sample, the unbound material was eluted with 3 mL of 5% acetic acid and then with 3 mL of methanol. The glycopeptides were eluted with a 5% NH4OH solution in 50% methanol. The desalted glycopeptides were then dried by vacuum centrifugation.
Deglycosylation of peptides
The dried glycopeptides were rehydrated in 30 μL of 50 mM ammonium bicarbonate in 95% H218O (Isotec through Sigma) 15. Recombinant PNGase F (0.2 U, a kind gift from Dr. Kelley Moremen), which had been suspended in H2 18O, was added to the glycopeptides, and deglycosylation was carried out overnight at 37 °C under nitrogen atmosphere. Formic acid (0.1%, 40 μL) was added and the PNGase F was removed by filtration over microcon YM-30 centrifugal filter. The filtrate was collected and the deglycosylated peptides were analyzed by LC-MS/MS.
LC-MS/MS and data analysis
Each of the deglycosylated peptides from the ConA bound fractions of ES and EB cells were analyzed on an Agilent 1100 capillary LC (Palo Alto, CA) interfaced directly to a LTQ linear ion trap mass spectrometer (Thermo Electron, San Jose, CA). Mobile phase A and B were H2O/0.1% formic acid and ACN/0.1% formic acid, respectively. Each fraction was loaded for 1 h onto a PicoFrit 8 cm 50 μm column (New Objective, Woburn, MA) packed with 5 μm diameter C18 beads using positive N2 pressure. The peptides were then desalted for 10 min with 0.1% formic acid using positive N2 pressure, and were eluted from the column into the mass spectrometer during a 70 min linear gradient from 5 to 45% B at a flow rate of 200 nL/min. The instrument was set to acquire MS/MS spectra on the 9 most abundant precursor ions from each MS scan with a repeat count of 3 and repeat duration of 15 s. Dynamic exclusion was enabled for 160 s. Raw tandem mass spectra were converted into mzXML format and then into peak list using ReAdW followed by mzXML2. The peak lists were then searched using Mascot 1.9 (Matrix Science, Boston, MA) against a target database composed of 34,966 Mus musculus protein sequences obtained from NCBI (www.ncbi.nih.gov) and a decoy database created by reversing the sequences in the target database. Database searches were performed against the target and decoy databases using the following parameters: fully tryptic enzymatic cleavage with 3 possible missed cleavages, peptide tolerance of 500 parts-per-million, fragment ion tolerance of 0.6 Da, and a variable modification due to carboxyamidomethylation (+57 Da). For identification of deglycosylated 18O labeled peptides the database search was performed using a variable modification of + 3 Da on asparagine residues, which accounts for the asparagine to aspartic acid transformations associated with deglycosylation and the incorporation of the isotope during this process 15. Following database searching, dataset organization and peptide statistical validation was performed using the PROVALT algorithm as integrated in the software package ProteoIQ (BioInquire, Athens GA)30. Statistical validation of peptide identifications was performed using the peptide false-discovery rate (PEP-FDR) approach by comparing the distribution of peptide identifications between the target and decoy database search results at each Mascot Ion Score31. Peptides were considered identified at a 1% PEP-FDR if they exceeded a Mascot Ion Score of 30. Further validation of glycopeptide identifications was performed to ensure that each contained an 18O label on asparagines only present in the N-glycosylation consensus sequence (N X S/T) where X is any amino acid other than proline.
For the identified peptide sequences and proteins Gene ID's and annotations were acquired from NCBI (http://www.ncbi.nlm.nih.gov). Transmembrane spanning domains were predicted with TMHMM 2.0.28, Signal peptide motifs were predicted with SignalP 3.0.29, subcellular localization and function was annotated using literature references, and the Ingenuity Pathways Analysis Software (http://www.ingenuity.com).
Exhaustive methylation of Glycans and MS Analysis
Dried glycans (30 μg aliquots) were permethylated with modifications of the procedure by Ciucanu and Kerek 32. Glycans were suspended in DMSO (0.1mL) and NaOH (20 mg in 0.1 mL of dry DMSO) was added. After strong mixing, 0.1 mL of 12C or 13C labeled methyl iodide (Aldrich) was added. According to the manufacturer, 13C labeled methyl iodide contained 99% of the 13C isotope. After 10 minutes incubation in a bath sonicator 1 mL of water was added, and the excess of methyl iodide was removed by bubbling with a stream of N2. One mL of methylene chloride was added with vigorous mixing, and after phase separation the upper aqueous layer was removed and discarded. The organic phase was then extracted three times with water. Methylene chloride was evaporated under a stream of N2, and the methylated glycans were dissolved in 25-50 μL of 50% methanol.
Before MALDI-TOF-MS analysis, mixtures of methylated oligosaccharides were redissolved 25 μL of 50% Methanol, 1mM NaOH. The MALDI-TOF-MS matrix was prepared by dissolving 13 mg of dihydrohybenzoic acid (DHB, Sigma) in 1 mL 50% acetonitrile in water. Then, 0.5μL of methylated glycan sample were mixed with 0.5μL of matrix solution and 0.5 μL of the mixture were applied to the MS probe and crystallized by evaporating solvents at room temperature. The samples were then analyzed in an Applied Biosystems 4700 Proteomics Analyzer working in TOF-reflector mode.
3. Results
Rationale of method
The purpose of this study was to develop a glycoproteomics protocol using lectin affinity chromatography of glycopeptides that would allow the identification of potential glycobiomarkers (proteins which express a specific glycan structure at a defined biological stage) whose expression could be potentially used to distinguish pluripotent murine embryonic stem cells (ES) from differentiated embryoid bodies (EB). Tryptic digests of glycoprotein extracts from ES and EB stages were subjected to lectin affinity chromatography, followed by proteomic identification by LC-MS/MS after enzymatic deglycosylation. To select an appropriate lectin for the glycoproteomic analysis, an initial glycan analysis was performed on N-linked glycan fractions purified from ES and EB stages. These oligosaccharides were subjected to exhaustive permethylation and were analyzed by MALDI-TOF-MS (Figure 1). Permethylation of glycans greatly enhances the ionization and detection of glycans by MS since methylated glycans ionize more efficiently than their native counterparts and due to their hydrophobic nature, are easily separated from salts and other impurities that may affect the MS analysis 33, 34. The glycans annotated for the MALDI-TOF-MS peaks in Figure 1 for ES and EB cells were inferred from the monosaccharide compositions derived from the masses represented by each peak and represent probable structures. These structures are consistent with those identified recently by total ion mapping and MSn fragmentation analysis using ESI-LTQ-MS in extracts from ES and EB cells (L. Wells and M. Tiemeyer, personal communication). The MS profiles of the permethylated glycans (Figure 1) indicated that N-linked oligosaccharides with high mannose and complex biantennary structures were abundant in both ES and ES cell stages. Glycans from EB cells however showed a larger proportion of complex biantennary and triantennary structures than ES cells. The presence of substantial amounts of N-linked glycans with high mannose, hybrid and biantennary structures in both ES and EB stages indicated that Concanavalin A (ConA) would be a suitable lectin to isolate and identify glycopeptide sequences from both cell stages.
Figure 1. MALDI-TOF spectrum of permethylated N-linked oligosaccharides obtained from glycoprotein extracts of ES(A) and EB(B) cells.

Glycoproteins were obtained from ES and EB cell pellets (ca 1×107 cells) after delipidation and N-linked oligosaccharides were released with PNGase F, permethylated and analyzed by MALDI-TOF/MS
 GlcNAc,
GlcNAc,
 Gal,
Gal,
 Man,
Man,
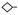 NeuNAc,
NeuNAc,
 Fuc.
Fuc.
ConA is a lectin purified from the seeds of the legume Canavalia ensiformis (jack bean). Several studies 29, 35-39 have shown that in affinity chromatography separations with this lectin, unbound glycopeptides (designated as ConA1 fraction) are eluted, followed by N-linked glycopeptides with hybrid or complex biantennary structures (designated as ConA2 fraction) when the column is eluted with 10mM α-methyl-glucopyranoside. The high mannose N-linked glycans for which this lectin has the highest affinity (designated as ConA3 fraction) are eluted with a solution that contains a high concentration (100 mM) of α-methyl mannopyranoside. Therefore, this lectin can be used to separate many of the N-linked glycopeptides present in ES and EB.
Lectin fractionation of glycopeptides from ES and EB cells and criteria for identification of N-glycosylation sites
A total of three cell pellets from each ES or EB cell stage (each with ca. 1×107 cells) were analyzed in this study. Proteins were isolated from ES and EB cell pellets after extraction of lipids with a mixture of chloroform and methanol 40, 41. The protein-enriched fraction was then reduced with DTT, carbamidomethylated with iodoacetamide, and proteolyzed with trypsin. The resulting mixture of peptides and glycopeptides was fractionated on a Concanavalin A lectin affinity column into the following fractions: ConA1 (lectin unbound, containing N-linked complex triantennary or tetraantennary glycopeptides), ConA2 (containing N-linked biantennary and hybrid) and ConA3 (containing high mannose). From the lectin fractionation described above a total of 12 samples were obtained, which represented the following lectin bound fractions from ES or EB cells: ES-ConA2, ES ConA3, EB-ConA2 and EB-ConA3. Each of these samples was prepared for LC-MS/MS (see below). The lectin-unbound fractions (ConA1) were not analyzed by LC-MS, but were saved for further fractionation with additional lectins.
The resulting ConA2 and ConA3 fractions were separately desalted and deglycosylated with PNGase F in the presence of H218O before being analyzed by LC-MS/MS. The deglycosylated peptides incorporated two 18O atoms in the aspartic acid residues that resulted after the hydrolytic deamidation of the glycosylated asparagine residues catalyzed by the glycoamidase. These labeled peptides with N-linked glycosylation sites were then identified by a 3 mass unit increase 12, 15. Peptide sequences were considered to be part of a glycoprotein if they contained a glycopeptide with a Mascot score above 30 and contained an 18O label on asparagines that were only present in the N-glycosylation consensus sequence (N X S/T where X is any amino acid other than proline). All of the peptide identification results for the lectin-bound fractions of ES and EB cells are presented in Supplementary Table 1. In Figure 2, the N-linked glycopeptide listings were organized by ES or EB cell type and the lectin fraction in which they were identified.
Figure 2. Venn diagram with the distribution of the glycopeptide sequences identified in the ConA bound fractions from ES and EB cells.
A total of three cell pellets from each ES or EB cell stage were subjected to lectin affinity fractionation and glycoproteomic analysis and a total of 12 samples were obtained, which represented the following lectin bound fractions (3 samples for each fraction): ES-ConA2, ES ConA3, EB ConA2, and EB ConA3. ConA2 represents the glycopeptides fraction eluted with 10 mM α-methyl-glucopyranoside (which contains biantennary and hybrid N-glycans) and ConA3 the glycopeptides eluted with 100 mM α-methyl-mannopyranoside which contain high mannose N-glycans.
For ES cells, 100 peptide sequences with N-linked glycosylation sites were identified in the ES-ConA2 fraction and 141 in the ES-ConA3 fraction. In the case of EB cells, 112 N-glycosylated peptides were identified in the EB-ConA2 fraction and 203 in the EB-ConA3 fraction. Considering that a substantial portion of the identified glycopeptide sequences were present in more than one of the samples analyzed by LC-MS/MS, a total of 293 unique N-linked glycopeptide sequences was identified when the results from all the fractions were combined in one data set. Figure 2 shows a Venn diagram with the distribution of the glycopeptide sequences identified among the ConA bound fractions from ES and EB cells. A total of 119 glycopeptides was identified exclusively in only one of the lectin bound fractions, (24 in the ES-ConA2, 15 in the ES-ConA3, 16 in the EB-ConA2 and 64 in the EB-ConA3), and 200 sequences were present in at least two of the fractions. Of these, a total of 21 N-linked glycopeptide sequences were identified in all the four lectin bound fractions from ES and EB.
The N-linked glycopeptide sequences detected in the lectin-bound samples from ES and EB cells were assigned to 180 glycoprotein-encoding genes which are shown in Supplementary Table 2. Most of the identified proteins were predicted to be processed through the post-translational N-linked glycosylation pathway and to be localized in extra-cellular or membrane associated subcellular compartments. Only 7 of these proteins had unknown subcellular locations (see below). When the complete sequences of the glycoproteins shown in Supplementary Table 2 were analyzed in silico for the presence of the consensus N-glycosylation sequon –N-X-S/T- (for practical purposes, the N-glycosylation sequons in each protein sequence were assigned regardless of their location in the lumenal, extracellular, transmembrane or cytoplasmic domains of the protein), most proteins had more than one potential N-glycosylation site. For example, 77 proteins possessed between 2 and 5 sequons, and 94 proteins had more than 5 N-glycosylation sequons. Only nine proteins in the data set reported in this study contained only one N-glycosylation sequon. However, when the sequences of the N-linked glycopeptides that were identified in ConA bound fractions from ES and EB cells were reviewed, the largest proportion of the glycoproteins (123) were represented by only one N-linked glycopeptide sequence (Supplementary Table 2, see below). There were 35 proteins that showed two N-linked glycosylated sequences, 20 proteins that gave between 3 and 4 glycopeptide sequences, and 4 proteins that gave more than 4 glycosylated sequences.
In this study, 3 samples for each ConA bound sample for ES and EB cells were analyzed by LC-MS/MS. As a result, 174, 161 and 198 N-glycosylated sequences were identified in Samples 1, 2 and 3 respectively (Supplementary Figure 1), when the distribution of identified glycopeptide sequences among the ConA bound samples for ES and EB stages was compared (Table 1), an average of 5% of these sequences were identified in the ES-ConA2 (range between 2 and 6%, with a standard deviation of 1.4); an average of 44% was identified in the ES-ConA3 (ranging from 35 to 55%, with a standard deviation of 8.6); an average of 32% was found in the EB-ConA2 fraction (ranging from 27 to 42%, with a standard deviation of 6.6); and average of 44% (ranging from 36 to58%, with a standard deviation of 10). These data suggest that the proportion of sequences found among the ConA2 and ConA3 fractions in the ES and EB stages is consistent between the three samples that were analyzed. Despite this consistency in the proportion of peptides in the ConA bound fractions in the three ES and EB samples, only 81 of the 293 identified sequences (28%) were present in all of the samples analyzed (where each sample represents the combination of sequences found in ES and EB cells, see Supplementary Figure 1), 134 sequences were present in only one sample (41, 35 and 58 in samples 1, 2 and 3 respectively, Supplementary Figure 1); and 78 sequences were present in two samples (19 in samples 1 and 2, 26 in samples 2 and 3 and 33 sequences in samples 1 and 3, see Supplementary Table 1). This lack of consistency in the sequences identified in the three samples is due to the fact that data acquisition during LC-MS/MS analysis of complex peptide mixtures never is comprehensive and the process is unable to collect tandem mass spectra from all eluting peptides. Peptide ions of low-abundance are often missed because (1) those ions are masked from the data-dependent acquisition process by more abundant ions, (2) the ions elute at the wrong point in the data-dependent acquisition cycle (e.g., during MS/ MS), and (3) chromatographic elution times are too short. Therefore, the acquisition of tandem mass spectra appears to be a “semi-random” process42 and often, several samples or repetitions are required to increase the number identified peptides or proteins. As a result, usually there are significant differences in the protein identifications when different samples of the same biological source are analyzed.
Table 1.
Distribution of identified glycopeptide sequences among the ConA bound samples for each of the three glycopeptide samples from ES and EB cells
| Total sequences | sequences in ES-2 | sequences in ES-3 | sequences in EB-2 | Sequences in EB-3 | |
|---|---|---|---|---|---|
| Repetition 1 | 174(100%) | 10(6%) | 97(55%) | 73(42%) | 102(58%) |
| Repetition 2 | 161(100%) | 4(2%) | 67(40%) | 47(28%) | 64(39%) |
| Repetition 3 | 198(100%) | 7(4%) | 70(35%) | 54(27%) | 71(36%) |
|
| |||||
| Average percentage | 4% | 44% | 32% | 44% | |
| Rel std. dev. | 1.4% | 8.6% | 6.6% | 10.0% | |
Characteristics of the identified ConA bound N-linked glycopeptides from ES and EB cells
The largest portion of the N-glycosylation sites that were identified in the ConA bound glycopeptide fractions from ES and EB cells in this study were represented by only one sequence in the data set (121 sequences). Only 25 protein identifications were made from one peptide identification with a single MS/MS spectrum Therefore, most of the single sequence identifications were derived from more than one spectrum (supplementary Tables 2 and 3). Moreover, the largest portion of these one-hit wonders (71 proteins) were obtained from four or more spectra, and many of these spectra were recorded in more than one of the ConA bound samples from both ES and EB stages that were analyzed by LC-MS/MS. Since our approach was to isolate glycopeptides rather than glycoproteins, it is not unlikely that a significant portion of the protein assignments would result from single peptide identification. To ensure that the most accurate dataset was presented, the peptide false discovery rate approach31 was utilized to remove peptide identifications resulting from potential random assignments. In addition, considering that the peptides reported in this study (Supplementary Table 3) were isolated by means of their posttranslational modifications (their carbohydrate moieties were bound by an immobilized lectin) and that most of them gave more than one spectra in more than one of the analyzed samples, the protein identifications derived from them were considered to be valid. Information of all the MS/MS spectra for the top scoring peptides (including those from the single hit proteins) is presented in Supplementary Table 3.
Of the glycoproteins found in the ConA bound fractions from ES and EB in this study, 59 of them were identified from 2 or more peptide sequences (Supplementary Table 2). However, there were several instances in which more than two sequences from one glycoprotein contained the same glycosylation site. Many of these peptides had overlapping sequences with different amino acid lengths that resulted from missed cleavages during the proteolytic treatment. Examples of these overlapping peptides are shown in the Table 2; one example is cubilin (intrinsic factor-cobalamin receptor, gi 82796190) a glycoprotein of 3591 amino acids with 42 N-glycosylation sequons (asn-X-Ser/thr). Ten glycosylated sequences from this protein were observed, distributed among the ConA bound fractions from ES and EB cells; however, two of the identified peptides contained the same glycosylation site (Asn781); one of these peptides was one amino acid longer due to a missed cleavage at Lys777. The second example, shown in Table 2, is Lamp-1 (Lysosomal associated membrane glycoprotein 1, gi: 7106339), a protein that contains 382 amino acid residues with 20 potential N-glycosylation sites. Analysis of ConA bound tryptic digests from both ES and EB resulted in the identification of 7 N-linked glycosylated sequences from Lamp-1. Two of these sequences contained glycosylation sites Asn248 and Asn252; however, because of missed cleavages in Lys241 and Arg245, one of these peptides possessed a longer sequence that also contained and additional glycosylation site at Asn240.
Table 2.
Examples of glycoproteins that were represented by more than one glycosylated sequence in the ConA bound fractions from ES and EB cells
| Glycopeptide sequencea | Mascot Score | ES | EB | ||
|---|---|---|---|---|---|
| CA2 | CA3 | CA2 | CA3 | ||
| gi 82796190: Cubilin (CUBN, intrinsic factor-cobalamin receptor) | |||||
| ICbGN781ETLFPIR | 55 | yes | yes | ||
| KICbGN781ETLFPIR | 66 | yes | yes | yes | |
| VLTESTGIIESPGHPNVYPSGVN957CbTWHIVVQR | 49 | yes | |||
| YCbGNSLPGN1819YSSIEGHNLWVR | 50 | yes | yes | ||
| FTSDGSVTGAGFN2085ASFQK | 128 | yes | yes | yes | |
| DFVEIWEN2400HTSGILLGR | 60 | yes | |||
| VN2531VTNEFK | 33 | yes | |||
| TFN2925SSTGDIVSPNFPK | 100 | yes | yes | ||
| FNDFEIVPSNLCbSHDYLEVFDGPSIGN3106R | 51 | yes | |||
| STN3125NSLTLLFK | yes | ||||
| gi 7106339: Lysosome-associated membrane glycoprotein 1 (LAMP-1) | |||||
| N70GSSCbGKEN78VSDPSLTITFGR | 75 | yes | yes | yes | |
| GYLLTLN97FTK | 35 | yes | yes | ||
| N159VTVVLR | 34 | yes | yes | yes | yes |
| DATIQAYLSSGN177FSK | 89 | yes | yes | ||
| AFN248ISPN252DTSSGSCbGINLVTLK | 88 | yes | yes | yes | yes |
| DN240KTVTRAFN248ISPN252DTSSGSCbGINLVTLK | 58 | yes | yes | ||
| LN296MTLPDALVPTFSISN310HSLK | 42 | yes | yes | ||
| gi 821389311: Intercellular adhesion molecule 1 (ICAM-1) | |||||
| EAFLPQGGSVQVN47CbSSSCK | 139 | yes | |||
| TELDLRPQGLALFSN204VSEAR | 63 | yes | yes | ||
| N388QTLELHVLYGPR | 77 | yes | |||
| LDETDCbLGN409WTWQEGSQQTLK | 128 | yes | |||
| QEMN456GTYVCbHAFSSHGN469VTR | 54 | yes | |||
| gi 63054837: Lysosome-associated membrane glycoprotein 2 (LAMP-2) | |||||
| CbNSVLTYN156LTPVVQK | 76 | yes | |||
| VPFIFNINPATTN265FTGSCbQPQSAQLR | 86 | yes | |||
| N322LSFWDAPLGSSYMCbNK | 56 | yes | |||
Position of N-linked glycosylation site is indicated after subtraction of signal peptide sequence
cysteine residue is carbamidomethylated
Table 2 shows four examples of glycoproteins that each yielded several glycopeptides in the lectin bound glycopeptides from ES and EB cells. The first two (cubilin and Lamp-1) were the glycoproteins that showed the largest number of glycopeptides in the analyzed data sets and were discussed above. The glycopeptides from both of these proteins were distributed in the different lectin bound fractions from both ES and EB cells; however, cubilin showed four glycopeptides that were present only in EB cells, indicating that glycosylation sites Asn2400, Asn2531, Asn3106 and Asn3125 possessed ConA binding glycans exclusively during this cell stage. All of the peptides from Lamp-1 were present in both cell stages. The third protein is Intercellular Adhesion Molecule 1 (ICAM-1, gi 821389311), a protein with 537 amino acids and 13 N-glycosylation sequons. ICAM-1 showed 6 N-glycosylation sites in 5 peptides in the ConA bound fractions, and four of them were found only in ES cells; only the peptide with Asn204 was identified in the fractions from both ES and EB cells. The fourth glycoprotein is Lysosome-Associated Membrane Glycoprotein 2 (gi 63054837, LAMP-2) a 415 residue protein with 17 potential N-glycosylation sites that showed 3 N-glycosylated sequences in the analyzed fractions, all of them found only in EB cells. The data described above indicate that the separation of glycopeptides from ES and EB cells with lectin affinity chromatography allowed the glycoproteomic identification of individual N-glycosylation sites of proteins that express specific types of glycan structures. The presence of these types of N-linked structures in glycopeptide sequences isolated only from ES or EB cells suggest that these glycan structures expressed at these sites may change as ES cells differentiate into EB.
Identification of specific glycopeptide sequences that expressed more than one N-linked glycan structure
Some of the N-glycosylated sequences identified in this study were found in both the ConA2 fraction (glycopeptides N-linked biantennary and hybrid structures) and the ConA3 fraction (glycopeptides with high mannose structures) in either ES or EB cells. For example, in cubilin, the peptides with Asn781 were identified in ConA2 and ConA3 fractions from both ES and EB cells, and the peptide with Asn2085 was found in the ConA2 and ConA3 fractions from EB cells. In LAMP-1, the peptide with N-glycosylation sites Asn70 and Asn78 was found in the ConA2 and ConA3 fractions from EB cells. The peptide that contains Asn159 and the peptides with N-glycosylation sites Asn248 and Asn252 were found in the ConA2 and ConA3 fractions from both ES and EB cells. These data indicate that the use of ConA allowed the identification of specific N-glycosylation sites that can express more than a single glycan structure and that in some instances, such as in the peptide with Asn2085 from cubilin and in the peptide with glycosylation sites Asn70 and Asn78 from LAMP-1, this micro heterogeneity may be cell stage specific.
Distribution of glycoproteins in the ConA bound fractions from ES and EB cells and identification of potential stage-specific glycobiomarkers
The proteins that were identified in the lectin-bound fractions were classified into three groups: the first group contained those that were present in both ES and EB fractions (117 proteins, 65% of the total number of proteins identified, Supplementary Table 2). Most of the proteins that yielded more than one glycosylated sequence (54 out of 59) were present in this group, including the proteins that gave the highest number of N-glycosylated sequences (cubilin and Lamp-1, Table 2). The second group comprised those proteins that were present exclusively in the ES cells (Table 3). This group contained 18 proteins (10%); most of them with only one identified N-glycosylated sequence (except Poliovirus receptor-related protein 2, which had two sequences). The third group, which comprised of 45 proteins (25%), was present only in the EB stage (Table 3); four of the proteins gave more than one N-linked glycopeptide sequence. Since the last two groups of ConA bound proteins were identified exclusively in only one of the cell stages (ES only or EB only, see Table 3 and Figure 3), these data suggest that there are distinct proteins in each cell stage that express specific N-glycan structures and, therefore, can be considered as potential differentiation stage-specific glycobiomarkers.
Table 3.
Glycoproteins found in the ConA bound fractions That were exclusive of ES or EB cell stages.
| Gi number | Mascot score | Protein ID | No of Sequences identified (Potential N-glycosylation sites) | Functional classification |
|---|---|---|---|---|
| Glycoproteins identified in ES cells Only | ||||
| 42558906 | 64 | CD97 antigen | 1(8) | cell membrane receptor |
| 118105 | 48 | peptidylprolyl isomerase A | 1(3) | enzyme |
| 31560574 | 53 | integrin alpha 5 | 1(14) | cell adhesion |
| 40556286 | 36 | endothelin converting enzyme 1 | 1(10) | enzyme |
| 31560781 | 110 | plexin domain containing 1 | 1(8) | cell membrane receptor |
| 14916479 | 33 | mannose-6-phosphate receptor, cation dependent | 1(5) | transporter |
| 31542362 | 39 | CD38 antigen | 1(4) | enzyme |
| 6678347 | 40 | thymus cell antigen 1, theta | 1(4) | cell adhesion |
| 94381789 | 100 | PREDICTED: similar to Poliovirus receptor-related protein 2 precursor (Murine herpesvirus entry protein B) (mHveB) (Nectin-2) (Poliovirus receptor homolog) (CD112 antigen) | 2(3) | cell membrane receptor |
| 37620147 | 46 | elastin microfibril interfacer 3 | 1(12) | other |
| 6680644 | 60 | a disintegrin and metalloprotease domain 9 (meltrin gamma) | 1(8) | enzyme |
| 6677897 | 73 | stromal cell derived factor receptor 1 | 1(7) | cell adhesion |
| 84875513 | 69 | Sel1 (suppressor of lin-12) 1 homolog isoform a | 1(6) | other |
| 9910138 | 43 | UDP-Gal:betaGlcNAc beta 1,3-galactosyltransferase, polypeptide 3 | 1(5) | enzyme |
| 33469043 | 31 | nicalin homolog | 1(2) | enzyme |
| 31981799 | 71 | SPFH domain family, member 1 | 1(1) | other |
| 30794452 | 48 | reticulocalbin 3 | 1(1) | cell membrane receptor |
| 6677905 | 66 | PREDICTED: similar to Golgi apparatus protein 1 precursor (Golgi sialoglycoprotein MG-160) (E-selectin ligand 1) (ESL-1) (Selel) | 1(5) | cell membrane receptor |
| Glycoproteins identified in EB cells Only | ||||
| 49274623 | 44 | semaphorin 4D | 1(8) | other |
| 74024915 | 56 | platelet/endothelial cell adhesion molecule 1 isoform 2 | 1(7) | cell adhesion |
| 46852189 | 73 | exostoses (multiple)-like 3 | 1(4) | enzyme |
| 41282044 | 67 | pantophysin isoform 2 | 1(4) | transporter |
| 19527236 | 48 | transmembrane emp24 protein transport domain containing 4 | 1(1) | transporter |
| 6679731 | 37 | coagulation factor V | 1(27) | other |
| 7549781 | 43 | odd Oz/ten-m homolog 4 | 1(18) | other |
| 63054837 | 75 | lysosomal membrane glycoprotein 2 isoform 1 | 3(17) | enzyme |
| 82958464 | 50 | PREDICTED: similar to Plexin-B2 precursor (MM1) isoform 21 | 1(16) | cell membrane receptor |
| 61656167 | 43 | sulfatase 2 | 1(13) | enzyme |
| 14389423 | 76 | scavenger receptor class B, member 1 | 1(11) | transporter |
| 6754622 | 114 | mannosidase 2, alpha B1 | 1(11) | enzyme |
| 31980636 | 40 | mannosidase, beta A, lysosomal | 1(10) | enzyme |
| 26986617 | 69 | sulfatase 1 | 1(10) | enzyme |
| 18702313 | 39 | protein tyrosine phosphatase, receptor type, F | 1(9) | enzyme |
| 19526900 | 37 | transmembrane protein 30A | 1(8) | other |
| 94406482 | 44 | PREDICTED: similar to Low-density lipoprotein receptor-related protein 5 precursor (LRP7) (Lr3) | 1(7) | cell membrane receptor |
| 94397735 | 85 | PREDICTED: similar to ceroid-lipofuscinosis, neuronal 5 | 1(7) | other |
| 6755112 | 65 | phospholipid transfer protein | 1(7) | cell adhesion |
| 45387933 | 76 | UDP-glucose ceramide glucosyltransferase-like 1 | 1(6) | enzyme |
| 45331202 | 72 | hyaluronidase 2 | 1(6) | enzyme |
| 31981425 | 86 | dipeptidylpeptidase 7 | 1(6) | enzyme |
| 30424573 | 62 | hypothetical protein LOC211499 | 1(6) | other |
| 7305299 | 81 | alpha-N-acetylglucosaminidase | 1(6) | enzyme |
| 82879262 | 76 | PREDICTED: GPI deacylase | 2(5) | enzyme |
| 9506985 | 31 | palmitoyl-protein thioesterase 2 | 1(5) | enzyme |
| 32189434 | 83 | immunoglobulin superfamily, member 8 | 1(4) | other |
| 13385482 | 107 | sarcoma amplified sequence | 1(4) | other |
| 6754186 | 119 | hexosaminidase B | 2(4) | enzyme |
| 6754098 | 93 | glucuronidase, beta | 1(4) | enzyme |
| 85701786 | 49 | glycosyltransferase 8 domain containing 3 | 1(3) | enzyme |
| 65301488 | 34 | D-glucuronyl C5-epimerase | 1(3) | enzyme |
| 41235747 | 48 | hypothetical protein LOC380967 | 1(3) | other |
| 30424569 | 60 | hypothetical protein LOC210035 | 1(3) | other |
| 28077083 | 82 | amnionless | 1(3) | cell membrane receptor |
| 94401936 | 49 | PREDICTED: similar to H-2 class I histocompatibility antigen, D-37 alpha chain precursor | 1(2) | cell membrane receptor |
| 75677587 | 44 | growth differentiation factor 3 | 1(2) | other |
| 31542965 | 39 | heparan sulfate 2-O-sulfotransferase 1 | 1(2) | enzyme |
| 15212492 | 106 | interferon gamma inducible protein 30 | 1(2) | enzyme |
| 6754970 | 41 | procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), alpha II polypeptide | 1(2) | enzyme |
| 6671678 | 89 | carbonic anhydrase 4 | 1(2) | enzyme |
| 94387741 | 79 | PREDICTED: similar to Cartilage-associated protein precursor | 1(1) | transporter |
| 31560607 | 46 | cathepsin C preproprotein | 1(3) | enzyme |
| 7949098 | 79 | neuronal pentraxin 2 | 1(3) | other |
| 6679451 | 75 | palmitoyl-protein thioesterase 1 | 2(3) | enzyme |
Figure 3. Distribution of the proteins identified in the present study according to cell stage or Con A bound fraction.
The proteins that were identified in the lectin bound fractions were classified in three groups according the cell stage in which they were present (ES only, EB only, or ES and EB or the lectin bound fraction (ConA2 only, ConA3 only, or ConA2 and ConA3). Arrows denote the number of proteins identified in the indicated fractions.
The glycoproteins described above were also classified in three groups, according to the ConA fraction in which they were identified. In the first group, 73 proteins (41% of the total number of proteins identified) were identified in both ConA2 and ConA3 fractions. In the second, 25 proteins (14%) were identified in the ConA2 fraction exclusively. In the third group 82 proteins (46%) were present only in the ConA3 fraction. When the glycoproteins that were classified in the ConA bound fractions were analyzed in the context of their distribution among the ES or EB cell stage groups (Figure 3), one half of the proteins that were exclusive to the ES cells were identified in the ConA2 glycopeptide fraction (9 out of 18 proteins), which is enriched in biantennary and hybrid structures. This proportion was larger than that of the proteins identified exclusively in the ConA3 fraction (8 proteins) or in both the ConA2 and ConA3 fractions (1 protein). The proteins that were exclusive to the EB stage, by contrast, were found predominantly in the ConA3 fraction (37 out of 45 proteins), which is enriched in high mannose structures. The proteins that were present in both ES and EB stages were found most frequently in both ConA2 and ConA3 fractions (69 out of 117 proteins). In summary, N-linked glycopeptides that were EB-specific were isolated mainly in the ConA3 fraction and a significantly larger proportion of the ES-specific glycopeptides were found only in the ConA2 fraction. These data indicate that some glycans are likely stage-specific.
Subcellular localization and physiological functions of the proteins from ES and EB cell stages
Information about the subcellular localization and physiological roles of proteins from ES and EB cells that were identified in the ConA bound fractions was obtained from databases and gene ontologies (http://www.ncbi.nlm.nih.gov/, and http://au.expasy.org/) or using bioinformatics tools at the Center for Biological Sequence Analysis, Technical University of Denmark (http://www.cbs.dtu.dk/), as well as the Ingenuity Pathways analysis software (http://www.ingenuity.com). Inspection of the predicted subcellular localization of each of the proteins identified in the data sets suggested that many are often present in more than one subcellular compartment (Supplementary Table 2). When the data on the subcellular localization of the proteins from all of the 180 proteins identified in the ConA bound fractions from ES or EB stages was examined (Figure 4A), these glycoproteins were identified more frequently as present in the extracellular space (73 hits), followed by the plasma membrane (64 hits), the endoplasmic reticulum (49 hits), lysosomes (38 hits), and the Golgi apparatus (21 hits). There was insufficient information in the databases or gene ontologies to determine the localization of 8 glycoproteins in the data set.
Figure 4. Predicted subcellular localization of the proteins identified in the ConA bound fractions from ES or EB stages.
(A) subcellular distribution of all 180 ConA bound proteins identified in ES or EB stages. (B) subcellular distribution of the 116 proteins that were shared by both ES and EB stages. (C) subcellular distribution of the 17 proteins identified only in the ES stage. (D) subcellular distribution of the 47 proteins identified only in the EB stage
When the subcellular localization of the 117 proteins that were shared by both ES and EB stages was examined (Figure 4B), the relative proportion of the proteins in each of the subcellular locations was similar to that of the whole data set (Figure 4A). However, this proportion was substantially different for the proteins that were present exclusively in either of the two cell stages. For example, the 18 proteins that were present only in the ES fraction (Figure 4C) are more frequently expressed in the plasma membrane (9 hits), followed by the extracellular space (5hits), the endoplasmic reticulum (4 hits), the Golgi apparatus (2 hits) and the lysosomes (1 hit), with 1 protein of unknown localization. On the other hand, the proteins identified in the EB stage only (45 proteins, Figure 4D), were more abundant in the extracellular space (20 hits), followed by the lysosomes (10 hits), endoplasmic reticulum (9 hits), plasma membrane (9 hits) and Golgi apparatus (7 hits). Three proteins in this set could not be assigned to a subcellular location. These data suggest that there are differences in the predicted subcellular localization of the glycoproteins that express high mannose, hybrid or complex biantennary glycan structures that are cell stage specific.
The glycoproteins identified from the ConA bound glycopeptides were also classified into several categories based on their predicted physiological role (Table 4). If this classification were applied to the complete data set of 180 glycoproteins identified in the ConA bound fractions from ES and EB, the categories in which the identified proteins were most frequently found were: cell adhesion (24 proteins), enzymes (72 proteins), cell membrane receptors (26 proteins), transporters (16 proteins), and ion channels (6 proteins). There were 38 proteins that were classified as “other” because either there was insufficient information to classify them into a defined functional category, or they had been classified into a protein type of which only a small number of members (less than three hits) were identified in the lectin bound samples from ES or EB.
Table 4.
Classification of glycoproteins identified in ConA bound fractions from ES and EB cells according to function, the portion of proteins that were assigned to a particular category is indicated as percentage of the number of protein hits in the fraction. The number of proteins in the functional category is indicated between the parentheses. The gene names of the identified proteins are listed in italics.
| Functional category | ES only (18 proteins) |
EB only (45 proteins) |
ES and EB (117 proteins) |
Total (180 proteins) |
|---|---|---|---|---|
| percentage of proteins identified in lectin bound fraction (No of proteins of functional category that were identified in fraction) Gene name of identified proteins |
||||
| cell adhesion | 17% (3) ITGA5, NPTN, THY1 |
4% (2) PECAM1, PLTP |
16% (19) APLP2, BSG, CADM1, COL18A1, EMB, EPDR1, FAT, HSPG2 (includes EG:3339), ITGA3, ITGA6, ITGAV, LAMA1, LAMA5, LAMB1, LAMC1, MCAM, NID2, TACSTD1, VTN |
13% (24) |
| enzymes | 33% (6) PPIA, B3GALNT1, CD38, ADAM9, ECE1, NCLN, |
51% (23) HEXB, MAN2B1, IFI30, GUSB, CA4, NAGLU, PGAP1, UGCGL1, LAMP2, PPT1, EXTL3, HYAL2, SULF1, GLT8D3, SULF2, P4HA2, MANBA, HS2ST1, GLCE, PPT2, DPP7, CTSC, PTPRF |
37% (43) ALPL, ANPEP, ASAH1, C5ORF14, CTSA, CTSD, CTSF, CTSL2, CTSZ (includes EG:1522), DPP4, EDEM3, ENPP3, ERO1L, FKBP10, FKBP9, FN1, GAA, GALNT1, GBA, GGH, GLA, GLT25D1, HEXA, IMPAD1, LIPA, LNPEP, LYPLA3, MINPP1, NCSTN, P4HA1, PCYOX1, PIGS, PLOD1, PLOD2, PLOD3, PSAP, PTK7, SIAE, SMPDL3B, STT3A, STT3B, TPP1, TXNDC10, |
40% (72) |
| cell membrane receptors | 28% (5) GLG1, PLXDC1, RCN3, PVRL2, CD97 |
9% (4) AMN, LRP5, PLXNB2, HLA-E |
15% (17) CD276, CUBN, HSP90B1, LAMP1, LOC196463, LY75, PTGFRN, SCARB2, CEACAM1, CNTFR, ICAM1, IGF2R, ITGB1, LRP1, LRPAP1, PLXNA1, MPZL1 |
14% (26) |
| transporters | 6% (1) M6PR |
9% (4) CRTAP, SCARB1, SYPL1, TMED4 |
19% (11) LRP2, NPC1, NUP210, SLC2A1, SLC2A3, SLC3A2, SORL1, SORT1, TFRC, TM9SF3, TMED9 |
9% (16) |
| ion channels | N. I. | N. I. | 5% (6) ATP1B1, ATP1B2, ATP1B3, ATP6AP1, ATP6V0E1, SLC12A7 |
3% (6) |
| other | 17% (3) MMRN2, SEL1L, ERLIN1 |
27% (12) GDF3, TSPAN31, CLN5, IGSF8, NPTX2, TMEM87A, KIAA0286, TMEM106C, SEMA4D, ODZ4 (includes EG:26011), TMEM30A, F5 |
18% (21) GRN, CLPTM1, NOMO1, C20ORF3, TMEM106B, TOR1AIP2, SUMF1, TSPAN13, SSR2, HYOU1, STCH, PTTG1IP, SSR1, KIAA0090, SPARC, KTELC1, TOR2A, GOLM1, 4932417I16RIK, CALU (includes EG:813), CALU (includes EG:813) |
20% (36) |
The proteins that were identified in only one cell stage (ES only or EB only, Table 3) were analyzed in the context of the functional category groups in which they were classified. The resulting data indicated that cell adhesion proteins represented 17% of the proteins found in ES cells only, by contrast to 4% of those found exclusively in EB cells; enzymes represented 33% and 51% of the hits identified exclusively in ES and EB cell stages, respectively, cell membrane receptors 15% and 9%, transporters 11% and 9%, and other proteins 18% and 27% respectively. These data suggest a possible connection between the N-linked glycan structures of the proteins that were identified in only one cell stage with the specific function of these proteins.
4. Discussion
The aim of this study was to develop a protocol for the identification of potential glycoprotein biomarkers that express specific N-linked glycans in specific glycosylation sites at defined developmental stages during embryonic stem cell differentiation. Lectin affinity chromatography of tryptic peptides extracted from cell pellets from the different cell stages was performed, allowing the isolation and identification of peptides with specific N-glycosylation sites. This protocol is in contrast to the lectin affinity fractionation of intact (non-trypsinized) glycoproteins that is often utilized 9, 10, 16, 43-51. In the latter approach, there is often found a substantial number of proteins that do not carry the glycan epitopes recognized by the lectin used for separation, but are present in the lectin-bound fractions because they form oligomeric complexes or aggregate with the glycoproteins that express the glycans that are directly bound by the lectin. These specific or non-specific associations often result in the identification of cytoplasmic or nuclear proteins of high abundance that are not predicted to be N-glycosylated 48.
Previous reports on the use of lectin affinity for the identification of biomarkers in tissues or cells obtained from different sources such as diseased vs. non-diseased models, genetically modified organisms, different developmental stages etc. have focused on the use of lectin affinity for the isolation of glycoproteins with glycan structures that are specific to a defined stage, developmental or physiological condition 8, 16, 45. The present study, by contrast, focused on the isolation of glycopeptides that expressed N-linked high-mannose, hybrid and complex biantennary structures abundant in both ES and EB stages (Figure 1) as a means to identify glycoproteins that express these structures at specific sites by proteomics techniques. Despite the abundance of these three types of structures in both cell stages, they were not expressed by the same set of proteins in ES and EB cells (Figure 2). For example, there were 40 sequences (out of 293) identified exclusively in the ES stage (24 sequences in the ConA2 and 16 in the ConA3); and 87 sequences that were identified only in the EB stage (16 in the ConA2, 64 in the ConA3; and 7 in both ConA2 and ConA3). The results presented in Figure 3 show that a significant number of glycopeptides extracted from ES or EB cells was bound by ConA. These glycopeptides represent a total of 180 glycoproteins, many of which express multiple glycosylation sites. Many of these glycoproteins (117, 65%) were found in the ConA bound fractions from both ES and EB cells. There were, however, 18 glycoproteins that were found exclusively in the ConA bound glycopeptides from ES cells, and 45 proteins that were found exclusively in the fractions from EB (Table 3). These results show that despite the high abundance of the high mannose, hybrid and complex N-linked glycan structures in both cell stages, there were sets of glycoproteins which contained these structures that were present only in one cell stage. The fact that sequences with specific glycan structures were identified in only cell stage (Table 3) suggests that these glycoproteins can be considered as potential stage-specific glycobiomarkers.
It is not known whether the proteins to which these sequences are attached were expressed exclusively at a defined developmental stage, or were expressed in both cell stages but carried the ConA-bound lectin in only one of them. Therefore, In order to verify further the stage specific biomarker candidates isolated by ConA from ES and EB cells in this study, other analytical techniques need to be applied. For example, these candidates can be detected and possibly quantified by immunodetection techniques 52, 53, or multiple reaction monitoring (MRM) LC-MS/MS experiments using tryptic peptide extracts from the ES and EB cell lines 54, 55. These techniques should allow us to distinguish whether the glycoproteins are expressed exclusively in once cell stage, or are expressed in both cell stages, but carry the targeted glycan structure in a specific cell stage.
The present study used lectin affinity chromatography on glycopeptide extracts from cultured cells to search for potential glycobiomarkers of stem cell differentiation. These glycobiomarkers can be proteins that are expressed exclusively in a cell stage or glycoproteins that are expressed in multiple cell stage, but carry a defined oligosaccharide structure during a specific developmental stage. Our results (Figure 1) show that N-linked oligosaccharides that bind to ConA (high mannose, hybrid and complex biantennary) are abundant in pluripotent (ES) stem cells and in embroid bodies (EB). Lectin affinity chromatography allowed the isolation of glycopeptides from glycoproteins that express these oligosaccharides in only one of the cell stages (ES or EB). Therefore, the results presented in this study indicate that the glycoproteomic strategy presented here can serve as a basis for identification of potential glycobiomarkers, which are, either the glycopeptides themselves, or the glycoproteins from which they are derived, that are cell-type specific.
Our results show a possible connection between the glycosylation of specific sites on glycoproteins and their putative function. For example, ConA bound peptides expressed exclusively in ES cells were derived from proteins whose functions were more frequently classified as enzymes, cell membrane receptors and cell adhesion molecules (33, 28 and 17% respectively). However, in the proteins identified exclusively in EB cells, the proportion of cell adhesion molecules and cell membrane receptors decreased drastically to 4% and 9%, respectively, and that of enzymes increased to 51%. These changes likely reflect the fact that ES cells are pluripotent and actively dividing and, therefore, are required to up-regulate cell-cell interactions for potential differentiation signals.
The physiological relevance of the specific glycans observed on these potential biomarkers is largely unknown, and for most of the proteins identified, specific experiments will be required to address these issues. Information on the significance of the glycosylation of some of the glycoproteins identified in this study is available, however. One example is the CD97 antigen identified exclusively in the ES cell stage in the three experimental replicates analyzed from this sample. CD97 is a G-protein coupled receptor potentially involved in both adhesion and signaling processes, plays an essential role in leukocyte migration56 and might be a differentiation marker for several types of carcinomas57. There are recent reports that N-linked glycosylation of this receptor may be essential for epitope binding58. Another example of a glycoprotein expressed exclusively in the ES fraction is the Golgi sialoglycoprotein MG-160 also known as E-selectin ligand 1 (ESL-1), which was detected in the three experimental replicates analyzed. This protein is a Single-pass type I membrane protein that has been identified as ligant for E-selectin, which is a cell-adhesion lectin on endothelial cells that mediates the binding of neutrophils59 Glycosylation on this protein is relevant since it has been demonstrated that ESL-1 requires N-linked carbohydrates for binding to E-selectin 60One more example is the cation dependent mannose 6-phosphate receptor (M6PR) that was also detected in two replicates only in the ES stage. M6PR is important in intracellular protein sorting 61 and the presence of this protein in the ES only fractions may indicate an up-regulation that may important, considering that pluripotent ES cells cultured for this study were actively dividing and would require up-regulation of lysosomal enzymes. One last example is the ADP-ribosyl cyclase 1 (CD38), which was also detected in two experimental replicates exclusively in ES cells. CD38 synthesizes cyclic ADP-ribose, a second messenger that regulates intracellular calcium which is important for glucose-induced insulin secretion 62 and requires N-linked for the stabilization of its structure in the cell membrane 63.
Our results suggest that specific oligosaccharide structures are expressed by different sets of glycoproteins at different developmental stages. For this reason, glycoproteomic analysis of the ConA unbound (ConA1) fractions with other lectins using serial lectin affinity approaches 29, 37 will likely result in the identification of additional stage specific glycobiomarkers.
Supplementary Material
Acknowledgments
We thank Dr. Will York for his critical review of the manuscript. This work was supported by funds from the National Center for Research Resources/NIH P41RR 018502.
Abbreviations
- ES
embryonic stem cells
- EB
embroid bodies
- PNGase F
peptide N-glycosidase F
- LC-MS/MS
Liquid chromatography coupled to tandem mass spectrometry
- a. m. u.
atomic mass units
- MALDI-TOF-MS
Matrix assisted laser desorption- time of flight mass spectrometry
- ConA
Concanavalin A
- PEP-FDR
peptide false-discovery rates
References
- 1.Budnik BA, Lee RS, Steen JAJ. Global methods for protein glycosylation analysis by mass spectrometry. Biochimica et Biophysica Acta (BBA) - Proteins & Proteomics. 2006;1764(12):1870–1880. doi: 10.1016/j.bbapap.2006.10.005. [DOI] [PubMed] [Google Scholar]
- 2.Ireland BS, Brockmeier U, Howe CM, Elliott T, Williams DB. Lectin-deficient Calreticulin Retains Full Functionality as a Chaperone for Class I Histocompatibility Molecules. Molecular biology of the cell. 2008;19(6):2413–23. doi: 10.1091/mbc.E07-10-1055. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 3.Drickamer K, Taylor ME. Evolving views of protein glycosylation. Trends Biochem Sci. 1998;23(9):321–4. doi: 10.1016/s0968-0004(98)01246-8. [DOI] [PubMed] [Google Scholar]
- 4.Dwek RA. Glycobiology: “towards understanding the function of sugars”. Biochem Soc Trans. 1995;23(1):1–25. doi: 10.1042/bst0230001. [DOI] [PubMed] [Google Scholar]
- 5.Hemmerich S. Glycomics: coming of age across the globe. Drug discovery today. 2005;10(5):307–9. doi: 10.1016/S1359-6446(04)03359-8. [DOI] [PubMed] [Google Scholar]
- 6.Lubner GC. Glycomics: an innovative branch of science. Boll Chim Farm. 2003;142(2):50. [PubMed] [Google Scholar]
- 7.Ludwig JA, Weinstein JN. Biomarkers in cancer staging, prognosis and treatment selection. Nature reviews. 2005;5(11):845–56. doi: 10.1038/nrc1739. [DOI] [PubMed] [Google Scholar]
- 8.Kim YS, Hwang SY, Kang HY, Sohn H, Oh S, Kim JY, Yoo JS, Kim YH, Kim CH, Jeon JH, Lee JM, Kang HA, Miyoshi E, Taniguchi N, Yoo HS, Ko JH. Functional proteomic study reveals that N-acetylglucosaminyltransferase V reinforces the invasive/metastatic potential of colon cancer through aberrant glycosylation on TIMP-1. Mol Cell Proteomics. 2007 doi: 10.1074/mcp.M700084-MCP200. [DOI] [PubMed] [Google Scholar]
- 9.Novotny MV, Mechref Y. New hyphenated methodologies in high-sensitivity glycoprotein analysis. J Sep Sci. 2005;28(15):1956–68. doi: 10.1002/jssc.200500258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Drake RR, Schwegler EE, Malik G, Diaz J, Block T, Mehta A, Semmes OJ. Lectin capture strategies combined with mass spectrometry for the discovery of serum glycoprotein biomarkers. Mol Cell Proteomics. 2006;5(10):1957–67. doi: 10.1074/mcp.M600176-MCP200. [DOI] [PubMed] [Google Scholar]
- 11.Zhao J, Qiu W, Simeone DM, Lubman DM. N-linked glycosylation profiling of pancreatic cancer serum using capillary liquid phase separation coupled with mass spectrometric analysis. J Proteome Res. 2007;6(3):1126–38. doi: 10.1021/pr0604458. [DOI] [PubMed] [Google Scholar]
- 12.Kaji H, Saito H, Yamauchi Y, Shinkawa T, Taoka M, Hirabayashi J, Kasai K, Takahashi N, Isobe T. Lectin affinity capture, isotope-coded tagging and mass spectrometry to identify N-linked glycoproteins. Nature biotechnology. 2003;21(6):667–72. doi: 10.1038/nbt829. [DOI] [PubMed] [Google Scholar]
- 13.Bunkenborg J, Pilch BJ, Podtelejnikov AV, Wisniewski JR. Screening for N-glycosylated proteins by liquid chromatography mass spectrometry. Proteomics. 2004;4(2):454–65. doi: 10.1002/pmic.200300556. [DOI] [PubMed] [Google Scholar]
- 14.Xiong L, Andrews D, Regnier F. Comparative proteomics of glycoproteins based on lectin selection and isotope coding. J Proteome Res. 2003;2(6):618–25. doi: 10.1021/pr0340274. [DOI] [PubMed] [Google Scholar]
- 15.Atwood JA, 3rd, Minning T, Ludolf F, Nuccio A, Weatherly DB, Alvarez-Manilla G, Tarleton R, Orlando R. Glycoproteomics of Trypanosoma cruzi trypomastigotes using subcellular fractionation, lectin affinity, and stable isotope labeling. J Proteome Res. 2006;5(12):3376–84. doi: 10.1021/pr060364b. [DOI] [PubMed] [Google Scholar]
- 16.Norton PA, Comunale MA, Krakover J, Rodemich L, Pirog N, D'Amelio A, Philip R, Mehta AS, Block TM. N-linked glycosylation of the liver cancer biomarker GP73. J Cell Biochem. 2007 doi: 10.1002/jcb.21610. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Siddiqui SF, Pawelek J, Handerson T, Lin CY, Dickson RB, Rimm DL, Camp RL. Coexpression of beta1,6-N-acetylglucosaminyltransferase V glycoprotein substrates defines aggressive breast cancers with poor outcome. Cancer Epidemiol Biomarkers Prev. 2005;14(11 Pt 1):2517–23. doi: 10.1158/1055-9965.EPI-05-0464. [DOI] [PubMed] [Google Scholar]
- 18.Ishibashi Y, Dosaka-Akita H, Miyoshi E, Shindoh M, Miyamoto M, Kinoshita I, Miyazaki H, Itoh T, Kondo S, Nishimura M, Taniguchi N. Expression of N-acetylglucosaminyltransferase V in the development of human esophageal cancers: immunohistochemical data from carcinomas and nearby noncancerous lesions. Oncology. 2005;69(4):301–10. doi: 10.1159/000089680. [DOI] [PubMed] [Google Scholar]
- 19.Nash R, Neves L, Faast R, Pierce M, Dalton S. The lectin Dolichos biflorus agglutinin recognizes glycan epitopes on the surface of murine embryonic stem cells: a new tool for characterizing pluripotent cells and early differentiation. Stem cells (Dayton, Ohio) 2007;25(4):974–82. doi: 10.1634/stemcells.2006-0224. [DOI] [PubMed] [Google Scholar]
- 20.Conley BJ, Young JC, Trounson AO, Mollard R. Derivation, propagation and differentiation of human embryonic stem cells. The international journal of biochemistry & cell biology. 2004;36(4):555–67. doi: 10.1016/j.biocel.2003.07.003. [DOI] [PubMed] [Google Scholar]
- 21.He X, Gonzalez V, Tsang A, Thompson J, Tsang TC, Harris DT. Differential gene expression profiling of CD34+ CD133+ umbilical cord blood hematopoietic stem progenitor cells. Stem Cells Dev. 2005;14(2):188–98. doi: 10.1089/scd.2005.14.188. [DOI] [PubMed] [Google Scholar]
- 22.Hemmoranta H, Satomaa T, Blomqvist M, Heiskanen A, Aitio O, Saarinen J, Natunen J, Partanen J, Laine J, Jaatinen T. N-glycan structures and associated gene expression reflect the characteristic N-glycosylation pattern of human hematopoietic stem and progenitor cells. Exp Hematol. 2007;35(8):1279–92. doi: 10.1016/j.exphem.2007.05.006. [DOI] [PubMed] [Google Scholar]
- 23.Nairn AV, York WS, Harris K, Hall EM, Pierce JM, Moremen KW. Regulation of Glycan Structures in Animal Tissues: Transcript Profiling of Glycan-Related Genes. The Journal of biological chemistry. 2008;283(25):17298–313. doi: 10.1074/jbc.M801964200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Atwood JA, 3rd, Cheng L, Alvarez-Manilla G, Warren NL, York WS, Orlando R. Quantitation by isobaric labeling: applications to glycomics. J Proteome Res. 2008;7(1):367–74. doi: 10.1021/pr070476i. [DOI] [PubMed] [Google Scholar]
- 25.Hadjantonakis AK, Gertsenstein M, Ikawa M, Okabe M, Nagy A. Generating green fluorescent mice by germline transmission of green fluorescent ES cells. Mechanisms of development. 1998;76(1-2):79–90. doi: 10.1016/s0925-4773(98)00093-8. [DOI] [PubMed] [Google Scholar]
- 26.Stead E, White J, Faast R, Conn S, Goldstone S, Rathjen J, Dhingra U, Rathjen P, Walker D, Dalton S. Pluripotent cell division cycles are driven by ectopic Cdk2, cyclin A/E and E2F activities. Oncogene. 2002;21(54):8320–33. doi: 10.1038/sj.onc.1206015. [DOI] [PubMed] [Google Scholar]
- 27.Lake J, Rathjen J, Remiszewski J, Rathjen PD. Reversible programming of pluripotent cell differentiation. Journal of cell science. 2000;113(Pt 3):555–66. doi: 10.1242/jcs.113.3.555. [DOI] [PubMed] [Google Scholar]
- 28.Svennerholm L, Fredman P. A procedure for the quantitative isolation of brain gangliosides. Biochimica et biophysica acta. 1980;617(1):97–109. doi: 10.1016/0005-2760(80)90227-1. [DOI] [PubMed] [Google Scholar]
- 29.Cummings RD, Kornfeld S. Fractionation of asparagine-linked oligosaccharides by serial lectin-Agarose affinity chromatography. A rapid, sensitive, and specific technique. The Journal of biological chemistry. 1982;257(19):11235–40. [PubMed] [Google Scholar]
- 30.Weatherly DB, Atwood JA, 3rd, Minning TA, Cavola C, Tarleton RL, Orlando R. A Heuristic method for assigning a false-discovery rate for protein identifications from Mascot database search results. Mol Cell Proteomics. 2005;4(6):762–72. doi: 10.1074/mcp.M400215-MCP200. [DOI] [PubMed] [Google Scholar]
- 31.Elias JE, Gygi SP. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat Methods. 2007;4(3):207–14. doi: 10.1038/nmeth1019. [DOI] [PubMed] [Google Scholar]
- 32.Ciucanu I, Kerek F. A simple and rapid method for the permethylation of carbohydrates. Carbohydrate Research. 1984;131(2):209–217. [Google Scholar]
- 33.Dell A, Reason AJ, Khoo KH, Panico M, McDowell RA, Morris HR. Mass spectrometry of carbohydrate-containing biopolymers. Methods in enzymology. 1994;230:108–32. doi: 10.1016/0076-6879(94)30010-0. [DOI] [PubMed] [Google Scholar]
- 34.Kang P, Mechref Y, Klouckova I, Novotny MV. Solid-phase permethylation of glycans for mass spectrometric analysis. Rapid Commun Mass Spectrom. 2005;19(23):3421–8. doi: 10.1002/rcm.2210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Krusius T, Finne J, Rauvala H. The structural basis of the different affinities of two types of acidic N-glycosidic glycopeptides for concanavalin A--sepharose. FEBS letters. 1976;72(1):117–20. doi: 10.1016/0014-5793(76)80911-8. [DOI] [PubMed] [Google Scholar]
- 36.Yamamoto K, Tsuji T, Osawa T. Analysis of Asparagine-Linked Oligosaccharides by Sequential Lectin Affinity-Chromatography. Molecular Biotechnology. 1995;3(1):25–36. doi: 10.1007/BF02821332. [DOI] [PubMed] [Google Scholar]
- 37.Merkle RK, Cummings RD. Lectin affinity chromatography of glycopeptides. Methods in enzymology. 1987;138:232–59. doi: 10.1016/0076-6879(87)38020-6. [DOI] [PubMed] [Google Scholar]
- 38.Varki A, Cummings RD, Esko JD, Freeze HH, Stanley P, Bertozzi CR, Hart GW. Essentials of Glycobiology. Cold Spring Harbor Laboratory Press; Plainview (NY): 2008. p. 784. [PubMed] [Google Scholar]
- 39.Osawa T, Tsuji T. Fractionation and Structural Assessment of Oligosaccharides and Glycopeptides by Use of Immobilized Lectins. Annual Review of Biochemistry. 1987;56(1):21–40. doi: 10.1146/annurev.bi.56.070187.000321. [DOI] [PubMed] [Google Scholar]
- 40.Aoki K, Perlman M, Lim JM, Cantu R, Wells L, Tiemeyer M. Dynamic developmental elaboration of N-linked glycan complexity in the Drosophila melanogaster embryo. The Journal of biological chemistry. 2007;282(12):9127–42. doi: 10.1074/jbc.M606711200. [DOI] [PubMed] [Google Scholar]
- 41.Cheng L, Atwood JAI, Alvarez-Manilla G, Warren NL, York W, Orlando R. Quantitation by Isobaric Labeling: Applications to Glycomics. J Proteome Res. 2007 doi: 10.1021/pr070476i. In Press. [DOI] [PubMed] [Google Scholar]
- 42.Liu H, Sadygov RG, Yates JR., 3rd A model for random sampling and estimation of relative protein abundance in shotgun proteomics. Anal Chem. 2004;76(14):4193–201. doi: 10.1021/ac0498563. [DOI] [PubMed] [Google Scholar]
- 43.Orazine CI, Hincapie M, Hancock WS, Hattersley M, Hanke JH. A proteomic analysis of the plasma glycoproteins of a MCF-7 mouse xenograft: a model system for the detection of tumor markers. J Proteome Res. 2008;7(4):1542–54. doi: 10.1021/pr7008516. [DOI] [PubMed] [Google Scholar]
- 44.Plavina T, Wakshull E, Hancock WS, Hincapie M. Combination of abundant protein depletion and multi-lectin affinity chromatography (M-LAC) for plasma protein biomarker discovery. J Proteome Res. 2007;6(2):662–71. doi: 10.1021/pr060413k. [DOI] [PubMed] [Google Scholar]
- 45.Block TM, Comunale MA, Lowman M, Steel LF, Romano PR, Fimmel C, Tennant BC, London WT, Evans AA, Blumberg BS, Dwek RA, Mattu TS, Mehta AS. Proceedings of the National Academy of Sciences of the United States of America. 3. Vol. 102. 2005. Use of targeted glycoproteomics to identify serum glycoproteins that correlate with liver cancer in woodchucks and humans; pp. 779–84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Madera M, Mechref Y, Klouckova I, Novotny MV. Semiautomated high-sensitivity profiling of human blood serum glycoproteins through lectin preconcentration and multidimensional chromatography/tandem mass spectrometry. J Proteome Res. 2006;5(9):2348–63. doi: 10.1021/pr060169x. [DOI] [PubMed] [Google Scholar]
- 47.Madera M, Mechref Y, Novotny MV. Combining lectin microcolumns with high-resolution separation techniques for enrichment of glycoproteins and glycopeptides. Anal Chem. 2005;77(13):4081–90. doi: 10.1021/ac050222l. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Madera M, Mechref Y, Klouckova L, Novotny MV. High-sensitivity profiling of glycoproteins from human blood serum through multiple-lectin affinity chromatography and liquid chromatography/tandem mass spectrometry. Journal of Chromatography B-Analytical Technologies in the Biomedical and Life Sciences. 2007;845(1):121–137. doi: 10.1016/j.jchromb.2006.07.067. [DOI] [PubMed] [Google Scholar]
- 49.Wang Y, Ao X, Vuong H, Konanur M, Miller FR, Goodison S, Lubman DM. Membrane Glycoproteins Associated with Breast Tumor Cell Progression Identified by a Lectin Affinity Approach. J Proteome Res. 2008 doi: 10.1021/pr8002547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Qiu Y, Patwa TH, Xu L, Shedden K, Misek DE, Tuck M, Jin G, Ruffin MT, Turgeon DK, Synal S, Bresalier R, Marcon N, Brenner DE, Lubman DM. Plasma glycoprotein profiling for colorectal cancer biomarker identification by lectin glycoarray and lectin blot. J Proteome Res. 2008;7(4):1693–703. doi: 10.1021/pr700706s. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kreunin P, Zhao J, Rosser C, Urquidi V, Lubman DM, Goodison S. Bladder cancer associated glycoprotein signatures revealed by urinary proteomic profiling. J Proteome Res. 2007;6(7):2631–9. doi: 10.1021/pr0700807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Guo HB, Lee I, Bryan BT, Pierce M. Deletion of mouse embryo fibroblast N-acetylglucosaminyltransferase V stimulates alpha5beta1 integrin expression mediated by the protein kinase C signaling pathway. The Journal of biological chemistry. 2005;280(9):8332–42. doi: 10.1074/jbc.M413532200. [DOI] [PubMed] [Google Scholar]
- 53.Guo HB, Lee I, Kamar M, Pierce M. N-acetylglucosaminyltransferase V expression levels regulate cadherin-associated homotypic cell-cell adhesion and intracellular signaling pathways. The Journal of biological chemistry. 2003;278(52):52412–24. doi: 10.1074/jbc.M308837200. [DOI] [PubMed] [Google Scholar]
- 54.Hulsmeier AJ, Paesold-Burda P, Hennet T. N-glycosylation site occupancy in serum glycoproteins using multiple reaction monitoring liquid chromatography-mass spectrometry. Mol Cell Proteomics. 2007;6(12):2132–8. doi: 10.1074/mcp.M700361-MCP200. [DOI] [PubMed] [Google Scholar]
- 55.Stahl-Zeng J, Lange V, Ossola R, Eckhardt K, Krek W, Aebersold R, Domon B. High sensitivity detection of plasma proteins by multiple reaction monitoring of N-glycosites. Mol Cell Proteomics. 2007;6(10):1809–17. doi: 10.1074/mcp.M700132-MCP200. [DOI] [PubMed] [Google Scholar]
- 56.Leemans JC, te Velde AA, Florquin S, Bennink RJ, de Bruin K, van Lier RAW, van der Poll T, Hamann J. The Epidermal Growth Factor-Seven Transmembrane (EGF-TM7) Receptor CD97 Is Required for Neutrophil Migration and Host Defense. J Immunol. 2004;172(2):1125–1131. doi: 10.4049/jimmunol.172.2.1125. [DOI] [PubMed] [Google Scholar]
- 57.Aust G, Eichler W, Laue S, Lehmann I, Heldin NE, Lotz O, Scherbaum WA, Dralle H, Hoang-Vu C. CD97: A Dedifferentiation Marker in Human Thyroid Carcinomas. Cancer Res. 1997;57(9):1798–1806. [PubMed] [Google Scholar]
- 58.Wobus Manja, V B, Schmücking Eike, Hamann Jörg, Aust Gabriela. N-glycosylation of CD97 within the EGF domains is crucial for epitope accessibility in normal and malignant cells as well as CD55 ligand binding. International Journal of Cancer. 2004;112(5):815–822. doi: 10.1002/ijc.20483. [DOI] [PubMed] [Google Scholar]
- 59.Willmroth F, Beaudet AL. Structure of the murine E-selectin ligand 1 (ESL-1) gene and assignment to Chromosome 8. Mammalian Genome. 1999;10(11):1085–1088. doi: 10.1007/s003359901166. [DOI] [PubMed] [Google Scholar]
- 60.Vesweber D, Blanks JE. Mechanisms That Regulate the Function of the Selectins and Their Ligands. Physiol Rev. 1999;79(1):181–213. doi: 10.1152/physrev.1999.79.1.181. [DOI] [PubMed] [Google Scholar]
- 61.Hille-Rehfeld A. Mannose 6-phosphate receptors in sorting and transport of lysosomal enzymes. Biochimica et biophysica acta. 1995;1241(2):177–94. doi: 10.1016/0304-4157(95)00004-b. [DOI] [PubMed] [Google Scholar]
- 62.Malavasi F, Deaglio S, Funaro A, Ferrero E, Horenstein AL, Ortolan E, Vaisitti T, Aydin S. Evolution and Function of the ADP Ribosyl Cyclase/CD38 Gene Family in Physiology and Pathology. Physiol Rev. 2008;88(3):841–886. doi: 10.1152/physrev.00035.2007. [DOI] [PubMed] [Google Scholar]
- 63.Gao Y, Mehta K. N-linked glycosylation of CD38 is required for its structure stabilization but not for membrane localization. Mol Cell Biochem. 2007;295(1-2):1–7. doi: 10.1007/s11010-006-9265-9. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.





