Abstract
The database of topographic mapping of Single Nucleotide Polymorphism (topoSNP) provides an online resource for analyzing non-synonymous SNPs (nsSNPs) that can be mapped onto known 3D structures of proteins. These include disease- associated nsSNPs derived from the Online Mendelian Inheritance in Man (OMIM) database and other nsSNPs derived from dbSNP, a resource at the National Center for Biotechnology Information that catalogs SNPs. TopoSNP further classifies each nsSNP site into three categories based on their geometric location: those located in a surface pocket or an interior void of the protein, those on a convex region or a shallow depressed region, and those that are completely buried in the interior of the protein structure. These unique geometric descriptions provide more detailed mapping of nsSNPs to protein structures. The current release also includes relative entropy of SNPs calculated from multiple sequence alignment as obtained from the Pfam database (a database of protein families and conserved protein motifs) as well as manually adjusted multiple alignments obtained from ClustalW. These structural and conservational data can be useful for studying whether nsSNPs in coding regions are likely to lead to phenotypic changes. TopoSNP includes an interactive structural visualization web interface, as well as downloadable batch data. The database will be updated at regular intervals and can be accessed at: http://gila.bioengr.uic.edu/snp/toposnp.
INTRODUCTION
Non-synonymous single nucleotide polymorphisms (nsSNPs) have been implicated in numerous disease processes because they may alter protein function (1), alter splice sites (2), destabilize protein core structure and reduce protein solubility (3). However, not all nsSNPs are associated with disease, and it is useful to explore general structural and conservational features of disease-associated nsSNPs versus non disease-associated nsSNPs. For example, solvent accessibility and other features such as experimental B-factors are found to be indicative of functional changes accompanying nsSNPs (4–6). Additionally, sequence conservation has been shown to be useful for predicting when a nsSNP is likely to have deleterious effects (7–9). In another recent study, it was found that disease-associated nsSNPs are more likely to be located in surface pockets or interior voids when compared with control nsSNPs (10). In addition, disease-associated nsSNPs buried in the protein interior are more likely to occur at conserved residue sites, whereas disease-associated nsSNPs located in surface pockets or interior voids do not have such propensity (10). Two data sets of nsSNPs were used in this study, one for disease-associated SNPs, which is derived from the Online Mendelian Inheritance in Man (OMIM) database (11), and one for non disease-associated or control nsSNPs, which is derived from dbSNP (dbSNP is a resource at the National Center for Biotechnology Information that catalogs SNPs as well as other genetic differences) (12). This study suggests that nsSNPs occurring in conserved and interior locations are likely to have dramatic effects. Although results obtained using this approach alone cannot lead to prediction of deleterious effects of nsSNP sites, these studies illustrate that structural characterization of nsSNP sites and their sequence conservation as measured by entropy scores are useful information that can be incorporated into studies addressing the fundamental problem of predicting when nsSNPs are likely to cause disease or significant phenotypic changes. Here we make the structural mapping of both disease- and non-disease-associated nsSNPs available through a web-accessible database topoSNP (http://gila.bioengr.uic.edu/snp/toposnp). In addition, we also provide structural characterization and entropy measurement of these nsSNP sites. TopoSNP also allows for convenient visualization of both disease-associated and non-disease-associated nsSNPs.
METHODS
Disease-associated nsSNPs were extracted from the OMIM database (http://www.ncbi.nlm.nih.gov/omim) (11). The control nsSNPs were extracted from dbSNP (12) (release 103) (ftp://ftp.ncbi.nlm.nih.gov/snp/human). While not a perfect source of negative control nsSNPs, it is reasonable to expect that a much smaller fraction of the nsSNPs from dbSNP would be associated with disease. Geometric locations and entropy calculations were performed as described previously (10).
DATABASE ACCESS
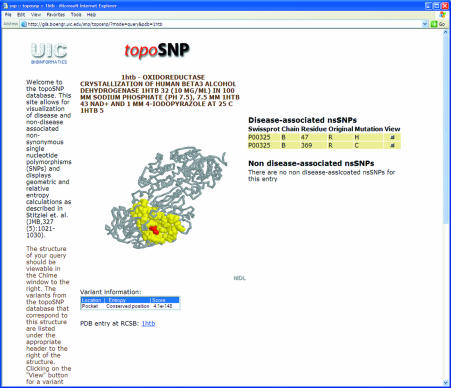
The database can be accessed at http://gila.bioengr.uic.edu/snp/toposnp. This site hosts an interactive session based on the CHIME plug-in (freely available from http://www.mdlchime.com, a plug-in that interactively displays 3D molecules), allowing for the visualization of the mutation along with its classification of geometric location and entropy score. After selecting a gene and a specific protein structure to explore, the 3D representation of the protein is displayed, along with a list of known nsSNPs. Selecting a SNP will highlight its position in the protein as well as its corresponding pocket or void if appropriate (Fig. 1). Selecting a SNP will also bring up its specific assignment of geometric class and relative entropy score, which are displayed below the protein visualization.
Figure 1.
Example of topoSNP visualization for a nsSNP from alcohol dehydrogenase (PDB code 1htb). The R47 mutation is highlighted along with the surface pocket where it is located.
There is an online help page as well as a walk-through example for familiarization with the database. The entire database is also downloadable in tar format from the topoSNP website (http://gila.bioengr.uic.edu/snp/toposnp).
DATABASE STATUS AND FUTURE WORK
The database currently contains 27 417 nsSNP mappings (26 859 from disease-associated nsSNPs as derived from the OMIM database and 558 control nsSNPs as derived from the dbSNP database) that correspond to 770 protein structures. These 770 protein structures are derived from 421 gene loci. This is a much larger data set than was previously published (10) as it was expanded to include redundant and homologous protein structures. The database will be updated at regular intervals as new SNP data and structure data become available.
DISCUSSION
We present here a resource for accessing both geometric location information and conservation information from a study of disease-associated nsSNPs and control nsSNPs. Conservation is assessed here by entropy calculated using a Hidden Markov Model (HMM). There are other approaches for assessing conservation at SNP sites. For example, recent studies demonstrated that position-specific scoring matrices (PSSMs) are quite effective for SNP analysis (7). A comparison of PSSM and HMM methods for remote homology detection showed that these two methods often obtain comparable results (13), although it is unclear exactly to what extent these two approaches differ when assessing conservation at individual sites. In addition, phylogenetic information ideally should be incorporated when assessing sequence conservation, e.g. into a codon or amino acid substitution model, together with a maximum likelihood estimator, or a Bayesian estimator (14–16). However, these methods are not easily scalable for a large set of proteins. Entropy calculation in this case provides an efficient, albeit less precise, assessment of sequence conservation. An area of ongoing work is the rapid construction of multiple alignment, which provides the basis for entropy calculation.
In this study, we do not differentiate missense mutations that cause Mendelian type diseases from nsSNPs that are associated with complex disease phenotypes. Our purpose is to examine nsSNPs that are clearly associated with disease. It is possible that these two sub-populations of nsSNPs may have different characteristics.
Acknowledgments
ACKNOWLEDGEMENTS
This work is supported by grants from the National Science Foundation (CAREER DBI0133856, DBI0078270, and KDI MCB998008) and the National Institutes of Health (GM68958). N.S. was supported in part by an NIH/NIDDK-funded predoctoral training program (T32 DK007739) in ‘Signal Transduction and Cellular Endocrinology’.
REFERENCES
- 1.Yoshida A., Huang,I.Y. and Ikawa,M. (1984) Molecular abnormality of an inactive aldehyde dehydrogenase variant commonly found in Orientals. Proc. Natl Acad. Sci. USA, 81, 258–261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Jaruzelska J., Abadie,V., d’Aubenton-Carafa,Y., Brody,E., Munnich,A. and Marie,J. (1995) In vitro splicing deficiency induced by a C to T mutation at position –3 in the intron 10 acceptor site of the phenylalanine hydroxylase gene in a patient with phenylketonuria. J. Biol. Chem., 270, 20370–20375. [DOI] [PubMed] [Google Scholar]
- 3.Proia R.L. and Neufeld,E.F. (1982) Synthesis of β-hexosaminidase in cell-free translation and in intact fibroblasts: an insoluble precursor α chain in a rare form of Tay–Sachs disease. Proc. Natl Acad. Sci. USA, 79, 6360–6364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Sunyaev S., Ramensky,V. and Bork,P. (2000) Towards a structural basis of human non-synonymous single nucleotide polymorphisms. Trends Genet., 16, 198–200. [DOI] [PubMed] [Google Scholar]
- 5.Chasman D. and Adams,R.M. (2001) Predicting the functional consequences of non-synonymous single nucleotide polymorphisms: Structure-based assessment of amino acid variation. J. Mol. Biol., 307, 683–706. [DOI] [PubMed] [Google Scholar]
- 6.Ramensky V., Bork,P. and Sunyaev,S. (2002) Human non-synonymous SNPs: server and survey. Nucleic Acids Res., 30, 3894–3900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ng P.C. and Henikoff,S. (2001) Predicting deleterious amino acid substitutions. Genome Res., 11, 863–874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ng P.C. and Henikoff,S. (2002) Accounting for human polymorphisms predicted to affect protein function. Genome Res., 12, 436–446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ng P.C. and Henikoff,S. (2003) SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res., 31, 3812–3814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Stitziel N.O., Tseng,Y.Y., Pervouchine,D., Goddeau,D., Kasif,S. and Liang,J. (2003) Structural location of disease-associated single-nucleotide polymorphisms. J. Mol. Biol., 327, 1021–1030. [DOI] [PubMed] [Google Scholar]
- 11.Wheeler D.L., Church,D.M., Federhen,S., Lash,A.E., Madden,T.L., Pontius,J.U., Schuler,G.D., Schriml,L.M., Sequeira,E., Tatusova,T.A. et al. (2003) Database resources of the National Center for Biotechnology Information. Nucleic Acids Res., 31, 28–33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Sherry S.T., Ward,M.H., Kholodov,M., Baker,J., Phan,L., Smigielski,E.M. and Sirotkin,K. (2001) dbSNP: the NCBI database of genetic variation. Nucleic Acids Res., 29, 308–311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Madera M. and Gough,J. (2002) A comparison of profile Hidden Markov Model procedures for remote homology detection. Nucleic Acids Res., 30, 4321–4328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Swofford D.L., Olsen,G.L., Waddell,P.J. and Hillis,D.L. (1996) Phylogenetic inference. In Hillis,D.M., Moritz,C. and Mable,B. (eds), Molecular Systematics. Sinauer Associates, Sunderland, MA, pp. 407–514. [Google Scholar]
- 15.Yang Z. (2001) Adaptive molecular evolution. In Balding,D., Bishop,M. and Cannings,C. (eds), Handbook of Statistical Genetics. Wiley, London, pp. 327–350. [Google Scholar]
- 16.Huelsenbeck J.P. and Ronquist,F. (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics, 17, 754–755. [DOI] [PubMed] [Google Scholar]