Abstract
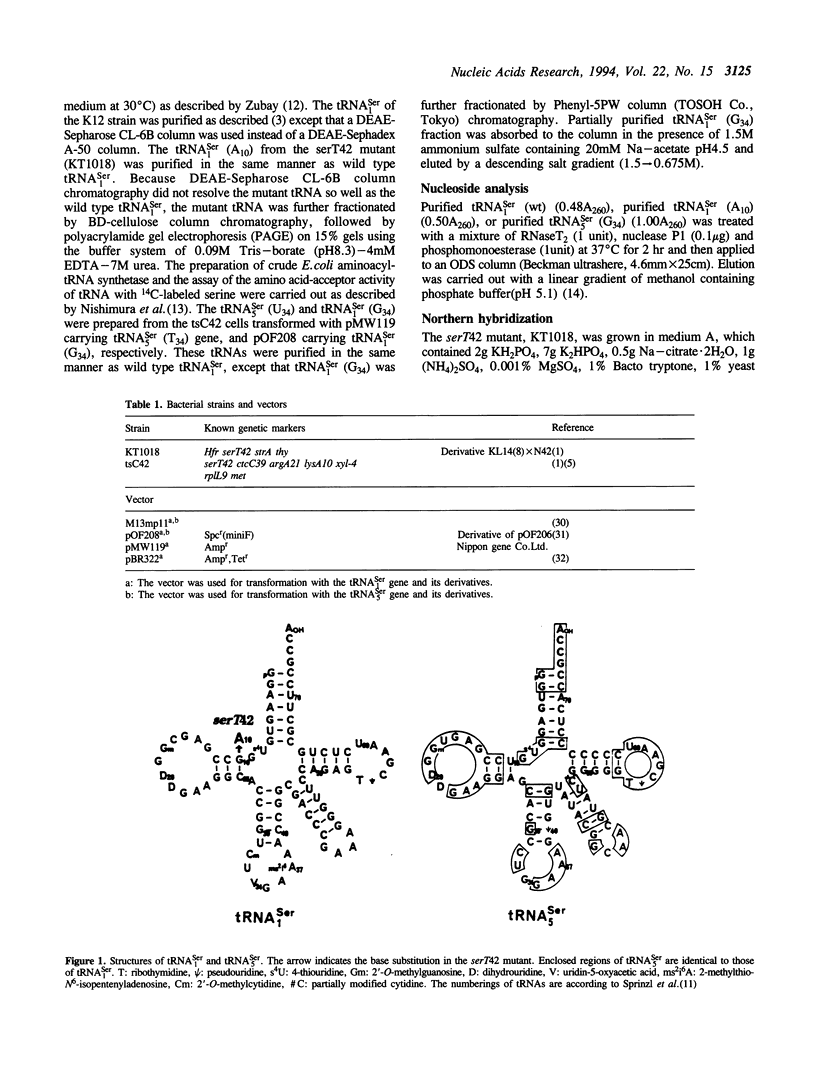
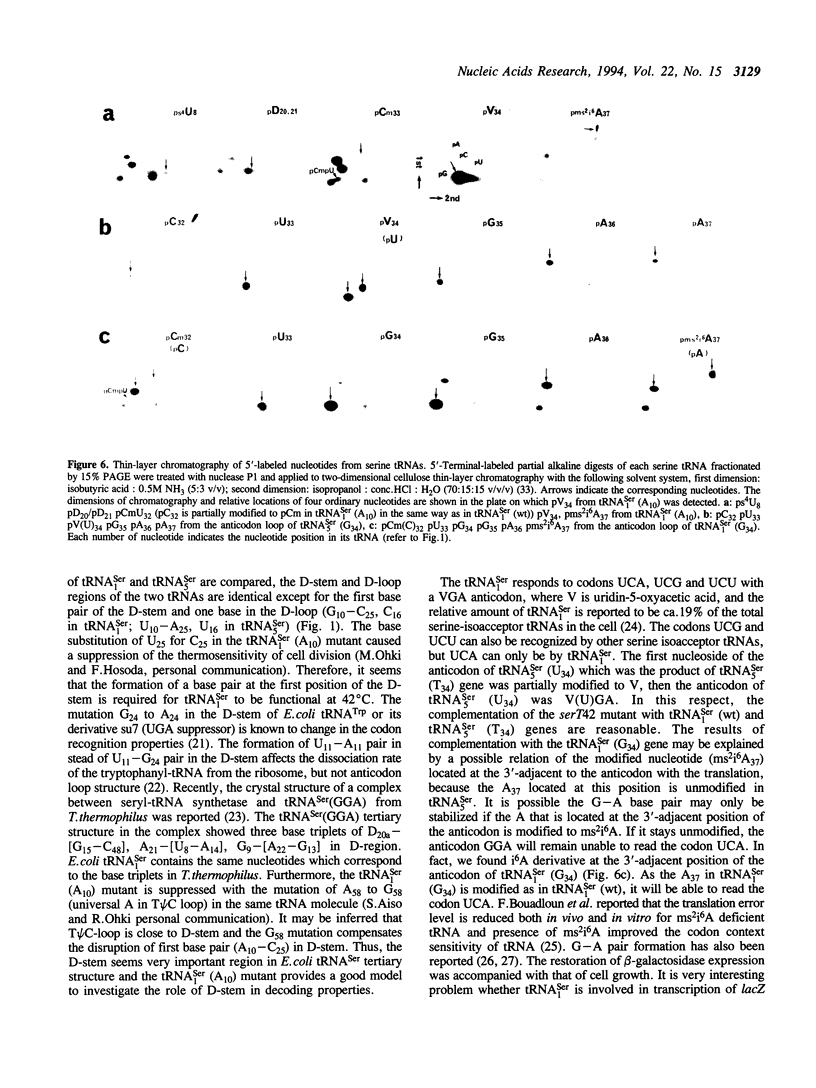
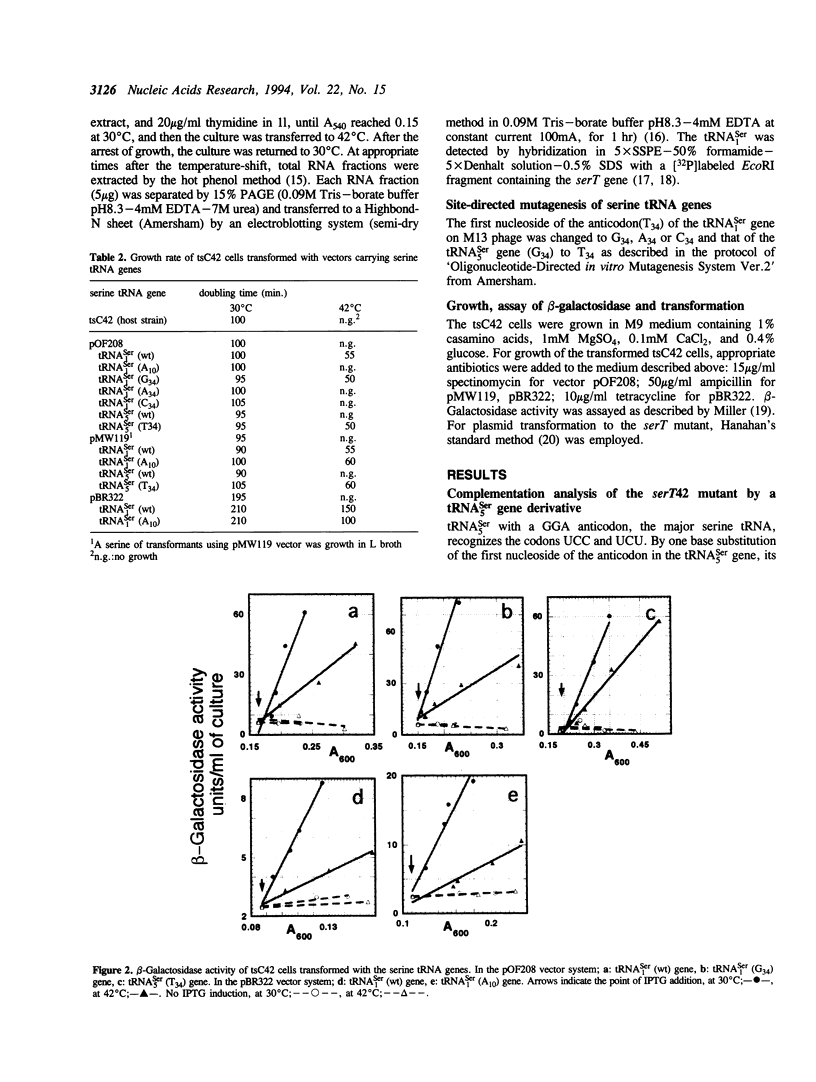
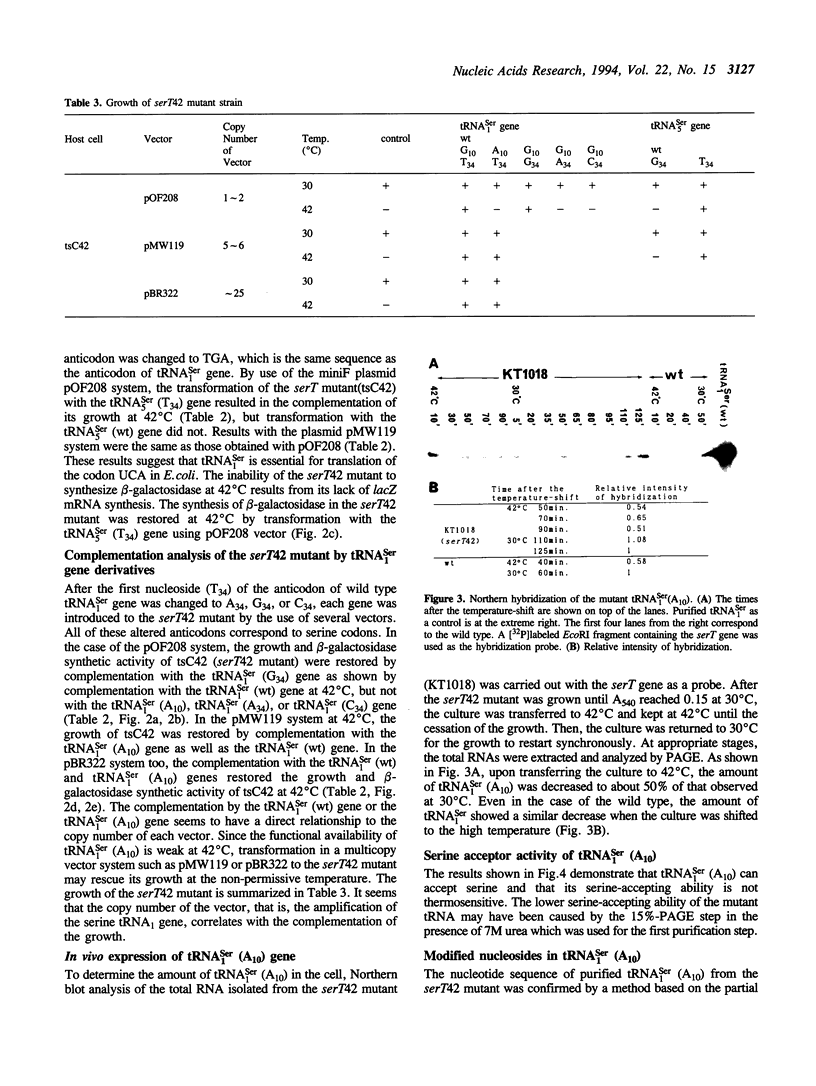
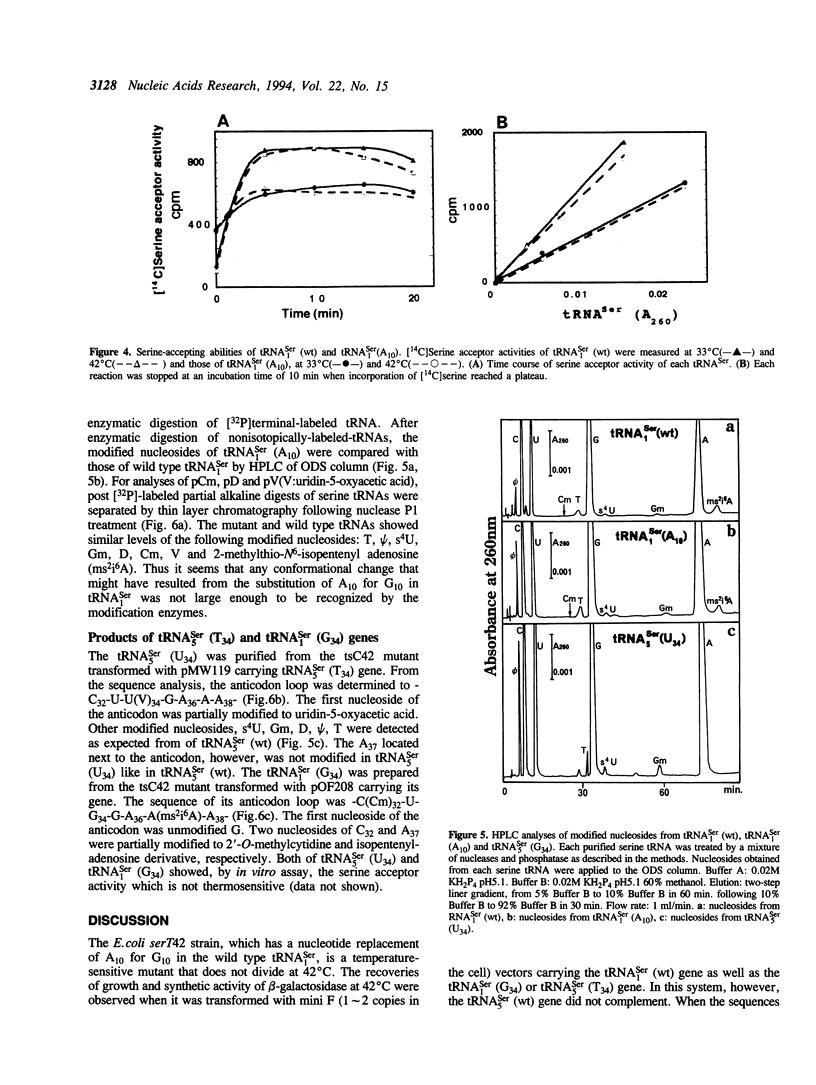
Serine tRNA gene derivatives with altered anticodons were introduced to the temperature-sensitive serT42 mutant, whose tRNA(1Ser) shows a base substitution of A10 for wild type G10. When a low copy number vector-system was used, the growth and beta-galactosidase synthetic activity of the serT42 mutant were restored by complementation with the tRNA(5Ser) (T34) gene or the tRNA(1Ser) (G34) gene as well as the tRNA(1Ser) (wt) gene, but not with tRNA(5Ser) (wt), tRNA(1Ser) (A34) or tRNA(1Ser) (C34) genes at 42 degrees C. When multicopy vectors were used, the transformation even with tRNA(1Ser) (A10) gene restored the growth and beta-galactosidase synthetic activity at 42 degrees C. The tRNA(1Ser) (A10) showed no thermosensitivity in serine acceptor activity by in vitro assay. At 42 degrees C, the amount of tRNA(1Ser) (A10) in the serT42 mutant was almost the same as those in the wild type. The nucleotides in the tRNA(1Ser) (A10) were found to be fully modified like those in the wild type tRNA(1Ser). Both of the tRNAs transcribed from tRNA(5Ser) (T34) and tRNA(1Ser) (G34) genes showed serine acceptor activity. Modified nucleosides of these tRNAs were also analyzed.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Biou V., Yaremchuk A., Tukalo M., Cusack S. The 2.9 A crystal structure of T. thermophilus seryl-tRNA synthetase complexed with tRNA(Ser). Science. 1994 Mar 11;263(5152):1404–1410. doi: 10.1126/science.8128220. [DOI] [PubMed] [Google Scholar]
- Bittner M., Kupferer P., Morris C. F. Electrophoretic transfer of proteins and nucleic acids from slab gels to diazobenzyloxymethyl cellulose or nitrocellulose sheets. Anal Biochem. 1980 Mar 1;102(2):459–471. doi: 10.1016/0003-2697(80)90182-7. [DOI] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Greene P. J., Betlach M. C., Heyneker H. L., Boyer H. W., Crosa J. H., Falkow S. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene. 1977;2(2):95–113. [PubMed] [Google Scholar]
- Bouadloun F., Srichaiyo T., Isaksson L. A., Björk G. R. Influence of modification next to the anticodon in tRNA on codon context sensitivity of translational suppression and accuracy. J Bacteriol. 1986 Jun;166(3):1022–1027. doi: 10.1128/jb.166.3.1022-1027.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grosjean H., Nicoghosian K., Haumont E., Söll D., Cedergren R. Nucleotide sequences of two serine tRNAs with a GGA anticodon: the structure-function relationships in the serine family of E. coli tRNAs. Nucleic Acids Res. 1985 Aug 12;13(15):5697–5706. doi: 10.1093/nar/13.15.5697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grundy F. J., Henkin T. M. tRNA as a positive regulator of transcription antitermination in B. subtilis. Cell. 1993 Aug 13;74(3):475–482. doi: 10.1016/0092-8674(93)80049-k. [DOI] [PubMed] [Google Scholar]
- Hirsh D. Tryptophan transfer RNA as the UGA suppressor. J Mol Biol. 1971 Jun 14;58(2):439–458. doi: 10.1016/0022-2836(71)90362-7. [DOI] [PubMed] [Google Scholar]
- Hosoda F., Nishimura S., Uchida H., Ohki M. An F factor based cloning system for large DNA fragments. Nucleic Acids Res. 1990 Jul 11;18(13):3863–3869. doi: 10.1093/nar/18.13.3863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishikura H., Yamada Y., Nishimura S. Structure of serine tRNA from Escherichia coli. I. Purification of serine tRNA's with different codon responses. Biochim Biophys Acta. 1971 Jan 28;228(2):471–481. doi: 10.1016/0005-2787(71)90052-9. [DOI] [PubMed] [Google Scholar]
- Ishikura H., Yamada Y., Nishimura S. The nucleotide sequence of a serine tRNA from Escherichia coli. FEBS Lett. 1971 Jul 15;16(1):68–70. doi: 10.1016/0014-5793(71)80688-9. [DOI] [PubMed] [Google Scholar]
- Komine Y., Adachi T., Inokuchi H., Ozeki H. Genomic organization and physical mapping of the transfer RNA genes in Escherichia coli K12. J Mol Biol. 1990 Apr 20;212(4):579–598. doi: 10.1016/0022-2836(90)90224-A. [DOI] [PubMed] [Google Scholar]
- Kuchino Y., Hanyu N., Nishimura S. Analysis of modified nucleosides and nucleotide sequence of tRNA. Methods Enzymol. 1987;155:379–396. doi: 10.1016/0076-6879(87)55026-1. [DOI] [PubMed] [Google Scholar]
- Messing J. New M13 vectors for cloning. Methods Enzymol. 1983;101:20–78. doi: 10.1016/0076-6879(83)01005-8. [DOI] [PubMed] [Google Scholar]
- Nishimura S., Harada F., Narushima U., Seno T. Purification of methionine-, valine-, phenylalanine- and tyrosine-specific tRNA from Escherichia coli. Biochim Biophys Acta. 1967 Jun 20;142(1):133–148. doi: 10.1016/0005-2787(67)90522-9. [DOI] [PubMed] [Google Scholar]
- Ohki M., Sato S. Regulation of expression of lac operon by a novel function essential for cell growth. Nature. 1975 Feb 20;253(5493):654–656. doi: 10.1038/253654a0. [DOI] [PubMed] [Google Scholar]
- Ohki M., Smith C. L. Tracking bacterial DNA replication forks in vivo by pulsed field gel electrophoresis. Nucleic Acids Res. 1989 May 11;17(9):3479–3490. doi: 10.1093/nar/17.9.3479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oki M., Mitsui H. Defective membrane synthesis in an E. coli mutant. Nature. 1974 Nov 1;252(5478):64–66. doi: 10.1038/252064a0. [DOI] [PubMed] [Google Scholar]
- SantaLucia J., Jr, Kierzek R., Turner D. H. Effects of GA mismatches on the structure and thermodynamics of RNA internal loops. Biochemistry. 1990 Sep 18;29(37):8813–8819. doi: 10.1021/bi00489a044. [DOI] [PubMed] [Google Scholar]
- Sato T., Ohki M., Yura T., Ito K. Genetic studies of an Escherichia coli K-12 temperature-sensitive mutant defective in membrane protein synthesis. J Bacteriol. 1979 May;138(2):305–313. doi: 10.1128/jb.138.2.305-313.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith D., Yarus M. Transfer RNA structure and coding specificity. I. Evidence that a D-arm mutation reduces tRNA dissociation from the ribosome. J Mol Biol. 1989 Apr 5;206(3):489–501. doi: 10.1016/0022-2836(89)90496-8. [DOI] [PubMed] [Google Scholar]
- Tamura F., Nishimura S., Ohki M. The E. coli divE mutation, which differentially inhibits synthesis of certain proteins, is in tRNASer1. EMBO J. 1984 May;3(5):1103–1107. doi: 10.1002/j.1460-2075.1984.tb01936.x. [DOI] [PMC free article] [PubMed] [Google Scholar]





