Abstract
Background
The wild barley Hordeum chilense Roem. et Schult. is a valuable source of genes for increasing carotenoid content in wheat. Tritordeums, the amphiploids derived from durum or common wheat and H. chilense, systematically show higher values of yellow pigment colour and carotenoid content than durum wheat. Phytoene synthase 1 gene (Psy1) is considered a key step limiting the carotenoid biosynthesis, and the correlation of Psy1 transcripts accumulation and endosperm carotenoid content has been demonstrated in the main grass species.
Methodology/Principal findings
We analyze the variability of Psy1 alleles in three lines of H. chilense (H1, H7 and H16) representing the three ecotypes described in this species. Moreover, we analyze Psy1 expression in leaves and in two seed developing stages of H1 and H7, showing mRNA accumulation patterns similar to those of wheat. Finally, we identify thirty-six different transcripts forms originated by alternative splicing of the 5′ UTR and/or exons 1 to 5 of Psy1 gene. Transcripts function is tested in a heterologous complementation assay, revealing that from the sixteen different predicted proteins only four types (those of 432, 370, 364 and 271 amino acids), are functional in the bacterial system.
Conclusions/Significance
The large number of transcripts originated by alternative splicing of Psy1, and the coexistence of functional and non functional forms, suggest a fine regulation of PSY activity in H. chilense. This work is the first analysis of H. chilense Psy1 gene and the results reported here are the bases for its potential use in carotenoid enhancement in durum wheat.
Introduction
Carotenoids are a diverse family of isoprenoid derived pigments found in plants, fungi and bacteria. They are involved in many critical plant processes such as photosynthesis, abscisic acid synthesis or prevention of photo-oxidative damage [1]. Regarding human nutrition, carotenoids are precursors of vitamin A and other nutritional factors, having essential roles in human health as antioxidants. Therefore, in the last years, increasing carotenoid content in seed endosperm of the main crop staples (rice, maize and wheat) has been a breeding objective and great efforts have been conducted to better understand carotenoid biosynthetic pathways and regulation [2], [3].
The first step of the carotenoid pathway is catalyzed by the enzyme phytoene synthase (PSY) which is encoded by three paralogous genes in grasses [4]. From the three paralogs, Psy1, Psy2 and Psy3, only Psy1 is related to endosperm carotenoid content [4]–[6] being considered a rate-limiting step in carotenoid accumulation.
Commonly, carotenoid content has not been a direct target for breeding in wheat. Instead, yellow pigment content (YPC) has been generally considered. A bright yellow colour is required for durum wheat (Triticum turgidum) for pasta production and, therefore, durum wheat breeding programs worldwide have selected high YPC varieties.
QTLs for YPC have been identified in several chromosomes in both durum and bread wheat, being those located on the chromosomes from the homoeologous group 7 the main responsible of the variation. Psy1 was considered a candidate gene to explain the high seed YPC of wheat grain since it plays a critical role in the carotenoid pathway and maps to chromosomes 7A and 7B of durum and bread wheat [7], [8]. From the first report confirming the association of Psy-B1 with variation in YPC in durum wheat seeds [8], several works have demonstrated co-segregation of Psy-A1 and Psy-B1 with QTLs for YPC in both durum and bread wheat [9]–[12]. Additionally, the role of Psy1 genes in YPC content has been also suggested using association studies and Psy1 allele-specific markers of high and low YPC wheat varieties have been developed [10], [13].
The wild barley Hordeum chilense (Roem. et Schultz.) is a valuable source of genes for increasing carotenoid content. The high compatibility of H. chilense with the genomes of Triticum species give raise to fertile and stable amphiploids and allows the transfer of traits to wheat [14]. The species shows a wide range of variation (at both morphological and molecular levels) distributed into two main groups plus an intermediate group, as revealed by molecular markers [15], [16]. Hexaploid tritordeums (xTritordeum Ascherson et Graebner) are the amphiploids derived from the cross between H. chilense and durum wheat. They exhibit a high YPC [17] and higher carotenoid content than their durum wheat parents (up to 8-fold) [18]. Using the series of addition lines of H. chilense into common wheat, it was shown that chromosome 7Hch was the main responsible of the high carotenoid content in tritordeums [19]. A further work mapped Psy1 gene of H. chilense to the α-arm of chromosome 7Hch with a diagnostic CAP marker [7]. Therefore, Psy1 gene of H. chilense is a good candidate for the improvement of YPC and has a great potential in durum wheat breeding. For this purpose, characterization of Psy1 gene in H. chilense is required for an effective use in durum wheat breeding.
Accordingly, the objectives of this work were to characterize Psy1 gene from H. chilense in four main aspects (1) allelic variability (2) transcript analysis (3) functional analysis of the transcripts and (4) expression analysis.
Results
Characterization of Psy1 genomic sequence
Lines H1, H7 and H16 of H. chilense were selected for this work, as they represent the three ecotypes described for the species [15], [16]. The amplification of the Psy1 5′ and 3′ ends from cDNA synthesized from line H1 was carried out by RACE-PCR using the primer pairs included in Table 1. The identity of the amplified fragments was confirmed with BLASTn. The resulting fragments shared high homology with Psy1 genes of different plant species. Considering the consensus sequence, new primers were designed to amplify both the complete genomic sequence and the cDNA coding sequence of Psy1 gene.
Table 1. Primers used for the characterization of Psy1 gene in H. chilense.
| Primer | Purpose | Sequence 5′→3′ |
| RvPsy1con1 | 5′ SP | GCCCTCTTGATCTGCCTCTTCATG |
| RvPsy1con3 | 5′ SP | CATATGGCGCGCCGCCGCTCCTC |
| NP for RvPsy1con1 | ||
| RvPsy1con2 | NP for RvPsy1con3 | ACGTCGTACACCTTCTGCTCCGA |
| FwPsy1con2 | 3′ SP | TCGGAGCAGAAGGTGTACGACGT |
| FwPsy1con1 | NP for FwPsy1con2 and FwPsy1con3 | CATGAAGAGGCAGATCAAGAGGGC |
| FwPsy1con3 | 3′ SP | CGGCGCGCCATATGGGCCATCTA |
| FwPsy1Hc2 | 3′ SP | TGTCGGAGAAGATGCAAGAAGAGGA |
| FwPsy1Hc1 | NP for FwPsy1Hch2 | CATCTCAAAGGAGTCGTCACCGAAA |
| HcPsy1-CDS3F | cDNA amplification | CACCACTCCGGCCGAGACAAAG |
| Genomic DNA amplification | ||
| Full-length DNA sequencing | ||
| HcPsy1-CDS4F | cDNA amplificationPhysical mapping | CATAGTGGTGAATCCATCCCTTGC |
| HcPsy1-5ER | Genomic DNA amplification | GTCGTTTGCCTCGATCTCAT |
| HcPsy1-CDS4R | cDNA amplification | CCTGACCATCTTCATCTTGCTTTG |
| HcPsy1-2EF | Genomic DNA amplification | CTTGCTGATGACGGAGGAG |
| Full-length DNA sequencing | ||
| HcPsy1-CDS5R | cDNA amplification | TGTCTTCAAACGCACATGTTCTAGC |
| Genomic DNA amplification | ||
| Full-length DNA sequencing | ||
| HcPsy1-3EF | Full-length DNA sequencing | GCCCCTAGACATGCTCGAA |
| HcPsy1-2ER | Physical mapping | GCCGCTCCTCCGTCATCAG |
| HcPsy1-4EF | Full-length DNA sequencing | CTCGCGAACCAGCTCACC |
| HcPsy1RT-F1 | RT-qPCR | CGAACCAGCTCACCAACATA |
| HcPsy1RT-R1 | RT-qPCR | GAGCTCGTCTTGTGGCAAAT |
SP: specific primer for RACE PCR; NP: nested primer for RACE PCR.
Using two primer pairs (HcPsy1-CDS3F/HcPsy1-5ER and HcPsy1-2EF/HcPsy1-CDS5R), two overlapping segments of Psy1 were amplified, cloned and sequenced in H1, H7 and H16 genomic DNA. The genomic DNA sequence of Psy1 contained 3,447 bp in H1; 3,400 bp in H7 and 3,396 bp in H16, including 173 and 265 bp of the 5′ UTR and 3′ UTR flanking regions, respectively (GenBank accession numbers HM598415, HM598416 and HM598417, respectively). Complete genomic sequence had a 97.3% homology between Psy1 alleles of H1 and H7; 98.2% between H1 and H16 and 98.7% between H7 and H16. The exon/intron structure of H. chilense Psy1 was inferred from the alignment of genomic sequence and the cDNA coding region and compared with its homologous from wheat (Figure 1). It consisted of six exons and five introns, being the exons of the same length in the three lines. The sequence of exons 2, 3 and 6 was identical in the three alleles and exons 1, 4 and 5 showed high sequence identities: 8 single nucleotide polymorphisms (SNPs) between H1-H7 and between H7-H16, and 2 SNPs between H1-H16. Otherwise, the introns differed in size among the three alleles, and showed a higher number of SNPs and insertion-deletion as expected for intronic regions. A 49 nucleotide insertion was identified in the intron 3 of H1, showing a partial inverted duplication of the sequences flanking the insertion.
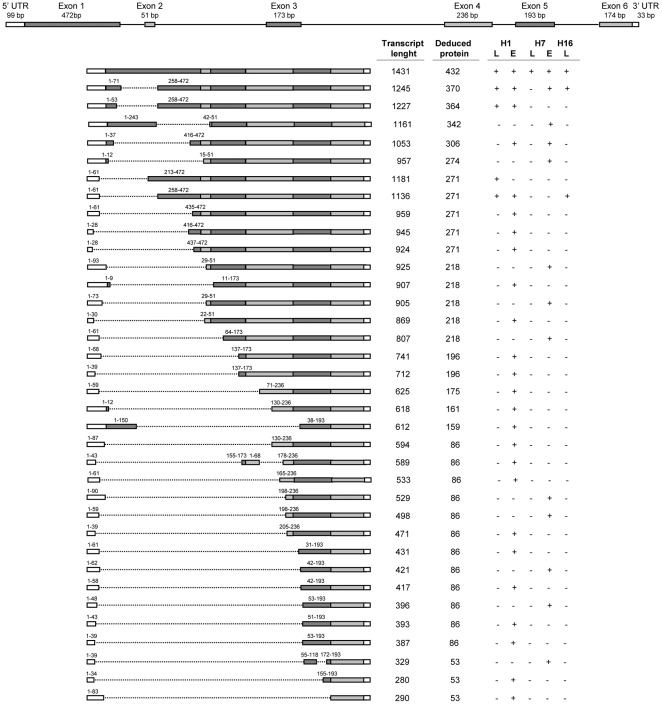
Figure 1. Schematic representation of Psy1 splicing forms in H. chilense.
Exons are represented by grey boxes and introns by a continuous line. White boxes represent 5′ and 3′ UTR. Discontinuous lines represent the untranscribed regions compared to the expected correct splicing form (1431 bp, 432 amino acids). For each transcript type, transcript length (bp), deduced protein length (amino acid residues) and the genotype/tissue where they were detected (L = leaf; E = endosperm; + = detected; - = not detected) is indicated. Exons length is indicated in the genomic form (top). When exons and/or 5′ UTR are not complete in a transcript form, it is indicated which part is included referring to the total exon and/or 5′ UTR length.
The molecular phylogenetic tree was constructed using the alignment of Psy1 coding sequences from Oryza sativa, Zea mays, Triticum aestivum, Triticum turgidum, Triticum urartu, Triticum monococcum, Aegilops tauschii and Aegilops speltoides. The Psy2 sequences of O. sativa and Z. mays, two partial sequences of Psy2 of Triticum turgidum, and Psy3 sequences from O. sativa and Z. mays were used like out-group. The sequences from H. chilense obtained in this work were grouped with the Psy1 sequences (Figure S1). Moreover, Psy1 sequences from H. chilense are closest to the Psy1 from wheat D genome by sharing a similarity of 97.5% (96.5% with A genome and 95.8% with B genome).
Quantitative real-time PCR
Once we had Psy1 sequence of H. chilense, we designed the primer pair HcPsy1RT-F1/HcPsy1RT-R1 to carry out a preliminary qPCR assay, in order to have an overview of the expression rates of Psy1 in H. chilense. A fragment of 80 bp of Psy1 gene located between exons 4 (197–236 bp) and 5 (1–40 bp) was amplified. The product obtained in the amplification of cDNA from H7 leaves (only one product expected) was sequenced. The identity of this fragment was confirmed by BLASTn (showing the highest homology with Psy1 genes of different species) and by the alignment with Psy1 sequences from H. chilense obtained in this work, confirming that the amplified fragment corresponds to the end of exon 4 and the beginning of exon 5, as expected.
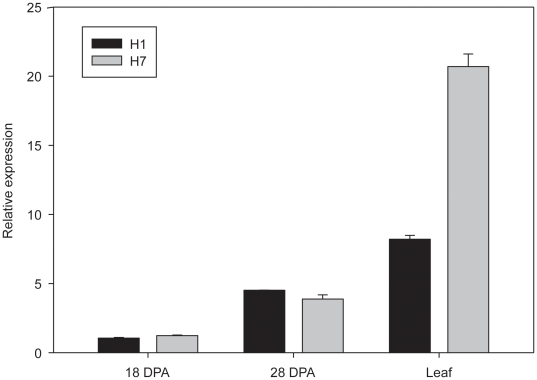
The expression of Psy1 was measured by qPCR in leaves and in seeds at 18 and 28 DPA in lines H1 and H7 (Figure 2). The expression of Psy1 was higher in leaves than in seeds, being 7.75 and 1.82 times higher in leaf than in seeds at 18 DPA and 28 DPA respectively in line H1, and 16.68 and 5.33 times higher in leaf than in seeds at 18 DPA and 28 DPA respectively in line H7. During seed developing, expression of Psy1 was increased, showing the highest value at 28 DPA in both lines. The Psy1 expression between lines H1 and H7 was only significantly different in leaf (LSD, p<0.05), being more than twice higher in H7.
Figure 2. Transcript levels of Psy1 in leaf and seeds in H.chilense lines H1 and H7.
Psy1 expression levels in seeds were measured at 18 and 28 days after anthesis (DPA) in both lines. Values in the Y axis are normalized and calibrated relative to reference genes. Error bars are SE of the means.
Characterization of Psy1 transcripts
The full coding sequence of Psy1 was amplified using primer pairs HcPsy1-CDS3F/HcPsy1-CDS5R (primer pair 1) or HcPsy1-CDS4F/HcPsy1-CDS4R (primer pair 2) in cDNA from leaves in lines H1, H7 and H16 and in two seed developing stages of H1 and H7 (at 18 and 28 DPA). Considering fragment sizes obtained with primer pair 2, a single product of 1431 bp was obtained in H7 leaves. However, more than one transcript was detected in H1 and H16 leaves and in endosperm samples from H1 and H7 lines. Regarding H1 leaves, we were able to identify five transcripts of 1431 bp, 1245 bp, 1227 bp, 1181 bp and 1136 bp. In H16 leaves, three PCR products of 1431 bp, 1245 bp and 1136 bp were obtained, being coincident with those found in H1 and H7.
Regarding endosperm, 25 different transcripts were detected in H1, being 21 of them exclusive (from tissue and genotype) and 4 of them the same as those of 1431 bp, 1245 bp, 1227 bp and 1136 bp found in H1 leaves. In H7 endosperm samples 13 transcripts were found, being 10 of them exclusive (from tissue and genotype), one in common with H1 endosperm (1053 bp) and 2 also found in H1 leaves and endosperm and H16 leaves (1431 bp and 1245 bp). Figure 1 describes all transcripts characterized in this work, indicating transcripts length (bp), deduced proteins length (aminoacid residues) and the genotype/tissue where they were detected. All cDNA sequences obtained in this work were deposited in the GenBank (accession numbers JF759623 to JF759664).
Alignment of the deduced PSY1 aminoacid sequences (those expected from the correctly spliced forms) revealed three residue differences in H7 when compared to H1 and H16, all them located in exon 1: an Ala to Ser substitution at position 22, an Arg to His substitution at position 32 and a Leu to Phe substitution at position 109. In total, the following 16 different proteins are deduced from the 36 transcript types detected (named PSY1Hc- followed by the number of amino acid residues): PSY1Hc-432a (in H1 and H16), PSY1Hc-432b (in H7), PSY1Hc-370a (in H1 and H16), PSY1Hc-370b (in H7), PSY1Hc-364, PSY1Hc-342, PSY1Hc-306, PSY1Hc-274, PSY1Hc-271, PSY1Hc-218, PSY1Hc-196, PSY1Hc-175, PSY1Hc-161, PSY1Hc-159, PSY1Hc-86 and PSY1Hc-53. Figure 3 shows the alignment of the sixteen deduced PSY1 proteins of H. chilense. Using the TargetP transit peptide (TP) predictor, the H. chilense PSY1 was predicted to have a plastid transit peptide. Exact length of H. chilense PSY1 TP was deduced from the alignment with the maize and Arabidopsis thaliana PSY1 protein sequences (AAR08445 and P37271, respectively). Thereby, H. chilense PSY1 was predicted to have a plastid TP of 77 amino acids from the N-terminal end (Figure 3). The complete TP sequence is present in PSY1Hc-432a, -432b and -342 deduced proteins; forms PSY1Hc-370a, -370b, -364, -306, -274, -161 and -159 show truncated TPs; and protein types PSY1Hc-271, -218, -196, -175, -83 and -53 do not show any region of TP (Figure 3).
Figure 3. Alignment of the deduced amino acid sequences from the Psy1 transcripts identified in H. chilense.
The plastid transit peptide (TP) and squalene-phytoene synthase 1 and 2 domains are indicated. The sixteen proteins are named PSY1Hc- followed by the number of amino acid residues. If the protein type is polymorphic between lines it is indicated: (a) H1/H16 form and (b) H7 form. Polymorphic positions are in bold.
Functional analysis of H. chilense Psy1 transcripts
To determine which transcripts were functional, transcripts coding for the sixteen different protein types were cloned and co-transformed with pAC-85b plasmid [20]. This plasmid contains all genes needed to produce β-carotene in Escherichia coli except for one encoding phytoene synthase. β-carotene production was detected by yellow color production of bacterial pellet and reversed-phase high-performance liquid chromatography (RP-HPLC) analysis of extracts from the co-transformed E. coli strains (Figure S2), showing that only PSY1Hc-432a, -432b, -370a, -364 and -271 proteins were functional in the bacterial system. All constructions produced similar rates β-carotene except for PSY1Hc-370a which produced a lower quantity (traces). Unfortunately, transcript type of 1245 bp detected in H7 endosperm and producing PSY1Hc-370b protein could not be tested after several attempts (see material and methods). Table 2 summarizes the results obtained in the functional analysis of Psy1 transcripts.
Table 2. Functional analysis of Psy1 transcripts of H. chilense.
| Function | |||
| Protein type | Transcript tested | Color1 | β-carotene2 |
| PSY1Hc-432a | 1431 | Yellow | Presence |
| PSY1Hc-432b | 1431 | Yellow | Presence |
| PSY1Hc-370a | 1245 | Pale Yellow | Traces |
| PSY1Hc-370b | - | - | - |
| PSY1Hc-364 | 1227 | Yellow | Presence |
| PSY1Hc-342 | 1161 | White | Absence |
| PSY1Hc-306 | 1053 | White | Absence |
| PSY1Hc-274 | 957 | White | Absence |
| PSY1Hc-271 | 1136 | Yellow | Presence |
| PSY1Hc-218 | 869 | White | Absence |
| PSY1Hc-196 | 712, 741 | White | Absence |
| PSY1Hc-175 | 625 | White | Absence |
| PSY1Hc-161 | 618 | White | Absence |
| PSY1Hc-159 | 612 | White | Absence |
| PSY1Hc-86 | 533, 529, 471, 417, 396 | White | Absence |
| PSY1Hc-53 | 329, 290 | White | Absence |
Color complementation observed in E. coli colonies co-transformed with pAc-85b plasmid and each of the expression vectors harboring H. chilense Psy1 transcripts.
β-carotene production detected by HPLC in co-transformed E. coli pellets.
Discussion
Psy1 gene of H. chilense contains six exons and five introns, showing the same structure as in other related species previously described [21], [22]. Nevertheless, the first exon of Psy1 gene in H. chilense is longer than Psy1 genes of maize, rice and wheat, and the sixth exon has the same length as in maize and rice and it is longer than in wheat. Compared to the three Psy1 genes in common wheat, Psy1 sequence in H. chilense is more related to Psy-D1 than to Psy-A1 or Psy-B1. Consistently, cytological characterizations [23] and marker transferability analyses [15] have also suggested that H. chilense is more related to wheat D-genome than to A and B genomes.
Regarding H. chilense intraspecific nucleotide variability, most of the polymorphisms are found in the intronic regions, as expected. SNPs are also found in exons one, four and five, but only three of them, located in exon one, lead to aminoacid changes in line H7 compared to H1 and H16.
The surprising abundance of different transcripts in H. chilense raises several questions. First, the possible mixture of transcripts of different Psy genes could be considered. In grasses, three paralogs (Psy1, Psy2 and Psy3) originated from duplication events, have been described [4], [24], [25]. The Psy sequences identified in this work are unequivocally included in the cluster comprising Psy1 sequences of wheat, maize and rice (Figure S1). Furthermore, when amplifying Psy1 using primer pair HcPsy1-CDS4F/HcPsy1-2ER in the set of addition lines of H. chilense in common wheat, a positive amplification is only observed in lines carrying the 7Hch complete chromosome or the α-arm of chromosome 7Hch (Figure S3). Psy1 has been located in this chromosomal position [7] whereas Psy2 and Psy3 are located in short and long arms of wheat chromosome 5 respectively, inferred by direct mapping or by synteny with rice chromosomes [4], [7], [26]. Additionally, a partial genomic sequence of Psy2 gene of H. chilense had been obtained in a previous work [7]. Once aligned with the equivalent region of Psy1 genomic and cDNA sequences of H. chilense, and considering the coding region, Psy1 and Psy2 have a sequence homology of only 74.5% (Figure S4). Thus, if any Psy2 form had been amplified, it could have been easily recognized. The same happens with Psy3 gene. Although Psy3 has not been characterized in many species, a putative sequence of Hordeum vulgare is available in GenBank as well as a putative Psy1 sequence (AK365521.1 and AK374031.1, respectively). When aligned with Psy1 from H. chilense there is no doubt of the identity of each of the sequences (Figure S5). Finally, the different Psy splicing forms described in this work have identical sequence, differences are found in sequence length and therefore, the most plausible hypothesis is that they correspond to the same gene. For all these reasons, we can conclude that the occurrence of Psy1, Psy2 and Psy3 transcripts mixture seems unlikely.
Another hypothesis would be the presence of an additional Psy duplication located in the α-arm of chromosome 7Hch. There are evidences of an additional locus in the long arm of chromosome 7A involved in YPC in durum wheat [11], the possibility of being a new Psy-like sequence cannot be discarded, although gene duplication cannot explain the abundance of different Psy1 transcripts in H. chilense.
On the other hand, it cannot be ruled out the possibility of that some of the transcripts considered as from endosperm come indeed from the embryo or other parts of the grain, as the small size of H. chilense seeds makes technically difficult to isolate it. Anyway, the abundance of different transcripts for Psy1 gene in H. chilense is evident and intriguing.
Alternative splicing in plants is an important post-transcriptional regulatory mechanism. This phenomenon results in gain or loss of protein function, changes in cellular localization, stability and activity among others [27]. In rice, alternative splicing forms of Psy1 gene have been described, resulting in four different transcripts (LOC_Os06g51290; http://rice.plantbiology.msu.edu). Besides, in Psy-A1 of wheat, Howitt et al. [9] identified a sequence duplication in exon 2 that creates a new splicing site and activates a cryptic exon. This alternative splicing produced four different transcripts: the wild type, two shorter transcripts originated by frame-shift mutations and early terminations, and a fourth transcript 35 aminoacids longer than the wild type. The same transcripts were observed in leaves and endosperm, and only the wild type produced an active protein. In our work, we identified 36 different Psy1 transcripts in H. chilense, being the largest spliced form (1431 bp) the size expected for correct splicing, and producing a protein of 432 amino acids. The rest of the forms are the result of different splicing events occurring in exons 1 to 5 and/or in the 5′ UTR and producing shorter proteins (Figure 1). Functionality of the predicted proteins was tested in a color complementation assay showing that only some of the transcripts lead to functional proteins in the bacterial system. As expected, both PSY1Hc-432a and -432b (predicted correctly spliced forms) complement the absence of crtB (bacterial phytoene synthase) gene in plasmid pAc-85b, producing β-carotene. The same happens with proteins PSY1Hc-370a, PSY1Hc-364 and PSY1Hc-271. Regarding protein PSY1Hc-370b, it is expectable to be a functional form as there is a single amino acid difference compared to the functional protein PSY1Hc-370a (an Ala to Ser substitution at position 22) and considering that both PSY1Hc-432a and PSY1Hc-432b are functional proteins despite their three amino acid differences, including that in position 22.
Several works have analyzed the activity of truncated forms of PSY proteins. Misawa et al. [28] demonstrated that tomato PSY proteins in which 114 amino acids from the N-terminal were eliminated had increased catalytic activity in E. coli compared to the complete form. In Narcissus, the full-length PSY enzyme showed reduced catalytic activity when compared with an N-truncated form with its transit sequence of 214 bp removed by site-directed mutagenesis [29]. More recently, in Citrus, the catalytic activity of PSY and a truncated form lacking 89 amino acids in its N-terminus was similar when analyzed their expression in E. coli cells [30]. Psy1 is a nuclear encoded gene that mediates the first step in the plastid-localized carotenoid biosynthetic pathway. A large protein complex containing enzymes from the isoprenoid pathway and carotenogenic enzymes as PSY has been hypothesized to occur in the plastid stroma [31]. It is well known that transit peptides direct proteins to specific organelles. Consequently, although PSY1Hc-370a, PSY1Hc-364 and PSY1Hc-271 proteins are functional in the bacterial system, their implication in carotenoid synthesis or accumulation could be questioned, as their transit peptide is truncated or totally absent.
Subfunctionalization of the three Psy genes in grasses is accepted as a mechanism providing a fine control of carotenogenesis in response to developmental and environmental signals [6]. The surprising abundance of Psy1 transcripts in H. chilense, only some of them leading to functional proteins, suggests a very accurate regulation of PSY1 active protein concentration and it would be possible to hypothesize that splicing to mature or alternative transcripts could also regulate the cellular concentrations of phytoene synthase, depending on the tissue, physiological or environmental conditions [9], [32].
Psy1 expression was analyzed in leaves and in seeds at 18 and 28 DPA in lines H1 and H7. Psy1 mRNA levels in leaves were higher than in seeds in both H1 and H7. Similar results were observed in common wheat, where Psy1 mRNA was about 50-fold lower in immature milky stage endosperm samples than in leaves [33]. High abundance of Psy1 transcripts has been also reported in maize leaves where they play an essential role in maintaining leaf carotenoid content, especially under heat stress conditions [5], [6]. Also in rice, PSY1 is the main enzyme responsible for carotenoid supply in chloroplasts [25]. Carotenoids are involved in photosynthetic processes like light harvesting and protection of the photosynthetic apparatus from photo-oxidation [34]. Regarding endosperm, in maize and rice it has been widely demonstrated the correlation of Psy1 transcript accumulation and endosperm carotenoid content [5], [24], [35]. The association of allelic variation of Psy1 with changes in grain pigment content has been also confirmed in both durum and bread wheat [8], [11], [36]. Psy1 mRNA levels are not significantly different between lines H1 and H7 in the two seed development stages analyzed, although differences in yellow pigment content have been previously described in mature seeds [37]. Assuming that in H. chilense the correlation Psy mRNA accumulation-carotenoid content is also valid, the absence of association between YPC and mRNA levels can be explained. Alvarez et al. [37] determined values of 27.2 and 62.0 µg/g of carotenoid in H1 and H7 seeds, respectively. But actually, these measures refer to YPC as inferred by spectrophotometric determinations after water-saturated butanol extraction method, as it has been usually accepted in wheat. Carotenoids represent about 30–50% of the YPC in wheat [38], [39], but there are other factors also influencing final flour colour [40], [41]. On the other hand, Psy1 mRNA levels are not in this case a direct estimation of functional PSY1 protein. In the pool of transcripts detected by qPCR only a part lead to functional proteins (Table 2). Additionally, 10 out of the 36 transcript types do not amplify with the selected primers (612 bp and 471–280 bp), although none of these transcripts have demonstrated to encode functional PSY1.
Moreover, other genes of the carotenoid synthesis pathway influence final carotenoid content and are also considered limiting steps, as it has been extensively studied in maize [35]. As an example, in the H1xH7 F2 mapping population, a QTL for YPC explaining the 14.8% of the phenotypic variance was identified in chromosome 2Hch [42]. The carotenogenic gene ζ-carotene desaturase (ZDS) has been located in durum wheat chromosomes 2A and 2B [43] and in H. chilense chromosome 2Hch (unpublished results), and it has been also associated with carotenoid accumulation in maize [44]. No QTLs for YPC were detected on chromosome 7Hch, probably due to the lack of polymorphism or saturation of this map region.
The characterization of H. chilense Psy1 means the first step in the potential use of this gene for increasing carotenoid content in wheat breeding. First, we describe the allelic variability found among the three ecotypes at the genomic level, and we focus in H1 and H7 lines for the following works as they represent two distant groups considering their phylogenetic relations and YPC content. Regarding these two lines, we show the first results of Psy1 expression in H. chilense and we evidence the surprising abundance of different Psy1 transcripts forms, not ever described for this gene in other species. Finally, functional analysis of the splicing forms has been addressed, showing that only some transcript forms produce functional proteins and suggesting a fine modulation of functional PSY1 concentration.
As shown in Golden Rice 2, a variety of rice engineered to produce β-carotene, the use of new Psy1 genes sources offers the potential to increase carotenoid content [45]. Accordingly, our results constitute an interesting basis for carotenoid enhancement in durum wheat either by chromosome engineering or by transgenesis. Functional analysis of Psy1 will be useful for future selection of the appropriate transcripts sequences to increase the PSY1 activity in wheat by transformation. Moreover, the development of substitution lines of chromosome 7Hch in durum wheat has been already started in order to test the effect of this chromosome in seeds carotenoid content.
Materials and Methods
Plant material
Accession lines of H1, H7 and H16 from the collection of the Instituto de Agricultura Sostenible, IAS-CSIC, Córdoba, were selected. Each line belongs to one of the three ecotypes described in H. chilense [16]. Leaf tissue of H1, H7 and H16 was harvested at tillering stage, frozen in liquid nitrogen, and stored at −80°C. Developing grains of H1 and H7 were harvested 18 and 28 DPA and conserved as described above. DNA from addition lines of H. chilense (line H1) into common wheat cv. Chinese Spring was used to confirm the location of the Psy fragment amplified.
Amplification of cDNA ends
Total RNA from H1, H7 and H16 leaves was isolated using the TRIzol® Reagent (Invitrogen, Carlsbad, CA) according to manufacturer's instructions. Total RNA from H1 was used to synthesize anchored cDNA at the 5′ and 3′ ends as described by the SMART™ RACE cDNA Amplification Kit (Clontech, Palo Alto, CA). The specific primers were designed in the conserved regions of Psy1 considering the complete coding sequences of maize (AY324431.1), rice (AY445521.1), common wheat A1b and D1a alleles (EF600064.1 and EU650397.1, respectively), durum wheat B1g allele (EU650396.1) and partial sequences of H1 and H7 lines of H. chilense (DU796677 and DU796679, respectively). All primers used in this work are included in Table 1.
RACE PCR reactions were carried out using Certamp complex enzyme mix (Biotools B&M Labs, Madrid) according to supplier's instructions and performed as follows: 25 cycles of 30 s at 94°C, 30 s at 64°C and 2 min at 72°C. PCR 5′ and 3′ products were reamplified using nested primers (see Table 1) and resolved on 1% agarose gels, stained with ethidium bromide and visualized under UV light. The resulting products were cloned in pGEMT-Easy vector (Promega, Madison, WI), and introduced into competent Escherichia coli (DH5α) cells by transformation. Plasmids were isolated and purified using Illustra plasmidPrep Mini Spin Kit (GE Healthcare, UK Ltd, UK) and used as template for sequencing.
Amplification of CDS and full-length genomic sequence
The sequences obtained from 5′ and 3′ ends were used for primer design to amplify the complete coding sequence of Psy1 in H. chilense. Total RNA was isolated from seeds at 18 and 28 DPA in lines H1 and H7 and from leaves of lines H1, H7 and H16 using the TRIzol® Reagent (Invitrogen, Carlsbad, CA) according to manufacturer's instructions. For the synthesis of first-strand cDNA, 4 µg of total RNA was reverse transcribed using oligo (dT) primer and M-MLV reverse transcriptase (Invitrogen, Carlsbad, CA) in 20 µl total volume according to the manufacturer's instructions. cDNA samples were diluted with additional 230 µl of water. cDNA PCR amplifications with primer combinations HcPsy1-CDS3F/HcPsy1-CDS5R (primer pair 1) or HcPsy1-CDS4F/HcPsy1-CDS4R (primer pair 2) were performed in 25 µl reactions consisting of 0.625 units of DNA polymerase (Biotools B&M Labs, Madrid), 1X PCR buffer, 2 mM MgCl2, 320 µM dNTPs (Promega, Madison, WI, USA), 1 M betaine (Sigma-Aldrich, St. Louis, MO), 0.6 µM of each primer and 1.2 µl of the cDNA dilution. PCR were carried out as follows: 5 min at 94°C, 28 cycles of 15 s at 94°C, 30 s at an annealing temperature progressively lowered from 68°C to 54°C by 0.5°C every cycle and 5 min at 72°C, 25 cycles of 15 s at 94°C, 30 s at 56°C and 5 min at 72°C, followed by 10 min at 72°C. Full-length Psy1 cDNA sequences were obtained by cloning and sequencing the obtained PCR fragments as specified before. For transcripts description we will refer to fragment sizes obtained by amplification with primer pair 2.
Genomic DNA was extracted from leaves using CTAB method according to [46]. Genomic DNA sequences of H1, H7 and H16 were obtained by amplifying two overlapping fragments using primer combinations HcPsy1-CDS3F/HcPsy1-5ER and HcPsy1-2EF/HcPsy1-CDS5R. PCR amplification, cloning and sequencing was performed as described above for cDNA. Psy1 full-length genomic sequences of H1, H7 and H16 were constructed by assembling the sequencing products obtained with primers HcPsy1-2EF, HcPsy1-3EF, HcPsy1-4EF, HcPsy1-CDS3F, HcPsy1-CDS5R (Table 1) and PSY1Rev2 [7]. In all cases, fragments were amplified twice to avoid artifacts in the sequencing stage.
Functional analyses of H. chilense Psy1 transcripts
The phytoene synthase gene from Adonis aestivalis, cloned into plasmid pAd-Psy [20], was cut and cloned into pGEMT-Easy vector to have a positive control of our co-transformation assays. It was noted that the insert was not expressed under T7 promoter and that only under SP6 promoter complementation was observed. All Psy1 transcripts had been cloned under T7 promoter, thus, the vectors harboring the transcripts selected for co-transformations were cut with EcoRI, filled with A-tailing reaction, cloned again into pGEMT-Easy vector and selected for the right orientation (under SP6). Unfortunately, we could not recover any plasmid harboring the transcript leading to PSY1Hc-370b under SP6 promoter after several unsuccessful attempts.
For color complementation, TOP10F' competent cells (Invitrogen, Carlsbad, CA) were transformed with pAC85-b [20] and each one of the expression constructs with Psy1 transcripts from H. chilense. Double transformants selected were grown overnight at 30°C in liquid LB medium containing ampicillin (150 mg/l), chloramphenicol (30 mg/l) and IPTG (50 mg/ml).
Cultured cells were harvested by centrifugation at 10,000×g, and the bacterial cell pellets were extracted with 3 ml of acetone containing 0.1% butylated hidroxy toluene (w/v), by the aid of sonication. The cell debris was separated by centrifugation at 5000 rpm at 4°C and the upper phase was dried under a nitrogen stream. The resulting residue was dissolved in 0.5 ml of acetone, centrifuged at 12,000×g and stored at −30°C until analyzed. Analysis of the carotenoid pigments was carried out by RP-HPLC as described by [47] with slight modifications.
Quantitative real time PCR (qPCR)
For qPCR, the FastStart Universal SYBR Green Master (Roche Applied Science, Mannheim, Germany) was used on the 7500 Real Time PCR System (Applied Biosystems, Foster City, USA) together with the gene-specific primers HcPsy1RT-F1 and HcPsy1RT-R1 (Table 1) and the reference genes ADP-RF and RLI [48]. The PCR conditions were 95°C for 10 min followed by 40 cycles of 95°C for 15 s and 60°C for 1 min. All reactions were replicated twice.
PCR efficiency of each primer pair was determined with the LinRegPCR (version 11.0) quantitative PCR data analysis program [49] using raw normalized fluorescence as input data. Expression of the genes for each sample (N0) was determined by using the equation N0 = 0.2/ECq, where E = PCR efficiency for each primer and Cq is the number of cycles needed to reach 0.2 arbitrary units of fluorescence. Levels of Psy1 expression were normalized relative to the geometric mean of the reference genes ADP-RF and RLI. The significance of the differences in transcript level between samples was determined using ANOVA and a subsequent means comparison with Least Significant Difference method (LSD). Statistical methods were performed using the Rcmdr package of R [50], [51].
Bioinformatic analyses
All primers were designed using Primer3plus software [52] (http://www.bioinformatics.nl/cgi-bin/primer3plus/primer3plus.cgi). Nucleotide and the deduced amino acid sequences were aligned using Edialign program (http://emboss.sourceforge.net/index.html) and edited using GeneDoc software (http://www.psc.edu/biomed/genedoc). Sequence identity searches were performed at the NCBI (http://www.ncbi.nlm.nih.gov) using BLAST.
The presence of a transit peptide was predicted by TargetP 1.1 Server (http://www.cbs.dtu.dk/services/TargetP/; [53]). The domains were predicted using InterProScan software (http://www.ebi.ac.uk/Tools/InterProScan/).
The coding sequence of Psy1 alleles of lines H1, H7 and H16, along with some Psy1 alleles previously described in wheat [11], [22], [36], [54], [55], partial sequences of wheat Psy2 and the coding sequences of Psy1, Psy2 and Psy3 of maize and rice, were used to construct the phylogenetic tree. Neighbor joining tree was generated using the complete deletion method in MEGA4 program [56], [57]. Bootstrap test was performed with 1000 replicates to calculate the confidence of the different nodes.
Supporting Information
Neighbor-joining tree generated from the alignment of the Psy1 coding sequences from different species. Neighbor-joining tree generated from the alignment of the Psy1 coding sequences from Oryza sativa, Zea mays, Triticum aestivum, Triticum turgidum, Triticum urartu, Triticum monococcum, Aegilops tauschii and Aegilops speltoides. The Psy2 coding sequences of O. sativa and Z. mays, two partial sequences of Psy2 of Triticum turgidum, and Psy3 sequences from O. sativa and Z. mays were used like out-group. Predicted correctly spliced forms of Psy1 gene in H. chilense lines H1, H7 and H16 were used. Numbers over the tree nodes are bootstrap confidence values based on 1,000 bootstrap iterations.
(TIF)
Color complementation in co-transformed E. coli pellets and the corresponding RP-HPLC chromatograms of the carotenoid extracts. (A) A yellow color is observed in E. coli pellets co-transformed with plasmid pAC-85b and pGEMT expressing proteins PSY1Hc-432a and PSY1Hc-432b, while no color complementation is observed when proteins PSY1Hc-86 or PSY1Hc-53 are expressed; (B) Absorption spectra in the HPLC mobile phase of β-carotene; (C) Corresponding RP-HPLC chromatograms of carotenoid extracts shown in (A).
(TIF)
Chromosomal localization of Psy1 gene in H. chilense . Amplification of genomic DNA with primer pair HcPsy1-CDS4F/HcPsy1-2ER in the following lines: (1) H. chilense line H1, (2) H. chilense line H7, (3) T. turgidum ssp. durum cv. Yavaros, (4) T. aestivum cv. Chinese Spring, (5) T. turgidum var. Kofa, (6) T. turgidum breeding line UC1113, (7) 1HchS monotelosomic addition line, (8) 1HchS ditelosomic addition line, (9) 2Hch-α ditelosomic addition line, (10) 4Hch disomic addition line (11) 5Hch disomic addition line, (12) 5HchL ditelosomic addition line, (13) 6Hch disomic addition line, (14) 7Hch disomic addition line, (15) 7Hch-α ditelosomic addition line, (16) 7Hch-β ditelosomic addition line and (17) 6Hch-S ditelosomic addition line.
(TIF)
Alignment of partial Psy1 and Psy2 sequences in H1 and H7 lines of H. chilense. Alignment of genomic (g) and cDNA (c) sequences of Psy1 (this work) and partial genomic sequence of Psy2 gene in H1 and H7 lines (DU796678 and DU796680, respectively).
(TIF)
Alignment of Psy3 gene from different species. Alignment of Psy3 sequences of Sorghum bicolor (Sb; AY705390.1), Zea mays (Zm; NM_001114653.1), Oryza sativa (Os; FJ214953.1) and Hordeum vulgare clone NIASHv2034M16 showing high homology with Psy3 gene of other species (AK365521.1), Hordeum vulgare clone NIASHv3051L01 showing high homology with Psy1 gene of other species (AK374031.1) and H. chilense Psy1 gene (this work).
(TIF)
Acknowledgments
We are grateful to Prof. Antonio Martin for providing the seeds used in this work and to E. León for her technical assistance. The authors thank Dr. D Hornero-Méndez for assistance with HPLC analysis and Prof. FX Cunningham for providing the plasmids pAC-85b and pAd-Psy.
Footnotes
Competing Interests: The authors have declared that no competing interests exist.
Funding: This work was supported by grants (to SGA) AGL2008-03720 and 200840I137 from the Spanish Ministry of Science and Innovation (MSI), CSIC and FEDER. FP acknowledges financial support from the MSI Juan de la Cierva program. CR-S and FP acknowledge financial support from JAE-Doc program (CSIC, co-funded by FSE) and the Ramón y Cajal program (Spanish Ministry of Science and Innovation) respectively. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Britton G. Functions of Intact Carotenoids. In: Britton G, Liaaen-Jensen S, Pfander H, editors. Carotenoids vol IV. Berlin: Birkhäuser Verlag; 2008. pp. 189–212. [Google Scholar]
- 2.Farré G, Sanahuja G, Naqvi S, Bai C, Capell T, et al. Travel advice on the road to carotenoids in plants. Plant Sci. 2010;179(1–2):28–48. [Google Scholar]
- 3.Rodríguez-Suárez C, Giménez MJ, Atienza SG. Progress and perspectives for carotenoid accumulation in selected Triticeae species. Crop Pasture Sci. 2010;61:743–751. [Google Scholar]
- 4.Li F, Vallabhaneni R, Wurtzel ET. PSY3, a new member of the phytoene synthase gene family conserved in the Poaceae and regulator of abiotic stress-induced root carotenogenesis. Plant Physiol. 2008;146:1333–1345. doi: 10.1104/pp.107.111120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Li F, Vallabhaneni R, Yu J, Rocheford T, Wurtzel ET. The maize phytoene synthase gene family: overlapping roles for carotenogenesis in endosperm, photomorphogenesis, and thermal stress tolerance. Plant Physiol. 2008;147:1334–1346. doi: 10.1104/pp.108.122119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Li F, Tsfadia O, Wurtzel ET. The phytoene synthase gene family in the Grasses. Plant Signal Behav. 2009;4:208–211. doi: 10.4161/psb.4.3.7798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Atienza SG, Avila CM, Martin A. The development of a PCR-based marker for PSY1 from Hordeum chilense, a candidate gene for carotenoid content accumulation in tritordeum seeds. Aust J Agric Res. 2007;58:767–773. [Google Scholar]
- 8.Pozniak CJ, Knox RE, Clarke FR, Clarke JM. Identification of QTL and association of a phytoene synthase gene with endosperm colour in durum wheat. Theor Appl Genet. 2007;114:525–537. doi: 10.1007/s00122-006-0453-5. [DOI] [PubMed] [Google Scholar]
- 9.Howitt CA, Cavanagh CR, Bowerman AF, Cazzonelli C, Rampling L, et al. Alternative splicing, activation of cryptic exons and amino acid substitutions in carotenoid biosynthetic genes are associated with lutein accumulation in wheat endosperm. Funct Integr Genomics. 2009;9:363–376. doi: 10.1007/s10142-009-0121-3. [DOI] [PubMed] [Google Scholar]
- 10.Singh A, Reimer S, Pozniak CJ, Clarke FR, Clarke JM, et al. Allelic variation at Psy1-A1 and association with yellow pigment in durum wheat grain. Theor Appl Genet. 2009;118:1539–1548. doi: 10.1007/s00122-009-1001-x. [DOI] [PubMed] [Google Scholar]
- 11.Zhang W, Dubcovsky J. Association between allelic variation at the Phytoene synthase 1 gene and yellow pigment content in the wheat grain. Theor Appl Genet. 2008;116:635–645. doi: 10.1007/s00122-007-0697-8. [DOI] [PubMed] [Google Scholar]
- 12.Zhang Y, Wu Y, Xiao Y, He Z, Zhang Y, et al. QTL mapping for flour and noodle colour components and yellow pigment content in common wheat. Euphytica. 2009;165:435–444. [Google Scholar]
- 13.Reimer S, Pozniak CJ, Clarke FR, Clarke JM, Somers DJ, et al. Association mapping of yellow pigment in an elite collection of durum wheat cultivars and breeding lines. Genome. 2008;51:1016–1025. doi: 10.1139/G08-083. [DOI] [PubMed] [Google Scholar]
- 14.Martín A, Martin LM, Cabrera A, Ramirez MC, Gimenez MJ, et al. The potential of Hordeum chilense in breeding Triticeae species. In: Jaradat AA, editor. Triticeae III. Enfield: Science publishers, Inc; 1998. pp. 377–386. [Google Scholar]
- 15.Castillo A, Budak H, Martín A, Dorado G, Börner A, et al. Interspecies and intergenus transferability of barley and wheat D-genome microsatellite markers. Ann Appl Biol. 2010;156:347–356. [Google Scholar]
- 16.Vaz Patto MC, Aardse A, Buntjer J, Rubiales D, Martín A, et al. Morphology and AFLP markers suggest three Hordeum chilense ecotypes that differ in avoidance to rust fungi. Can J Botany. 2001;79:204–213. [Google Scholar]
- 17.Atienza S, Avila C, Ramírez M, Martín A. Application of near infrared reflectance spectroscopy to the determination of carotenoid content in tritordeum for breeding purposes. Aust J Agric Res. 2005;56:85–89. [Google Scholar]
- 18.Atienza SG, Ballesteros J, Martín A, Hornero-Méndez D. Genetic variability of carotenoid concentration and degree of esterification among tritordeum (xTritordeum Ascherson et Graebner) and durum wheat accessions. J Agric Food Chem. 2007;55:4244–4251. doi: 10.1021/jf070342p. [DOI] [PubMed] [Google Scholar]
- 19.Alvarez JB, Martin LM, Martin A. Chromosomal localization of genes for carotenoid pigments using addition lines of Hordeum chilense in wheat. Plant Breed. 1998;117:287–289. [Google Scholar]
- 20.Cunningham FX, Gantt E. A portfolio of plasmids for identification and analysis of carotenoid pathway enzymes: Adonis aestivalis as a case study. Photosynth Res. 2007;92:245–259. doi: 10.1007/s11120-007-9210-0. [DOI] [PubMed] [Google Scholar]
- 21.Fu Z, Yan J, Zheng Y, Warburton ML, Crouch JH, et al. Nucleotide diversity and molecular evolution of the PSY1 gene in Zea mays compared to some other grass species. Theor Appl Genet. 2010;120:709–720. doi: 10.1007/s00122-009-1188-x. [DOI] [PubMed] [Google Scholar]
- 22.Wang J, He X, He Z, Wang H, Xia X. Cloning and phylogenetic analysis of phytoene synthase 1 (Psy1) genes in common wheat and related species. Hereditas. 2009;146:208–256. doi: 10.1111/j.1601-5223.2009.02132.x. [DOI] [PubMed] [Google Scholar]
- 23.Cabrera A, Friebe B, Jiang J, Gill BS. Characterization of Hordeum chilense chromosomes by C-banding and in situ hybridization using highly repeated DNA probes. Genome. 1995;38:435–442. doi: 10.1139/g95-057. [DOI] [PubMed] [Google Scholar]
- 24.Gallagher CE, Matthews PD, Li F, Wurtzel ET. Gene duplication in the carotenoid biosynthetic pathway preceded evolution of the grasses. Plant Physiol. 2004;135:1776. doi: 10.1104/pp.104.039818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Welsch R, Wust F, Bar C, Al-Babili S, Beyer P. A third Phytoene synthase is devoted to abiotic stress-induced abscisic acid formation in rice and defines functional diversification of phytoene synthase genes. Plant Physiol. 2008;147:367–380. doi: 10.1104/pp.108.117028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ahn S, Anderson JA, Sorrells ME, Tanksley SD. Homoeologous relationships of rice, wheat and maize chromosomes. Mol Gen Genet. 1993;241–241:483–490. doi: 10.1007/BF00279889. [DOI] [PubMed] [Google Scholar]
- 27.Reddy AS. Alternative splicing of pre-messenger RNAs in plants in the genomic era. Annu Rev Plant Biol. 2007;58:267–294. doi: 10.1146/annurev.arplant.58.032806.103754. [DOI] [PubMed] [Google Scholar]
- 28.Misawa N, Truesdale MR, Sandmann G, Fraser PD, Bird C, et al. Expression of a tomato cDNA coding for phytoene synthase in Escherichia coli, phytoene formation in vivo and in vitro, and functional analysis of the various truncated gene products. J Biochem. 1994;116:980–985. doi: 10.1093/oxfordjournals.jbchem.a124656. [DOI] [PubMed] [Google Scholar]
- 29.Schledz M, Al-Babili S, Lintig JV, Haubruck H, Rabbani S, et al. Phytoene synthase from Narcissus pseudonarcissus: functional expression, galactolipid requirement, topological distribution in chromoplasts and induction during flowering. Plant J. 1996;10:781–792. doi: 10.1046/j.1365-313x.1996.10050781.x. [DOI] [PubMed] [Google Scholar]
- 30.Kim IJ, Ko KC, Nam TS, Kim YW, Chung WI, et al. Expression and activity of citrus phytoene synthase and beta-carotene hydroxylase in Escherichia coli. J Microbiol. 2003;41:212–218. [Google Scholar]
- 31.Cunningham FX, Jr, Gantt E. Genes and enzymes of carotenoid biosynthesis in plants. Annu Rev Plant Biol. 1998;49:557–583. doi: 10.1146/annurev.arplant.49.1.557. [DOI] [PubMed] [Google Scholar]
- 32.Cazzonelli CI, Pogson BJ. Source to sink: regulation of carotenoid biosynthesis in plants. Trends Plant Sci. 2010;15:266–274. doi: 10.1016/j.tplants.2010.02.003. [DOI] [PubMed] [Google Scholar]
- 33.Cong L, Wang C, Chen L, Liu H, Yang G, et al. Expression of phytoene synthase1 and carotene desaturase crtI genes result in an increase in the total carotenoids content in transgenic elite wheat (Triticum aestivum L.). J Agric Food Chem. 2009;57:8652–8660. doi: 10.1021/jf9012218. [DOI] [PubMed] [Google Scholar]
- 34.Hirschberg J. Carotenoid biosynthesis in flowering plants. Curr Opin Plant Biol. 2001;4:210–218. doi: 10.1016/s1369-5266(00)00163-1. [DOI] [PubMed] [Google Scholar]
- 35.Vallabhaneni R, Wurtzel ET. Timing and biosynthetic potential for carotenoid accumulation in genetically diverse germplasm of maize. Plant Physiol. 2009;150:562–572. doi: 10.1104/pp.109.137042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.He XY, Zhang YL, He ZH, Wu YP, Xiao YG, et al. Characterization of phytoene synthase 1 gene (Psy1) located on common wheat chromosome 7A and development of a functional marker. Theor Appl Genet. 2008;116:213–221. doi: 10.1007/s00122-007-0660-8. [DOI] [PubMed] [Google Scholar]
- 37.Alvarez JB, Martin LM, Martin A. Genetic variation for carotenoid pigment content in the amphiploid Hordeum chilense x Triticum turgidum conv. durum. Plant Breed. 1999;118:187–189. [Google Scholar]
- 38.Digesų AM, Platani C, Cattivelli L, Mangini G, Blanco A. Genetic variability in yellow pigment components in cultivated and wild tetraploid wheats. J Cer Sci. 2009;50:210–218. [Google Scholar]
- 39.Hentschel V, Kranl K, Hollmann J, Lindhauer MG, Böhm V, et al. Spectrophotometric determination of yellow pigment content and evaluation of carotenoids by high-performance liquid chromatography in durum wheat grain. J Agric Food Chem. 2002;50:6663–6668. doi: 10.1021/jf025701p. [DOI] [PubMed] [Google Scholar]
- 40.Borrelli GM, De Leonardis AM, Fares C, Platani C, Di Fonzo N. Effects of modified processing conditions on oxidative properties of semolina dough and pasta. Cereal Chem. 2003;80:225–231. [Google Scholar]
- 41.Panfili G, Fratianni A, Irano M. Improved normal-phase high-performance liquid chromatography procedure for the determination of carotenoids in cereals. J Agric Food Chem. 2004;52:6373–6377. doi: 10.1021/jf0402025. [DOI] [PubMed] [Google Scholar]
- 42.Atienza SG, Ramirez CM, Hernandez P, Martin A. Chromosomal location of genes for carotenoid pigments in Hordeum chilense. Plant Breed. 2004;123:303–304. [Google Scholar]
- 43.Cenci A, Somma S, Chantret N, Dubcovsky J, Blanco A. PCR identification of durum wheat BAC clones containing genes coding for carotenoid biosynthesis enzymes and their chromosome localization. Genome. 2004;47:911–917. doi: 10.1139/g04-033. [DOI] [PubMed] [Google Scholar]
- 44.Wong JC, Lambert RJ, Wurtzel ET, Rocheford TR. QTL and candidate genes phytoene synthase and zeta-carotene desaturase associated with the accumulation of carotenoids in maize. Theor Appl Genet. 2004;108:349–359. doi: 10.1007/s00122-003-1436-4. [DOI] [PubMed] [Google Scholar]
- 45.Paine JA, Shipton CA, Chaggar S, Howells RM, Kennedy MJ, et al. Improving the nutritional value of Golden Rice through increased pro-vitamin A content. Nat Biotechnol. 2005;23:482–487. doi: 10.1038/nbt1082. [DOI] [PubMed] [Google Scholar]
- 46.Murray MG, Thompson WF. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 1980;8:4321. doi: 10.1093/nar/8.19.4321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Mínguez-Mosquera MI, Hornero-Méndez D. Separation and quantification of the carotenoid pigments in red peppers (Capsicun annuum L.), paprika and oleoresin by reversed-phase HPLC. J Agric Food Chem. 1993;41:1616–1620. [Google Scholar]
- 48.Giménez MJ, Atienza SG, Pistón F. Identification of suitable reference genes for normalization of qPCR data in comparative transcriptomics analyses in the Triticeae. Planta. 2010;233:163–173. doi: 10.1007/s00425-010-1290-y. [DOI] [PubMed] [Google Scholar]
- 49.Ruijter JM, Ramakers C, Hoogaars WMH, Karlen Y, Bakker O, et al. Amplification efficiency: linking baseline and bias in the analysis of quantitative PCR data. Nucl Acids Res. 2009;37(6):e45. doi: 10.1093/nar/gkp045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Ihaka R, Gentleman R. R: A language for data analysis and graphics. J Comput Graph Stat. 1996;5:299–314. [Google Scholar]
- 51.Fox J. The R commander: A basic statistics graphical user interface to R. J Stat Softw. 2005;14:1–42. [Google Scholar]
- 52.Untergasser A, Nijveen H, Rao X, Bisseling T, Geurts R, et al. Primer3Plus, an enhanced web interface to Primer3. Nucl Acids Res. 2007;35:W71–74. doi: 10.1093/nar/gkm306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Emanuelsson O, Nielsen H, Brunak S, von Heijne G. Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J Mol Biol. 2000;300:1005–1016. doi: 10.1006/jmbi.2000.3903. [DOI] [PubMed] [Google Scholar]
- 54.He X, Wang J, Ammar K, Peña RJ, Xia X, et al. Allelic variants at the and loci in durum wheat and their associations with grain yellowness. Crop Sci. 2009;49:2058–2064. [Google Scholar]
- 55.He XY, He ZH, Ma W, Appels R, Xia XC. Allelic variants of phytoene synthase 1 (Psy1) genes in Chinese and CIMMYT wheat cultivars and development of functional markers for flour colour. Mol Breed. 2009;23:553–563. [Google Scholar]
- 56.Tamura K, Nei M, Kumar S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci USA. 2004;101:11030–11035. doi: 10.1073/pnas.0404206101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Tamura K, Dudley J, Nei M, Kumar S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0. Mol Biol Evol. 2007;24:1596–1599. doi: 10.1093/molbev/msm092. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Neighbor-joining tree generated from the alignment of the Psy1 coding sequences from different species. Neighbor-joining tree generated from the alignment of the Psy1 coding sequences from Oryza sativa, Zea mays, Triticum aestivum, Triticum turgidum, Triticum urartu, Triticum monococcum, Aegilops tauschii and Aegilops speltoides. The Psy2 coding sequences of O. sativa and Z. mays, two partial sequences of Psy2 of Triticum turgidum, and Psy3 sequences from O. sativa and Z. mays were used like out-group. Predicted correctly spliced forms of Psy1 gene in H. chilense lines H1, H7 and H16 were used. Numbers over the tree nodes are bootstrap confidence values based on 1,000 bootstrap iterations.
(TIF)
Color complementation in co-transformed E. coli pellets and the corresponding RP-HPLC chromatograms of the carotenoid extracts. (A) A yellow color is observed in E. coli pellets co-transformed with plasmid pAC-85b and pGEMT expressing proteins PSY1Hc-432a and PSY1Hc-432b, while no color complementation is observed when proteins PSY1Hc-86 or PSY1Hc-53 are expressed; (B) Absorption spectra in the HPLC mobile phase of β-carotene; (C) Corresponding RP-HPLC chromatograms of carotenoid extracts shown in (A).
(TIF)
Chromosomal localization of Psy1 gene in H. chilense . Amplification of genomic DNA with primer pair HcPsy1-CDS4F/HcPsy1-2ER in the following lines: (1) H. chilense line H1, (2) H. chilense line H7, (3) T. turgidum ssp. durum cv. Yavaros, (4) T. aestivum cv. Chinese Spring, (5) T. turgidum var. Kofa, (6) T. turgidum breeding line UC1113, (7) 1HchS monotelosomic addition line, (8) 1HchS ditelosomic addition line, (9) 2Hch-α ditelosomic addition line, (10) 4Hch disomic addition line (11) 5Hch disomic addition line, (12) 5HchL ditelosomic addition line, (13) 6Hch disomic addition line, (14) 7Hch disomic addition line, (15) 7Hch-α ditelosomic addition line, (16) 7Hch-β ditelosomic addition line and (17) 6Hch-S ditelosomic addition line.
(TIF)
Alignment of partial Psy1 and Psy2 sequences in H1 and H7 lines of H. chilense. Alignment of genomic (g) and cDNA (c) sequences of Psy1 (this work) and partial genomic sequence of Psy2 gene in H1 and H7 lines (DU796678 and DU796680, respectively).
(TIF)
Alignment of Psy3 gene from different species. Alignment of Psy3 sequences of Sorghum bicolor (Sb; AY705390.1), Zea mays (Zm; NM_001114653.1), Oryza sativa (Os; FJ214953.1) and Hordeum vulgare clone NIASHv2034M16 showing high homology with Psy3 gene of other species (AK365521.1), Hordeum vulgare clone NIASHv3051L01 showing high homology with Psy1 gene of other species (AK374031.1) and H. chilense Psy1 gene (this work).
(TIF)