Abstract
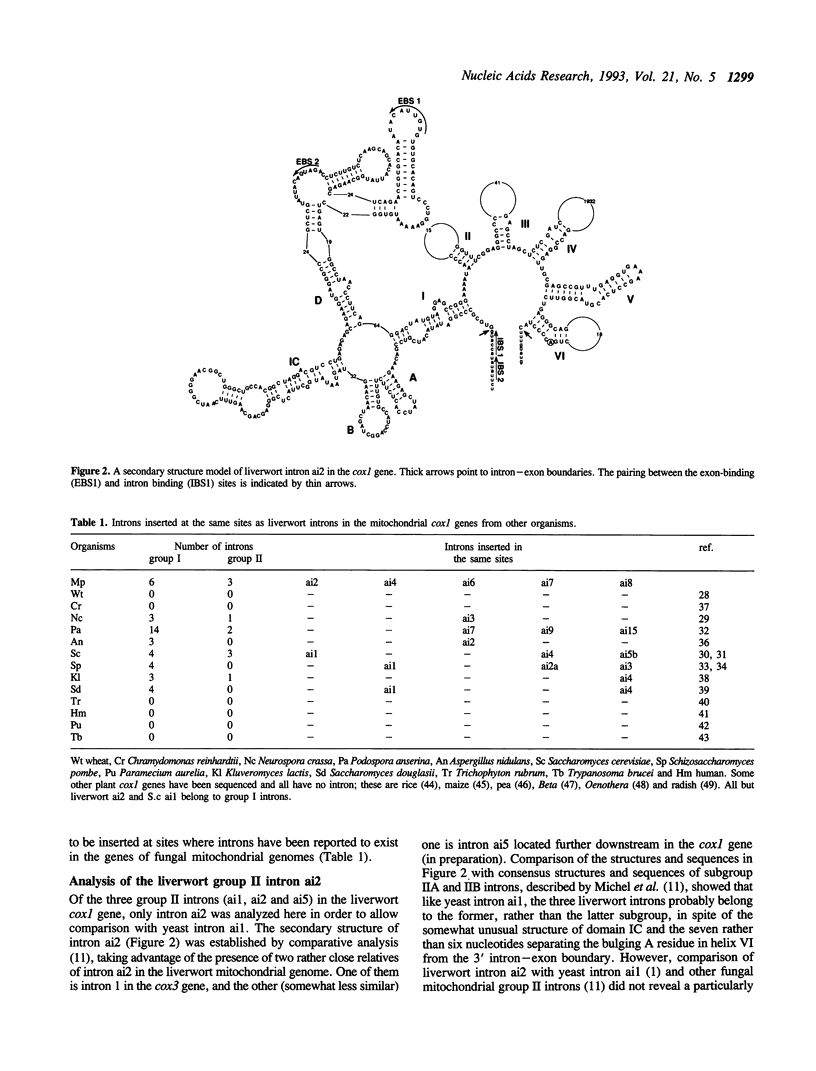
The complete nucleotide sequence of the mitochondrial DNA (mtDNA) from a liverwort, Marchantia polymorpha, contains thirty-two introns. Twenty-five of these introns possess the characteristic secondary structures and consensus sequences of group II introns. The remaining seven are group I introns, six of which happen to interrupt the gene coding for subunit 1 of cytochrome oxidase (cox1). Interestingly, the insertion sites of one group II and four group I introns in the cox1 gene coincide with those of the respective fungal mitochondrial interns. Moreover, comparison of the four group I introns with their fungal counterparts shows that group I introns inserted at identical genomic sites in different organisms are indeed related to one another, in terms of the peptide sequences generated from the complete or fragmental ORFs encoded by these introns. At the same time, the liverwort introns turned out to be more divergent from their fungal cognates than the latter are from one another. We therefore conclude that vertical transmission from a common ancestor organism is the simplest explanation for the presence of cognate introns in liverwort and fungal mitochondrial genomes.
Full text
PDF






Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Anziano P. Q., Hanson D. K., Mahler H. R., Perlman P. S. Functional domains in introns: trans-acting and cis-acting regions of intron 4 of the cob gene. Cell. 1982 Oct;30(3):925–932. doi: 10.1016/0092-8674(82)90297-5. [DOI] [PubMed] [Google Scholar]
- Belfort M. Phage T4 introns: self-splicing and mobility. Annu Rev Genet. 1990;24:363–385. doi: 10.1146/annurev.ge.24.120190.002051. [DOI] [PubMed] [Google Scholar]
- Boer P. H., Gray M. W. Nucleotide sequence of a protein coding region in Chlamydomonas reinhardtii mitochondrial DNA. Nucleic Acids Res. 1986 Sep 25;14(18):7506–7507. doi: 10.1093/nar/14.18.7506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonen L., Boer P. H., McIntosh J. E., Gray M. W. Nucleotide sequence of the wheat mitochondrial gene for subunit I of cytochrome oxidase. Nucleic Acids Res. 1987 Aug 25;15(16):6734–6734. doi: 10.1093/nar/15.16.6734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonitz S. G., Coruzzi G., Thalenfeld B. E., Tzagoloff A., Macino G. Assembly of the mitochondrial membrane system. Structure and nucleotide sequence of the gene coding for subunit 1 of yeast cytochrme oxidase. J Biol Chem. 1980 Dec 25;255(24):11927–11941. [PubMed] [Google Scholar]
- Burke J. M., Belfort M., Cech T. R., Davies R. W., Schweyen R. J., Shub D. A., Szostak J. W., Tabak H. F. Structural conventions for group I introns. Nucleic Acids Res. 1987 Sep 25;15(18):7217–7221. doi: 10.1093/nar/15.18.7217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cech T. R. Conserved sequences and structures of group I introns: building an active site for RNA catalysis--a review. Gene. 1988 Dec 20;73(2):259–271. doi: 10.1016/0378-1119(88)90492-1. [DOI] [PubMed] [Google Scholar]
- Chapdelaine Y., Bonen L. The wheat mitochondrial gene for subunit I of the NADH dehydrogenase complex: a trans-splicing model for this gene-in-pieces. Cell. 1991 May 3;65(3):465–472. doi: 10.1016/0092-8674(91)90464-a. [DOI] [PubMed] [Google Scholar]
- Colleaux L., d'Auriol L., Betermier M., Cottarel G., Jacquier A., Galibert F., Dujon B. Universal code equivalent of a yeast mitochondrial intron reading frame is expressed into E. coli as a specific double strand endonuclease. Cell. 1986 Feb 28;44(4):521–533. doi: 10.1016/0092-8674(86)90262-x. [DOI] [PubMed] [Google Scholar]
- Cummings D. J., Michel F., McNally K. L. DNA sequence analysis of the 24.5 kilobase pair cytochrome oxidase subunit I mitochondrial gene from Podospora anserina: a gene with sixteen introns. Curr Genet. 1989 Dec;16(5-6):381–406. doi: 10.1007/BF00340719. [DOI] [PubMed] [Google Scholar]
- De La Salle H., Jacq C., Slonimski P. P. Critical sequences within mitochondrial introns: pleiotropic mRNA maturase and cis-dominant signals of the box intron controlling reductase and oxidase. Cell. 1982 Apr;28(4):721–732. doi: 10.1016/0092-8674(82)90051-4. [DOI] [PubMed] [Google Scholar]
- Delahodde A., Goguel V., Becam A. M., Creusot F., Perea J., Banroques J., Jacq C. Site-specific DNA endonuclease and RNA maturase activities of two homologous intron-encoded proteins from yeast mitochondria. Cell. 1989 Feb 10;56(3):431–441. doi: 10.1016/0092-8674(89)90246-8. [DOI] [PubMed] [Google Scholar]
- Delahodde A., Goguel V., Becam A. M., Creusot F., Perea J., Banroques J., Jacq C. Site-specific DNA endonuclease and RNA maturase activities of two homologous intron-encoded proteins from yeast mitochondria. Cell. 1989 Feb 10;56(3):431–441. doi: 10.1016/0092-8674(89)90246-8. [DOI] [PubMed] [Google Scholar]
- Dujon B. Group I introns as mobile genetic elements: facts and mechanistic speculations--a review. Gene. 1989 Oct 15;82(1):91–114. doi: 10.1016/0378-1119(89)90034-6. [DOI] [PubMed] [Google Scholar]
- Field D. J., Sommerfield A., Saville B. J., Collins R. A. A group II intron in the Neurospora mitochondrial coI gene: nucleotide sequence and implications for splicing and molecular evolution. Nucleic Acids Res. 1989 Nov 25;17(22):9087–9099. doi: 10.1093/nar/17.22.9087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fox T. D., Leaver C. J. The Zea mays mitochondrial gene coding cytochrome oxidase subunit II has an intervening sequence and does not contain TGA codons. Cell. 1981 Nov;26(3 Pt 1):315–323. doi: 10.1016/0092-8674(81)90200-2. [DOI] [PubMed] [Google Scholar]
- Hardy C. M., Clark-Walker G. D. Nucleotide sequence of the COX1 gene in Kluyveromyces lactis mitochondrial DNA: evidence for recent horizontal transfer of a group II intron. Curr Genet. 1991 Jul;20(1-2):99–114. doi: 10.1007/BF00312772. [DOI] [PubMed] [Google Scholar]
- Hensgens L. A., Bonen L., de Haan M., van der Horst G., Grivell L. A. Two intron sequences in yeast mitochondrial COX1 gene: homology among URF-containing introns and strain-dependent variation in flanking exons. Cell. 1983 Feb;32(2):379–389. doi: 10.1016/0092-8674(83)90457-9. [DOI] [PubMed] [Google Scholar]
- Hensgens L. A., Brakenhoff J., De Vries B. F., Sloof P., Tromp M. C., Van Boom J. H., Benne R. The sequence of the gene for cytochrome c oxidase subunit I, a frameshift containing gene for cytochrome c oxidase subunit II and seven unassigned reading frames in Trypanosoma brucei mitochondrial maxi-circle DNA. Nucleic Acids Res. 1984 Oct 11;12(19):7327–7344. doi: 10.1093/nar/12.19.7327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hiesel R., Schobel W., Schuster W., Brennicke A. The cytochrome oxidase subunit I and subunit III genes in Oenothera mitochondria are transcribed from identical promoter sequences. EMBO J. 1987 Jan;6(1):29–34. doi: 10.1002/j.1460-2075.1987.tb04714.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunt M. D., Newton K. J. The NCS3 mutation: genetic evidence for the expression of ribosomal protein genes in Zea mays mitochondria. EMBO J. 1991 May;10(5):1045–1052. doi: 10.1002/j.1460-2075.1991.tb08043.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Isaac P. G., Jones V. P., Leaver C. J. The maize cytochrome c oxidase subunit I gene: sequence, expression and rearrangement in cytoplasmic male sterile plants. EMBO J. 1985 Jul;4(7):1617–1623. doi: 10.1002/j.1460-2075.1985.tb03828.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kadowaki K., Suzuki T., Kazama S., Oh-fuchi T., Sakamoto W. Nucleotide sequence of the cytochrome oxidase subunit I gene from rice mitochondria. Nucleic Acids Res. 1989 Sep 25;17(18):7519–7519. doi: 10.1093/nar/17.18.7519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kemmerer E. C., Kao T. H., Deng G. R., Wu R. Isolation and nucleotide sequence of the pea cytochrome oxidase subunit I gene. Plant Mol Biol. 1989 Jul;13(1):121–124. doi: 10.1007/BF00027340. [DOI] [PubMed] [Google Scholar]
- Knoop V., Schuster W., Wissinger B., Brennicke A. Trans splicing integrates an exon of 22 nucleotides into the nad5 mRNA in higher plant mitochondria. EMBO J. 1991 Nov;10(11):3483–3493. doi: 10.1002/j.1460-2075.1991.tb04912.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuhsel M. G., Strickland R., Palmer J. D. An ancient group I intron shared by eubacteria and chloroplasts. Science. 1990 Dec 14;250(4987):1570–1573. doi: 10.1126/science.2125748. [DOI] [PubMed] [Google Scholar]
- Lamattina L., Grienenberger J. M. RNA editing of the transcript coding for subunit 4 of NADH dehydrogenase in wheat mitochondria: uneven distribution of the editing sites among the four exons. Nucleic Acids Res. 1991 Jun 25;19(12):3275–3282. doi: 10.1093/nar/19.12.3275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lang B. F. The mitochondrial genome of the fission yeast Schizosaccharomyces pombe: highly homologous introns are inserted at the same position of the otherwise less conserved cox1 genes in Schizosaccharomyces pombe and Aspergillus nidulans. EMBO J. 1984 Sep;3(9):2129–2136. doi: 10.1002/j.1460-2075.1984.tb02102.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lazowska J., Jacq C., Slonimski P. P. Sequence of introns and flanking exons in wild-type and box3 mutants of cytochrome b reveals an interlaced splicing protein coded by an intron. Cell. 1980 Nov;22(2 Pt 2):333–348. doi: 10.1016/0092-8674(80)90344-x. [DOI] [PubMed] [Google Scholar]
- Makaroff C. A., Apel I. J., Palmer J. D. The role of coxI-associated repeated sequences in plant mitochondrial DNA rearrangements and radish cytoplasmic male sterility. Curr Genet. 1991 Mar;19(3):183–190. doi: 10.1007/BF00336485. [DOI] [PubMed] [Google Scholar]
- Michel F., Cummings D. J. Analysis of class I introns in a mitochondrial plasmid associated with senescence of Podospora anserina reveals extraordinary resemblance to the Tetrahymena ribosomal intron. Curr Genet. 1985;10(1):69–79. doi: 10.1007/BF00418495. [DOI] [PubMed] [Google Scholar]
- Michel F., Jacquier A., Dujon B. Comparison of fungal mitochondrial introns reveals extensive homologies in RNA secondary structure. Biochimie. 1982 Oct;64(10):867–881. doi: 10.1016/s0300-9084(82)80349-0. [DOI] [PubMed] [Google Scholar]
- Michel F., Lang B. F. Mitochondrial class II introns encode proteins related to the reverse transcriptases of retroviruses. Nature. 1985 Aug 15;316(6029):641–643. doi: 10.1038/316641a0. [DOI] [PubMed] [Google Scholar]
- Michel F., Umesono K., Ozeki H. Comparative and functional anatomy of group II catalytic introns--a review. Gene. 1989 Oct 15;82(1):5–30. doi: 10.1016/0378-1119(89)90026-7. [DOI] [PubMed] [Google Scholar]
- Michel F., Westhof E. Modelling of the three-dimensional architecture of group I catalytic introns based on comparative sequence analysis. J Mol Biol. 1990 Dec 5;216(3):585–610. doi: 10.1016/0022-2836(90)90386-Z. [DOI] [PubMed] [Google Scholar]
- Mota E. M., Collins R. A. Independent evolution of structural and coding regions in a Neurospora mitochondrial intron. Nature. 1988 Apr 14;332(6165):654–656. doi: 10.1038/332654a0. [DOI] [PubMed] [Google Scholar]
- Nobrega F. G., Tzagoloff A. Assembly of the mitochondrial membrane system. DNA sequence and organization of the cytochrome b gene in Saccharomyces cerevisiae D273-10B. J Biol Chem. 1980 Oct 25;255(20):9828–9837. [PubMed] [Google Scholar]
- Nomiyama H., Kuhara S., Kukita T., Otsuka T., Sakaki Y. Nucleotide sequence of the ribosomal RNA gene of Physarum polycephalum: intron 2 and its flanking regions of the 26S rRNA gene. Nucleic Acids Res. 1981 Nov 11;9(21):5507–5520. doi: 10.1093/nar/9.21.5507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nomiyama H., Sakaki Y., Takagi Y. Nucleotide sequence of a ribosomal RNA gene intron from slime mold Physarum polycephalum. Proc Natl Acad Sci U S A. 1981 Mar;78(3):1376–1380. doi: 10.1073/pnas.78.3.1376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oda K., Yamato K., Ohta E., Nakamura Y., Takemura M., Nozato N., Akashi K., Kanegae T., Ogura Y., Kohchi T. Gene organization deduced from the complete sequence of liverwort Marchantia polymorpha mitochondrial DNA. A primitive form of plant mitochondrial genome. J Mol Biol. 1992 Jan 5;223(1):1–7. doi: 10.1016/0022-2836(92)90708-r. [DOI] [PubMed] [Google Scholar]
- Pritchard A. E., Seilhamer J. J., Mahalingam R., Sable C. L., Venuti S. E., Cummings D. J. Nucleotide sequence of the mitochondrial genome of Paramecium. Nucleic Acids Res. 1990 Jan 11;18(1):173–180. doi: 10.1093/nar/18.1.173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reinhold-Hurek B., Shub D. A. Self-splicing introns in tRNA genes of widely divergent bacteria. Nature. 1992 May 14;357(6374):173–176. doi: 10.1038/357173a0. [DOI] [PubMed] [Google Scholar]
- Sanger F., Coulson A. R., Barrell B. G., Smith A. J., Roe B. A. Cloning in single-stranded bacteriophage as an aid to rapid DNA sequencing. J Mol Biol. 1980 Oct 25;143(2):161–178. doi: 10.1016/0022-2836(80)90196-5. [DOI] [PubMed] [Google Scholar]
- Senda M., Harada T., Mikami T., Sugiura M., Kinoshita T. Genomic organization and sequence analysis of the cytochrome oxidase subunit II gene from normal and male-sterile mitochondria in sugar beet. Curr Genet. 1991 Mar;19(3):175–181. doi: 10.1007/BF00336484. [DOI] [PubMed] [Google Scholar]
- Shub D. A., Gott J. M., Xu M. Q., Lang B. F., Michel F., Tomaschewski J., Pedersen-Lane J., Belfort M. Structural conservation among three homologous introns of bacteriophage T4 and the group I introns of eukaryotes. Proc Natl Acad Sci U S A. 1988 Feb;85(4):1151–1155. doi: 10.1073/pnas.85.4.1151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trinkl H., Wolf K. The mosaic cox1 gene in the mitochondrial genome of Schizosaccharomyces pombe: minimal structural requirements and evolution of group I introns. Gene. 1986;45(3):289–297. doi: 10.1016/0378-1119(86)90027-2. [DOI] [PubMed] [Google Scholar]
- Turmel M., Boulanger J., Lemieux C. Two group I introns with long internal open reading frames in the chloroplast psbA gene of Chlamydomonas moewusii. Nucleic Acids Res. 1989 May 25;17(10):3875–3887. doi: 10.1093/nar/17.10.3875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turmel M., Boulanger J., Schnare M. N., Gray M. W., Lemieux C. Six group I introns and three internal transcribed spacers in the chloroplast large subunit ribosomal RNA gene of the green alga Chlamydomonas eugametos. J Mol Biol. 1991 Mar 20;218(2):293–311. doi: 10.1016/0022-2836(91)90713-g. [DOI] [PubMed] [Google Scholar]
- Wahleithner J. A., MacFarlane J. L., Wolstenholme D. R. A sequence encoding a maturase-related protein in a group II intron of a plant mitochondrial nad1 gene. Proc Natl Acad Sci U S A. 1990 Jan;87(2):548–552. doi: 10.1073/pnas.87.2.548. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waring R. B., Davies R. W., Scazzocchio C., Brown T. A. Internal structure of a mitochondrial intron of Aspergillus nidulans. Proc Natl Acad Sci U S A. 1982 Oct;79(20):6332–6336. doi: 10.1073/pnas.79.20.6332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss-Brummer B., Rödel G., Schweyen R. J., Kaudewitz F. Expression of the split gene cob in yeast: evidence for a precursor of a "maturase" protein translated from intron 4 and preceding exons. Cell. 1982 Jun;29(2):527–536. doi: 10.1016/0092-8674(82)90169-6. [DOI] [PubMed] [Google Scholar]
- Wenzlau J. M., Saldanha R. J., Butow R. A., Perlman P. S. A latent intron-encoded maturase is also an endonuclease needed for intron mobility. Cell. 1989 Feb 10;56(3):421–430. doi: 10.1016/0092-8674(89)90245-6. [DOI] [PubMed] [Google Scholar]
- Wild M. A., Gall J. G. An intervening sequence in the gene coding for 25S ribosomal RNA of Tetrahymena pigmentosa. Cell. 1979 Mar;16(3):565–573. doi: 10.1016/0092-8674(79)90030-8. [DOI] [PubMed] [Google Scholar]
- Wissinger B., Schuster W., Brennicke A. Trans splicing in Oenothera mitochondria: nad1 mRNAs are edited in exon and trans-splicing group II intron sequences. Cell. 1991 May 3;65(3):473–482. doi: 10.1016/0092-8674(91)90465-b. [DOI] [PubMed] [Google Scholar]
- Xu M. Q., Kathe S. D., Goodrich-Blair H., Nierzwicki-Bauer S. A., Shub D. A. Bacterial origin of a chloroplast intron: conserved self-splicing group I introns in cyanobacteria. Science. 1990 Dec 14;250(4987):1566–1570. doi: 10.1126/science.2125747. [DOI] [PubMed] [Google Scholar]
- de Bièvre C., Dujon B. Mitochondrial DNA sequence analysis of the cytochrome oxidase subunit I and II genes, the ATPase9 gene, the NADH dehydrogenase ND4L and ND5 gene complex, and the glutaminyl, methionyl and arginyl tRNA genes from Trichophyton rubrum. Curr Genet. 1992 Sep;22(3):229–234. doi: 10.1007/BF00351730. [DOI] [PubMed] [Google Scholar]



