Abstract
Background
Bacille Calmette-Guerin (BCG) is currently the only available vaccine against tuberculosis (TB) and comprises a heterogeneous family of sub-strains with genotypic and phenotypic differences. The World Health Organization (WHO) affirms that the characterization of BCG sub-strains, both on genomic and proteomic levels, is crucial for a better comprehension of the vaccine. In addition, these studies can contribute in the development of a more efficient vaccine against TB. Here, we combine two-dimensional electrophoresis (2DE) and mass spectrometry to analyse the proteomic profile of culture filtrate proteins (CFPs) from M. bovis BCG Moreau, the Brazilian vaccine strain, comparing it to that of BCG Pasteur. CFPs are considered of great importance given their dominant immunogenicity and role in pathogenesis, being available for interaction with host cells since early infection.
Results
The 2DE proteomic map of M. bovis BCG Moreau CFPs in the pH range 3 - 8 allowed the identification of 158 spots corresponding to 101 different proteins, identified by MS/MS. Comparison to BCG Pasteur highlights the great similarity between these BCG strains. However, quantitative analysis shows a higher expression of immunogenic proteins such as Rv1860 (BCG1896, Apa), Rv1926c (BCG1965c, Mpb63) and Rv1886c (BCG1923c, Ag85B) in BCG Moreau when compared to BCG Pasteur, while some heat shock proteins, such as Rv0440 (BCG0479, GroEL2) and Rv0350 (BCG0389, DnaK), show the opposite pattern.
Conclusions
Here we report the detailed 2DE profile of CFPs from M. bovis BCG Moreau and its comparison to BCG Pasteur, identifying differences that may provide relevant information on vaccine efficacy. These findings contribute to the detailed characterization of the Brazilian vaccine strain against TB, revealing aspects that may lead to a better understanding of the factors leading to BCG's variable protective efficacy against TB.
Background
Tuberculosis (TB) remains a major cause of morbidity and mortality, particularly in developing countries, and is considered a serious public health problem worldwide, killing almost 2 million people every year [1]. According to the WHO, one-third of the world's population is infected with Mycobacterium tuberculosis (Mtb). The incidence of new cases of TB has increased mainly due to the impact of the HIV epidemic [2] and the emergence of resistance to anti-TB drugs [3]. The currently available vaccine, Mycobacterium bovis bacillus Calmette-Guérin (BCG), is one of the oldest and most commonly administered vaccines worldwide [4]. It was obtained in the early 1920's by Albert Calmette and Camille Guérin at the Pasteur Institute, Lille, France, after 231 serial passages of a clinical isolate of M. bovis in glycerinated medium containing ox bile [5]. Attenuation during in vitro passages is believed to have resulted from the loss and/or reorganization of genomic regions, some of which have been recently identified [6-9]. M. bovis BCG Moreau is the strain used in Brazil for vaccine production since the 1930's [10]. According to recent molecular studies [11], it is considered an "old" strain, more similar to the original BCG derived by Calmette and Guérin. Vaccination with BCG has many advantages, yielding efficient protection against severe childhood forms of TB, and also against leprosy [12]. In addition, it is recognized as a safe and inexpensive vaccine that can be administered shortly after birth [13,14]. On the other hand, it shows variable protection against the most common form of the disease, pulmonary tuberculosis in adults, and it does not prevent the establishment of latent TB. It has been reported that different M. bovis BCG strains, including BCG Moreau, induce varying levels of protection against M. tuberculosis infection in animal models [15]. Comparative genetic analysis of BCG strains has revealed that each vaccine currently in use is unique [11], and providing several clues for the failure of BCG as an effective vaccine. Proteomic studies have shown that BCG strains with similar genomic content, such as BCG Denmark and Phipps, exhibit phenotypic differences that can be particularly important for pathogenesis, immune response and variable efficacy of BCG vaccine [16]. Other factors have also been implicated in its unpredictable efficacy: (i) the genetic variability amongst vaccinated individuals; (ii) cross-reactivity of the immune response to BCG due to environmental mycobacteria [17]; (iii) differences in vaccine production procedures, variable doses, and bacterial viability, amongst others [18,19]. New vaccination strategies are therefore urgently needed, particularly against pulmonary forms of TB.
The modulation of cellular functions of the host cell is a dynamic process that requires viable mycobacteria, supporting the idea that the components actively secreted by the living bacteria are the main players involved in this process [20]. Membrane and membrane associated proteins also play an important role in this process [21]. Subunit vaccines based on mixtures of culture filtrate proteins have resulted in protective immunity in animal models of TB [22-26]. These molecules are also strongly recognized during Mtb infection in various animal models, as well as in early stages of pulmonary TB in humans [27,28]. Culture filtrate is therefore an attractive source of potential candidate antigens for the development of new vaccines and diagnostic reagents. In this report, we have employed a combination of 2DE and mass spectrometry analysis in order to generate a proteomic map of CFPs from the Brazilian M. bovis BCG Moreau strain, comparing it to the reference strain, M. bovis BCG Pasteur. The data presented may contribute to the identification of useful markers for quality control of the BCG Moreau vaccine production, and yield possible clues regarding the variable effectiveness of these vaccine strains.
Results
Protein separation, identification and sub-cellular localization
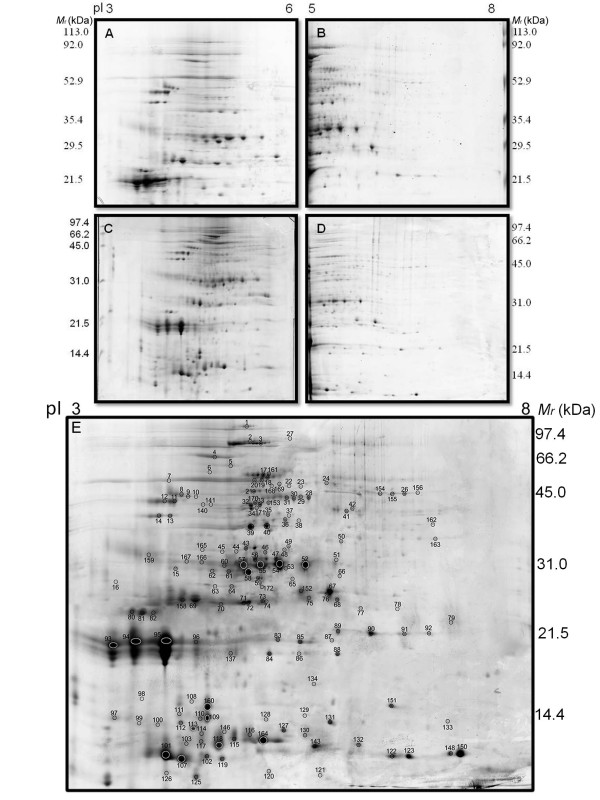
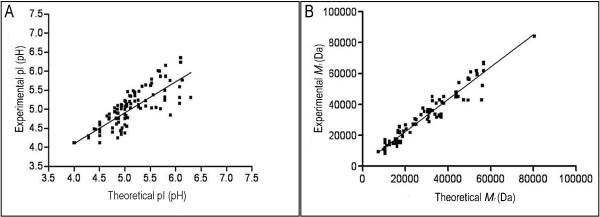
The BCG strains were grown in static cultures, as surface pellicles, for 15 days, with no apparent difference in growth. The genetic profile of the 2 strains was confirmed by PCR (Additional file 1, Figure S1), corroborating with previous reports [29] The preliminary separation of BCG Moreau CFPs by 2DE revealed that most protein spots clustered in the pH range 3-8 (data not shown). To generate proteomic maps, samples were therefore applied to immobilized pH gradient (IPG) strips in the pH intervals 3-6 (Figure 1A and 1C) and 5-8 (Figure 1B and 1D) and subsequently separated in the second dimension across 12% (Figure 1A and 1B) and 15% SDS-PAGE (Figure 1C and 1D). To aid in visualization, gel images were merged to produce an artificial map representative of the pH range 3-8 (Figure 1E) comprising all the 280 spots resolved in the individual gels. These spots were excised and digested with trypsin. The resulting peptides were submitted to mass spectrometry analysis leading to the putative identification of 158 protein spots corresponding to 101 different proteins (Additional file 2, Table S1). For clarity, we have adopted the standard protein nomenclature used for M. tuberculosis strain H37Rv (http://genolist.pasteur.fr/tuberculist) and M. bovis BCG Pasteur 1173P2 (http://genolist.pasteur.fr/BCGList/). Identified proteins showed a pI variation between 3-8 and a molecular mass (Mr) range between 9 and 120 kDa. The comparison of experimentally determined and theoretical Mr and pI values of the identified protein spots from BCG Moreau against the predicted values for M. tuberculosis strain H37Rv proteins, obtained from the search with Mascot version 2.2, showed a positive correlation according to the Spearman coefficient (Figure 2A and 2B) Considering the fact that the proteins identified in this study were obtained from the culture filtrate, we analyzed the presence of possible signals that could direct these proteins to the extracellular fraction (Additional file 3, Table S2), using Signal P (for sec-dependent secretion; [30]), LipoP (lipoproteins; [31]), TatP (for secretion through the twin-arginine translocation system; [32]) and SecretomeP (for non-classical secretion of leaderless proteins; [33]). Of the 101 proteins, 67 (66%) have no extracellular prediction. However, when we compare our data to 2 previous reports on the culture filtrate proteome of M. tuberculosis H37Rv - the 2DE database at the Max Planck Institute (http://web.mpiib-berlin.mpg.de/) and a recent work by de Souza and collaborators [34] - 93 proteins (92%) have been previously reported in one or both studies, including 60 of the proteins with no extracellular prediction. We also evaluated the number of potential transmembrane (TM) domains using TMHMM ([35]; Additional file 3, Table S2). Thirteen proteins were found to contain 1 predicted TM domain which, although coinciding in all cases with the signal peptide region predicted by SignalP, does not exclude a possible membrane localization for some of these proteins [36]. For the 22 proteins with a predicted signal peptide, the theoretical pI and Mr were calculated for the full protein and for the mature protein, after removal of the signal peptide region predicted by SignalP (Additional file 4, Table S3).
Figure 1.
2DE proteomic profile of CFPs from M. bovis BCG Moreau. Proteins (500 ug) were applied to 17 cm IPG strips in the pH intervals of 3 - 6 (panels A and C) and 5 - 8 (panels B and D) and separated in the second dimension across 12% (panels A and B) and 15% (panels C and D) SDS-PAGE. The images were merged to obtain a composite map in the pH range 3 - 8 (panel E). Protein spots were visualized by colloidal CBB-G250 staining. Identified proteins are numbered in panel E and detailed in Additional file 2, Table S1. Molecular weight standards indicated in kDa.
Figure 2.
Correlation between experimentally determined and theoretical pI and Mr and distribution of predicted cellular localization of the identified proteins. The experimental and theoretical pI (panel A) and Mr (panel B) values for the identified protein were compared. Overall, a positive Spearman correlation coefficient r = 0.75, p < 0.0001 and r = 0.95, p < 0.0001, is observed for the data in the pI range between 3 and 8 and Mr range of 9 to 120 kDa, respectively.
Predicted biological functions for the identified proteins
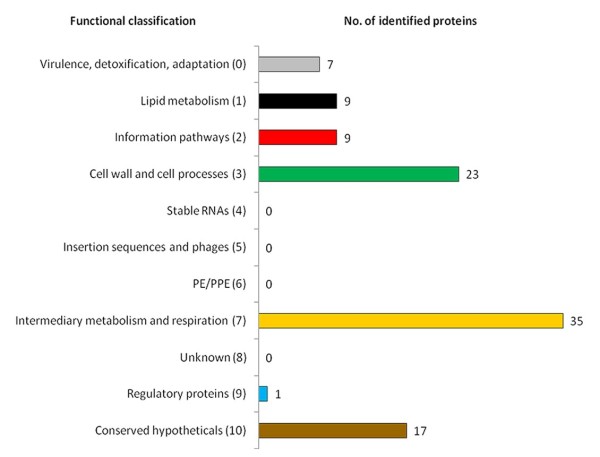
The assignment of the identified CFPs into functional categories was based on the functional classification tree from BCGList (http://genolist.pasteur.fr/BCGList/). The 101 proteins identified by MS/MS are distributed across 7 of those functional groups (Figure 3). The largest groups were "intermediary metabolism and respiration" (35%), "cell wall and cell processes" (23%) and "conserved hypotheticals" (17%).
Figure 3.
Functional classification of the identified M. bovis BCG Moreau CFPs. Identified proteins were classified into functional categories according to BCGList (http://genolist.pasteur.fr/BCGList/).
Differential CFP proteomic profiles between M. bovis BCG strains Moreau and Pasteur
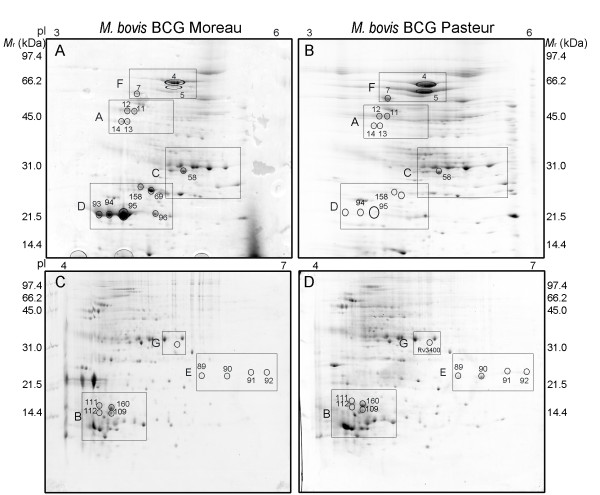
The 2DE profiles from M. bovis BCG strains Moreau and Pasteur were compared to identify differences that could provide relevant information about the Brazilian vaccine strain. For quantification analyses of the protein spots derived from both strains, the PDQuest software was used, comparing the optical densities of the matched spots in 2DE gel images. The experiments were repeated at least 3 times, and only the differences confirmed in all comparisons were accepted as strain specific. As expected, the proteomic profiles of CFPs from BCG strains Moreau and Pasteur were very similar (Figure 4 A-D); however, some variations in relative protein quantifications were observed. A total of 9 proteins represented by 18 spots showed a differential expression pattern between the two BCG strains (Table 1, Figure 5 and Additional file 5, Figure S2). In addition, 2 proteins were found exclusively in BCG Moreau and one protein exclusively in BCG Pasteur (Figure 4 A-D and Additional file 6, Figure S3).
Figure 4.
Comparative 2DE profiles of CFPs from M. bovis BCG strains Moreau and Pasteur. Proteins (500 ug) were applied to IPG strips in the pH intervals of 3 - 6 (panels A and B) and 4 - 7 (panels C and D) and separated in the second dimension in 12% (panels A and B) and 15% (panels C and D) SDS-PAGE. Protein spots were visualized by colloidal CBB-G250 staining and the gels images compared with PDQuest (Bio-Rad). Molecular weight standards indicated in kDa. The sectors shown in more detail in Additional files 5 and 6, Figures S2 and S3, are indicated in the figure (sectors A - G).
Table 1.
CFPs differentially expressed between BCG strains Moreau and Pasteur
| Spot number | Mtb ortholog | BCG Pasteur ortholog | Protein | Ratio# | Fold Increase## ± SD | p-value |
|---|---|---|---|---|---|---|
| 11### | Rv1860 | BCG1896 | Apa | M/P | 2.31 ± 0.22 | 0.09 |
| 12### | M/P | 2.01 ± 0.71 | 0.27 | |||
| 13 | M/P | 3.42 ± 1.06 | 0.02 | |||
| 14 | M/P | 3.05 ± 0.11 | 0.009 | |||
| 95 | Rv2875 | BCG2897 | Mpt70 | M/P | 39.50 ± 4.52 | 0.0004 |
| 94 | Rv2875/Rv2873 | BCG2897/BCG2895 | Mpt70/Mpt83 | M/P | 185.27 ± 30.35 | 0.004 |
| 109### | BCG1965c | M/P | 4.45 ± 1.59 | 0.19 | ||
| 111 | M/P | 2.54 ± 0.39 | 0.03 | |||
| 112### | Rv1926c | Mpt63 | M/P | 3.50 ± 0.48 | 0.41 | |
| 160 | M/P | 3.68 ± 0.23 | 0.03 | |||
| 58 | Rv1886c | BCG1923c | FbpB | M/P | 2.46 ± 0.034 | 0.01 |
| 7 | Rv2462c | BCG2482c | Tig | P/M | 3.42 ± 0.13 | 0.001 |
| 89### | BCG0009 | P/M | 2.81 ± 1.24 | 0.07 | ||
| 90 | Rv0009 | PPIase A | P/M | 2.01 ± 0.87 | 0.008 | |
| 91 | P/M | 23.28 ± 0.87 | 0.005 | |||
| 92 | P/M | 55.21 ± 12.61 | 0.05 | |||
| 4 | Rv0350 | BCG0389 | DnaK | P/M | 2.04 ± 0.21 | 0.03 |
| 5 | Rv0440 | BCG0479 | GroEL2 | P/M | 15.66 ± 0.93 | 0.00005 |
#In order to report values as fold increase, ratio was calculated for BCG Moreau (M) in relation to Pasteur (P) or vice-versa, as specified
##Ratio of mean pixel intensity value (±SD) for the specified protein spot in one BCG strain vs. the other
###Protein spots that did not show statistically significant change (p > 0.05)
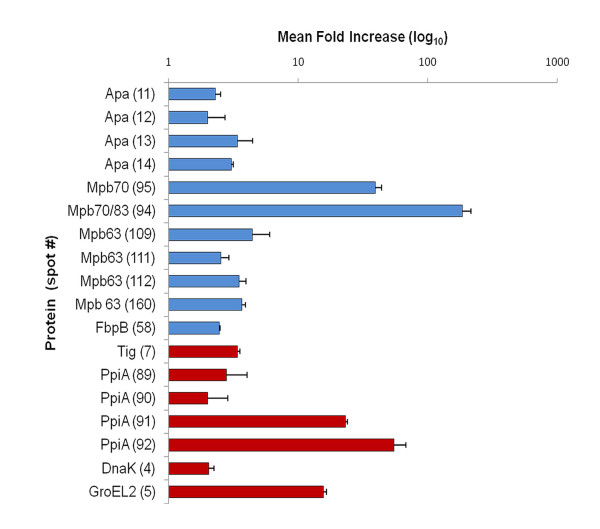
Figure 5.
CFPs differentially expressed between BCG strains Moreau and Pasteur. Bars represent fold increase (mean ± SD of the pixel intensity ratios for each specified protein spot between strains). Protein spots more expressed in BCG Moreau compared to Pasteur are represented by blue bars while those more expressed in BCG Pasteur compared to Moreau are represented by red bars. Individual values are detailed in Table 1.
Quantitative analysis revealed that 5 proteins were present in at least 2-fold higher concentration in BCG Moreau when compared to BCG Pasteur (Additional file 5, Figure S2): the Apa glycoprotein (Rv1860/BCG1896; spots 11, 12, 13 and 14); the immunogenic protein MPB63 (Rv1926c/BCG1965c; spots 109,111, 112 and 160); the secreted antigen 85B (Ag85B, FbpB, Rv1886c/BCG1923c; spot 58); and proteins MPB70 and MPB83 (Rv2875/BCG2897 and Rv2873/BCG2985; spots 94 and 95, respectively) (Table 1 and Figure 5). Spot 93 was also identified as MPB70 but was observed only in BCG Moreau (Figure 4).
Four proteins were more expressed in BCG Pasteur when compared to Moreau (Additional file 5, Figure S2): the heat shock proteins Hsp70 (DnaK, Rv0350/BCG0389; spot 4) and Hsp65 (GroEL2, Cpn60.2, Rv0440/BCG0479; spot 5); the presumed trigger factor (Tig, Rv2462c/BCG2482c; spot 7) and the probable iron-regulated peptidyl-prolyl cis-trans isomerase A (PPIaseA, Rv0009/BCG0009; spots 89, 90, 91 and 92) (Table 1 and Figure 5).
As expected, MPB64 (Rv1980c, spots 69 and 158) and CFP21 (Rv1984c; spot 96) were identified in BCG Moreau but were not present in BCG Pasteur (Figure 4 and Additional file 6, Figure S3) due to the loss of genomic region RD2 in the more recent BCG strains [7]. On the other hand, BCG Moreau contains a genomic deletion (RD16) encompassing genes rv3400-rv3405c (bcg3470-bcg3475c). In this study we identified only one protein present in BCG Pasteur and absent in BCG Moreau: a probable hydrolase encoded by rv3400 (bcg3470) (Figure 4 and Additional file 6, Figure S3). This difference is consistent with previous reports [7].
Discussion
The main goal of this study was to perform a comprehensive proteomic analysis of CFPs from M. bovis BCG strains Moreau and Pasteur by combining 2DE and mass spectrometry, allowing for the evaluation of phenotypic differences between the two strains. The 2DE patterns were highly similar, presenting numerous prominent common spots that could be used as landmarks. From 2DE gels of CFP preparations from M. bovis BCG Moreau, 158 spots were identified. The Mr and pI values estimated by 2DE showed a good correlation with expected values; however 34% of the identified proteins were detected in 2 or more spots with different Mr and/or pI. This is probably due to post-translational modifications (PTMs) such as glycosylation, phosphorylation or other modifications already described for several of the identified proteins [37-40]. For example, Rv1827 (BCG1862, Cfp17, GarA; spots 80, 81, 82) and Rv0020c (BCG0050c, TB39.8. FhaA; spots 8, 9 and 10) possess FHA domains that bind phosphothreonine [40], and Rv0685 (BCG0734, Tuf; spots 28, 29, 30 and 31) is also described as being phosphorylated in the same amino acid residues [38].
Protein modifications in prokaryotes are of great biological interest but are not yet well understood. In this work we observed several deaminated proteins (approximately 22%), possibly associated with important biological processes such as protein turnover, molecular aging and cell adhesion [41]. In addition, deamination may be useful for the refinement of protein searches by MS/MS as well as tryptophan oxidation and N-terminal pyroglutamylation [42], which are also observed in several peptides identified in this study (Additional file 2, Table S1). Interestingly, formylation was only observed for one N-terminal methionine residue in Rv1827 (BCG1862, Cfp17, GarA), a FHA domain-containing protein that constitutes the major substrate for an essential kinase, PknB, in Mtb cell extracts [43]. Formylated peptides and proteins are specific signatures of bacterial metabolism, and attractive targets to the innate immune system, serving as potent chemoattractants for mammalian phagocytic leukocytes [44]. The lack of other proteins showing this particular PTM could also indicate that peptide deformylases are operating with high efficiency. Another chemical modification observed was threonine acetylation. Although N-terminal acetylation is common in eukaryotic proteins, it has been reported to be rare in prokaryotes [45]. This PTM is present in 2 proteins identified as putative ESAT-6 like proteins, EsxI (Rv1037c, BCG1095c) and EsxN (Rv1793, BCG1825) (Additional file 2, Table S1). The N-terminal acetylation may not always alter function, but in Mtb it has been shown that antigen ESAT-6, which normally interacts with the protein CFP-10, fails to do so when acetylated [46], possibly hindering its secretion via the mycobacterial-specific type VII secretion system [47].
In the current study, only 21 (21%) of the identified proteins were found to have a predicted signal peptide. Of these, 13 have one predicted TM segment coinciding with the predicted signal peptide region. Since current methods are not fully efficient in predicting membrane retention of proteins with typical Sec-type signal peptides [36], we cannot discard the possibility that the extracellular localization of some of these proteins may result from the gradual proteolytic release of membrane proteins containing a single amino-terminal membrane anchor.
Several mycobacterial proteins that do not present a canonical signal peptide can be secreted by alternative secretion mechanisms, such as the twin-arginine translocation system, the alternative SecA2 pathway or the recently described Type VII (Esx) secretion system [48-50]. Other studies on the culture filtrate proteome of mycobacteria have also reported the presence of numerous leaderless proteins [51-53]. Some of the proteins identified by us are also reported in the membrane proteome of BCG Moreau [54] and the cell wall proteome of M. smegmatis [55]. The abundance in the culture filtrate of M. bovis BCG Moreau of proteins without signal peptide prediction may also result from bacterial lysis, in combination with high levels of protein expression and extracellular stability, as described for several Mtb proteins [56]. Nevertheless, the precise mechanism by which these proteins are exported is still unknown.
Approximately 24% of the CFPs identified in the present study have no defined function (conserved hypotheticals); among these we can highlight the conserved hypothetical proteins TB27.3 (Rv0577, BCG0622), TB18.6 (Rv2140c, BCG2175c), Rv2626c (BCG2653c) and TB15.3 (Rv1636, BCG1674) this last, recently described as being differentially expressed in the secretome of a recombinant BCG strain [57]. Although their functions have not been established, these proteins have been considered as antigens, able to distinguish between tuberculosis clinical states, or attractive targets for the development of new drugs, vaccines and diagnostic strategies for TB [58-60].
Several other mycobacterial antigens, previously described as important for vaccine development and TB diagnosis, have also been identified in the present study, including the ESAT-6 like protein EsxG (Rv0287, BCG0327), recognized by multiple T-cell lines and able to boost IFN-γ levels in mouse and guinea pig models of TB [61], and the secreted MPT51 protein (Rv3803c, BCG3865c), described as being able to induce higher levels of antigen-specific CD8+ T-cell responses [62].
Proteins involved in biosynthesis and degradation of fatty acids were also identified, such as the members of the antigen-85 complex, FbpA (Rv3804c, BCG3866c), FbpB (Rv1886c, BCG1923c), FbpC (Rv0129c, BCG0163) and FbpD (Rv3803c, BCG3865c; Mpb51), essential for the biosynthesis of mycolic acids [63]. In this work, Ag85B (FbpB) was found to be more abundant in the culture filtrate of BCG Moreau than in that of BCG Pasteur. The protein has been shown to induce partial protection against TB in animal models, and is considered an important immunodominant antigen and a promising vaccine candidate [64]. MPT64 (Rv1980) and CFP21 (Rv1984c) were found only in BCG Moreau; both are described in the literature as immunogenic antigens and potential targets for the development of new vaccines against TB [65,66]. The genes encoding these proteins are located in RD2, a genomic region deleted in a number of more recent BCG strains, including M. bovis BCG Pasteur, but present in BCG Moreau [6,7].
The low levels of MPB70 and MPB83 in M. bovis BCG Pasteur were also confirmed in our study. Their reduced expression is due to a point mutation in the translational start codon of the sigK gene [67], observed in many BCG strains, but absent in BCG Moreau. Immunologic studies have shown that both proteins induce cellular and humoral immune responses in experimental models of infection and in natural infection in humans [68,69]. MPB63 is a protein only found in species within the Mtb complex [70], shown to be immunodominant both in humans and animal models [71] and a promising candidate for serodiagnosis of active TB as well as for vaccine development. MPB63 was identified in four different spots (109,111,112 and 160), 2 of which (111 and 160) showed statistically significant differences in expression, with an increase of more than 3-fold in BCG Moreau as compared to BCG Pasteur (Table 1 and Figure 5). These 4 protein spots are likely to represent isoforms, probably differing due to the presence of PTMs, known to cause changes in pI resulting in slightly different migration. Moreover, MPB63 contains an N-terminal signal sequence as predicted by the SignalP software, which was experimentally verified [72].
The alanine-proline rich protein (Apa, Rv1860, BCG1896, spots 11, 12, 13 and 14) is known to present a high content of proline and carbohydrate groups [37] that interferes with its migratory behavior in SDS-PAGE. Although we have not identified modifications such as glycosylation, this protein displays a characteristic four-spot pattern (doublet of 2 horizontally dispersed spots) on 2DE [39] (Additional file 5, Figure S2). The isoforms of lowest molecular mass (spots 13 and 14) showed a statistically significant 3-fold increase in expression in BCG Moreau (Table 1 and Figure 5). This protein seems to be restricted to the Mtb complex and has been shown to be a target for immune recognition in animals immunized with live BCG [73]. In addition to its high immunogenicity, it has also been described as a potential adhesin involved in the colonization of target cells [39]. Its higher expression could contribute to an increase in the immunogenicity of BCG Moreau.
Four proteins were found to be at least 2-fold more expressed in BCG Pasteur compared to Moreau: a peptidyl-prolyl cis-trans isomerase (PPIaseA, Rv0009, BCG0009), the trigger factor (TF, Rv2462c, BCG2482c), Hsp65 (GroEL2, Rv0440, BCG0479) and Hsp70 (DnaK, Rv0350, BCG0389), all described as participating in protein folding and response to stress, among other functions. The gene ppiA is located in the Pasteur-specific DU1 duplicated region. The duplication of this gene alone may be responsible for the observed increased expression of PPIaseA in BCG Pasteur. Comparative transcriptome analysis has shown that bcg0009, bcg0389, bcg0479 and bcg2482c are all up-regulated in BCG Pasteur when compared to BCG Tokyo [11]. Considering the genealogy of BCG vaccines [7], BCG Moreau, Tokyo and Russia belong to the same group of "older" strains, closer to the original attenuated strain derived by Calmette and Guérin in the early 1920's, and all lack the DU1 duplication. The genome of BCG Pasteur, unlike the older strains, carries 2 copies of sigH, due to a second genomic duplication (DU2), and its expression is at least 2-fold higher [11]. SigH is an alternative extra-cytoplasmic sigma factor involved in the response to heat shock and oxidative stress, positively regulating the expression of other genes, including dnaK and possibly groEL2 [74]. GroEL2 (Rv0440, BCG0479; Hsp65) and DnaK (Rv0350, BCG0389; Hsp70) are chaperones involved in protein-folding, and have been associated with the induction of protection against TB infection in mice by immunization with experimental DNA vaccines [75,76]. Recently, these mycobacterial chaperones have been described as having vital moonlighting functions when present outside the cell: GroEL2 acts as a major adhesin, mediating binding of Mtb to monocytes and the soluble protein is capable of competing for this binding, reducing bacterial association to macrophages [77]. DnaK stimulates the secretion of chemokines required for granuloma formation [78] and its overexpression was found to favor the host over the pathogen during chronic Mtb infection [79]. All in all, subtle variations in the balance of expression and/or localization of these proteins may have profound impacts on the interaction between the bacteria (in this case, different BCG vaccine strains) and the host's immune system, impacting vaccine efficacy.
Conclusions
The findings reported here provide new information about the proteomic characteristics of the BCG Moreau vaccine strain and contribute to shed more light on the differentiated immune response and the variable effectiveness of the different BCG vaccines. In Brazil, approximately 90,000 new cases of TB are reported annually by the health system [80]. The BCG Moreau vaccine has been used since 1925, and its production by Fundação Ataulpho de Paiva (FAP) currently represents 5% of the BCG vaccine production in the world [10]. According to recent data from the WHO, global BCG immunization increased since the 1980's and Brazil, with a population close to 200 million, shows over 99% coverage for BCG vaccination [81]. Despite the genetic differences accumulated in BCG strains, the originally described protective efficacy of BCG Moreau was not reduced, and the Brazilian strain is regarded as one of the most immunogenic among the vaccine preparations that are currently available [82,83]. The exploration of the secreted sub-proteome of M. bovis BCG Moreau provides valuable information regarding specific proteins, many of which have been implicated in protective immune responses, and helps defining candidates for future vaccination strategies.
Methods
Bacterial strains and growth conditions
Mycobacterium bovis BCG Pasteur 1173P2 was obtained from the Pasteur Institute (Paris, France) culture collection, and stocks were maintained at -80°C. Mycobacterium bovis BCG Moreau was provided by Fundação Ataulpho de Paiva (FAP). Both strains were cultured as surface pellicles, for 2 weeks at 37°C, in 100 ml of Sauton vaccine production medium, provided by FAP.
Sample preparation
Culture filtrate proteins (CFPs) were obtained after separation of culture supernatants from the bacterial pellicles and subsequent centrifugation at 2,500 × g for 10 min at 4°C. The resulting supernatant was filtered through a 0.22 μm low protein binding membrane (Millipore Express; Millipore, Bedford, MA, USA) in order to remove any remaining bacteria. CFPs (on average 5.5 mg total protein) were precipitated with 17% (v/v) TCA and washed with cold acetone. Finally, proteins were dissolved in 1.5 ml of IEF buffer (8 M urea, 2% CHAPS, 4 mM tributylphosphine [TBP], 0.4% ampholytes pH 3-10) for 1 h at room temperature. Protein concentration was determined using the RC-DC Kit (Bio-Rad). Proteins were stored at -80°C until analysis.
Two dimensional gel electrophoresis (2DE)
IPG strips and all 2DE reagents were purchased from Bio-Rad (Hercules, CA, USA). Isoelectric focusing was performed at 20°C on 17 cm IPG strips, using 500 μg of CFPs diluted in a final volume of 300 μl in rehydration buffer (8 M urea, 2% CHAPS, 4 mM TBP, 0.4% ampholytes pH 3-10). Samples were applied to IPG strips (pH intervals of 3-6, 4-7 and 5-8) by in-gel rehydration and incubated for 1 h at room temperature. Isoelectric focusing was performed on a Protean® IEF cell (Bio-Rad) with maximum current of 50 μA/strip. Focusing parameters used for IPG strips in the pH range 4-7 and 5-8 were: active rehydration (50 V) for 11 h; step 1- linear gradient from 1 to 250 V over 20 min; step 2 - linear gradient from 250 to 10,000 V over 2 h; step 3- constant 10,000 V until 80,000 Vh was achieved. For IPG strips in the pH range 3-6, step 3 was constant 10,000 V until 60,000 Vh was achieved. After isoelectric focusing, proteins were reduced in 130 mM DTT and alkylated in 270 mM iodoacetamide, both in equilibration buffer (6 M urea, 2% SDS, 375 mM Tris-HCl pH 8.8, 20% glycerol). Second dimension separation was done in 17 cm, 12% or 15% SDS-PAGE gels, 1.0 mm thick, using a vertical system (Bio-Rad) in standard Laemli buffer [84] at 40 mA/gel, 10°C, until the tracking dye left the gel.
Protein visualization and image analysis
Gels were stained with colloidal Coomassie Brilliant Blue G-250 essentially as described [85], and documented using a GS-800™ auto-calibrating imaging densitometer (Bio-Rad). Image analysis was performed using PDQuest™ software version 8.0.1 (Bio-Rad). Comparative 2DE data were derived from 4 separate protein preparations, each one obtained from independent cultures. The spots were quantified on the basis of their relative 'volume': the amount of a protein spot was expressed as the sum of the intensities of all the pixels that made up the spot. To compensate for subtle differences in sample loading, gel staining and de-staining, the volume of each spot was normalized in relation to the total density of valid spots present in the gel image. After automated detection and matching, manual editing was carried out. To determine the experimental pI and Mr coordinates for each single protein spot, 2DE gels were calibrated using a selected set of five protein landmarks distributed throughout the gel.
Protein digestion, peptide extraction and MS/MS analysis
In-gel digestion of 2DE separated protein spots was carried out essentially as described [86]. Briefly, protein spots were excised and the gel pieces washed 3 times with 50% (v/v) acetonitrile (ACN) in 25 mM ammonium bicarbonate for 15 min each, dehydrated in ACN, and dried in a vacuum centrifuge. Gel pieces were rehydrated in 15 μl of 50 mM ammonium bicarbonate containing 200 ng of sequencing grade modified trypsin (Promega). This step was performed for 40 minutes at 4°C and, after that, 20 μl of 50 mM ammonium bicarbonate were added to keep the gel pieces wet during tryptic digestion (37°C, 16 h). To extract peptides, 20 μl of 0.5% (v/v) trifluoroacetic acid (TFA) in 50% (v/v) ACN were added and samples were sonicated 3 times for 10 min each in a sonicator bath. The supernatant was recovered and concentrated under vacuum to a volume of approximately 10 μl. The resulting peptides were extracted, partially dried, and salts were removed using C18 ZipPlate (Millipore, Bedford, MA) following the manufacturer's instructions.
The tryptic peptides were analyzed in a 4700-Proteomics Analyzer MALDI-TOF/TOF (Applied Biosystems, Foster City, CA). All mass spectra were acquired on positive ion reflector mode with 2,000 shots per spot and externally mass calibrated with a peptide mixture. The 10 most intense ion peaks from the peptide mass fingerprinting (or MS run) were further submitted to fragmentation using PSD mode with CID gas off and 1 keV collision energy.
Protein identification
Following MS acquisition, each spectrum was submitted to a peptide mass fingerprinting search, in the case of MS/MS spectra, using Mascot version 2.2 (Matrix Science - http://www.matrixscience.com/ ). For protein identification, the search was performed against the NCBI-nr non-redundant database (NCBI-nr200709, National Center for Biotechnology Information, http://www.ncbi.nlm.nih.gov/) without taxonomy restriction. When necessary, further searches were performed against the Mycobacterium tuberculosis database (http://genolist.pasteur.fr/tuberculist). Decoy search was performed automatically on the search form from Mascot using the same database described above with taxonomy restriction to the Mycobacterium tuberculosis complex to determine the false discovery rate (FDR). A FDR < 3.0% to peptide matches above homology or identity threshold was considered significant. For Mascot searches, the parameters used were trypsin as the enzyme of choice and one missed cleavage, ± 1 Da for the precursor mass, ± 0.5 Da for the fragment ion mass. Oxidation of methionines along with N-terminal acetylation of proteins, N-terminal formylation, deamidation and cyclization of glutamine (pyro-glutamate) were allowed as possible modifications whereas alkylation of cysteines (carbamidomethylcysteines) was set as constant modification. Identification was considered valid for Mascot protein scores greater than 30 and a significance threshold of p < 0.05. If a protein 'hit' was identified by only one peptide, the MS/MS data was to exhibit a clear spectrum with sequence tags that matched at least three consecutive y or b fragment ion series. Finally, a good correlation between the experimental and theoretical molecular mass and pI was also considered for positive identifications.
Putative signals for protein export were predicted using SignalP 3.0 (http://www.cbs.dtu.dk/services/SignalP/), LipoP 1.0 (http://www.cbs.dtu.dk/services/ LipoP/), TatP 1.0 (http://www.cbs.dtu.dk/services/TatP/) and SecretomeP 2.0 (http://www.cbs.dtu.dk/services/SecretomeP/). Potential transmembrane domais were predicted with TMHMM 2.0 (http://www.cbs.dtu.dk/services/TMHMM/).
Molecular weight (Mr) and pI of secreted proteins was calculated with the Expasy compute pI/Mw tool (http://www.expasy.ch/tools/pi_tool.html).
Statistical analysis
Spot intensity differences obtained from comparative 2DE gel images of M. bovis BCG strains Moreau and Pasteur were statistically analyzed by one-way ANOVA with Student's t-test to determine significant differences among group means. Statistical analysis was carried out using the data obtained from 4 different sets of independent biological samples. A p-value ≤0.05 was considered as statistically significant.
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
MBP contributed in the experimental design, data acquisition and interpretation and was involved in writing the manuscript. DEK carried out the mass spectrometry analysis and protein identification. PCR and MPP contributed to data acquisition. LHFG carried out the PCR assays. RFS provided technical assistance. LRRCB contributed to data interpretation and manuscript revision. WMD took part in supervision, data interpretation and writing the manuscript. LML was responsible for the experimental design, supervision, data interpretation and writing the manuscript. All authors have read and approved the final manuscript.
Supplementary Material
Figure S1 - PCR confirmation of the genetic identity of the BCG strains used.
Table S1 - M. bovis BCG Moreau culture filtrate proteins identified by MS/MS
Table S2 - Predicted localization of identified proteins.
Table S3 - Mr and pI of secreted proteins.
Figure S2 - Magnified 2DE gel regions showing protein spots differentially expressed between BCG strains Moreau and Pasteur. Panels A - F represent the magnified gel regions indicated in Figure 4. Protein spot numbering is the same as in Figure 1.
Figure S3 - Magnified 2DE gel regions showing protein spots expressed exclusively in BCG strains Moreau or Pasteur. Panels A and B represent the magnified gel regions as indicated in Figure 4. Protein spot numbering is the same as in Figure 1. MPT64 (spots 69 and 158) and CFP21 (spot 96) are only found in BCG Moreau culture filtrate (panel A), while Rv3400 (BCG3470) was only found in BCG Pasteur (panel B).
Contributor Information
Marcia Berrêdo-Pinho, Email: mberredo@ioc.fiocruz.br.
Dario E Kalume, Email: kalume@ioc.fiocruz.br.
Paloma R Correa, Email: pacorrea@ioc.fiocruz.br.
Leonardo HF Gomes, Email: leonardoh@ioc.fiocruz.br.
Melissa P Pereira, Email: melissa@ioc.fiocruz.br.
Renata F da Silva, Email: renatafs@ioc.fiocruz.br.
Luiz RR Castello-Branco, Email: branco@ioc.fiocruz.br.
Wim M Degrave, Email: wdegrave@fiocruz.br.
Leila Mendonça-Lima, Email: lmlima@ioc.fiocruz.br.
Acknowledgements
We thank Rodrigo Mexas (Laboratório de Produção e Tratamento de Imagem, IOC/FIOCRUZ) for his precious contributions and the FIOCRUZ/PDTIS 2DE and Mass Spectrometry platform facilities (Dr. J. Perales and André Ferreira). Carolina Zavareze (FAP) kindly provided the Sauton culture medium and the BCG Moreau vaccine strain. This work received financial support from the WHO/TDR Special Programme for Research Training in Tropical Diseases and the following Brazilian agencies: CNPq, FAPERJ and PDTIS/FIOCRUZ.
References
- WHO. Global Tuberculosis Control, Surveillance, Planning, Financing. Geneva: World Health Organization; 2008. [Google Scholar]
- Dye C. Global epidemiology of tuberculosis. Lancet. 2006;367:938–940. doi: 10.1016/S0140-6736(06)68384-0. [DOI] [PubMed] [Google Scholar]
- Aziz MA, Wright A, Laszlo A, De Muynck A, Portaels F, Van Deun A, Wells C, Nunn P, Blanc L, Raviglione M. Epidemiology of antituberculosis drug resistance (the Global Project on Anti-tuberculosis Drug Resistance Surveillance): an updated analysis. Lancet. 2006;368:2142–2154. doi: 10.1016/S0140-6736(06)69863-2. [DOI] [PubMed] [Google Scholar]
- Ritz N, Curtis N. Mapping the global use of different BCG vaccine strains. Tuberculosis (Edinb) 2009;89:248–251. doi: 10.1016/j.tube.2009.03.002. [DOI] [PubMed] [Google Scholar]
- Calmette A, Guerin C, Negre L, Bocquet A. Sur la vaccination preventive des enfants nouveau-nés contre la tuberculose par le BCG. Ann Inst Pasteur (Paris) 1927;3:201–208. [Google Scholar]
- Mahairas GG, Sabo PJ, Hickey MJ, Singh DC, Stover CK. Molecular analysis of genetic differences between Mycobacterium bovis BCG and virulent M. bovis. J Bacteriol. 1996;178:1274–1282. doi: 10.1128/jb.178.5.1274-1282.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Behr MA, Wilson MA, Gill WP, Salamon H, Schoolnik GK, Rane S, Small PM. Comparative genomics of BCG vaccines by whole-genome DNA microarray. Science. 1999;284:1520–1523. doi: 10.1126/science.284.5419.1520. [DOI] [PubMed] [Google Scholar]
- Gordon SV, Brosch R, Billault A, Garnier T, Eiglmeier K, Cole ST. Identification of variable regions in the genomes of tubercle bacilli using bacterial artificial chromosome arrays. Mol Microbiol. 1999;32:643–655. doi: 10.1046/j.1365-2958.1999.01383.x. [DOI] [PubMed] [Google Scholar]
- Brosch R, Pym AS, Gordon SV, Cole ST. The evolution of mycobacterial pathogenicity: clues from comparative genomics. Trends Microbiol. 2001;9:452–458. doi: 10.1016/S0966-842X(01)02131-X. [DOI] [PubMed] [Google Scholar]
- Benevolo-de-Andrade TC, Monteiro-Maia R, Cosgrove C, Castello-Branco LR. BCG Moreau Rio de Janeiro: an oral vaccine against tuberculosis--review. Mem Inst Oswaldo Cruz. 2005;100:459–465. doi: 10.1590/S0074-02762005000500002. [DOI] [PubMed] [Google Scholar]
- Brosch R, Gordon SV, Garnier T, Eiglmeier K, Frigui W, Valenti P, Dos Santos S, Duthoy S, Lacroix C, Garcia-Pelayo C, Inwald JK, Golby P, Garcia JN, Hewinson RG, Behr MA, Quail MA, Churcher C, Barrell BG, Parkhill J, Cole ST. Genome plasticity of BCG and impact on vaccine efficacy. Proc Natl Acad Sci USA. 2007;104:5596–5601. doi: 10.1073/pnas.0700869104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodrigues LC, Kerr-Pontes LR, Frietas MV, Barreto ML. Long lasting BCG protection against leprosy. Vaccine. 2007;25:6842–6844. doi: 10.1016/j.vaccine.2007.07.032. [DOI] [PubMed] [Google Scholar]
- Andersen P, Doherty TM. The success and failure of BCG - implications for a novel tuberculosis vaccine. Nat Rev Microbiol. 2005;3:656–662. doi: 10.1038/nrmicro1211. [DOI] [PubMed] [Google Scholar]
- Antas PR, Castello-Branco LR. New vaccines against tuberculosis: lessons learned from BCG immunisation in Brazil. Trans R Soc Trop Med Hyg. 2008;102:628–630. doi: 10.1016/j.trstmh.2008.03.014. [DOI] [PubMed] [Google Scholar]
- Castillo-Rodal AI, Castanon-Arreola M, Hernandez-Pando R, Calva JJ, Sada-Diaz E, Lopez-Vidal Y. Mycobacterium bovis BCG substrains confer different levels of protection against Mycobacterium tuberculosis infection in a BALB/c model of progressive pulmonary tuberculosis. Infect Immun. 2006;74:1718–1724. doi: 10.1128/IAI.74.3.1718-1724.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodriguez-Alvarez M, Mendoza-Hernandez G, Encarnacion S, Calva JJ, Lopez-Vidal Y. Phenotypic differences between BCG vaccines at the proteome level. Tuberculosis (Edinb) 2009;89:126–135. doi: 10.1016/j.tube.2008.12.001. [DOI] [PubMed] [Google Scholar]
- Brandt L, Feino Cunha J, Weinreich Olsen A, Chilima B, Hirsch P, Appelberg R, Andersen P. Failure of the Mycobacterium bovis BCG vaccine: some species of environmental mycobacteria block multiplication of BCG and induction of protective immunity to tuberculosis. Infect Immun. 2002;70:672–678. doi: 10.1128/IAI.70.2.672-678.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Colditz GA, Brewer TF, Berkey CS, Wilson ME, Burdick E, Fineberg HV, Mosteller F. Efficacy of BCG vaccine in the prevention of tuberculosis. Meta-analysis of the published literature. JAMA. 1994;271:698–702. doi: 10.1001/jama.271.9.698. [DOI] [PubMed] [Google Scholar]
- Fine PE, Carneiro IA, Milstien JB, Clements CJ. Issues Relating to the Use of BCG in Immunisation Programmes. A discussion document. Geneva: World Health Organisation. Department of Vaccines and Biologicals; 1999. pp. 1–45. 1-45. [Google Scholar]
- Trajkovic V, Natarajan K, Sharma P. Immunomodulatory action of mycobacterial secretory proteins. Microbes Infect. 2004;6:513–519. doi: 10.1016/j.micinf.2003.12.015. [DOI] [PubMed] [Google Scholar]
- Malen H, Berven FS, Fladmark KE, Wiker HG. Comprehensive analysis of exported proteins from Mycobacterium tuberculosis H37Rv. Proteomics. 2007;7:1702–1718. doi: 10.1002/pmic.200600853. [DOI] [PubMed] [Google Scholar]
- Hubbard RD, Flory CM, Collins FM. Immunization of mice with mycobacterial culture filtrate proteins. Clin Exp Immunol. 1992;87:94–98. doi: 10.1111/j.1365-2249.1992.tb06419.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andersen P. Effective vaccination of mice against Mycobacterium tuberculosis infection with a soluble mixture of secreted mycobacterial proteins. Infect Immun. 1994;62:2536–2544. doi: 10.1128/iai.62.6.2536-2544.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horwitz MA, Harth G, Dillon BJ, Maslesa-Galic S. Recombinant bacillus calmette-guerin (BCG) vaccines expressing the Mycobacterium tuberculosis 30-kDa major secretory protein induce greater protective immunity against tuberculosis than conventional BCG vaccines in a highly susceptible animal model. Proc Natl Acad Sci USA. 2000;97:13853–13858. doi: 10.1073/pnas.250480397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamath AT, Rochat AF, Valenti MP, Agger EM, Lingnau K, Andersen P, Lambert PH, Siegrist CA. Adult-like anti-mycobacterial T cell and in vivo dendritic cell responses following neonatal immunization with Ag85B-ESAT-6 in the IC31 adjuvant. PLoS One. 2008;3:e3683. doi: 10.1371/journal.pone.0003683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Badell E, Nicolle F, Clark S, Majlessi L, Boudou F, Martino A, Castello-Branco L, Leclerc C, Lewis DJ, Marsh PD, Gicquel B, Winter N. Protection against tuberculosis induced by oral prime with Mycobacterium bovis BCG and intranasal subunit boost based on the vaccine candidate Ag85B-ESAT-6 does not correlate with circulating IFN-gamma producing T-cells. Vaccine. 2009;27:28–37. doi: 10.1016/j.vaccine.2008.10.034. [DOI] [PubMed] [Google Scholar]
- Boesen H, Jensen BN, Wilcke T, Andersen P. Human T-cell responses to secreted antigen fractions of Mycobacterium tuberculosis. Infect Immun. 1995;63:1491–1497. doi: 10.1128/iai.63.4.1491-1497.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Målen H, Softeland T, Wiker HG. Antigen analysis of Mycobacterium tuberculosis H37Rv culture filtrate proteins. Scand J Immunol. 2008;67:245–252. doi: 10.1111/j.1365-3083.2007.02064.x. [DOI] [PubMed] [Google Scholar]
- Liu J, Tran V, Leung AS, Alexander DC, Zhu B. BCG vaccines: their mechanisms of attenuation and impact on safety and protective efficacy. Hum Vaccin. 2009;5:70–78. doi: 10.4161/hv.5.2.7210. [DOI] [PubMed] [Google Scholar]
- Bendtsen JD, Nielsen H, von Heijne G, Brunak S. Improved prediction of signal peptides: SignalP 3.0. J Mol Biol. 2004;340:783–795. doi: 10.1016/j.jmb.2004.05.028. [DOI] [PubMed] [Google Scholar]
- Juncker AS, Willenbrock H, Von Heijne G, Brunak S, Nielsen H, Krogh A. Prediction of lipoprotein signal peptides in Gram-negative bacteria. Protein Sci. 2003;12:1652–1662. doi: 10.1110/ps.0303703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bendtsen JD, Nielsen H, Widdick D, Palmer T, Brunak S. Prediction of twin-arginine signal peptides. BMC Bioinformatics. 2005;6:167. doi: 10.1186/1471-2105-6-167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bendtsen JD, Kiemer L, Fausboll A, Brunak S. Non-classical protein secretion in bacteria. BMC Microbiol. 2005;5:58. doi: 10.1186/1471-2180-5-58. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Souza GA, Malen H, Softeland T, Saelensminde G, Prasad S, Jonassen I, Wiker HG. High accuracy mass spectrometry analysis as a tool to verify and improve gene annotation using Mycobacterium tuberculosis as an example. BMC Genomics. 2008;9:316. doi: 10.1186/1471-2164-9-316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krogh A, Larsson B, von Heijne G, Sonnhammer EL. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol. 2001;305:567–580. doi: 10.1006/jmbi.2000.4315. [DOI] [PubMed] [Google Scholar]
- Tjalsma H, van Dijl JM. Proteomics-based consensus prediction of protein retention in a bacterial membrane. Proteomics. 2005;5:4472–4482. doi: 10.1002/pmic.200402080. [DOI] [PubMed] [Google Scholar]
- Horn C, Namane A, Pescher P, Riviere M, Romain F, Puzo G, Barzu O, Marchal G. Decreased capacity of recombinant 45/47-kDa molecules (Apa) of Mycobacterium tuberculosis to stimulate T lymphocyte responses related to changes in their mannosylation pattern. J Biol Chem. 1999;274:32023–32030. doi: 10.1074/jbc.274.45.32023. [DOI] [PubMed] [Google Scholar]
- Archambaud C, Gouin E, Pizarro-Cerda J, Cossart P, Dussurget O. Translation elongation factor EF-Tu is a target for Stp, a serine-threonine phosphatase involved in virulence of Listeria monocytogenes. Mol Microbiol. 2005;56:383–396. doi: 10.1111/j.1365-2958.2005.04551.x. [DOI] [PubMed] [Google Scholar]
- Ragas A, Roussel L, Puzo G, Riviere M. The Mycobacterium tuberculosis cell-surface glycoprotein apa as a potential adhesin to colonize target cells via the innate immune system pulmonary C-type lectin surfactant protein A. J Biol Chem. 2007;282:5133–5142. doi: 10.1074/jbc.M610183200. [DOI] [PubMed] [Google Scholar]
- England P, Wehenkel A, Martins S, Hoos S, Andre-Leroux G, Villarino A, Alzari PM. The FHA-containing protein GarA acts as a phosphorylation-dependent molecular switch in mycobacterial signaling. FEBS Lett. 2009;583:301–307. doi: 10.1016/j.febslet.2008.12.036. [DOI] [PubMed] [Google Scholar]
- Weintraub SJ, Manson SR. Asparagine deamidation: a regulatory hourglass. Mech Ageing Dev. 2004;125:255–257. doi: 10.1016/j.mad.2004.03.002. [DOI] [PubMed] [Google Scholar]
- Gupta N, Tanner S, Jaitly N, Adkins JN, Lipton M, Edwards R, Romine M, Osterman A, Bafna V, Smith RD, Pevzner PA. Whole proteome analysis of post-translational modifications: applications of mass-spectrometry for proteogenomic annotation. Genome Res. 2007;17:1362–1377. doi: 10.1101/gr.6427907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villarino A, Duran R, Wehenkel A, Fernandez P, England P, Brodin P, Cole ST, Zimny-Arndt U, Jungblut PR, Cervenansky C, Alzari PM. Proteomic identification of M. tuberculosis protein kinase substrates: PknB recruits GarA, a FHA domain-containing protein, through activation loop-mediated interactions. J Mol Biol. 2005;350:953–963. doi: 10.1016/j.jmb.2005.05.049. [DOI] [PubMed] [Google Scholar]
- Schiffmann E, Corcoran BA, Wahl SM. N-formylmethionyl peptides as chemoattractants for leucocytes. Proc Natl Acad Sci USA. 1975;72:1059–1062. doi: 10.1073/pnas.72.3.1059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Polevoda B, Sherman F. The diversity of acetylated proteins. Genome Biol. 2002;3:reviews0006. doi: 10.1186/gb-2002-3-5-reviews0006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okkels LM, Muller EC, Schmid M, Rosenkrands I, Kaufmann SH, Andersen P, Jungblut PR. CFP10 discriminates between nonacetylated and acetylated ESAT-6 of Mycobacterium tuberculosis by differential interaction. Proteomics. 2004;4:2954–2960. doi: 10.1002/pmic.200400906. [DOI] [PubMed] [Google Scholar]
- Abdallah AM, Gey van Pittius NC, Champion PA, Cox J, Luirink J, Vandenbroucke-Grauls CM, Appelmelk BJ, Bitter W. Type VII secretion--mycobacteria show the way. Nat Rev Microbiol. 2007;5:883–891. doi: 10.1038/nrmicro1773. [DOI] [PubMed] [Google Scholar]
- Posey JE, Shinnick TM, Quinn FD. Characterization of the twin-arginine translocase secretion system of Mycobacterium smegmatis. J Bacteriol. 2006;188:1332–1340. doi: 10.1128/JB.188.4.1332-1340.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bitter W, Houben EN, Bottai D, Brodin P, Brown EJ, Cox JS, Derbyshire K, Fortune SM, Gao LY, Liu J, Liu J, Gey van Pittius NC, Pym AS, Rubin EJ, Sherman DR, Cole ST, Brosch R. Systematic genetic nomenclature for type VII secretion systems. PLoS Pathog. 2009;5:e1000507. doi: 10.1371/journal.ppat.1000507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rigel NW, Braunstein M. A new twist on an old pathway--accessory Sec [corrected] systems. Mol Microbiol. 2008;69:291–302. doi: 10.1111/j.1365-2958.2008.06294.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosenkrands I, King A, Weldingh K, Moniatte M, Moertz E, Andersen P. Towards the proteome of Mycobacterium tuberculosis. Electrophoresis. 2000;21:3740–3756. doi: 10.1002/1522-2683(200011)21:17<3740::AID-ELPS3740>3.0.CO;2-3. [DOI] [PubMed] [Google Scholar]
- Rosenkrands I, Weldingh K, Jacobsen S, Hansen CV, Florio W, Gianetri I, Andersen P. Mapping and identification of Mycobacterium tuberculosis proteins by two-dimensional gel electrophoresis, microsequencing and immunodetection. Electrophoresis. 2000;21:935–948. doi: 10.1002/(SICI)1522-2683(20000301)21:5<935::AID-ELPS935>3.0.CO;2-P. [DOI] [PubMed] [Google Scholar]
- Mattow J, Schaible UE, Schmidt F, Hagens K, Siejak F, Brestrich G, Haeselbarth G, Muller EC, Jungblut PR, Kaufmann SH. Comparative proteome analysis of culture supernatant proteins from virulent Mycobacterium tuberculosis H37Rv and attenuated M. bovis BCG Copenhagen. Electrophoresis. 2003;24:3405–3420. doi: 10.1002/elps.200305601. [DOI] [PubMed] [Google Scholar]
- Målen H, Berven FS, Softeland T, Arntzen MO, D'Santos CS, De Souza GA, Wiker HG. Membrane and membrane-associated proteins in Triton X-114 extracts of Mycobacterium bovis BCG identified using a combination of gel-based and gel-free fractionation strategies. Proteomics. 2008;8:1859–1870. doi: 10.1002/pmic.200700528. [DOI] [PubMed] [Google Scholar]
- He Z, De Buck J. Cell wall proteome analysis of Mycobacterium smegmatis strain MC2 155. BMC Microbiol. p. 121. [DOI] [PMC free article] [PubMed]
- Tullius MV, Harth G, Horwitz MA. High extracellular levels of Mycobacterium tuberculosis glutamine synthetase and superoxide dismutase in actively growing cultures are due to high expression and extracellular stability rather than to a protein-specific export mechanism. Infect Immun. 2001;69:6348–6363. doi: 10.1128/IAI.69.10.6348-6363.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodriguez-Alvarez M, Palomec-Nava ID, Mendoza-Hernandez G, Lopez-Vidal Y. The secretome of a recombinant BCG substrain reveals differences in hypothetical proteins. Vaccine. pp. 3997–4001. [DOI] [PubMed]
- Benabdesselem C, Fathallah DM, Huard RC, Zhu H, Jarboui MA, Dellagi K, Ho JL, Barbouche RM. Enhanced patient serum immunoreactivity to recombinant Mycobacterium tuberculosis CFP32 produced in the yeast Pichia pastoris compared to Escherichia coli and its potential for serodiagnosis of tuberculosis. J Clin Microbiol. 2006;44:3086–3093. doi: 10.1128/JCM.02672-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roupie V, Romano M, Zhang L, Korf H, Lin MY, Franken KL, Ottenhoff TH, Klein MR, Huygen K. Immunogenicity of eight dormancy regulon-encoded proteins of Mycobacterium tuberculosis in DNA-vaccinated and tuberculosis-infected mice. Infect Immun. 2007;75:941–949. doi: 10.1128/IAI.01137-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weldingh K, Hansen A, Jacobsen S, Andersen P. High resolution electroelution of polyacrylamide gels for the purification of single proteins from Mycobacterium tuberculosis culture filtrate. Scand J Immunol. 2000;51:79–86. doi: 10.1046/j.1365-3083.2000.00655.x. [DOI] [PubMed] [Google Scholar]
- Coler RN, Dillon DC, Skeiky YA, Kahn M, Orme IM, Lobet Y, Reed SG, Alderson MR. Identification of Mycobacterium tuberculosis vaccine candidates using human CD4+ T-cells expression cloning. Vaccine. 2009;27:223–233. doi: 10.1016/j.vaccine.2008.10.056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uchijima M, Nagata T, Koide Y. Chemokine receptor-mediated delivery of mycobacterial MPT51 protein efficiently induces antigen-specific T-cell responses. Vaccine. 2008;26:5165–5169. doi: 10.1016/j.vaccine.2008.03.059. [DOI] [PubMed] [Google Scholar]
- Belisle JT, Vissa VD, Sievert T, Takayama K, Brennan PJ, Besra GS. Role of the major antigen of Mycobacterium tuberculosis in cell wall biogenesis. Science. 1997;276:1420–1422. doi: 10.1126/science.276.5317.1420. [DOI] [PubMed] [Google Scholar]
- Qie YQ, Wang JL, Zhu BD, Xu Y, Wang QZ, Chen JZ, Wang HH. Evaluation of a new recombinant BCG which contains mycobacterial antigen ag85B-mpt64(190-198)-mtb8.4 in C57/BL6 mice. Scand J Immunol. 2008;67:133–139. doi: 10.1111/j.1365-3083.2007.02048.x. [DOI] [PubMed] [Google Scholar]
- Luo Y, Wang B, Hu L, Yu H, Da Z, Jiang W, Song N, Qie Y, Wang H, Tang Z, Xian Q, Zhang Y, Zhu B. Fusion protein Ag85B-MPT64(190-198)-Mtb8.4 has higher immunogenicity than Ag85B with capacity to boost BCG-primed immunity against Mycobacterium tuberculosis in mice. Vaccine. 2009;27:6179–6185. doi: 10.1016/j.vaccine.2009.08.018. [DOI] [PubMed] [Google Scholar]
- Lv H, Gao Y, Wu Y, Zhai M, Li L, Zhu Y, Liu W, Wu Z, Chen F, Qi Y. Identification of a novel cytotoxic T lymphocyte epitope from CFP21, a secreted protein of Mycobacterium tuberculosis. Immunol Lett. [DOI] [PubMed]
- Charlet D, Mostowy S, Alexander D, Sit L, Wiker HG, Behr MA. Reduced expression of antigenic proteins MPB70 and MPB83 in Mycobacterium bovis BCG strains due to a start codon mutation in sigK. Mol Microbiol. 2005;56:1302–1313. doi: 10.1111/j.1365-2958.2005.04618.x. [DOI] [PubMed] [Google Scholar]
- Lyashchenko KP, Wiker HG, Harboe M, McNair J, Komissarenko SV, Pollock JM. Novel monoclonal antibodies against major antigens of Mycobacterium bovis. Scand J Immunol. 2001;53:498–502. doi: 10.1046/j.1365-3083.2001.00914.x. [DOI] [PubMed] [Google Scholar]
- Wiker HG. Liberation of soluble proteins from live and dead mycobacterial cells and the implications for pathogenicity of tubercle bacilli hypothesis. Scand J Immunol. 2001;54:82–86. doi: 10.1046/j.1365-3083.2001.00946.x. [DOI] [PubMed] [Google Scholar]
- Manca C, Lyashchenko K, Wiker HG, Usai D, Colangeli R, Gennaro ML. Molecular cloning, purification, and serological characterization of MPT63, a novel antigen secreted by Mycobacterium tuberculosis. Infect Immun. 1997;65:16–23. doi: 10.1128/iai.65.1.16-23.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagai S, Wiker HG, Harboe M, Kinomoto M. Isolation and partial characterization of major protein antigens in the culture fluid of Mycobacterium tuberculosis. Infect Immun. 1991;59:372–382. doi: 10.1128/iai.59.1.372-382.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Målen H, Berven FS, Fladmark KE, Wiker HG. Comprehensive analysis of exported proteins from Mycobacterium tuberculosis H37Rv. Proteomics. 2007;7:1702–1718. doi: 10.1002/pmic.200600853. [DOI] [PubMed] [Google Scholar]
- Romain F, Laqueyrerie A, Militzer P, Pescher P, Chavarot P, Lagranderie M, Auregan G, Gheorghiu M, Marchal G. Identification of a Mycobacterium bovis BCG 45/47-kilodalton antigen complex, an immunodominant target for antibody response after immunization with living bacteria. Infect Immun. 1993;61:742–750. doi: 10.1128/iai.61.2.742-750.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mehra S, Kaushal D. Functional genomics reveals extended roles of the Mycobacterium tuberculosis stress response factor sigmaH. J Bacteriol. 2009;191:3965–3980. doi: 10.1128/JB.00064-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferraz JC, Stavropoulos E, Yang M, Coade S, Espitia C, Lowrie DB, Colston MJ, Tascon RE. A heterologous DNA priming-Mycobacterium bovis BCG boosting immunization strategy using mycobacterial Hsp70, Hsp65, and Apa antigens improves protection against tuberculosis in mice. Infect Immun. 2004;72:6945–6950. doi: 10.1128/IAI.72.12.6945-6950.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Souza PR, Zarate-Blades CR, Hori JI, Ramos SG, Lima DS, Schneider T, Rosada RS, Torre LG, Santana MH, Brandao IT, Masson AP, Coelho-Castelo AA, Bonato VL, Galetti FC, Gonçalves ED, Botte DA, Machado JB, Silva CL. Protective efficacy of different strategies employing Mycobacterium leprae heat-shock protein 65 against tuberculosis. Expert Opin Biol Ther. 2008;8:1255–1264. doi: 10.1517/14712598.8.9.1255. [DOI] [PubMed] [Google Scholar]
- Hickey TB, Thorson LM, Speert DP, Daffe M, Stokes RW. Mycobacterium tuberculosis Cpn60.2 and DnaK are located on the bacterial surface, where Cpn60.2 facilitates efficient bacterial association with macrophages. Infect Immun. 2009;77:3389–3401. doi: 10.1128/IAI.00143-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henderson B, Lund PA, Coates AR. Multiple moonlighting functions of mycobacterial molecular chaperones. Tuberculosis (Edinb) pp. 119–124. [DOI] [PubMed]
- Stewart GR, Snewin VA, Walzl G, Hussell T, Tormay P, O'Gaora P, Goyal M, Betts J, Brown IN, Young DB. Overexpression of heat-shock proteins reduces survival of Mycobacterium tuberculosis in the chronic phase of infection. Nat Med. 2001;7:732–737. doi: 10.1038/89113. [DOI] [PubMed] [Google Scholar]
- WHO. WHO REPORT Global Tuberculosis control Brazil. World Health Organization; 2009. [Google Scholar]
- Immunization W-EPo. WHO vaccine-preventable diseases:monitoring system - 2009 global summary. 2009. pp. 243–380.
- Hayashi D, Takii T, Fujiwara N, Fujita Y, Yano I, Yamamoto S, Kondo M, Yasuda E, Inagaki E, Kanai K, Fujiwara A, Kawarazaki A, Chiba T, Onozaki K. Comparable studies of immunostimulating activities in vitro among Mycobacterium bovis bacillus Calmette-Guerin (BCG) substrains. FEMS Immunol Med Microbiol. 2009;56:116–128. doi: 10.1111/j.1574-695X.2009.00559.x. [DOI] [PubMed] [Google Scholar]
- Ladefoged A, Bunch-Christensen K, Guld J. Tuberculin sensitivity in guinea-pigs after vaccination with varying doses of BCG of 12 different strains. Bull World Health Organ. 1976;53:435–443. [PMC free article] [PubMed] [Google Scholar]
- Laemmli UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227:680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Neuhoff V, Arold N, Taube D, Ehrhardt W. Improved staining of proteins in polyacrylamide gels including isoelectric focusing gels with clear background at nanogram sensitivity using Coomassie Brilliant Blue G-250 and R-250. Electrophoresis. 1988;9:255–262. doi: 10.1002/elps.1150090603. [DOI] [PubMed] [Google Scholar]
- Shevchenko A, Tomas H, Havlis J, Olsen JV, Mann M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat Protoc. 2006;1:2856–2860. doi: 10.1038/nprot.2006.468. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figure S1 - PCR confirmation of the genetic identity of the BCG strains used.
Table S1 - M. bovis BCG Moreau culture filtrate proteins identified by MS/MS
Table S2 - Predicted localization of identified proteins.
Table S3 - Mr and pI of secreted proteins.
Figure S2 - Magnified 2DE gel regions showing protein spots differentially expressed between BCG strains Moreau and Pasteur. Panels A - F represent the magnified gel regions indicated in Figure 4. Protein spot numbering is the same as in Figure 1.
Figure S3 - Magnified 2DE gel regions showing protein spots expressed exclusively in BCG strains Moreau or Pasteur. Panels A and B represent the magnified gel regions as indicated in Figure 4. Protein spot numbering is the same as in Figure 1. MPT64 (spots 69 and 158) and CFP21 (spot 96) are only found in BCG Moreau culture filtrate (panel A), while Rv3400 (BCG3470) was only found in BCG Pasteur (panel B).