Abstract
A prospective in vivo assay was used to identify cells with potential for multiple lineage differentiation. With this assay, it was first determined that the 5-fluorouracil resistant cells capable of osseous tissue formation in vivo also migrated toward stromal derived factor-1 (SDF-1) in vitro. In parallel, an isolation method based on fluorescence-activated cell sorting was employed to identify a very small cell embryonic-like Lin−Sca-1+CD45− cell that with as few as 500 cells was capable of forming bone-like structures in vivo. Differential marrow fractionation studies determined that the majority of the Lin−Sca-1+CD45− cells reside in the subendosteal regions of marrow. To determine whether these cells were capable of differentiating into multiple lineages, stromal cells harvested from Col2.3ΔTK mice were implanted with a gelatin sponge into SCID mice to generate thymidine kinase sensitive ossicles. At 1.5 months, 2,000 green fluorescent protein (GFP)+ Lin−Sca-1+CD45− cells were injected into the ossicles. At harvest, colocalization of GFP-expressing cells with antibodies to the osteoblast-specific marker Runx-2 and the adipocyte marker PPARγ were observed. Based on the ability of the noncultured cells to differentiate into multiple mesenchymal lineages in vivo and the ability to generate osseous tissues at low density, we propose that this population fulfills many of the characteristics of mesenchymal stem cells.
Introduction
Hematopoietic stem cell (HSC) and progenitor cell transplantation has emerged as an important therapeutic modality for the treatment of many hematopoietic and malignant diseases [1,2]. Until recently, however, less attention has been devoted to optimizing and determining the therapeutic benefits of the transplantation bone marrow stromal cells (BMSCs). Consequently, the promise of bone marrow mesenchymal stem cells for tissue repair and immune modulation makes identifying the cells a scientific priority [3–5].
Marrow stromal cells have been largely defined operationally based on their in vitro activity. When bone marrow is cultured in vitro, adherent fibroblast-like cells proliferate and display many of the characteristics attributed to bone marrow stroma in vivo. Within these adherent cell populations, at least some of the cells are capable of self-renewal and can differentiate into several phenotypes including bone, cartilage, adipocytes, and hematopoiesis-supporting stroma [6,7]. The majority of work in this area has focused on the ability of stromal cells to differentiate into bone-like tissues. Thus, in vitro expanded stromal cells may be a rich source of osteogenic progenitor cells [8–12] that are capable of promoting the repair or regeneration of skeletal defects [7,13,14].
The ability to move forward with therapies to repair or regenerate skeletal defects and our ability to understand the biology of stem and progenitor cells are impeded by our inability to identify and characterize these cells. If the goal is to expand bone marrow progenitor cells that may regenerate bone in vivo, then the study of these heterogeneous stromal populations may be sufficient. However, if the goal is to understand how a stem cell functions to give rise to mesenchymal derivatives and whether it contributes to a functional niche for HSCs, then strategies must be developed to isolate pure populations to determine their function in vivo. For identification of HSCs, the approach has been to eliminate the functional hematopoietic cells using lethal levels of radiation. For mesenchymal cells with stem-like activities and/or their immediate progeny, this approach is not feasible, because an assay comparable to the competitive reconstitution assay has not been developed.
In a previous report, we described an in vivo assay that could be used to identify cells with stem-like activities. It was found that cells with mesenchymal stem cell-like activity are present in a murine marrow fraction that is of low density and resistant in vivo to 5-fluorouracil (5-FU) [15]. 5-FU is a nucleotide analog that is incorporated into DNA during the S-phase of the cell cycle, leads to the death of cycling cells, and has been described as having the ability to enhance the osteogenic potential of stromal cells in vitro [16,17]. Here, further characterization of these cells is presented. First, it was found that the 5-FU resistant cells that are capable of osseous tissue formation also migrated toward stromal derived factor-1 (SDF-1) (CXCL12) in vitro. In parallel, an isolation method based on fluorescence-activated cell sorting (FACS) was employed to identify a very small cell type that expressed the Sca-1 antigen but did not express CD45 (pan-hematopoietic) or other hematopoietic lineage markers (Lin−). This Lin−Sca-1+CD45− population has been previously described as having embryonic-like features and is, therefore, referred to as very small embryonic-like (VSEL) cells [18–21]. Using immunohistochemistry, it was demonstrated that Lin−Sca-1+CD45− cells express the SDF-1 receptor CXCR4. Freshly isolated Lin−Sca-1+CD45− cells when used in our in vivo model demonstrated that as few as 500 cells are able to generate bone-like tissues [22]. Importantly, when transplanted to a bone marrow environment, the cells are able to differentiate into multiple mesenchymal lineages. Together these studies demonstrate the utility of an in vivo approach to validate isolation of cells with primitive phenotypes and to provide an essential prerequisite for the study of these cells for the development of regenerative cell-based therapies.
Methods
Isolation of donor cells
Eight-week-old Balb/c mice from Charles River Laboratories (Wilmington, MA) and 7- to 8-week-old C57BL/6-Tg(ACTB-EGFP)1Osb/J mice (Jackson Laboratory, Bar Harbor, ME) were used as the source of marrow donor cells. In some cases, transgenic mice bearing a fusion gene composed of the 2.3-kb fragment of the rat type I collagen α1 promoter whose activity is largely restricted to differentiated osteoblasts and HSV-TK (Col2.3ΔTK) were provided by Dr. David W. Rowe (University of Connecticut Health Center, Farmington, CT) [23]. Animals were caged under standard conditions and fed a laboratory diet and water ad libitum. Care and use of the laboratory animals followed the guidelines established by the University of Michigan Committee for the Use and Care of Animals.
Donor animals were euthanized by CO2 inhalation, followed by cervical dislocation; and the bone marrow was harvested under sterile conditions. Bone marrow cells were flushed from the femora, tibia, and humera of the donor animals in Hank's Buffer Salt Solution (HBSS; Gibco, Grand Island, NY) delivered from a 5 mL syringe fitted with a 23-gauge needle. Single cell suspensions were obtained by passage of the marrow cells through a 70-mesh Nylon cell strainer (BD Biosciences, St. Jose, CA).
In some cases, the donor mice were treated with 5-FU (American Pharmaceutical Partners, Schaumburg, IL) intravenously administrated via the lateral tail vein at a dose of 150 mg/kg body weight in 100 μL phosphate-buffered saline 5 days before marrow harvest. Control animals were injected with an equal volume of 0.9% sodium chloride vehicle solution.
In other studies, Balb/c mice were treated with 5-FU or vehicle. Five days later, the animals were sacrificed and the femurs were removed, cleaned of muscle and connective tissues, and treated with 0.125% trypsin/0.1% collagenase (Sigma, St. Louis, MO) to remove the periosteal cells for 2 h. After vigorous washing in HBSS, the epiphyses were removed and the femurs were flushed; and the bone marrow cells were collected and termed whole bone marrow (WBM). After flushing, the femurs were sectioned into 3–5 mm long fragments, extensively washed in balanced saline solution, and the resulting cells were discarded. The remaining bone fragments were treated twice for 30 min with 0.1% collagenase at 37°C. The cells isolated during the first digestion were termed fraction of subendosteal cells. Cells isolated during the second digestion were designated fraction of endosteal cells. Both the subendosteal and endosteal cells were washed twice in ice-cold HBSS and resuspended in Dulbecco's modified Eagle's medium (DMEM; Invitrogen, Carlsbad, CA) with antibiotics for further analysis. To establish BMSCs, flushed marrow cells were incubated in DMEM that was supplemented with 15% fetal calf serum, 100 U/mL penicillin, 100 μg/mL streptomycin, 12 mM l-glutamine, and 1 × 10−6 M dexamethasone (Sigma). The media was changed twice per week for 3 weeks.
Chemotactic assays
WBM cells recovered from vehicle or 5-FU treated animals were resuspended in serum-free DMEM containing antibiotics (100 U/mL penicillin, 100 μg/mL streptomycin; Invitrogen) and equilibrated for 10 min at 37°C. Marrow cells were then seeded onto 12-μM TransWell® microporous membranes (Costar Corp., Cambridge, MA) 24-well plates containing 650 μL serum-free DMEM containing 0.5% bovine serum albumin. In some cases, Lin−Sca-1+CD45− cells were seeded onto 5-μM TransWell microporous membranes 24-well plates. Migration of the cells was supported by an SDF-1 gradient (0–200 ng/mL) achieved by adding SDF-1 to the lower chamber. Random migration (chemokinesis) was assessed by adding SDF-1 to both the upper and lower chambers. The plates were incubated at 37°C in 95% humidity and 5% CO2. After 4 h, the number of cells that had migrated into the lower chambers was determined. Subsequently, cells isolated from both chambers were separately collected for in vivo assessment of their tissue generating capacity.
RNA isolation and reverse transcription
Total cellular RNA was isolated using an RNeasy Mini kit (Qiagen, Valencia, CA). The resulting RNA was treated with the RNase-free DNase Set (Qiagen) during isolation to ensure DNA free preparations. Two micrograms of total RNA was used for cDNA synthesis with the TaqMan Reverse Transcription Reagents (Applied Biosystems, Foster City, CA) in a 100-μL reaction volume.
Quantitative real-time polymerase chain reaction (PCR) analysis was performed using an ABI 7700 sequence detector (Applied Biosystems). The assay-on-demand gene products, which contain 2 unlabeled PCR primers and an FAM™ dye-labeled TaqMan® minor groove binder probes for CXCR4, Runx-2, and PPARγ, were used for the quantitative detection of the following gene markers. Real-time PCR was performed using the TaqMan Universal PCR Master Mix Kit (Applied Biosystems). A 30 μL PCR was prepared with 1 μL cDNA and a 1.5 μL mixture of each of the specific minor groove binder probe and primer sets. Each reaction was incubated at 50°C for 2 min, 95°C for 10 min followed by 45 cycles of 95°C for 15 s and 60°C for 1 min. Target gene expression was normalized to β-actin housekeeping gene levels. Relative gene expression values were calculated using a standard curve method.
In vivo assessment of tissue generating activities
Test cells were incorporated into a gelatin sponge/scaffold implants for in vivo assessment of tissue generating activity (Gelfoam™; Pharmacia and Upjohn, Kalamazoo, MI). The cell/scaffold constructs were subcutaneously transplanted (s.c.) into immune deficient 5-week-old male recipient mice (NIH-bg-nu-xid BR; Harlan Sprague Dawley, Indianapolis, IN). Recipient mice were anesthetized by intraperitoneal injection (75 mg/kg ketamine, 10 mg/kg xylazine of body weight), shaved, and the skin was washed and cleaned with Betadine® solution. Two 1 cm skin incisions were made on the dorsal surfaces of each mouse, and 4 s.c. pockets per mouse were created by blunt dissection. A single implant was placed into each pocket, and the incisions were closed with surgical staples.
Sample harvest and evaluations
The implants were harvested 5 weeks postimplantation and fixed in aqueous buffered zinc formalin for 24 h at 4°C. For microcomputed tomography (μCT) analysis, the specimens were scanned at a 8.93 μm voxel resolution on an EVS Corp. μCT scanner (London, Ontario, Canada), with a total of 667 slices per scan. GEMS MicroView® software was used to make a 3-dimensional reconstruction from the set of scans. A fixed threshold (1,500) was used to quantify the mineralized bone phase, actual bone volume, and bone mineral density was calculated. For histologic analyses, the specimens were decalcified for 3 days in 10% formic acid, embedded in paraffin, and 5-μm serial sections were prepared and stained with hematoxylin and eosin.
Isolation of Lin−Sca-1+CD45− cells
Lin−Sca-1+CD45− and Lin−Sca-1+CD45+ cells were isolated from a suspension of murine bone marrow mononuclear cells by multiparameter, live sterile cell sorting (FACSVantage SE; Becton Dickinson, Mountainview, CA or MoFlo; Dako A/S, Fort Collins, CO). Briefly, bone marrow mononuclear cells (100 × 106 cells/mL) were resuspended in cell-sort medium, containing 1 × HBSS without phenol red, 2% heat-inactivated fetal calf serum, 10 mM HEPES buffer, and 30 U/mL of Gentamicin (Invitrogen). The following mAbs were used to stain these cells: biotin-conjugated rat anti-mouse Ly-6A/E (Sca-1) (clone E13-161.7) after streptavidin-PE-Cy5 conjugate, anti-CD45-APC [(pan hematopoietic antigen) clone 30-F11], anti-CD45R/B220-PE (clone RA3-6B2), anti-Gr-1-PE [(granulocyte-differentiation antigen-1) clone RB6-8C5], anti-TCRαβ PE (T-cell receptor, clone H57-597), anti-TCRγζ PE (clone GL3), anti-CD11b PE [(monocytes/macrophages) clone M1/70], and anti-Ter-119 PE [(eyrthroid cells), clone TER-119]; all mAbs were from BD Biosciences. All mAbs were added at saturating concentrations, and the cells were incubated for 30 min on ice and washed twice, then resuspended for cell sorting in cell-sort medium at a concentration of 5 × 106 cells/mL. In some cases, HSCs were isolated based on the SLAM family of receptors (CD150+ Lin−CD48−CD41−Sca1+c-Kit+) using isolation methods previously described [24].
In vivo assessment of multilineage activities of VSEL cells
BMSCs derived from the Col2.3ΔTK mice were transplanted with a gelatin sponge/scaffold (ossicles) into male 5- to 7-week-old SCID mice anesthetized with isofluorane inhalation. Two 1 cm incisions were made along the back of each mouse. Pouches made on either side of the incision by blunt dissection served as implant recipient beds. Ossicles were implanted in the dorsal region of each mouse (n = 4). The surgical sites were closed with surgical staples. At 1 month, ganciclovir was administered once a day for 2 weeks (3–8 mg/kg) to ablate the mature osteoblasts. At 1.5 months, the ossicles were surgically exposed and injected with 2,000 Lin−Sca-1+CD45− cells isolated from mice that express green fluorescent protein (GFP) [C57BL/6-Tg(ACTB-EGFP)1Osb/J mice] as previously described [22]. After an additional 3 months, the tissues were recovered, embedded, and frozen tissue sections prepared for immunohistochemistry.
On recovery of the ossicles containing Lin−Sca-1+CD45− cells, the tissues were snap frozen in liquid nitrogen. The specimens were embedded in water-soluble optimal cutting temperature solution (Tissue-Tek, Sekura Finetek, Torrance, CA), and 5 μm-thick serial frozen sections were cut and mounted on glass slides. Tissue sections were blocked with Image-iT FX signal enhancer (Invitrogen) for 30 min before fluorescent-labeled primary antibodies were applied. Rabbit polyclonal antibodies to osteoblasts (Runx-2) and adipocytes (PPARγ) (Abcam, Cambridge, MA) were combined with Zenon rabbit IgG labeling Alexa Fluor555 reagents (Invitrogen). Nonimmune rabbit antiserum was used to control for nonspecific binding. Images were acquired with an Olympus FV 500 confocal microscope. The images of fluorescence emission of 461 nm (DAPI, blue), 509 nm (GFP), and 568 nm (Alexa555, red) were simultaneously saved in 24 bit single TIFF files. Individual images and merged files are presented. Cells appearing double positive for GFP and either Runx-2 or PPARγ appeared as either yellow or white and were considered as evidence of multiple lineages. Image Pro Plus 5.1 software was used to quantify cells that were double positive for Runx-2 or PPARγ and GFP. To identify mineralized nodules, the implant sections were stained with 1% silver nitrate for 20 min at room temperature (von Kossa's staining). In some cases, the slides were hybridized with digoxin-labeled osteocalcin antisense or sense osteocalcin riboprobes [25].
Statistical analyses
Numerical data are expressed as mean ± standard deviation. Statistical analysis was performed by analysis of variance or unpaired 2-tailed Student's t-test using the GraphPad Instat statistical program (GraphPad Software, San Diego, CA) with significance at P < 0.05. For the quantitative PCR assays, a Kruskal–Wallis test and Dunn's multiple comparisons tests were utilized with the level of significance set at P < 0.05.
Results
Mesenchymal stem-like cells are responsive to SDF-1
Cells with stem cell-like properties must be able to rapidly migrate into wound sites for regeneration to occur and to prevent the onset of scaring. One molecule that is thought to play a critical role in recruiting HSCs into wounds after tissue injury is SDF-1 [26,27]. Therefore, it was hypothesized that cells with early mesenchymal features would also be able to rapidly migrate in response to SDF-1. Accordingly, in vitro migration assays were performed to determine whether cells with stem cell-like properties migrate toward SDF-1. In the first set of studies, WBM cells were isolated and placed into the top chamber of dual chambered culture dishes. Chemotaxtic gradients were established by adding SDF-1 (200 ng/mL) or vehicle into the bottom chamber of the dishes. In these experiments, significantly more cells migrated in response to SDF-1 than in response to control (Fig. 1A). This activity was not due to enhanced random movement, because when SDF-1 was placed into both the top and bottom chambers the number of cells migrating into the bottom chambers were not different from the nontreated SDF-1(-) controls (not presented). In previous studies, we found that cells with mesenchymal stem cell-like properties were present in the low-density marrow fractions that were resistant to 5-FU treatment [15]. When the chemotaxis studies were repeated using low-density murine bone marrow cells collected from animals treated with 5-FU, it was also observed that cells preferentially migrated toward the SDF-1 containing chamber (Fig. 1A).
FIG. 1.
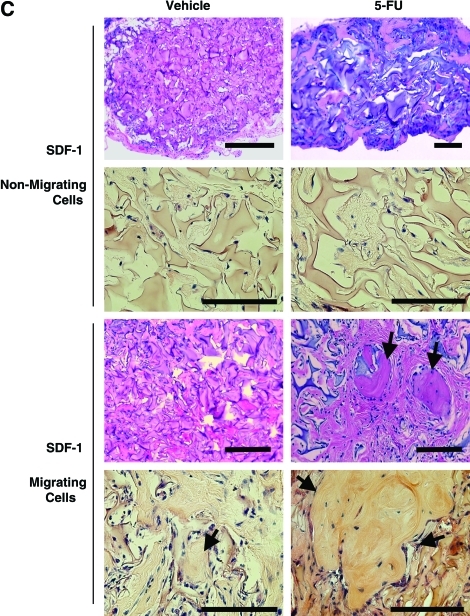
Stromal derived factor-1 (SDF-1) responsive marrow cells recovered from 5-fluorouracil (5-FU) treated animals form discrete bone tissues in vivo. (A) Migration assay of vehicle or 5-FU or in vivo treated marrow cells in response to SDF-1. Bone marrow cells were harvested from Balb/c mice treated in vivo 5 days earlier with vehicle or 5-FU. In vitro, 1 × 106 cells were placed in the top chamber of chemotactic wells in the presence (SDF-1) or absence (control) of 200 ng/mL SDF-1 in the bottom chamber. After 4 h, the cells that had migrated into the bottom chamber of the chemotactic TransWell plates were quantified. Significantly more cells from both the in vivo vehicle and 5-FU treated animals migrated in response to SDF-1. *P < 0.05. (B) mRNA expression of CXCR4 by marrow cells recovered from vehicle or 5-FU treated animals that had migrated into the bottom chambers of chemotactic TransWell plates in response to SDF-1. As in (A), marrow cells were recovered from in vivo vehicle or 5-FU treated animals. After in vitro migration assays in dual chambered plates, the migrating cells were recovered and prepared for real-time reverse-transcriptase polymerase chain reaction (PCR) determination of CXCR4 mRNA levels. As controls, in some cases whole bone marrow (WBM) cells were placed into the upper chambers with no chemoattractant in the bottom chamber. At the conclusion of the assays, the cells were recovered from the top chamber and prepared for mRNA analyses by real-time reverse transcriptase (RT)-PCR. CXCR4 mRNA levels were normalized to β-actin levels. *P < 0.05. (C) Histology of tissues generated by SDF-1-migrating marrow cells (50,000 cells) recovered from 5-FU treated animals. As in (A), bone marrow cells were harvested from Balb/c mice and treated in vivo with 5-FU. Recovered WBM cells were placed in the top chamber of chemotactic apparatus with 200 ng/mL SDF-1 in the bottom chamber. After 4 h, the SDF-1-migrating (recovered from the bottom chambers) and nonmigrating cells (top chambers) were collected, counted, and 50,000 cells/implant were loaded in gelatin sponges and implanted into SCID mice. After 5 weeks, the implants were harvested, imaged by microcomputed tomography (μCT), and sectioned for histology [hematoxylin and eosin (H&E)]. The data demonstrate that cells isolated from animals treated with 5-FU which migrated in response to SDF-1 formed discrete bone tissues in vivo. Arrows depict a typical bone foci, H&E staining, 100 ×; Bar = 50 μm. (D) μCT analysis for bone mineral density (BMD). μCT analysis of tissues generated by SDF-1-migrating marrow cells (50,000 cells) recovered from 5-FU treated animals. As in (A), bone marrow cells were harvested from Balb/c mice and treated in vivo with 5-FU. Recovered WBM cells were placed in the top chamber of chemotactic apparatus with 200 ng/mL SDF-1 in the bottom chamber. After 4 h, the SDF-1-migrating (recovered from the bottom chambers) and nonmigrating cells (top chambers) were collected, counted, and 50,000 cells/implant were loaded in gelatin sponges and implanted into SCID mice. After 5 weeks, the implants were harvested, imaged by μCT. The data demonstrate that marrow cells isolated from 5-FU treated mice which had migrated in response to SDF-1 produced tissues that were most significantly mineralized. *Significant differences from control at P < 0.01. Color images available online at www.liebertonline.com/scd.
SDF-1 binds exclusively to CXCR4, and molecules that inhibit SDF-1 binding to CXCR4 are useful for mobilizing HSCs or preventing their engraftment in vivo [26,27]. To determine the extent to which CXCR4 is expressed in the low-density marrow fractions recovered from 5-FU treated animals, mRNA was recovered from WBM or cells after the in vitro SDF-1 migration assays. CXCR4 mRNA levels were significantly enhanced in whole marrow derived from 5-FU treated animals compared with vehicle treated animals (Fig. 1B). As expected, CXCR4 mRNA level in cells that had migrated in response to the chemokine were significantly increased relative to controls (Fig. 1B).
Next, the SDF-1-migrating, 5-FU resistant low-density bone marrow cells were evaluated for their ability to form bone-like tissues in vivo on a per cells basis. Equal numbers of migrating and nonmigrating cells were collected after SDF-1 migration assays. These cells were then incorporated into gelatin sponge/scaffolds (Gelfoam; Pharmacia and Upjohn) and transplanted s.c. into 5-week-old male SCID mice. Five weeks after implantation, the tissues were recovered and prepared for μCT and histologic analyses. Histologic analysis demonstrated that tissues generated by cells isolated from animals treated with either vehicle or 5-FU and that had not migrated in vitro in response to SDF-1 produced only amorphous and fibrous tissues (Fig. 1C). Similarly, when implants were established with cells derived from animals treated with vehicle alone and that had migrated in response to SDF-1, no bone formation was observed in 5/5 implants (Fig. 1C). In contrast, when implants were established from cells isolated from 5-FU treated animals that had migrated in response to SDF-1, isolated foci of woven bone formation were observed in 3/5 implants (Fig. 1C). Quantitative μCT analyses confirmed the histologic observations and demonstrated that the 5-FU resistant, SDF-1-migrating cells generated more mineralized tissue than control treated cells or even WBM recovered from 5-FU treated mice (Fig. 1D).
Prospective FACS isolation of cells with mesenchymal stem cell-like activities
Recently, a set of reports have suggested that cells with stem-like activities express SDF-1 receptors [18,28,29]. These cells have been further characterized by FACS and represent a population of Lin−Sca-1+CD45− cells that are extremely small and have embryonic characteristics (VSEL) [21,29,30].
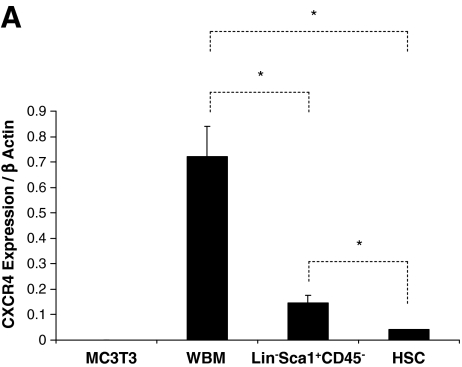
FACS isolation of small Lin−Sca-1+CD45− cells was performed, and the expression of CXCR4 was verified by immunohistochemistry and real-time reverse transcriptase (QRT)-PCR. Murine CXCR4 mRNA expression was detected in WBM cells but not in MC3T3-E1 osteoblasts or adherent BMSCs (Fig. 2A). Both Lin−Sca-1+CD45− cells and HSCs isolated based on the SLAM receptors expressed CXCR4 mRNA; however, higher levels were expressed by the Lin−Sca-1+CD45− cells. To validate that the Lin−Sca-1+CD45− cells synthesize CXCR4 protein, immunohistochemistry was performed using C2C12 mouse myoblasts as negative and WBM as positive cells that express CXCR4. As demonstrated in Fig. 2B, Lin−Sca-1+CD45− cells express CXCR4 protein. As expected, when Lin−Sca-1+CD45− cells were isolated and placed into the top chamber of dual chambered culture dishes, significantly more Lin−Sca-1+CD45− cells migrated in response to SDF-1 than in response to controls (Fig. 2C).
FIG. 2.
CXCR4 expression and SDF-1 responsiveness of Lin−Sca1+CD45− cells. (A) Expression of CXCR4 mRNA CXCR4 by Lin−Sca1+CD45− cells as determined by quantitative RT-PCR (QRT-PCR). Freshly isolated Lin−Sca1+CD45− very small embryonic-like (VSEL) cells, hematopoietic stem cells (HSCs) were isolated based on the SLAM family of receptors (CD150+ Lin−CD48−CD41−Sca1+c-Kit+), isolation using methods previously described [24]. WBM, and the murine osteoblastic cell line MC3T3E1 (MC3T3) cells were prepared for evaluation of CXCR4 mRNA levels. CXCR4 mRNA levels were normalized against β-actin. *P < 0.05. (B) Immunohistochemistry for CXCR4 expression by Lin−Sca1+CD45− cells. WBM (positive control cells), C2C12 cells (negative controls), and Lin−Sca1+CD45− cells isolated by fluorescence-activated cell sorting (FACS) were fixed on microscope slides and stained for CXCR4 and DAPI. Differential interference contrast (DIC) imaging, immune florescence (IF) for the detection of CXCR4, and merged images are presented. Bar = 5 μm. (C) Migration assay of Lin−Sca1+CD45− cells in response to SDF-1. Two thousand Lin−Sca1+CD45− cells were placed in the top chamber of chemotactic wells in the presence [SDF-1 (+)] or absence [SDF-1 (−)] of 200 ng/mL SDF-1 in the bottom chamber. After 2 h, the migrating cells were quantified. Significantly more cells from SDF-1 (−) in response to SDF-1. *P < 0.05. Color images available online at www.liebertonline.com/scd.
Next, we evaluated the frequency of Lin−Sca-1+CD45− cells in cells recovered from the 5-FU treated marrow. The numbers of Lin−Sca-1+CD45− cells were very low in the bone marrow of vehicle treated animals, representing only 0.03% of all mononuclear cells. However, the number of these cells increased ∼400% in response to 5-FU treatment (Table 1). Recently, Balduino et al. developed a technique that facilitates the fractionation of marrow using differential digestion [25]. By this method, 2 distinct stromal-cell populations may be isolated from periosteal-free fragments of murine femurs using a 2-step collagenase-digestion method. One fraction contains subendosteal cells and the other endosteal fraction consists predominantly of osteoblasts [25]. Therefore, this fractionation method was combined with our 5-FU treatment studies to identify the location of the Lin−Sca-1+CD45− cells in marrow (Table 1). The number of Lin−Sca-1+CD45− cells were very low in WBM (Table 1), whereas the frequency of the Lin−Sca-1+CD45− VSEL cells was enriched ∼80 and 200-fold in the endosteal and subendosteal marrow fractions, respectively (Table 1). Surprisingly, the location of the Lin−Sca-1+CD45− cells shifted from the subendosteal to the endosteal regions after 5-FU treatment, although overall the number of Lin−Sca-1+CD45− cells were not altered (not shown).
Table 1.
Location of Lin−Sca-1+CD45− Cells In Vivo
| Frequency (% MNC) | Control | 5-FU |
|---|---|---|
| WBM | 0.03 ± 0.15 | 0.13 ± 0.04a |
| Subendosteal | 5.97 ± 7.11 | 0.48 ± 0.30a |
| Endosteal | 2.50 ± 2.50 | 6.78 ± 0.33a |
Lin−Sca-1+CD45− cell frequency in various bone marrow fractions by FACS. Bone marrow was serially digested to harvest WBM cells, subendosteal and endosteal from animals with/without 5-FU treatment. Lin−Sca-1+CD45− cells were sorted and counted by FACS.
P < 0.05 compared with control.
Abbreviations: FACS, fluorescence-activated cell sorting; 5-FU, 5-fluorouracil; WBM, whole bone marrow; MNC, mononuclear cells.
To verify that the Lin−Sca-1+CD45− cells represent a population of cells with stem cell-like activities, the cells were isolated from mice that express GFP [C57BL/6-Tg(ACTB-EGFP)1Osb/J mice] and transplanted with a gelatin sponge/scaffold into SCID mice. At 4 weeks, the regenerative potential of the cells was evaluated. A large number of GFP+ cells were identified within the implants, indicating that the transplanted cells maintain the ability to contribute to the generation of new tissue (Fig. 3A). Mineralized tissue was formed by the Lin−Sca-1+CD45− cells with as few as 500 cells/implant, whereas 5,000 Lin−Sca-1+CD45− cells generated significantly larger bone volume, which was confirmed by μCT, von Kossa staining, and in situ hybridization for osteocalcin mRNA (Fig. 3B–E). As controls, Lin−Sca-1+CD45+ cells were also implanted. Compared with the results generated with Lin−Sca-1+CD45− cells, when 2 × 105 Lin−Sca-1+CD45+ cells were implanted, only nonmineralized amorphous tissues were identified and few, if any, GFP-expressing cells were found, suggesting the largely hematopoietic populations had either undergone apoptosis or migrated away from the implant site (data not presented).
FIG. 3.
Transplantation of Lin−Sca-1+CD45− cell generates osseous tissues in vivo. Transplantation of 500 or 5,000 Lin−Sca-1+CD45− (VSEL) cells and 20,000 Lin−Sca-1+CD45+ hematopoietic progenitor cells isolated from green fluorescent protein (GFP)+ mice into gelatin sponges subcutaneously in SCID mice. The transplants were harvested at 5 weeks and frozen-sections were prepared. (A) H&E staining of the transplanted tissues demonstrating trabeculae-like structures in both groups [3/4 in the 500 Lin−Sca-1+CD45− (VSEL) cells group, whereas 4/4 in the 5,000 cell group]. Tissues recovered from the 20,000 Lin−Sca-1+CD45+ hematopoietic progenitor cell implants demonstrate predominately collagen transplant vehicle. Size indicators; for 500 cell implants the bar represents 100 μm, for 5,000 cell implants the bar represents 50 μm, for 20,000 cell implants the bar represents 25 μm. (B) Fluorescent microscopy revealing retention of GFP expression (arrows), (C) μCT demonstrating high-density mineralized tissues in both the 500 and 5,000 cell groups; however, the 5,000 cell transplants produced more mineralized tissue. (D) The presence of mineralized tissues were confirmed by von Kossa staining (arrows). (E) In situ hybridization of osteocalcin (OCN) (arrows). Color images available online at www.liebertonline.com/scd.
To evaluate the potential of the Lin−Sca-1+CD45− cells to undergo multilineage differentiation, intramarrow transplantation studies were performed [22]. First, BMSCs were harvested from Col2.3ΔTK mice, expanded in culture, and implanted into SCID mice to generate a recipient site in which thymidine kinase tissues would be established (Experimental outline; Fig. 4A). Our rationale for using this strategy was to be able to ablate osteoblast numbers in the resulting marrow of the implant to clear space for injection of the GFP marked cells to undergo lineage progression. As controls, mRNA from freshly isolated Lin−Sca-1+CD45− was evaluated for the expression of the osteoblast-specific marker Runx-2 and the adipocyte marker PPARγ. Compared with the osteoblastic cell line MC3T3-E1 and BMSCs treated with 1 × 10−6 M dexamethasone, Lin−Sca-1+CD45− cells expressed little, if any, mRNA for Runx-2 (Fig. 4B) and PPARγ (Fig. 4C). At 1.5 months, the ossicles were surgically exposed and injected with Lin−Sca-1+CD45− cells isolated from mice that express GFP. After an additional 1.5 months, the implants were harvested, and differentiation of the injected GFP cells was determined by cell surface markers and confocal microscopy. Colocalization of GFP-expressing cells with an antibody to the osteoblast-specific marker Runx-2 and the adipocyte marker PPARγ was performed (Fig. 4D, E) and data quantified (Fig. 4F and G). Approximately half of the Runx-2-expressing cells also expressed GFP (Fig. 4C and D). Similarly, nearly half of the PPARγ-expressing cells were marked with GFP (Fig. 4F and G). These data suggest that lineage progression down at least 2 lineages from the transplanted Lin−Sca-1+CD45− cells had occurred in vivo.
FIG. 4.
Lin−Sca-1+CD45− cells differentiate into 2 lineages in vivo. (A) Experimental scheme. Bone ossicles were established using stromal donor cells (bone marrow stromal cells, BMSCs) derived from Col2.3ΛTK mice in host SCID mice. At 1 month, ganciclovir was administered for 2 weeks to ablate mature osteoblasts. At 1.5 months, the ossicles were surgically exposed and injected with 2,000 Lin− Sca-1+CD45− cells isolated from mice that express GFP [(C57BL/6-Tg(ACTB- EGFP)1Osb/J mice]. After an additional 3 months, the tissues were recovered, embedded, and frozen tissue sections were prepared for immunohistochemistry. (B) Lin−Sca-1+CD45− cells do not express Runx-2 mRNA. mRNA was harvested from mixed BMSC, the osteoblastic MC3T3-E1 (MC3T3) cell line (a positive control), and freshly isolated Lin−Sca-1+CD45− cells. QRT-PCR was performed to determine expression levels of the bone specific transcription factor Runx-2. mRNA levels were normalized to β-actin levels. *Significance differences at P < 0.01. (C) Lin−Sca-1+CD45− cells do not express mRNA for peroxisome proliferator-activated receptor (PPAR-Gamma)γ. mRNA was harvested from dexamethasone treated BMSC (positive control), the osteoblastic MC3T3-E1 (MC3T3) cell line, and freshly isolated Lin−Sca-1+CD45− cells. QRT-PCR was performed to determine expression levels of PPARγ. mRNA levels were normalized to β-actin levels. *Significant difference at P < 0.01. (D) Tissue sections demonstrating multilineage differentiation of GFP-expressing cells. Rabbit polyclonal antibodies to osteoblasts (Runx-2) were used to demonstrate colocalization of with GFP. Four individual images and merged files are presented. Cells appearing double positive for GFP and Runx-2 appear as either yellow or white (arrows). The bar indicates 10 μm. (E) Quantification of cells double positive for GFP and Runx-2 as shown in (D). The data are presented as total cells positive for Runx-2 and double positive (Runx-2 and GFP). *Significant difference from Runx-2 only at P < 0.05. (F) As in (E), tissue sections demonstrating multilineage differentiation of GFP-expressing cells. Rabbit polyclonal antibodies to adipocytes (PPARγ) were used to demonstrate colocalization of with GFP. Cells appearing double positive for GFP and PPARγ appear as either yellow or white (arrowheads). The bar indicates 10 μm. (G) Quantification of cells double positive for GFP and PPARγ as shown in (F). The data are presented as total cells positive for PPARγ and double positive (PPARγ and GFP). *Significant difference from PPARγ only at P < 0.05. Color images available online at www.liebertonline.com/scd.
Discussion
We adapted and modified an in vivo model in which it is possible to functionally characterize prospectively isolated cells with stem-like properties using noncultured cells [22]. With this assay, we demonstrated that prospectively harvested and purified Lin−Sca-1+CD45− cells differentiate into multiple mesenchymal lineages in vivo. We observed at least 4 different examples of each merged image of marked mesenchymal stem-like cells (GFP+ cells) with 2 different lineages: (1) osteoblasts-Runx-2 and (2) adipocytes-PPARγ. These data demonstrate that cells within the small Lin−Sca-1+CD45− cells exhibit mesenchymal stem cell-like activities in vivo.
An important aspect of this work is the development of a physiologically relevant model for prospective identification of cells with mesenchymal stem-like capacity that is much more flexible than conventional, whole animal studies, because: (1) several implants can be generated in a single animal, (2) the implants can be taken out sequentially leaving others to more fully develop without sacrificing the animal, (3) marked stem cells can be directly injected into the implants of living animals, (4) cells from different transgenic lines can be transplanted and compared with wild type activity in the same animal, and (5) both bone formation and the development of the marrow can be monitored and manipulated over time. By combining the study of our tissue-engineered ossicle model with the purified Lin−Sca-1+CD45− cells or other populations, we will be in a position to expand our understanding of mesenchymal stem cell biology and their interactions with HSC interactions in vivo.
The data also demonstrate that noncultured bone-marrow-derived small Lin−Sca-1+CD45− cells exhibit the capacity to form bone in vivo with as low as 500 cells/implant. Moreover, this population increases in frequency in marrow fractions that we demonstrate have an enhanced ability to regenerate a bone marrow organ in vivo. Lin−Sca-1+CD45− cells are also highly responsive to SDF-1 and resistant to chemotherapeutic agents (5-FU), suggesting that under basal conditions these cells are quiescent or slowly cycling through the cell cycle. Previous characterization of these cells demonstrated that they represent ∼0.02% of bone marrow mononuclear cells and express markers of pluripotent stem cells such as SSEA-1, Oct-4, Nanog, and Rex-1 by real-time PCR and immunohistochemistry [31]. In addition, these cells are quite small (∼2–4 μm), and they contain open-type chromatin (euchromatin) that is typical of stem cells [21,29,30].
It is also clear that Lin−Sca-1+CD45− cells are not capable of regenerating an entire marrow organ alone, as transplantation of 500 cells was not sufficient to form an intact ossicle. One possibility for this modest bone forming capacity is that in postfetal life, more cells are required for organ regeneration than are required for similar activities during development [29]. Another possibility is that accessory cells are necessary to regenerate the marrow organ. In fact, it has been demonstrated that subpopulations of nonadherent, low-density human bone marrow cells regulate the development of human bone precursor cells [32]. In this study, feeder-layer cultures were used to demonstrate that osteopoietic accessory cells (OACs) do not require cell–cell interaction to promote bone precursor cell development but, rather, produce soluble molecules responsible for their effects. FACS analyses reveal that bone-marrow-derived B cells, T cells, macrophages, natural killer cells, and endothelial cells do not produce this stimulatory factor [32]. Moreover, the factor could be replaced by addition of exogenous cytokines that were evaluated, and the isolation of human transforming growth factor beta receptor type II-positive cells increases OAC-specific activity in bone cell ex vivo expansion cultures [32]. Interestingly, the OAC bone marrow cells were characterized by high transforming growth factor beta receptor type II expression, relatively low cellular complexity, and small size. In an other study, nonadherent bone marrow cells were not only able to give rise to multiple mesenchymal phenotypes including (fibroblastic, osteoblastic, chondrocytic, and adipogenic), but these cells could be used to rescue mice which were lethally irradiated suggesting the possibility, but not proving, that bone marrow cell can differentiate along both hematopoietic and mesenchymal lineages [33].
The endosteal/subendosteal environment is known to be comprised of bone-lining and reticular cells and contribute to the regulation of hematopoiesis. Previous work has led to the isolation of 2 distinct stroma-cell populations containing subendosteal reticulocytes (F-RET) and endosteal osteoblasts (F-OST) from periosteum-free fragments of murine femurs by a 2-step collagenase-digestion procedure. Both populations produce similar extracellular matrix (collagen I, laminin, fibronectin, decorin), except for collagen IV, which is low in endosteal cells (F-OST) [34]. They also express several osteogenic markers including osteopontin, osteonectin, bone sialoprotein, and alkaline phosphatase; however, the subendosteal (F-RET) population exhibit a less mature phenotype [34]. Therefore, a surprising aspect of these studies was the observation that the Lin−Sca-1+CD45− cells shifted location from the subendosteal to the endosteum after 5-FU treatment. This observation is in line with the notion that repopulation of an osseous surface would require the rapid influx of cells with regenerative potential. This concept also fits well with a model of cells that are responsive to SDF-1, as endosteal surfaces respond to physiologic stimuli including parathyroid hormone (PTH) [35] and marrow injury [36]. Recently, it was demonstrated that the Lin−Sca-1+CD45− population under steady state conditions circulate at very low levels in peripheral blood (∼100–200 cells/mL), but they can be mobilized during pharmacological granulocyte-colony stimulating factor (G-CSF)-induced or stress-related mobilization after tissue damage. In fact, reports suggest that mesenchymal stem-like cells may be capable of generating osseous in a limited capacity under basal conditions and circulate in the peripheral blood at low levels [36,37]. Moreover, the number of circulating Lin−Sca-1+CD45− cells in 2-month-old animals are 5 times higher as compared with 1-year-old mice [28].
In summary, current transplantation systems require the expansion of BMSCs in culture to generate enough cells to form bone in vitro. We have isolated and initially characterized an enriched mesenchymal stem cell-like population that maintains its multipotential function in vivo without the need to expand the cells in culture where it is possible to acquire a nonphysiologic phenotype. These methods and model systems represent significant strides forward in the mesenchymal biology field as we are able to demonstrate that a prospectively isolated population can generate bone in vivo before cell culture with as few as 500 mononucleated cells (Lin−Sca-1+CD45−). Although we have not definitively isolated a single stem cell, the combination of methods described here may serve as a platform to prospectively isolate stem cells and functionally characterize their phenotype in vivo.
Acknowledgments
This study was supported by grants from the National Institutes of Health (DK082481, R43AR056893, DE020721, and RC1DE020721).
Author Disclosure Statement
No competing financial interests exist.
References
- 1.Lansdorp PM. Stem cell biology for the transfusionist. Vox Sang. 1998;74(Suppl. 2):91–94. doi: 10.1111/j.1423-0410.1998.tb05402.x. [DOI] [PubMed] [Google Scholar]
- 2.Lapidot T. Dar A. Kollet O. How do stem cells find their way home? Blood. 2005;106:1901–1910. doi: 10.1182/blood-2005-04-1417. [DOI] [PubMed] [Google Scholar]
- 3.Aggarwal S. Pittenger MF. Human mesenchymal stem cells modulate allogeneic immune cell responses. Blood. 2005;105:1815–1822. doi: 10.1182/blood-2004-04-1559. [DOI] [PubMed] [Google Scholar]
- 4.Le Blanc K. Mesenchymal stromal cells: tissue repair and immune modulation. Cytotherapy. 2006;8:559–561. doi: 10.1080/14653240601045399. [DOI] [PubMed] [Google Scholar]
- 5.Yamaza T. Miura Y. Akiyama K. Bi Y. Sonoyama W. Gronthos S. Chen W. Le A. Shi S. Mesenchymal stem cell-mediated ectopic hematopoiesis alleviates aging-related phenotype in immunocompromised mice. Blood. 2009;113:2595–2604. doi: 10.1182/blood-2008-10-182246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Alhadlaq A. Mao JJ. Mesenchymal stem cells: isolation and therapeutics. Stem Cells Dev. 2004;13:436–448. doi: 10.1089/scd.2004.13.436. [DOI] [PubMed] [Google Scholar]
- 7.Krebsbach PH. Kuznetsov SA. Satomura K. Emmons RV. Rowe DW. Robey PG. Bone formation in vivo: comparison of osteogenesis by transplanted mouse and human marrow stromal fibroblasts. Transplantation. 1997;63:1059–1069. doi: 10.1097/00007890-199704270-00003. [DOI] [PubMed] [Google Scholar]
- 8.Mankani MH. Kuznetsov SA. Avila NA. Kingman A. Robey PG. Bone formation in transplants of human bone marrow stromal cells and hydroxyapatite-tricalcium phosphate: prediction with quantitative CT in mice. Radiology. 2004;230:369–376. doi: 10.1148/radiol.2302011529. [DOI] [PubMed] [Google Scholar]
- 9.Mankani MH. Kuznetsov SA. Wolfe RM. Marshall GW. Robey PG. In vivo bone formation by human bone marrow stromal cells: reconstruction of the mouse calvarium and mandible. Stem Cells. 2006;24:2140–2149. doi: 10.1634/stemcells.2005-0567. [DOI] [PubMed] [Google Scholar]
- 10.Mankani MH. Kuznetsov SA. Robey PG. Formation of hematopoietic territories and bone by transplanted human bone marrow stromal cells requires a critical cell density. Exp Hematol. 2007;35:995–1004. doi: 10.1016/j.exphem.2007.01.051. [DOI] [PubMed] [Google Scholar]
- 11.Bianco P. Robey PG. Simmons PJ. Mesenchymal stem cells: revisiting history, concepts, and assays. Cell Stem Cell. 2008;2:313–319. doi: 10.1016/j.stem.2008.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kuznetsov SA. Mankani MH. Bianco P. Robey PG. Enumeration of the colony-forming units-fibroblast from mouse and human bone marrow in normal and pathological conditions. Stem Cell Res. 2009;2:83–94. doi: 10.1016/j.scr.2008.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kaigler D. Krebsbach PH. West ER. Horger K. Huang YC. Mooney DJ. Endothelial cell modulation of bone marrow stromal cell osteogenic potential. FASEB J. 2005;19:665–667. doi: 10.1096/fj.04-2529fje. [DOI] [PubMed] [Google Scholar]
- 14.Jiang Y. Jahagirdar BN. Reinhardt RL. Schwartz RE. Keene CD. Ortiz-Gonzalez XR. Reyes M. Lenvik T. Lund T. Blackstad M. Du J. Aldrich S. Lisberg A. Low WC. Largaespada DA. Verfaillie CM. Pluripotency of mesenchymal stem cells derived from adult marrow. Nature. 2002;418:41–49. doi: 10.1038/nature00870. [DOI] [PubMed] [Google Scholar]
- 15.Wang Z. Song J. Taichman RS. Krebsbach PH. Ablation of proliferating marrow with 5-fluorouracil allows partial purification of mesenchymal stem cells. Stem Cells. 2006;24:1573–1582. doi: 10.1634/stemcells.2005-0399. [DOI] [PubMed] [Google Scholar]
- 16.Koller MR. Manchel I. Smith AK. Quantitative long-term culture-initiating cell assays require accessory cell depletion that can be achieved by CD34-enrichment or 5-fluorouracil exposure. Blood. 1998;91:4056–4064. [PubMed] [Google Scholar]
- 17.Van Vlasselaer P. Falla N. Snoeck H. Mathieu E. Characterization and purification of osteogenic cells from murine bone marrow by two-color cell sorting using anti-Sca-1 monoclonal antibody and wheat germ agglutinin. Blood. 1994;84:753–763. [PubMed] [Google Scholar]
- 18.Ratajczak MZ. Kucia M. Reca R. Majka M. Janowska-Wieczorek A. Ratajczak J. Stem cell plasticity revisited: CXCR4-positive cells expressing mRNA for early muscle, liver and neural cells “hide out” in the bone marrow. Leukemia. 2004;18:29–40. doi: 10.1038/sj.leu.2403184. [DOI] [PubMed] [Google Scholar]
- 19.Ratajczak MZ. Majka M. Kucia M. Drukala J. Pietrzkowski Z. Peiper S. Janowska-Wieczorek A. Expression of functional CXCR4 by muscle satellite cells and secretion of SDF-1 by muscle-derived fibroblasts is associated with the presence of both muscle progenitors in bone marrow and hematopoietic stem/progenitor cells in muscles. Stem Cells. 2003;21:363–371. doi: 10.1634/stemcells.21-3-363. [DOI] [PubMed] [Google Scholar]
- 20.Ratajczak MZ. Machalinski B. Wojakowski W. Ratajczak J. Kucia M. A hypothesis for an embryonic origin of pluripotent Oct-4(+) stem cells in adult bone marrow and other tissues. Leukemia. 2007;21:860–867. doi: 10.1038/sj.leu.2404630. [DOI] [PubMed] [Google Scholar]
- 21.Kucia M. Zuba-Surma EK. Wysoczynski M. Wu W. Ratajczak J. Machalinski B. Ratajczak MZ. Adult marrow-derived very small embryonic-like stem cells and tissue engineering. Expert Opin Biol Ther. 2007;7:1499–1514. doi: 10.1517/14712598.7.10.1499. [DOI] [PubMed] [Google Scholar]
- 22.Song J. Kiel MJ. Wang Z. Wang J. Taichman RS. Morrison SJ. Krebsbach PH. An in vivo model to study and manipulate the hematopoietic stem cell niche. Blood. 2010;115:2592–2600. doi: 10.1182/blood-2009-01-200071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Visnjic D. Kalajzic I. Gronowicz G. Aguila HL. Clark SH. Lichtler AC. Rowe DW. Conditional ablation of the osteoblast lineage in Col2.3deltatk transgenic mice. J Bone Miner Res. 2001;16:2222–2231. doi: 10.1359/jbmr.2001.16.12.2222. [DOI] [PubMed] [Google Scholar]
- 24.Jung Y. Wang J. Song J. Shiozawa Y. Wang J. Havens A. Wang Z. Sun YX. Emerson SG. Krebsbach PH. Taichman RS. Annexin II expressed by osteoblasts & endothelial cells regulates stem cell adhesion, homing & engraftment following transplantation. Blood. 2007;110:82–90. doi: 10.1182/blood-2006-05-021352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Balduino A. Hurtado SP. Frazão P. Takiya CM. Alves LM. Nasciutti LE. El-Cheikh MC. Borojevic R. Bone marrow subendosteal microenvironment harbours functionally distinct haemosupportive stromal cell populations. Cell Tissue Res. 2005;319:255–266. doi: 10.1007/s00441-004-1006-3. [DOI] [PubMed] [Google Scholar]
- 26.Petit I. Szyper-Kravitz M. Nagler A. Lahav M. Peled A. Habler L. Ponomaryov T. Taichman RS. Arenzana-Seisdedos F. Fujii N. Sandbank J. Zipori D. Lapidot T. G-CSF induces stem cell mobilization by decreasing bone marrow SDF-1 and up-regulating CXCR4. Nat Immunol. 2002;3:687–694. doi: 10.1038/ni813. [DOI] [PubMed] [Google Scholar]
- 27.Ponomaryov T. Peled A. Peled I. Taichman RS. Habler L. Sandbank J. Arenzana-Seisdedos G. Nagler A. Lahav M. Szyper-Kravitz M. Zipori D. Lapidot T. Increased production of SDF-1 following treatment with DNA damaging agents: relevance for human stem cell homing and repopulation of NOD/SCID mice. J Clin Invest. 2000;106:1331–1339. doi: 10.1172/JCI10329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kucia M. Halasa M. Wysoczynski M. Baskiewicz-Masiuk M. Moldenhawer S. Zuba-Surma E. Czajka R. Wojakowski W. Machalinski B. Ratajczak MZ. Morphological and molecular characterization of novel population of CXCR4+ SSEA-4+ Oct-4+ very small embryonic-like cells purified from human cord blood: preliminary report. Leukemia. 2007;21:297–303. doi: 10.1038/sj.leu.2404470. [DOI] [PubMed] [Google Scholar]
- 29.Kucia M. Reca R. Campbell FR. Zuba-Surma E. Majka M. Ratajczak J. Ratajczak MZ. A population of very small embryonic-like (VSEL) CXCR4(+)SSEA-1(+)Oct-4+ stem cells identified in adult bone marrow. Leukemia. 2006;20:857–869. doi: 10.1038/sj.leu.2404171. [DOI] [PubMed] [Google Scholar]
- 30.Kucia M. Wu W. Ratajczak MZ. Bone marrow-derived very small embryonic-like stem cells: their developmental origin and biological significance. Dev Dyn. 2007;236:3309–3320. doi: 10.1002/dvdy.21180. [DOI] [PubMed] [Google Scholar]
- 31.Jung Y. Wang J. Schneider A. Sun YX. Koh-Paige AJ. Osman NI. McCauley LK. Taichman RS. Regulation of SDF-1 (CXCL12) production by osteoblasts; a possible mechanism for stem cell homing. Bone. 2006;38:497–508. doi: 10.1016/j.bone.2005.10.003. [DOI] [PubMed] [Google Scholar]
- 32.Eipers PG. Kale S. Taichman RS. Pipia GG. Swords NA. Mann KG. Long MW. Bone marrow accessory cells regulate human bone precursor cell development. Exp Hematol. 2000;28:815–825. doi: 10.1016/s0301-472x(00)00183-1. [DOI] [PubMed] [Google Scholar]
- 33.Zhang ZL. Tong J. Lu RN. Scutt AM. Goltzman D. Miao DS. Therapeutic potential of non-adherent BM-derived mesenchymal stem cells in tissue regeneration. Bone Marrow Transplant. 2009;43:69–81. doi: 10.1038/bmt.2008.260. [DOI] [PubMed] [Google Scholar]
- 34.Balduino A. Hurtado SP. Frazão P. Takiya CM. Alves LM. Nasciutti LE. El Cheikh MC. Borojevic R. Bone marrow subendosteal microenvironment harbours functionally distinct haemosupportive stromal cell populations. Cell Tissue Res. 2005;319:255–266. doi: 10.1007/s00441-004-1006-3. [DOI] [PubMed] [Google Scholar]
- 35.Schneider A. Taboas JM. McCauley LK. Krebsbach PH. Skeletal homeostasis in tissue-engineered bone. J Orthop Res. 2003;21:859–864. doi: 10.1016/S0736-0266(03)00042-1. [DOI] [PubMed] [Google Scholar]
- 36.Rochefort GY. Delorme B. Lopez A. Herault O. Bonnet P. Charbord P. Eder V. Domenech J. Multipotential mesenchymal stem cells are mobilized into peripheral blood by hypoxia. Stem Cells. 2006;24:2202–2208. doi: 10.1634/stemcells.2006-0164. [DOI] [PubMed] [Google Scholar]
- 37.Paczkowska E. Kucia M. Koziarska D. Halasa M. Safranow K. Masiuk M. Karbicka A. Nowik M. Nowacki P. Ratajczak MZ. Machalinski B. Clinical evidence that very small embryonic-like stem cells are mobilized into peripheral blood in patients after stroke. Stroke. 2009;40:1237–1244. doi: 10.1161/STROKEAHA.108.535062. [DOI] [PubMed] [Google Scholar]