Abstract
Summary: We developed a Python package, ProDy, for structure-based analysis of protein dynamics. ProDy allows for quantitative characterization of structural variations in heterogeneous datasets of structures experimentally resolved for a given biomolecular system, and for comparison of these variations with the theoretically predicted equilibrium dynamics. Datasets include structural ensembles for a given family or subfamily of proteins, their mutants and sequence homologues, in the presence/absence of their substrates, ligands or inhibitors. Numerous helper functions enable comparative analysis of experimental and theoretical data, and visualization of the principal changes in conformations that are accessible in different functional states. ProDy application programming interface (API) has been designed so that users can easily extend the software and implement new methods.
Availability: ProDy is open source and freely available under GNU General Public License from http://www.csb.pitt.edu/ProDy/.
Contact: ahb12@pitt.edu; bahar@pitt.edu
1 INTRODUCTION
Protein dynamics plays a key role in a wide range of molecular events in the cell, including substrate/ligand recognition, binding, allosteric signaling and transport. For a number of well-studied proteins, the Protein Data Bank (PDB) hosts multiple high-resolution structures. Typical examples are drug targets resolved in the presence of different inhibitors. These ensembles of structures convey information on the structural changes that are physically accessible to the protein, and the delineation of these structural variations provides insights into structural mechanisms of biological activity (Bakan and Bahar, 2009; Yang et al., 2008).
Existing computational tools and servers for characterizing protein dynamics are suitable for single structures [e.g. Anisotropic Network Model (ANM) server (Eyal et al., 2006), elNémo (Suhre and Sanejouand, 2004), FlexServ (Camps et al., 2009)], pairs of structures [e.g. open and closed forms of enzymes; MolMovDB (Gerstein and Krebs, 1998)], or nucleic magnetic resonance (NMR) models [e.g. PCA_NEST (Yang et al., 2009)]. Tools for systematic retrieval and analyses of ensembles of structures are not publicly accessible. Ensembles include X-ray structures for a given protein and its complexes; families and subfamilies of proteins that belong to particular structural folds; or a protein and its mutants resolved in the presence of different inhibitors, ligands or substrates. The analysis of structural variability in these ensembles could open the way to gain insights into rearrangements selected/stabilized in different functional states (Bahar et al., 2007, 2010), or into the structure-encoded dynamic features shared by protein family or subfamily members (Marcos et al., 2010; Raimondi et al., 2010; Velazquez-Muriel et al., 2009). The lack of software for performing such operations is primarily due to the non-uniform content of structural datasets such as sequence variations at particular regions, including missing or substituted residues, short segments or loops. We developed ProDy to analyze and retrieve biologically significant information from such heterogeneous structural datasets. ProDy delivers information on the structural variability of target systems and allows for systematic comparison with the intrinsic dynamics predicted by theoretical models and methods, thus helping gain insight into the relation between structure, dynamics and function.
2 DESCRIPTION AND FUNCTIONALITY
2.1 Input for ProDy
The input for ProDy is the set of atomic coordinates in PDB format for the protein of interest, or simply the PDB id or sequence of the protein. Given a query protein, fast and flexible ProDy parsers are used to Blast search the PDB, retrieve the corresponding files (e.g. mutants, complexes or sequence homologs with user-defined minimal sequence identity) from the PDB FTP server and extract their coordinates and other relevant data. Additionally, the program can be used to analyze a series of conformers from molecular dynamics (MD) trajectories inputted in PDB file format or programmatically through Python NumPy arrays. More information on the input format is given at the ProDy web site tutorial and examples.
2.2 Protein ‘dynamics’ from experiments
The experimental data refer to ensembles of structures, X-ray crystallographic or NMR. These are usually heterogeneous datasets, in the sense that they have disparate coordinate data arising from sequence dissimilarities, insertions/deletions or missing data due to unresolved disordered regions. In ProDy, we implemented algorithms for optimal alignment of such heterogeneous datasets and building corresponding covariance matrices. Covariance matrices describe the mean-square deviations in atomic coordinates from their mean position (diagonal elements) or the correlations between their pairwise fluctuations (off-diagonal elements). The principal modes of structural variation are determined upon principal component analysis (PCA) of the covariance matrix, as described previously (Bakan and Bahar, 2009).
2.3 Protein dynamics from theory and simulations
We have implemented classes for Gaussian network model (GNM) analysis and for normal mode analysis (NMA) of a given structure using the ANM (Eyal et al., 2006). Both models have been widely used in recent years for analyzing and visualizing biomolecular systems dynamics (Bahar et al., 2010). The implementation is generic and flexible. The user can (i) build the models for any set of atoms, e.g. the substrate or inhibitor can be explicitly included to study the perturbing effect of binding on dynamics, and (ii) utilize user-defined or built-in distance-dependent or residue-specific force constants (Hinsen et al., 2000; Kovacs et al., 2004). ProDy also offers the option to perform essential dynamics analysis (EDA; Amadei et al., 1993) of MD snapshots, which is equivalent to the singular value decomposition of trajectories to extract principal variations (Velazquez-Muriel et al., 2009).
2.4 Dynamics analysis example
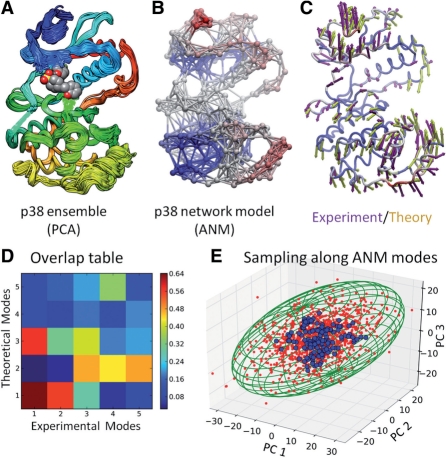
Figure 1 illustrates the outputs generated by ProDy in a comparative analysis of experimental and computational data for p38 kinase (Bakan and Bahar, 2011). Figure 1A displays the dataset of 150 X-ray crystallographically resolved p38 structures retrieved from the PDB and optimally overlaid by ProDy. The ensemble contains the apo and inhibitor-bound forms of p38, thus providing information on the conformational space sampled by p38 upon inhibitor binding. Parsing structures, building and diagonalizing the covariance matrix to determine the principal modes of structural variations takes only 38 s on Intel CPU at 3.20 GHz. Figure 1C illustrate the first principal mode of structural variation (PC1; violet arrows) based exclusively on experimental structural dataset for p38.
Fig. 1.
Comparative analysis of p38 dynamics from experiments (PCA) and theory (ANM). (A) Overlay of 150 p38 X-ray structures using ProDy. An inhibitor is shown in space-filling representation. (B) Network model (ANM) representation of p38 (generated using NMWiz and VMD). (C) Comparison of the principal mode PC1 (from experiments; violet arrows) and the softest mode ANM1 from theory (green arrows) and (D) overlap of the top five modes. (E) Distribution of X-ray structures (blue) and ANM-generated conformers (red) in the subspace spanned by PC1-3. The green ellipsoid is an analytical solution predicted by the ANM.
As to generating computational data, two approaches are taken in ProDy: NMA of a representative structure using its ANM representation (Figure 1B; color-coded such that red/blue regions refer to largest/smallest conformational mobilities); and EDA of MD trajectories provided that an ensemble of snapshots is provided by the user. The green arrows in Figure 1C describe the first (lowest frequency, most collective) mode predicted by the ANM, shortly designated as ANM1. The heatmap in Figure 1D shows the overlap (Marques and Sanejouand, 1995) between top-ranking PCA and ANM modes. The cumulative overlap between the top three pairs of modes is 0.73.
An important aspect of ProDy is the sampling of a representative set of conformers consistent with experiments—a feature expected to find wide utility in flexible docking and structure refinement. Figure 1E displays the conformational space sampled by experimental structures (blue dots), projected onto the subspace spanned by the top three PCA directions, which accounts for 59% of the experimentally observed structural variance. The conformations generated using the softest modes ANM1-ANM3 predicted to be intrinsically accessible to p38 in the apo form, are shown by the red dots. The sizes of the motions along these modes obey a Gaussian distribution with variance scaling with the inverse square root of the corresponding eigenvalues. ANM conformers cover a subspace (green ellipsoidal envelope) that encloses all experimental structures. Detailed information on how to generate such plots and figures using ProDy is given in the online documentation, along with several examples of downloadable scripts.
2.5 Graphical interface
We have designed a graphical interface, NMWiz, to enable users to perform ANM and PCA calculations from within a molecular visualization program. NMWiz is designed as a VMD (Humphrey et al., 1996) plugin, and is distributed within the ProDy installation package. It is used to do calculations for molecules loaded into VMD; and results are visualized on the fly. The plug-in allows for depicting color-coded network models and normal mode directions (Fig. 1B and C), displaying animations of various PCA and ANM modes, generating trajectories, and plotting square fluctuations.
2.6 Supporting features
ProDy comes with a growing library of functions to facilitate comparative analysis. Examples are functions to calculate, print and plot the overlaps between experiment, theory and computations (Fig. 1D) or to view the spatial dispersion of conformers (Fig. 1E).
For rapid and flexible analysis of large numbers of PDB structures, we designed a fast PDB parser. The parser can handle alternate locations and multiple models, and read specified chains or atom subsets selected by the user. We evaluated the performance of ProDy relative to Biopython PDB module (Hamelryck and Manderick, 2003) using 4701 PDB structures listed in the PDB SELECT dataset (Hobohm and Sander, 1994): we timed parsers for reading the PDB files and returning Cα-coordinates to the user (see documentation). The Python standard Biopython PDB parser evaluated the dataset in 52 min; and ProDy in 11 min. In addition, we implemented an atom selector using Pyparsing module for rapid access to subsets of atoms in PDB files. This feature reduces the user programming effort to access any set of atoms down to a single line of code from several lines composed of nested loops and comparisons required with the current Python packages for handling PDB data. The implementation of atom selections follows that in VMD. Full list of selection keywords and usage examples are provided in the documentation. ProDy also offers functions for structural alignment and comparison of multiple chains.
3 DISCUSSION
Several web servers have been developed for characterizing protein dynamics, including elNémo (Suhre and Sanejouand, 2004), ANM (Eyal et al., 2006) and FlexServ (Camps et al., 2009). These servers perform coarse-grained ENM–based NMA calculations, and as such are useful for elucidating structure-encoded dynamics of proteins. FlexServ also offers the option to use distance-dependent force constants (Kovacs et al., 2004), in addition to protocols for coarse-grained generation and PCA of trajectories. ProDy differs from these as it allows for systematic retrieval and comparative analysis of ensembles of heterogeneous structural datasets. Such datasets includes structural data collected for members of a protein family in complex with different substrates/inhibitors. ProDy further allows for the quantitative comparison of the results from experimental datasets with theoretically predicted conformational dynamics. In addition, ProDy offers the following advantages: (i) it is extensible, interoperable and suitable for use as a toolkit for developing new software; (ii) it provides scripts for automated tasks and batch analyses of large datasets; (iii) it has a flexible API suitable for testing new methods and hypotheses, and benchmarking them against existing methods with minimal effort and without the need to modify the source code; (iv) it allows for producing publication quality figures when used with Python plotting library Matplotlib; and (v) it provides the option to input user-defined distance-dependent force function or utilize elaborate classes that return force constants based on the type and properties of interacting residues [e.g. based on their secondary structure or sequence separation (Lezon and Bahar, 2010)].
4 CONCLUSION
ProDy is a free, versatile, easy-to-use and powerful tool for inferring protein dynamics from both experiments (i.e. PCA of ensembles of structures) and theory (i.e. GNM, ANM and EDA of MD snapshots). ProDy complements existing tools by allowing the systematic retrieval and analysis of heterogeneous experimental datasets, leveraging on the wealth of structural data deposited in the PDB to gain insights into structure-encoded dynamics. In addition, ProDy allows for comparison of the results from experimental datasets with theoretically predicted conformational dynamics. Finally, through a flexible Python-based API, ProDy can be used to quickly test and implement new methods and ideas, thus lowering the technical barriers to apply such methods in more complex computational analyses.
Funding: National Institutes of Health (1R01GM086238-01 to I.B. and UL1 RR024153 to A.B.).
Conflict of Interest: none declared.
REFERENCES
- Amadei A., et al. Essential dynamics of proteins. Proteins. 1993;17:412–425. doi: 10.1002/prot.340170408. [DOI] [PubMed] [Google Scholar]
- Bahar I., et al. Intrinsic dynamics of enzymes in the unbound state and relation to allosteric regulation. Curr. Opin. Struct. Biol. 2007;17:633–640. doi: 10.1016/j.sbi.2007.09.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bahar I., et al. Normal mode analysis of biomolecular structures: functional mechanisms of membrane proteins. Chem. Rev. 2010;110:1463–1497. doi: 10.1021/cr900095e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bakan A., Bahar I. Computational generation of inhibitor-bound conformers of p38 MAP kinase and comparison with experiments. Pac. Symp. Biocomput. 2011;16:181–192. doi: 10.1142/9789814335058_0020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bakan A., Bahar I. The intrinsic dynamics of enzymes plays a dominant role in determining the structural changes induced upon inhibitor binding. Proc. Natl Acad. Sci. USA. 2009;106:14349–14354. doi: 10.1073/pnas.0904214106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Camps J., et al. FlexServ: an integrated tool for the analysis of protein flexibility. Bioinformatics. 2009;25:1709–1710. doi: 10.1093/bioinformatics/btp304. [DOI] [PubMed] [Google Scholar]
- Eyal E., et al. Anisotropic network model: systematic evaluation and a new web interface. Bioinformatics. 2006;22:2619–2627. doi: 10.1093/bioinformatics/btl448. [DOI] [PubMed] [Google Scholar]
- Gerstein M., Krebs W. A database of macromolecular motions. Nucleic Acids Res. 1998;26:4280–4290. doi: 10.1093/nar/26.18.4280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamelryck T., Manderick B. PDB file parser and structure class implemented in Python. Bioinformatics. 2003;19:2308–2310. doi: 10.1093/bioinformatics/btg299. [DOI] [PubMed] [Google Scholar]
- Hinsen K., et al. Harmonicity in slow protein dynamics. Chem. Phys. 2000;261:25–37. [Google Scholar]
- Hobohm U., Sander C. Enlarged representative set of protein structures. Protein Sci. 1994;3:522–524. doi: 10.1002/pro.5560030317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Humphrey W., et al. VMD: visual molecular dynamics. J. Mol. Graph. 1996;14:33–38. doi: 10.1016/0263-7855(96)00018-5. [DOI] [PubMed] [Google Scholar]
- Kovacs J.A., et al. Predictions of protein flexibility: first-order measures. Proteins. 2004;56:661–668. doi: 10.1002/prot.20151. [DOI] [PubMed] [Google Scholar]
- Lezon T.R., Bahar I. Using entropy maximization to understand the determinants of structural dynamics beyond native contact topology. PLoS. Comput. Biol. 2010;6:e1000816. doi: 10.1371/journal.pcbi.1000816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marcos E., et al. On the conservation of the slow conformational dynamics within the amino acid kinase family: NAGK the paradigm. PLoS Comput. Biol. 2010;6:e1000738. doi: 10.1371/journal.pcbi.1000738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marques O., Sanejouand Y.H. Hinge-bending motion in citrate synthase arising from normal mode calculations. Proteins. 1995;23:557–560. doi: 10.1002/prot.340230410. [DOI] [PubMed] [Google Scholar]
- Raimondi F., et al. Deciphering the deformation modes associated with function retention and specialization in members of the Ras superfamily. Structure. 2010;18:402–414. doi: 10.1016/j.str.2009.12.015. [DOI] [PubMed] [Google Scholar]
- Suhre K., Sanejouand Y.H. ElNémo: a normal mode web server for protein movement analysis and the generation of templates for molecular replacement. Nucleic Acids Res. 2004;32:W610–W614. doi: 10.1093/nar/gkh368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Velazquez-Muriel J.A., et al. Comparison of molecular dynamics and superfamily spaces of protein domain deformation. BMC Struct. Biol. 2009;9:6. doi: 10.1186/1472-6807-9-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang L., et al. Close correspondence between the motions from principal component analysis of multiple HIV-1 protease structures and elastic network modes. Structure. 2008;16:321–330. doi: 10.1016/j.str.2007.12.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang L.W., et al. Principal component analysis of native ensembles of biomolecular structures (PCANEST): insights into functional dynamics. Bioinformatics. 2009;25:606–614. doi: 10.1093/bioinformatics/btp023. [DOI] [PMC free article] [PubMed] [Google Scholar]