Abstract
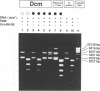
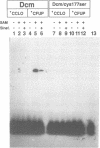
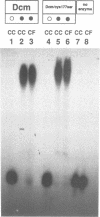
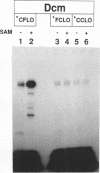
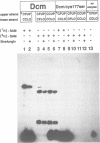
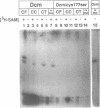
The product of the dcm gene is the only DNA cytosine-C5 methyltransferase of Escherichia coli K-12; it catalyses transfer of a methyl group from S-adenosyl methionine (SAM) to the C-5 position of the inner cytosine residue of the cognate sequence CCA/TGG. Sequence-specific, covalent crosslinking of the enzyme to synthetic oligonucleotides containing 5-fluoro-2'-deoxycytidine is demonstrated. This reaction is abolished if serine replaces the cysteine at residue #177 of the enzyme. These results lend strong support to a catalytic mechanism in which an enzyme sulfhydryl group undergoes Michael addition to the C5-C6 double bond, thus activating position C-5 of the substrate DNA cytosine residue for electrophilic attack by the methyl donor SAM. The enzyme is capable of self-methylation in a DNA-independent reaction requiring SAM and the presence of cysteine at position #177.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Barbés C., Sánchez J., Yebra M. J., Robert-Geró M., Hardisson C. Effects of sinefungin and S-adenosylhomocysteine on DNA and protein methyltransferases from Streptomyces and other bacteria. FEMS Microbiol Lett. 1990 Jun 1;57(3):239–243. [PubMed] [Google Scholar]
- Bellisario R. L., Maley G. F., Guarino D. U., Maley F. The primary structure of Lactobacillus casei thymidylate synthetase. II. The complete amino acid sequence of the active site peptide, CNBr 4. J Biol Chem. 1979 Feb 25;254(4):1296–1300. [PubMed] [Google Scholar]
- Chen L., MacMillan A. M., Chang W., Ezaz-Nikpay K., Lane W. S., Verdine G. L. Direct identification of the active-site nucleophile in a DNA (cytosine-5)-methyltransferase. Biochemistry. 1991 Nov 19;30(46):11018–11025. doi: 10.1021/bi00110a002. [DOI] [PubMed] [Google Scholar]
- Demple B., Sedgwick B., Robins P., Totty N., Waterfield M. D., Lindahl T. Active site and complete sequence of the suicidal methyltransferase that counters alkylation mutagenesis. Proc Natl Acad Sci U S A. 1985 May;82(9):2688–2692. doi: 10.1073/pnas.82.9.2688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dev I. K., Yates B. B., Leong J., Dallas W. S. Functional role of cysteine-146 in Escherichia coli thymidylate synthase. Proc Natl Acad Sci U S A. 1988 Mar;85(5):1472–1476. doi: 10.1073/pnas.85.5.1472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edelmann P., Gallant J. Mistranslation in E. coli. Cell. 1977 Jan;10(1):131–137. doi: 10.1016/0092-8674(77)90147-7. [DOI] [PubMed] [Google Scholar]
- Friedman S., Ansari N. Binding of the EcoRII methyltransferase to 5-fluorocytosine-containing DNA. Isolation of a bound peptide. Nucleic Acids Res. 1992 Jun 25;20(12):3241–3248. doi: 10.1093/nar/20.12.3241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friedman S. Binding of the EcoRII methylase to azacytosine-containing DNA. Nucleic Acids Res. 1986 Jun 11;14(11):4543–4556. doi: 10.1093/nar/14.11.4543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friedman S. The effect of 5-azacytidine on E. coli DNA methylase. Biochem Biophys Res Commun. 1979 Aug 28;89(4):1328–1333. doi: 10.1016/0006-291x(79)92154-5. [DOI] [PubMed] [Google Scholar]
- Friedman S. The irreversible binding of azacytosine-containing DNA fragments to bacterial DNA(cytosine-5)methyltransferases. J Biol Chem. 1985 May 10;260(9):5698–5705. [PubMed] [Google Scholar]
- Fritz H. J., Belagaje R., Brown E. L., Fritz R. H., Jones R. A., Lees R. G., Khorana H. G. High-pressure liquid chromatography in polynucleotide synthesis. Biochemistry. 1978 Apr 4;17(7):1257–1267. doi: 10.1021/bi00600a020. [DOI] [PubMed] [Google Scholar]
- Hardy L. W., Finer-Moore J. S., Montfort W. R., Jones M. O., Santi D. V., Stroud R. M. Atomic structure of thymidylate synthase: target for rational drug design. Science. 1987 Jan 23;235(4787):448–455. doi: 10.1126/science.3099389. [DOI] [PubMed] [Google Scholar]
- Jones P. A., Taylor S. M. Cellular differentiation, cytidine analogs and DNA methylation. Cell. 1980 May;20(1):85–93. doi: 10.1016/0092-8674(80)90237-8. [DOI] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Langenbach R. J., Danenberg P. V., Heidelberger C. Thymidylate synthetase: mechanism of inhibition by 5-fluoro-2'-deoxyuridylate. Biochem Biophys Res Commun. 1972 Sep 26;48(6):1565–1571. doi: 10.1016/0006-291x(72)90892-3. [DOI] [PubMed] [Google Scholar]
- Lauster R., Trautner T. A., Noyer-Weidner M. Cytosine-specific type II DNA methyltransferases. A conserved enzyme core with variable target-recognizing domains. J Mol Biol. 1989 Mar 20;206(2):305–312. doi: 10.1016/0022-2836(89)90480-4. [DOI] [PubMed] [Google Scholar]
- Osterman D. G., DePillis G. D., Wu J. C., Matsuda A., Santi D. V. 5-Fluorocytosine in DNA is a mechanism-based inhibitor of HhaI methylase. Biochemistry. 1988 Jul 12;27(14):5204–5210. doi: 10.1021/bi00414a039. [DOI] [PubMed] [Google Scholar]
- Pósfai J., Bhagwat A. S., Pósfai G., Roberts R. J. Predictive motifs derived from cytosine methyltransferases. Nucleic Acids Res. 1989 Apr 11;17(7):2421–2435. doi: 10.1093/nar/17.7.2421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubin R. A., Modrich P. EcoRI methylase. Physical and catalytic properties of the homogeneous enzyme. J Biol Chem. 1977 Oct 25;252(20):7265–7272. [PubMed] [Google Scholar]
- Santi D. V., Garrett C. E., Barr P. J. On the mechanism of inhibition of DNA-cytosine methyltransferases by cytosine analogs. Cell. 1983 May;33(1):9–10. doi: 10.1016/0092-8674(83)90327-6. [DOI] [PubMed] [Google Scholar]
- Santi D. V., Hardy L. W. Catalytic mechanism and inhibition of tRNA (uracil-5-)methyltransferase: evidence for covalent catalysis. Biochemistry. 1987 Dec 29;26(26):8599–8606. doi: 10.1021/bi00400a016. [DOI] [PubMed] [Google Scholar]
- Santi D. V., Norment A., Garrett C. E. Covalent bond formation between a DNA-cytosine methyltransferase and DNA containing 5-azacytosine. Proc Natl Acad Sci U S A. 1984 Nov;81(22):6993–6997. doi: 10.1073/pnas.81.22.6993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt S., Pein C. D., Fritz H. J., Cech D. Chemical synthesis of 2'-deoxyoligonucleotides containing 5-fluoro-2'-deoxycytidine. Nucleic Acids Res. 1992 May 25;20(10):2421–2426. doi: 10.1093/nar/20.10.2421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schneider R., Gander I., Müller U., Mertz R., Winnacker E. L. A sensitive and rapid gel retention assay for nuclear factor I and other DNA-binding proteins in crude nuclear extracts. Nucleic Acids Res. 1986 Feb 11;14(3):1303–1317. doi: 10.1093/nar/14.3.1303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sinha N. D., Biernat J., McManus J., Köster H. Polymer support oligonucleotide synthesis XVIII: use of beta-cyanoethyl-N,N-dialkylamino-/N-morpholino phosphoramidite of deoxynucleosides for the synthesis of DNA fragments simplifying deprotection and isolation of the final product. Nucleic Acids Res. 1984 Jun 11;12(11):4539–4557. doi: 10.1093/nar/12.11.4539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Som S., Friedman S. Identification of a highly conserved domain in the EcoRII methyltransferase which can be photolabeled with S-adenosyl-L-[methyl-3H]methionine. Evidence for UV-induced transmethylation of cysteine 186. J Biol Chem. 1991 Feb 15;266(5):2937–2945. [PubMed] [Google Scholar]
- Stanssens P., Opsomer C., McKeown Y. M., Kramer W., Zabeau M., Fritz H. J. Efficient oligonucleotide-directed construction of mutations in expression vectors by the gapped duplex DNA method using alternating selectable markers. Nucleic Acids Res. 1989 Jun 26;17(12):4441–4454. doi: 10.1093/nar/17.12.4441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Starzyk R. M., Koontz S. W., Schimmel P. A covalent adduct between the uracil ring and the active site of an aminoacyl tRNA synthetase. Nature. 1982 Jul 8;298(5870):136–140. doi: 10.1038/298136a0. [DOI] [PubMed] [Google Scholar]
- Subbaramaiah K., Simms S. A. Photolabeling of CheR methyltransferase with S-adenosyl-L-methionine (AdoMet). Studies on the AdoMet binding site. J Biol Chem. 1992 Apr 25;267(12):8636–8642. [PubMed] [Google Scholar]
- Wilke K., Rauhut E., Noyer-Weidner M., Lauster R., Pawlek B., Behrens B., Trautner T. A. Sequential order of target-recognizing domains in multispecific DNA-methyltransferases. EMBO J. 1988 Aug;7(8):2601–2609. doi: 10.1002/j.1460-2075.1988.tb03110.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu J. C., Santi D. V. Kinetic and catalytic mechanism of HhaI methyltransferase. J Biol Chem. 1987 Apr 5;262(10):4778–4786. [PubMed] [Google Scholar]
- Wyszynski M. W., Gabbara S., Bhagwat A. S. Substitutions of a cysteine conserved among DNA cytosine methylases result in a variety of phenotypes. Nucleic Acids Res. 1992 Jan 25;20(2):319–326. doi: 10.1093/nar/20.2.319. [DOI] [PMC free article] [PubMed] [Google Scholar]