Abstract
The angiotensin II peptide analog [Sar1,Ile4,Ile8]AngII (SII) is a biased AT1A receptor agonist that stimulates receptor phosphorylation, β-arrestin recruitment, receptor internalization, and β-arrestin-dependent ERK1/2 activation without activating heterotrimeric G-proteins. To determine the scope of G-protein-independent AT1A receptor signaling, we performed a gel-based phosphoproteomic analysis of AngII and SII-induced signaling in HEK cells stably expressing AT1A receptors. A total of 34 differentially phosphorylated proteins were detected, of which 16 were unique to SII and eight to AngII stimulation. MALDI-TOF/TOF mass fingerprinting was employed to identify 24 SII-sensitive phosphoprotein spots, of which three (two peptide inhibitors of protein phosphatase 2A (I1PP2A and I2PP2A) and prostaglandin E synthase 3 (PGES3)) were selected for validation and further study. We found that phosphorylation of I2PP2A was associated with rapid and transient inhibition of a β-arrestin 2-associated pool of protein phosphatase 2A, leading to activation of Akt and increased phosphorylation of glycogen synthase kinase 3β in an arrestin signalsome complex. SII-stimulated PGES3 phosphorylation coincided with an increase in β-arrestin 1-associated PGES3 and an arrestin-dependent increase in cyclooxygenase 1-dependent prostaglandin E2 synthesis. These findings suggest that AT1A receptors regulate a robust G protein-independent signaling network that affects protein phosphorylation and autocrine/paracrine prostaglandin production and that these pathways can be selectively modulated by biased ligands that antagonize G protein activation.
Keywords: Akt PKB, G-protein Coupled Receptors (GPCR), PP2A, Prostaglandins, Signal Transduction, AT1 Receptor, I2PP2A, PGES3, Arrestin, Biased Agonism
Introduction
Many of the effects of the vasopressor hormone angiotensin II (AngII)2 are mediated by the type 1A angiotensin receptor (AT1AR), a heptahelical G-protein-coupled receptor (GPCR) that activates Gq/11 proteins leading to phospholipase Cβ activation and intracellular calcium accumulation. Although many of its functions are attributable to the activation of heterotrimeric G-proteins, the AT1AR is known to affect a number of other intracellular pathways through less well defined signal transduction mechanisms. These include transactivation of receptor tyrosine kinases (1), Janus kinase-signal transducer and activator of transcription pathway activation (2), and activation of ERK1/2 through β-arrestin-dependent signalsome formation (3). The AT1AR plays a pathophysiologic role in several cardiovascular diseases, such as essential hypertension, atherosclerosis, and maladaptive cardiac hypertrophy. Little is known, however, about the role of non-classical AT1R signaling in the genesis of these conditions.
The term “functional selectivity” refers to the property of some ligands to selectively couple a GPCR to different effectors, producing a ligand-specific signaling profile that is distinct from that of the native agonist. This phenomenon is hypothesized to result from the ligand-dependent stabilization of receptor conformations that favor coupling to only a subset of all possible downstream effectors. Such “biased” agonists have been described for several GPCRs, among them the type 1 parathyroid hormone receptor, δ-opioid receptor, β2 adrenergic receptor, and the AT1AR. The synthetic congener of AngII, [Sar1,Ile4,Ile8]AngII (SII), is an arrestin pathway-selective biased agonist for the AT1AR. SII was originally described as an AT1AR antagonist that possessed the novel property of promoting receptor internalization without activating G-protein signaling (4). It was later demonstrated that SII activates ERK1/2 using β-arrestin as a ligand-regulated scaffold for the assembly of an ERK1/2 activation complex (5). Unlike G-protein-mediated ERK1/2 activation, ERK1/2 activated by SII remains associated with the receptor-arrestin signalsome complex in the cytosol, where it presumably performs distinct functions. This reversal of efficacy, wherein SII behaves as a competitive antagonist of AT1AR-G protein coupling but as an agonist for arrestin-dependent receptor internalization and signaling, defines it as an arrestin pathway-selective biased AT1AR agonist. Not surprisingly, the in vivo efficacy of SII appears to differ from both AngII and conventional AT1AR antagonists. Despite its antagonism of AngII-mediated calcium signaling, SII has been reported to increase cardiomyocyte inotropy and lusotropy (6), increase protein translation (7), and stimulate adrenal aldosterone synthesis and release (8).
Although it is now recognized that β-arrestins bind a diverse set of catalytically active proteins, enabling them to recruit protein and lipid kinase, phosphatase, phosphodiesterase, and ubiquitin ligase activity into the receptor-arrestin complexes (9, 10), there is limited information about how these signalsome functions integrate into the broader scope of cellular metabolism. The goal of the present study was to employ SII as a tool to investigate the G-protein-independent AT1AR signal transduction network using gel-based proteomics to identify downstream targets of AT1AR-arrestin signalsomes. Using a limited two-dimensional gel-based phosphoproteomic screen coupled with MALDI-TOF/TOF mass spectroscopy, we demonstrated that SII stimulates a robust signaling response with surprisingly limited overlap with AngII. We found that two previously unreported SII-induced protein phosphorylation events, phosphorylation of I2PP2A and PGES3, are associated with arrestin-dependent changes in the activity of these proteins and their downstream targets. These results indicate that the AT1AR regulates a robust G-protein-independent signaling network that affects protein phosphorylation and autocrine/paracrine prostaglandin production, and that these pathways can be selectively modulated by biased ligands that induce signalsome formation.
EXPERIMENTAL PROCEDURES
Materials
Pro Q Diamond resin, buffers and fluorescent stain, Sypro Ruby protein stain, ZOOM® (pH 4–7) isoelectric focusing strips, 4–20% Tris-glycine gels, Fluo-3 acetoxymethyl ester, cell culture medium, and cell culture additives were from Invitrogen. 35-mm glass bottom culture dishes were from MatTek Co. (Ashland, MA). Double-stranded siRNAs were from Qiagen (Germantown, MD). GeneSilencerTM transfection reagent was from Genelantis (San Diego, CA). FuGENE 6 was from Roche Diagnostics (Indianapolis, IN). AngII, losartan, rabbit polyclonal anti-FLAG IgG, EZ view red M2 anti-FLAG affinity resin, and indomethacin were from Sigma-Aldrich (St. Louis, MO). The eight amino acid SII peptide (Sar-Arg-Val-Ile-Ile-His-Pro-Ile) was custom-synthesized by the Cleveland Clinic Protein Core Facility (Cleveland, OH). Okadaic acid was purchased from EMD Biochemicals (Gibbstown, NJ). Biolytes (pH 3–10) and DCTM protein assay kits were from Bio-Rad Laboratories (Hercules, CA). Protein G plus/protein A-agarose and EDTA-free protease inhibitors (Set III) were from Calbiochem (San Diego, CA). Rabbit polyclonal anti-I1PP2A and -I2PP2A IgG were from Santa Cruz Biotechnology (Santa Cruz, CA). Monoclonal anti-PGES3 IgG and HRP-conjugated anti-phospho-Ser and anti-phospho-Ser/Thr were from Abcam (Cambridge, MA). Phosphorylation state-specific anti-ERK1/2, anti-Akt, anti-GSK3β and total anti-ERK1/2, anti-Akt, anti-GSK3β and anti-GAPDH were from Cell Signaling Technology (Beverly, MA). Rabbit polyclonal anti-PP2A IgG and PP2A activity assay kits were from Millipore (Billerica, MA). Rabbit polyclonal anti-β-arrestin was a gift from Robert J. Lefkowitz (Duke University, Durham, NC). Horseradish peroxidase-conjugated donkey anti-rabbit and anti-mouse IgG were from Jackson ImmunoResearch Laboratories, Inc. (West Grove, PA). Prostaglandin E2 monoclonal enzyme-linked immunoassay kits were from Cayman Chemical Co. (Ann Arbor, MI).
cDNA Expression Plasmids
The expression plasmids encoding the influenza HA epitope-tagged AT1AR (11), C-terminal GFP-tagged AT1AR (12), and FLAG epitope-tagged rat β-arrestin 2 (13) were prepared as described. Expression plasmids encoding PGES3 and cyclooxygenase (Cox)1 were purchased from Origene (Rockville, MD).
Cell Culture and Transfection
HEK293 cells stably expressing the HA epitope-tagged AT1AR (HEK-AT1AR) or GFP-tagged AT1AR were maintained in DMEM, 10% fetal bovine serum, and 1× antibiotic/antimycotic solution. Transient transfection of HEK293 cells was performed at 50% confluence in 10-cm dishes using FuGENE 6 according to the manufacturer's instructions with 10 μg of total plasmid DNA per dish and 3 μl FuGENE per μg DNA. Proteins were allowed to express for 72 h before the assay. Prior to each experiment, transfected cells were seeded into multi-well plates as appropriate, and monolayers were incubated for 2–24 h in serum-free DMEM supplemented with 10 mm Hepes (pH 7.4), 0.1% bovine serum albumin, and antibiotic/antimycotic. Primary rat aortic vascular smooth muscle cells were prepared and cultured as described previously (12). Vascular smooth muscle cells were used between the fifth and eighth passages and were serum-deprived for 72 h before experimentation.
Silencing β-Arrestin Expression by RNA Interference
Simultaneous silencing of β-arrestin 1 and 2 was performed using 19-nucleotide duplex RNA with 3′ dTdT overhangs. The siRNA sequence was 5′-ACCUGCGCCUUCCGCUAUGTT-3′. The non-silencing RNA duplex 5′-AAUUCUCCGAACGUGUCAC GU-3′ was used for the negative control. HEK293 cells at 50% confluence in 10-cm plates were transfected with 20 μg of control or β-arrestin 1/2-targeted siRNA using 50 μl of GeneSilencer transfection reagent according to the manufacturer's instructions. Assays were performed 72 h after gene silencing. Silencing of β-arrestin expression was confirmed by immunoblotting whole cell lysates using rabbit polyclonal anti-β-arrestin IgG with horseradish peroxidase-conjugated donkey anti-rabbit IgG as secondary antibody. For experiments requiring both siRNA silencing and transient transfection, the medium was replaced 24 h after siRNA transfection, and cells were transfected with plasmid cDNA encoding PGES3 and/or Cox1 using FuGENE 6. Assays were performed 72 h after FuGENE transfection.
Ligand Stimulation
Stock solutions (100×) of SII were prepared in 10 mm acetic acid, 154 mm NaCl, and 0.1% bovine serum albumin. The Kd of SII for the AT1AR is ∼0.3 μm compared with 0.1 nm for AngII. To ensure full receptor occupancy, stimulations were performed using SII and AngII at ∼100-fold greater than Kd (50 μm SII and 100 nm AngII). Non-stimulated controls were exposed to vehicle only for the equivalent times.
Intracellular Calcium Influx
HEK-AT1AR cells were seeded in 96-well clear-bottom black plates at a density of 6 × 105 cells/well and allowed to grow for 24 h before overnight serum starvation. Prior to assay, the cells were incubated for 60 min at 37 °C with 2 μm calcium-sensitive fluorescent dye, Fluo-3 acetoxymethyl ester, in Hanks' balanced salt solution (pH 7.4) containing 2.5 mm probenecid and 0.1% bovine serum albumin. After incubation, the cells were washed with Hanks' balanced salt solution and exposed to agonists. Intracellular calcium flux was measured using a fluorometric imaging plate reader (FLIPR1TM, Molecular Devices, Sunnyvale, CA). Acquired data were analyzed using system software.
Confocal Microscopy
To visualize AT1AR internalization, HEK293 stably expressing GFP-tagged AT1AR were plated onto collagen-coated 35-mm glass-bottom culture dishes, serum-starved overnight, stimulated as described in the figure legends, washed with Hank's balanced salt solution, and fixed with 4% paraformaldehyde for 30 min at room temperature. Confocal microscopy was performed on a Zeiss LSM510 laser scanning microscope using a Zeiss 63× 1.4 numerical aperture oil immersion lens and 488 nm excitation and 515–540 nm emission filter sets.
Phosphoprotein Enrichment
Five confluent 10-cm plates of HEK-AT1AR cells were used for each sample. Cells were serum-starved for 2 h before a 5-min stimulation with SII, AngII, or vehicle. Monolayers were washed once with ice-cold Hank's balanced salt solution and lysed with 0.4 ml/plate ice-cold Pro Q lysis buffer containing protease inhibitors and a phosphatase inhibitor mixture composed of 1 mm Na3VO4, 10 mm NaF, and 0.2 mm vanadium oxide. The lysates were sonicated, incubated on ice for 30 min with vortexing for 1 min every 10 min, clarified by centrifugation at 13,000 × g for 20 min at 4 °C, and then diluted to 10 ml with ice-cold Pro Q Diamond wash buffer. The diluted lysates were applied 1 ml at a time to a washed and equilibrated Pro Q Diamond column containing 1.5 ml resin. The column was washed with 4.5 ml ice-cold wash buffer before elution with 1.75 ml Pro Q Diamond elution buffer. The eluates were concentrated, the buffer was exchanged with 3-[(3-Cholamidopropyl)dimethyl-ammonio]-1-propanesulfonate buffer, and enriched phosphoproteins were precipitated according to the Pro Q Diamond protocol. Phosphoproteins were resuspended in 60 μl of DIGE buffer (7 M urea, 2 m thiourea, 4% CHAPS, 10 mm Tris (pH 7)) plus 1 mm Na3VO4 and 1 mm NaF. Sample protein concentration was determined using the DC protein assay, and samples were stored unfrozen at 4 °C until electrophoresis.
Two-dimensional Gel Electrophoresis
Enriched phosphoprotein samples were diluted to the desired concentration with DIGE buffer without Tris, supplemented with 0.2% Biolytes (pH 3–10) and 50 mm DTT, and clarified by ultracentrifugation at 100,000 × g for 30 min. 200 μg/gel of sample proteins were separated by isoelectric focusing using ZOOM® (pH 4–7) strips according to the manufacturer's instructions. The isoelectric focusing strips were equilibrated in sodium dodecyl sulfate electrophoresis buffer and loaded onto 4–20% Tris-glycine ZOOM® gels for sodium dodecyl sulfate-polyacrylamide gel electrophoresis in the second dimension. The two-dimensional gels were stained with the fluorescent phosphoprotein-specific stain Pro Q Diamond according to the manufacturer's protocol. Spot matching and image analysis were performed using PD Quest software. Spots with a 2-fold or greater difference relative to the non-stimulated sample were selected for identification by tandem mass spectrometry. Tryptic peptides were analyzed using a 4800 MALDI-TOF-TOF mass spectrometer (Applied Biosystems, Carlsbad, CA). The top 20 ions were selected for MS/MS, and the data were queried in MASCOT (Matrix Science, Boston, MA) against the NCBI mammalian database. The protein modifications that were allowed included phosphorylation of serine, threonine, and tyrosine; oxidation of methionine; carbamidomethylation of cysteine; acetylation of lysine; and one missed tryptic cleavage.
Immunoprecipitation and Immunoblotting
To detect endogenous β-arrestin 2-binding proteins, HEK-AT1AR cells were transiently transfected with the FLAG epitope-tagged β-arrestin 2 expression plasmid. 10-cm plates of subconfluent cells were serum-starved for 2 h; stimulated with SII, AngII, or vehicle for 5 min; and lysed with 0.4 ml ice-cold glycerol lysis buffer (50 mm Hepes (pH 7.5), 0.5% Nonidet P-40, 250 mm NaCl, 2 mm EDTA, and 10% glycerol) containing 1 mm Na3VO4, 10 mm NaF, and protease inhibitor mixture. Lysates were clarified by centrifugation and FLAG-β-arrestin 2 was immunoprecipitated using 30 μl of a 50% slurry of EZ view red M2 anti-FLAG affinity resin with 2 h of tumbling at 4 °C. The affinity resin was washed twice in lysis buffer, and proteins were eluted in Laemmli sample buffer for resolution by sodium dodecyl sulfate polyacrylamide gel electrophoresis. Immunoprecipitation of I1PP2A, I2PP2A, and PGES3 was performed similarly, using rabbit polyclonal primary IgG and collecting immune complexes on 50 μl of a 50% slurry of protein-G plus/protein A-agarose. Immunoblots of phospho-ERK1/2, ERK1/2, GAPDH, I2PP2A, and β-arrestin 1/2 were performed using rabbit polyclonal IgG with HRP-conjugated goat anti-rabbit IgG secondary antibody. Immunoblots of FLAG-epitope, PP2A, I1PP2A, and PGES3 were performed using mouse monoclonal IgG with HRP-conjugated goat anti-mouse IgG secondary antibody. Immunoblots of phospho-Ser and phospho-Ser/Thr were performed using HRP-conjugated primary mouse monoclonal IgG.
PP2A Activity
Whole cell and β-arrestin 2-associated PP2A activity was assayed using the Millipore PP2A activity assay kit. For whole cell activity, appropriately stimulated subconfluent 10-cm plates of serum-starved HEK-AT1AR cells were washed in Hank's balanced salt solution, lysed with 500 μl of hypotonic lysis buffer (20 mm imidazole (pH 7.0), 2 mm ethylenediaminetetra-acetic acid, 2 mm ethylene glycol tetraacetic acid, 1 mm phenylmethylsulfonyl fluoride and protease inhibitor mixture) and sonicated. The PP2A catalytic subunit was immunoprecipitated as described, and phosphatase activity was determined following the kit manufacturer's protocol. β-Arrestin 2-associated PP2A activity was determined by assaying phosphatase activity in FLAG-β-arrestin 2 immunoprecipitates. To confirm the source of FLAG-β-arrestin 2-associated phosphatase activity, some assays were performed in the presence of 2 nm okadaic acid, a specific PP2A inhibitor.
Prostaglandin E2 Synthesis
Appropriately transfected HEK-AT1AR or primary rat aortic vascular smooth muscle cells in 6-well plates were serum starved for 2 h and then exposed to SII, AngII, or vehicle for 30 min in fresh serum starvation media. Prostaglandin synthase activity was determined by measuring PGE2 release into the culture medium using the Cayman prostaglandin E2 monoclonal enzyme-linked immunoassay kit according to the manufacturer's protocol.
RESULTS
Distinct Efficacy Profiles of SII and AngII
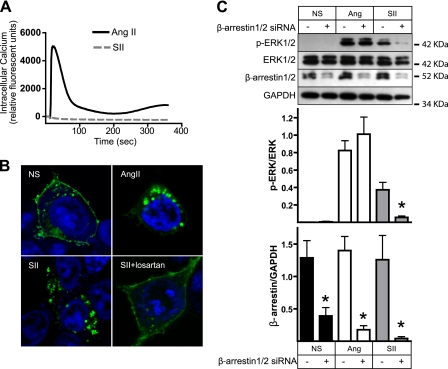
The AT1AR couples primarily to Gq/11 proteins that stimulate a PLCβ-inositol 3,4,5 trisphosphate pathway, leading to release of calcium from intracellular stores. As shown in Fig. 1A, AngII, but not SII, provokes a rapid increase in intracellular calcium in HEK-AT1AR cells. Despite its inability to produce Gq/11 signaling, SII has been shown to promote β-arrestin binding, AT1AR receptor internalization, and β-arrestin-dependent ERK1/2 activation (4–5). Fig. 1B demonstrates the effects of AngII and SII on the cellular distribution of AT1AR-GFP. Both ligands promote receptor internalization, as demonstrated by the loss of cell surface receptors after 30 min of agonist exposure. The effect of SII is reversed in the presence of the neutral AT1AR antagonist losartan, demonstrating that its actions are mediated through binding to the AT1AR orthosteric ligand binding site. As shown in Fig. 1C, both SII and AngII stimulate ERK1/2 phosphorylation in HEK-AT1AR cells. As reported previously (5, 11), this effect of SII is β-arrestin-dependent because it is blocked by siRNA silencing of β-arrestin 1/2 expression. In contrast, AngII-stimulated ERK1/2 activation was not significantly affected by β-arrestin silencing, likely because of the preservation of G-protein-dependent signaling pathways (11). Thus, AngII acts as a conventional GPCR agonist in our system, activating G-protein signaling and promoting β-arrestin-dependent receptor internalization, whereas SII behaves as an arrestin pathway-selective biased agonist, producing receptor internalization and β-arrestin-dependent signaling without detectable Gq/11 activation.
FIGURE 1.
Distinct efficacy of SII and AngII in HEK-AT1AR cells. A, effect of SII and AngII on intracellular calcium flux. Serum-deprived HEK-AT1AR cells were stimulated with SII or AngII, and time-dependent changes in intracellular calcium were measured using a fluorescence imaging plate reader. Representative traces are shown from one of three separate experiments that produced identical results. B, effect of SII and AngII on internalization of GFP-tagged AT1AR. Serum-deprived HEK cells transiently expressing AT1AR-GFP were treated for 30 min with vehicle (NS), AngII, or SII prior to fixation. Losartan (10 μm) was added to some dishes 10 min before SII stimulation. The cellular distribution of GFP was visualized by confocal fluorescence microscopy. Representative fields are shown from one of three separate experiments that produced identical results. C, effect of down-regulating β-arrestin expression on SII- and AngII-stimulated ERK1/2 phosphorylation. HEK-AT1AR cells were transfected with β-arrestin 1/2 targeted or control scrambled siRNA prior to serum starvation and stimulation for 5 min with SII or AngII. The upper panel depicts representative immunoblots of phospho- and total ERK1/2, β-arrestin 1/2, and GAPDH from one of three experiments. The upper bar graph depicts the mean ± S.E. of phospho-ERK1/2 normalized to the total ERK1/2. The lower bar graph depicts the mean ± S.E. of β-arrestin 1/2 expression normalized to GADPH. *, t test p < 0.05, less than cells treated with control siRNA (n = 3).
Gel-based Proteomic Characterization of SII-induced Protein Phosphorylation
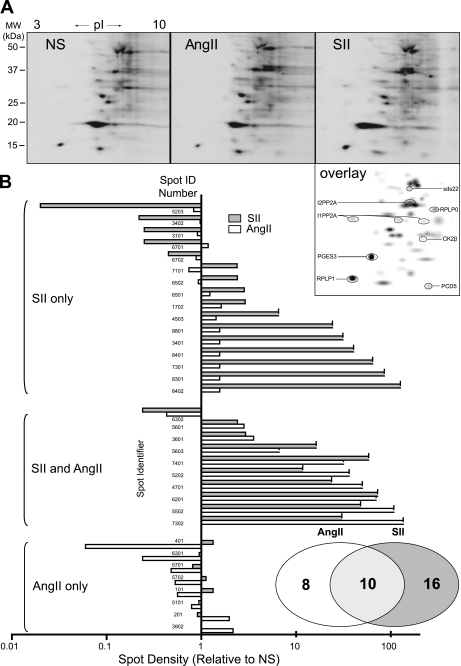
To gain insight into the G-protein-independent AT1AR signaling network, we performed a two-dimensional gel-based analysis of changes in protein phosphorylation upon SII stimulation. HEK-AT1AR cells were stimulated for 5 min with either AngII or SII, and whole cell phosphoproteins were enriched by column chromatography on Pro Q Diamond resin. Equal protein amounts of enriched sample were resolved by two-dimensional gel electrophoresis, and phosphoproteins were visualized using a Pro Q Diamond fluorescent stain with Sypro Ruby counterstaining for total protein. Fig. 2A depicts representative Pro Q Diamond-stained two-dimensional gels from vehicle-, SII-, and AngII-stimulated cells. Protein spots that differed in Pro Q Diamond staining intensity by a minimum of 2-fold were selected for identification by MALDI-TOF/TOF peptide mass fingerprinting. Of 63 individual phospho-protein spots matched across gels, 34 displayed a 2-fold or greater increase or decrease in intensity in response to either SII or AngII. Of these, 28 were identified by mass spectroscopy on the basis of at least six individual tryptic peptide fragments per protein. Fig. 2A (overlay) depicts the position of selected identified proteins on a gel overlay generated using PD Quest. Table 1 lists all protein identifications, along with the relative magnitude of change observed in SII- or AngII-treated samples.
FIGURE 2.
Two-dimensional gel-based screen of AT1AR-mediated protein phosphorylation. A, representative Pro Q Diamond-stained two-dimensional gels showing resolved phosphoproteins. Serum-deprived HEK-AT1AR cells were exposed to vehicle (NS), SII, or AngII for 5 min prior to phosphoprotein enrichment and two-dimensional electrophoresis as described. The overlay panel was derived from spot matching using PD Quest two-dimensional analysis software. The positions of selected phosphoproteins identified by MALDI-TOF/TOF mass spectroscopy are indicated. B, comparison of the phosphorylation profiles of SII and AngII-stimulated samples displaying the magnitude and direction of change in phosphoprotein staining spot intensity relative to NS samples. Note that a decrease in spot intensity may reflect either dephosphorylation or additional phosphorylation, leading to a shift in pI. A total of 34 spots changed a minimum of 2-fold relative to NS samples. The vertical axis lists individually resolved spots by arbitrary spot number assignment; the horizontal axis indicates normalized spot intensity on a log scale. The spots are grouped by SII-unique responses, changes common to SII and AngII, and AngII-unique responses. The Venn diagram displays distribution of spots in each category.
TABLE 1.
Phosphoprotein quantification
| Protein | Spot intensity |
|||||
|---|---|---|---|---|---|---|
| Non-stimulated | Ratio | Ang | Ratio | SII | Ratio | |
| ANP32B | 105.9 | 1 | 164.9 | 1.6 | 13363 | 126.18a |
| RBM8B | 105.9 | 1 | 7402.2 | 69.9a | 7692.9 | 72.64a |
| Prohibitin (PHB1) | 105.9 | 1 | 164.9 | 1.6 | 6807.6 | 64.28a |
| RPLP0 | 105.9 | 1 | 3352.2 | 31.7a | 6191.4 | 58.46a |
| Actin | 105.9 | 1 | 11,377.9 | 107.4a | 5076.5 | 47.93a |
| HBGB1 | 105.9 | 1 | 164.9 | 1.6 | 4277.6 | 40.39a |
| I1PP2A | 105.9 | 1 | 164.9 | 1.6 | 3307.2 | 31.23a |
| Calreticulin | 105.9 | 1 | 5299.7 | 50.0a | 2508.3 | 23.68a |
| CK2β | 105.9 | 1 | 3843.1 | 36.3a | 1243.3 | 11.74a |
| I2PP2A | 3266.2 | 1 | 4663.3 | 1.4 | 21,299.7 | 6.52a |
| Vimentin | 1112 | 1 | 3956.1 | 3.6a | 3257.1 | 2.93a |
| NAP1L1 | 1064.5 | 1 | 1719.8 | 1.6 | 3086.6 | 2.9a |
| HNRPC | 4838.1 | 1 | 5972 | 1.2 | 13,761.1 | 2.84a |
| Sds22 | 806.9 | 1 | 2276.9 | 2.8a | 1933.4 | 2.4a |
| β-Actin | 955.5 | 1 | 887.8 | 0.9 | 2291.4 | 2.4a |
| α-Tubulin 2 | 4858.6 | 1 | 3613.6 | 0.7 | 11,613.2 | 2.39a |
| RPLP1 | 15,492 | 1 | 8692.9 | 0.6 | 36094 | 2.33a |
| PDCD5 | 2407.1 | 1 | 6346.3 | 2.6a | 5577.7 | 2.32a |
| I1PP2A | 2629.4 | 1 | 164.9 | 0.1b | 3509.5 | 1.33 |
| HSP90α | 9082.6 | 1 | 19,608.4 | 2.2a | 9019 | 0.99 |
| PGES3 | 42,875 | 1 | 84,582.9 | 2.0a | 20,539.8 | 0.91 |
| β-Tubulin | 14,983 | 1 | 7147.2 | 0.5b | 12,146.2 | 0.81 |
| HSP70-2 | 3534 | 1 | 3125.3 | 0.9 | 1581.4 | 0.45b |
| I1PP2A | 13,332 | 1 | 12,096.4 | 0.9 | 3390.4 | 0.25b |
| HSP90β | 20,406 | 1 | 24,046.8 | 1.2 | 5054.3 | 0.25b |
| I1PP2A | 9354.5 | 1 | 4009.6 | 0.4b | 2256 | 0.24b |
| α-Tubulin | 7281.1 | 1 | 7041.3 | 1.0 | 1628.2 | 0.22b |
| NDUFS3 | 7613.7 | 1 | 6331.1 | 0.8 | 117 | 0.02b |
a Spots with >2-fold increased intensity relative to non-stimulated.
b Spots with >2-fold decreased intensity relative to non-stimulated.
Two-dimensional gel electrophoresis offers only limited coverage of transmembrane proteins, proteins with high molecular weight, or proteins with a highly basic pI. Despite this limited window, comparison of the global changes in protein phosphorylation revealed surprising differences between AngII and SII. As shown in Fig. 2B, only 10 of 34 phosphoprotein spots changed in response to both AngII and SII, whereas an additional 16 appeared to be responsive only to SII. It is unclear whether this is due to limitations in the detection threshold of our assay or to qualitative differences in signaling by SII-bound AT1AR. Nonetheless, it is clear that SII elicited a robust protein phosphorylation response despite its failure to activate G-protein signaling.
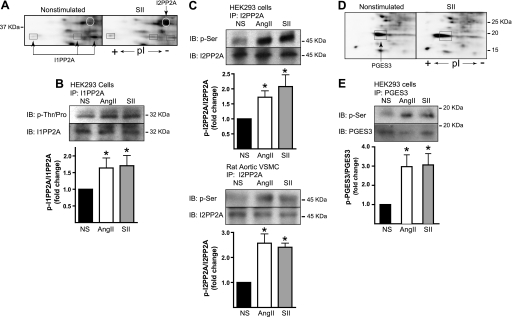
The list of 28 identified phosphostained proteins featured two inhibitors of the Ser/Thr phosphatase PP2A, I1PP2A, and I2PP2A. I1PP2A appeared as four discrete spots that shifted to a more acidic pI with stimulation, consistent with additional phosphorylation. I2PP2A appeared as a robust de novo phosphorylation response to SII (Fig. 3A). The activities of both proteins are reportedly regulated by phosphorylation. Phosphorylation of I1PP2A on Ser-9 disrupts its interaction with the PP2A catalytic subunit resulting in an increase in PP2A activity (14), whereas phosphorylation of I2PP2A increases its interaction with PP2A and the inhibition of PP2A activity (15). As shown in Fig. 3, B and C, immunoprecipitates of endogenous I1PP2A and I2PP2A from HEK-AT1AR cells demonstrated a net increase in Ser/Thr phosphorylation upon stimulation with either SII or AngII. SII-stimulated phosphorylation of I2PP2A was also detected in primary aortic vascular smooth muscle cells, indicating that the response occurs at endogenous levels of AT1AR expression (Fig. 3C).
FIGURE 3.
Validation of SII- and AngII-stimulated phosphorylation of I1PP2A, I2PP2A, and PGES3. A, representative Pro Q Diamond-stained two-dimensional gel regions showing SII-induced changes in the appearance of spots identified as I1PP2A and I2PP2A. I2PP2A is present as a single spot that appears upon stimulation, whereas I1PP2A appears as three spots of similar mass but shifting pI consistent with phosphorylation of additional sites. B, phospho-amino acid-specific immunoblot (IB) of immunoprecipitated (IP) I1PP2A. C, phospho-amino acid-specific immunoblots of endogenous I2PP2A immunoprecipitated from HEK293 cells (upper panel) and vascular smooth muscle cells (lower panel). In B and C, serum-deprived HEK-AT1AR or primary rat aortic vascular smooth muscle cells (VSMC) were stimulated for 5 min with SII or AngII before lysis and immunoprecipitation as described. Representative immunoblots are depicted above bar graphs indicating the fold change in phosphorylation relative to vehicle treated cells (NS). The intensity of the phospho-specific immunoblot was normalized to the total amount of I1PP2A or I2PP2A in each immunoprecipitate. *, t test p < 0.05, greater than NS (n = 5 for I1PP2A, n = 3–6 for I2PP2A). Pro Q Diamond-stained two-dimensional gel regions showing the SII-induced change in the appearance of the spot identified as PGES3. PGES3 appears as a single phosphoprotein that undergoes an acidic pI shift consistent with phosphorylation of additional sites. E, phospho-Ser-specific immunoblot of immunoprecipitated PGES3. HEK-AT1AR cells were stimulated for 5 min with SII or AngII before lysis and immunoprecipitation. Representative immunoblots are depicted above bar graphs indicating the fold change in phosphorylation relative to NS cells. The intensity of the phospho-specific immunoblot was normalized to the total amount of PGES3 in each immunoprecipitate. *, t test p < 0.05, greater than NS (n = 5).
Another phosphostained protein that exhibited an acidic isoelectric shift upon SII stimulation was identified by mass spectroscopy as the soluble prostaglandin synthase isoform, PGES3 (Fig. 3D). PGES3 mediates PGE2 synthesis by coupling to the constitutively expressed cyclooxygenase isoform, Cox1 (16), and is reportedly activated by Ser phosphorylation (17). As shown in Fig. 3E, immunoprecipitates of endogenous PGES3 showed a net increase in Ser phosphorylation in response to a 5-min SII or AngII stimulation.
G-protein-independent Regulation of β-Arrestin 2-associated PP2A Activity
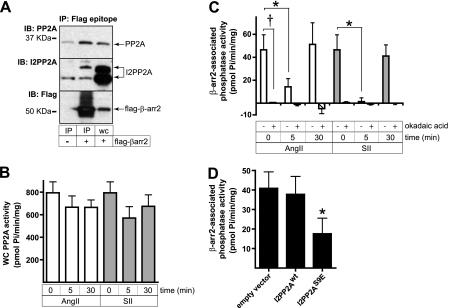
Recent data suggests that β-arrestin 2 controls PP2A activity in dopamine D2 receptor-rich striatum via a protein complex containing the catalytic subunit of PP2A, Akt, and GSK3β (18). I2PP2A has also been implicated in GPCR signal regulation, as it reportedly binds the muscarinic M3 receptor and attenuates calcium signaling (19). We therefore tested whether the AT1AR-mediated phosphorylation of I2PP2A we observed in response to SII could be involved in the regulation of a β-arrestin-associated pool of PP2A. As shown in Fig. 4A, endogenous PP2A and I2PP2A coprecipitated with FLAG-epitope-tagged β-arrestin 2 transiently expressed in HEK-AT1AR cells. Agonist treatmen had no significant effect on the abundance of either protein in the immunoprecipitate, suggesting that the β-arrestin 2-I2PP2A-PP2A complex was preformed and did not assemble or dissociate upon stimulation (data not shown). Although SII and AngII had no significant effect on whole cell PP2A catalytic activity (Fig. 4B), both agonists strongly inhibited β-arrestin 2-associated PP2A activity. As shown in Fig. 4C, the effect was transient, with near complete inhibition of phosphatase activity after 5 min and recovery to baseline levels within 30 min of stimulation. The identity of the β-arrestin 2-bound phosphatase activity was confirmed as PP2A on the basis of its sensitivity to the PP2A-specific inhibitor, okadaic acid (20). Because phosphorylation of I2PP2A leads to inhibition of PP2A (14), we tested whether expression of a phosphomimetic mutant of I2PP2A, in which Ser-9 was mutated to Glu, was sufficient to suppress β-arrestin 2-bound PP2A activity. As shown in Fig. 4D, transient overexpression of I2PP2A-S9E, but not wild-type I2PP2A, inhibited β-arrestin 2-associated PP2A activity. These data suggest that phosphorylation of I2PP2A in response to SII is a plausible mechanism for regulating β-arrestin 2-associated PP2A activity.
FIGURE 4.
Transient inhibition of β-arrestin 2-bound PP2A in response to SII and AngII treatment. A, coimmunoprecipitation (IP) of the endogenous PP2A catalytic subunit and I2PP2A with FLAG-tagged β-arrestin 2. HEK-AT1AR cells were transiently transfected with either empty vector or FLAG-β-arrestin 2 and FLAG immunoprecipitates performed as described. The presence of coprecipitating PP2A and I2PP2A was demonstrated by immunoblotting (IB). A whole cell lysate lane (WC) is shown for comparison. B, effect of SII and AngII treatment on whole cell PP2A activity. HEK-AT1AR cells were stimulated with SII or AngII for the indicated times, after which the PP2A catalytic subunit was immunoprecipitated and PP2A activity determined as described. Neither ligand produced a significant change in global PP2A activity (n = 5). C, transient inhibition of FLAG-β-arrestin 2-bound PP2A following SII or AngII stimulation. HEK-AT1AR cells transiently expressing FLAG-tagged β-arrestin 2 were stimulated for the indicated times with SII or AngII, after which FLAG-β-arrestin 2-associated Ser phosphatase activity was determined as described. Background phosphatase activity (average 24 pmol Pi/min/mg protein) was measured in FLAG immunoprecipitates performed on vector-transfected control cells and subtracted from that measured in FLAG-β-arrestin 2 immunoprecipitates. Parallel samples were incubated for 10 min with okadaic acid (2 nm) after immunoprecipitation to determine the extent to which PP2A contributed to FLAG-β-arrestin 2-bound phosphatase activity. Data shown are the mean ± S.E. of FLAG-β-arrestin 2-bound phosphatase activity determined under each condition. *, t test p < 0.05, less than NS (n = 5). †, p < 0.05, okadaic acid treated less than untreated (n = 5). D, inhibition of FLAG-β-arrestin 2-bound phosphatase activity by the phosphomimetic I2PP2A-S9E mutant. HEK-AT1AR cells were transiently transfected with plasmids encoding FLAG-tagged β-arrestin 2 and either empty vector, wild type I2PP2A, or I2PP2A-S9E. FLAG immunoprecipitates were isolated, and β-arrestin 2-associated PP2A activity was determined as described. The bar graph depicts mean ± S.E. of phosphatase activity measured under each condition. *, t test p < 0.05, less than NS (n = 3).
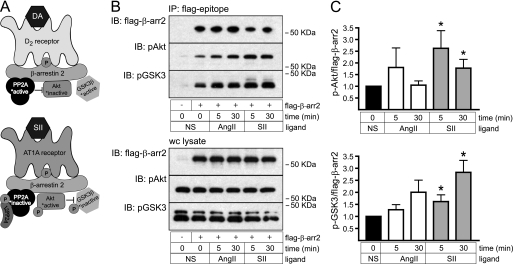
In striatum, dopamine D2 receptors have been reported to both positively (21) and negatively (18) regulate the prosurvival kinase Akt. Although the mechanism of D2 receptor-mediated PI3K-independent Akt activation is not understood, Akt inhibition involves the assembly of a β-arrestin 2-PP2A-Akt-GSK3β complex. Akt is activated by phosphorylation of Thr-308, a site that is phosphorylated by proline-directed kinase 1 and dephosphorylated by PP2A (22). Within the complex, tonic dephosphorylation of Thr-308 by PP2A keeps Akt inactive. Because Akt inactivates GSK3β by phosphorylating it on Ser-9, complex assembly maintains GSK3β in its active dephosphorylated state (Fig. 5A, top panel). Our finding that SII produces transient inhibition of PP2A suggests a mechanism for receptor-mediated activation of β-arrestin 2-bound Akt (Fig. 5A, bottom panel). As shown in Fig. 5, B and C, SII treatment increases phosphorylation of Akt Thr-308 in β-arrestin 2 immunoprecipitates, leading to Akt activation as reflected in a corresponding increase in GSK3β Ser-9 phosphorylation.
FIGURE 5.
Effect of SII and AngII on β-arrestin 2-bound Akt and GSK3β phosphorylation. A, schematic depicting the previously reported model for steady-state suppression of Akt activity in dopamine D2 receptor-β-arrestin 2 signalsomes (18) compared with the proposed model for Akt activation resulting from transient inhibition of signalsome PP2A activity (see text for explanation). B, representative immunoblots (IB) showing SII-induced phosphorylation of Akt-Thr-308 and its downstream target GSK3β-Ser-9 measured in FLAG-β-arrestin 2 immunoprecipitates (IP) from HEK-AT1AR cells isolated after stimulation with SII or AngII for the indicated times. For comparison, the lack of detectable ligand effect on the total cellular pools of phospho-Akt-Thr-308 and phospho-GSK3β-Ser-9 is shown in the lower panel. C, bar graphs depicting phospho-Akt-Thr-308 and phospho-GSK3β-Ser-9 normalized to the amount of FLAG-β-arrestin 2 in the immunoprecipitate. *, t test p < 0.05, greater than NS (n = 8 for Akt, n = 5 for GSK3β).
Arrestin-dependent Stimulation of PGE2 Synthesis
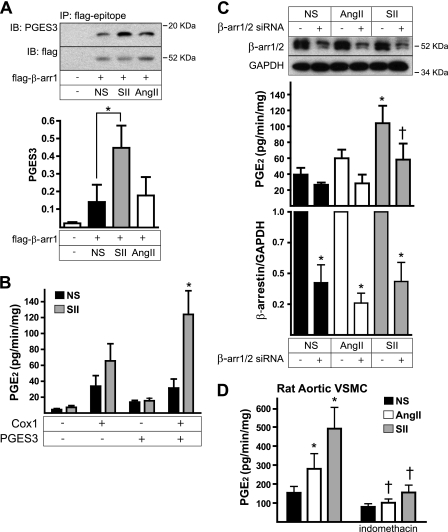
Our finding that SII and AngII stimulate phosphorylation of PGES3, a cytosolic prostaglandin synthase isoform that is coupled to the constitutively expressed Cox1 isoform (16), suggested a mechanism for rapid-onset effects on prostaglandin synthesis. As shown in Fig. 6A, endogenous PGES3 was detectable in FLAG-β-arrestin 1 immunoprecipitates, and the association was significantly increased by SII treatment, suggesting that SII promotes the assembly of a β-arrestin-PGES3 complex. HEK293 cells lack the components necessary to produce PGE2 endogenously, but prostaglandin production can be reconstituted in an HEK293 cell background by overexpressing both Cox 1 and PGES3 (16). As shown in Fig. 6B, neither Cox1 nor PGES3, expressed alone, was sufficient to confer SII-stimulated PGE2 synthesis on HEK-AT1AR cells. However, expressing both components together reconstituted SII-stimulated PGE2 synthesis, suggesting that SII is able to activate the Cox1-PGES3 pathway. To determine whether this response was β-arrestin-dependent, we repeated the experiment using siRNA to down-regulate β-arrestin expression. As shown in Fig. 6C, β-arrestin 1/2 knockdown significantly reduced SII-stimulated PGE2 synthesis in HEK-AT1AR cells with reconstituted Cox1-PGES3-dependent PGE2 synthesis. As shown in Fig. 6D, SII stimulated indomethacin-sensitive PGE2 production in non-transfected primary rat aortic vascular smooth muscle cells, indicating that G protein-independent acute regulation of PGE2 synthesis can occur under physiologic conditions. These data suggest that regulation of PGES3 phosphorylation is a plausible mechanism by which SII mediates arrestin-dependent activation of PGE2 synthesis.
FIGURE 6.
β-Arrestin-dependent activation of Cox1-PGES3 dependent PGE2 synthesis by SII. A, SII stimulation increases the association of endogenous PGES3 with FLAG-β-arrestin 1. HEK-AT1AR cells transiently expressing FLAG epitope-tagged β-arrestin 1 were stimulated for 5 min with SII or AngII, after which FLAG immunoprecipitates (IP) were probed for the presence of coimmunoprecipitating PGES3. A representative immunoblot (IB) is shown above a bar graph depicting the amount of PGES3 protein normalized to the amount of FLAG-β-arrestin 1 in the immunoprecipitate. *, t test p < 0.05, greater than NS (n = 3). B, reconstitution of SII-induced PGE2 synthesis in HEK-AT1AR cells. Cells were transfected with plasmids encoding Cox 1, PGES3, or both and then stimulated with SII for 30 min before enzyme-linked immunoassay of PGE2 released into the culture medium. Data shown are the mean ± S.E. of the rate of PGE2 release determined under each condition. *, p < 0.05, greater than NS (n = 7). C, effect of silencing β-arrestin 1/2 expression on SII-induced PGE2 synthesis. Down-regulation of β-arrestin 1/2 expression using siRNA was performed in HEK-AT1AR transiently coexpressing Cox1 and PGES3, after which the effects of SII and AngII on PGE2 synthesis was determined. The upper panel immunoblot and lower panel bar graph show the effect of siRNA treatment on endogenous β-arrestin 1/2 expression. *, t test p < 0.05, less than control siRNA-transfected cells (n = 4). The upper panel bar graph shows the effect of siRNA treatment on SII and AngII stimulated PGE2 release into the culture medium. *, t test p < 0.05, greater than NS; †, p < 0.05, less than control siRNA (n = 4). D, effect of SII and AngII on PGE2 synthesis by primary rat aortic vascular smooth muscle cells (VSMC). Some wells were preincubated for 30 min with the Cox 1/2 inhibitor indomethacin (10 μm) prior to stimulation. *, p < 0.05, greater than NS; †, p < 0.05, less than without indomethacin (n = 3).
DISCUSSION
Although the existence of β-arrestin-dependent signaling has been recognized for over a decade, comparatively little is known about the manner in which these signals affect cellular metabolism (9, 10). To explore the broader role of β-arrestins in AT1AR signaling, we employed a gel-based proteomic screen to identify changes in protein phosphorylation occurring in response to SII, an arrestin pathway-selective biased AT1AR agonist (4, 5). Despite the limited nature of our proteomic survey, our results demonstrate that the AT1AR regulates a robust G-protein-independent signaling network. In addition, we report two previously unknown arrestin-mediated signaling events: PP2A-dependent activation of Akt in a β-arrestin 2-PP2A-Akt-GSK3β complex, and rapid Cox1-dependent stimulation of prostaglandin synthesis via a β-arrestin 1-PGES3 complex.
Our two-dimensional gel separation of enriched phosphoprotein samples identified several proteins whose phosphorylation state changes upon AT1AR activation. Looking at the overall response pattern, one of the more striking observations is the degree to which SII appears to cause changes in protein phosphorylation that were not observed in response to AngII. This might simply reflect limited assay sensitivity, i.e. differences in signal strength rather than qualitative differences in signal transduction. Conversely, there is no a priori reason that the AT1AR conformations adopted upon binding SII would necessarily couple the receptor only to a subset of the effectors engaged upon binding AngII. Notably, a recent proteomic comparison of SII- and AngII-induced protein phosphorylation performed using a non-gel-based methodology capable of detecting substantially larger numbers of phosphopeptides also found significant non-overlap between the AngII- and SII-induced phosphoproteomes (23).
The Pro Q resin used for phosphoprotein enrichment in our study is a proprietary product, but on the basis of published patents is likely to be an immobilized and chelated gallium resin (24). Like most metal resins used to enrich phosphoproteins, protein contaminants are common. Proteins containing highly acidic domains may coelute with phosphoproteins (25). However, we are unaware of any publications that suggest molecular weight or other biases ascribed to phosphoprotein enrichment by immobilized gallium. The issue of non-phosphorylated contaminants is a concern, but the use of a phosphoprotein-specific stain to detect phosphoproteins in the two-dimensional gel minimized the likelihood of detecting non-phosphorylated proteins. It is possible that proteins with multiple phosphorylated residues are retained with greater efficiency than those that contain a single phosphorylated residue. This bias is not likely to affect the results presented here unless highly abundant, high molecular weight proteins that could not be resolved by two-dimensional gel electrophoresis were to shift from a multiple to a singly or non-phosphorylated state, which would artificially elevate the fractional abundance of the visualized proteins in the sample. However, total protein abundance was normalized within each gel by correcting for total spot volume, which should negate any bias resulting from differential phosphorylation of the unseen high molecular weight fraction. The major limitation of two-dimensional gel studies arises from the limited capacity of high molecular weight, membrane, and basic proteins to enter the second dimension. Therefore, neutral and acidic proteins of molecular weights between 10 and 100 kDa are detected most often. In addition, the phosphoprotein stain has a limit of detection of 1–16 ng, meaning only relatively high-abundance proteins are detectable.
Several lines of evidence suggest that regulation of phosphatidylinositol 3-kinase/Akt signaling is a key component of arrestin-mediated GPCR signaling (10). A recent analysis of the SII-regulated phosphoproteome identified several kinases downstream of Akt, among a total of 38 protein kinases, that were regulated in an arrestin-dependent manner (26). Functionally, β-arrestins have been implicated in the control of PP2A-Akt-GSK3β signaling by dopamine D2 receptors in striatum (18). PP2A is a ubiquitously expressed Ser/Thr protein phosphatase with broad substrate specificity. Its activity is constrained by binding to targeting subunits that confer spatial localization and limit access to substrates. One such PP2A regulatory complex appears to be a β-arrestin 2-PP2A-Akt-GSK3β complex originally isolated from brain. Within the complex, PP2A acts as a negative regulator of Akt by maintaining the Thr-308 site in the dephosphorylated state. In β-arrestin 2 null mice, the loss of this tonic inhibition leads to increased Akt activity, Akt-dependent inhibition of GSK3β, and β-catenin stabilization (27).
Although the steady-state effect of β-arrestin 2-PP2A-Akt-GSK3β complex assembly is to dampen global Akt signaling, our finding that two inhibitors of PP2A, I1PP2A and I2PP2A, are phosphorylated in response to SII points to a possible mechanism for arrestin-dependent Akt activation within the signalsome complex. I1PP2A and I2PP2A are known regulators of cytosolic and nuclear PP2A. I1PP2A is a tumor suppressor. In the dephosphorylated state it binds PP2A, inhibiting its activity. Phosphorylation of I1PP2A disrupts the interaction, resulting in an increase in PP2A activity (15). Conversely, I2PP2A, also known as the SET nuclear oncogene, binds and inhibits PP2A when phosphorylated on Ser-9 (14). Within the nucleus, I1PP2A and I2PP2A work together as components of the inhibitor of histone acetyl transferase complex (28). In the cytosol, I2PP2A has been reported to move to the plasma membrane and stimulate Rac1-dependent cell migration (15) and to bind the M3 muscarinic receptor and inhibit calcium signaling (19). Although we did not detect endogenous I1PP2A bound to β-arrestin 2 (data not shown), our findings are consistent with a model wherein phosphorylation of I2PP2A regulates the β-arrestin-bound PP2A pool. Specifically, these findings are that 1) I2PP2A is present along with PP2A, Akt, and GSK3β in FLAG-β-arrestin 2 immunoprecipitates; 2) SII-induced phosphorylation of I2PP2A correlates with a transient inhibition of PP2A in the complex; and 3) expression of a phospho-mimetic S9E mutant of I2PP2A reproduces the effect of SII treatment. The corresponding increases we observe in Akt Thr-308 and GSK3β Ser-9 phosphorylation within the complex demonstrate that inhibition of PP2A activates Akt, which in turn phosphorylates its downstream targets. Interestingly, I2PP2A has two splice variants, of which the species identified in our proteomic dataset, isoform 2, is reportedly antiapoptotic (29).
Our proteomic analysis also identified SII-induced phosphorylation of three proteins involved in prostaglandin synthesis: PGES3, heat shock protein 90 (HSP90), and casein kinase II (CK2) (17). Unlike PGES1 and PGES2, which associate with microsomes, PGES3 is cytosolic. It was originally discovered as a cochaperone of HSP90 that is required for HSP90 chaperone activity (16, 30). Conversely, HSP90 is required for PGES3 activity (16). PGES3 is activated by CK2 phosphorylation (17) and selectively couples to Cox1 (14). We have found that SII stimulates PGES3 phosphorylation and Cox1-dependent PGE2 synthesis. In addition, SII promotes binding of PGES3 to β-arrestin 1, and silencing β-arrestin expression inhibits SII-induced PGE2 synthesis, suggesting that β-arrestins play a role in rapid AT1AR regulation of prostaglandin synthesis. The recent report that cytosolic phospholipase A2, the rate-limiting enzyme in PGE2 biosynthesis, is part of an inducible β-arrestin complex formed in response to stimulation of the heptahelical receptor GPR109A, similarly supports a role for a arrestin-based signalsomes in regulating prostanoid metabolism (31).
Angiotensin II has long been known to stimulate PGE2 release in vascular tissues and the kidney (32–34). Cox1, which couples to PGES3, mediates immediate PGE2 production (14), whereas the inducible Cox2 enzyme mediates long-term PGE2 production and inflammation. Cox1- and Cox2-dependent PGE2 signaling reportedly have opposite effects in the kidney, with Cox1 increasing and Cox2 opposing the AngII-induced pressor response (34). We have previously reported that SII induces Cox2 expression in rat vascular smooth muscle cells (12), suggesting that G-protein-independent signals may affect both short-term prostaglandin synthesis through a Cox1-PGES3 pathway and long-term prostaglandin synthesis through transcriptional regulation of Cox2 expression.
Along with other recent proteomic surveys of SII action (23, 26), our results support the existence of a robust and still largely unexplored G-protein-independent signaling network regulated by the AT1AR. Although unrecognized until just over a decade ago (35), it is now apparent that many GPCR signals arise from “coupling” to β-arrestins independent of heterotrimeric G-protein activation. The fact that biased GPCR agonists, like SII, can dissociate G-protein- and arrestin-dependent signaling raises the prospect of functionally selective drugs whose biological activity is distinct from that of conventional agonist or antagonist ligands. At the same time, limitations in our current understanding of the complexity of arrestin-dependent signaling makes it difficult to predict whether the selective activation of arrestin signaling will produce a therapeutic benefit, and provide a strong impetus for further proteomic and genomic studies of biased GPCR efficacy.
Acknowledgments
We thank Ms. Alison Bland for excellent technical assistance.
This work was supported, in whole or in part, by National Institutes of Health Grants R01 DK055524 (to L. M. L.) and T32 HL07260 (to R. T. K.). This work was also supported by the Research Service of the Ralph H. Johnson Veterans Affairs Medical Center (to L. M. L. and M. G. J.) and the Nephcure Young Investigator and Department of Veterans Affairs Career Development Awards (to M. G. J.). Imaging facilities for this research were supported, in part, by Cancer Center Support Grant P30 CA138313 (to the Hollings Cancer Center, Medical University of South Carolina).
- AngII
- angiotensin II
- SII
- [Sar1,Ile4,Ile8]AngII
- AT1AR
- angiotensin II receptor type 1A
- GPCR
- G protein-coupled receptor
- PGES3
- prostaglandin E synthase type 3
- GSK3β
- glycogen synthase kinase 3β
- PP2A
- protein phosphatase 2A
- Cox
- cyclooxygenase
- PGE2
- prostaglandin E2
- I1PP2A
- inhibitor 1 of PP2A
- I2PP2A
- inhibitor 2 of PP2A
- CHAPS
- 3-[(3-Cholamidopropyl)dimethyl-ammonio]-1-propanesulfonate.
REFERENCES
- 1. Eguchi S., Numaguchi K., Iwasaki H., Matsumoto T., Yamakawa T., Utsunomiya H., Motley E. D., Kawakatsu H., Owada K. M., Hirata Y., Marumo F., Inagami T. (1998) J. Biol. Chem. 273, 8890–8896 [DOI] [PubMed] [Google Scholar]
- 2. Ali M. S., Sayeski P. P., Dirksen L. B., Hayzer D. J., Marrero M. B., Bernstein K. E. (1997) J. Biol. Chem. 272, 23382–23388 [DOI] [PubMed] [Google Scholar]
- 3. Luttrell L. M., Roudabush F. L., Choy E. W., Miller W. E., Field M. E., Pierce K. L., Lefkowitz R. J. (2001) Proc. Natl. Acad. Sci. U.S.A. 98, 2449–2454 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Holloway A. C., Qian H., Pipolo L., Ziogas J., Miura S., Karnik S., Southwell B. R., Lew M. J., Thomas W. G. (2002) Mol. Pharmacol. 61, 768–777 [DOI] [PubMed] [Google Scholar]
- 5. Wei H., Ahn S., Shenoy S. K., Karnik S. S., Hunyady L., Luttrell L. M., Lefkowitz R. J. (2003) Proc. Natl. Acad. Sci. U.S.A. 100, 10782–10787 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Rajagopal K., Whalen E. J., Violin J. D., Stiber J. A., Rosenberg P. B., Premont R. T., Coffman T. M., Rockman H. A., Lefkowitz R. J. (2006) Proc. Natl. Acad. Sci. U.S.A. 103, 16284–16289 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. DeWire S. M., Kim J., Whalen E. J., Ahn S., Chen M., Lefkowitz R. J. (2008) J. Biol. Chem. 283, 10611–10620 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Lymperopoulos A., Rengo G., Zincarelli C., Kim J., Soltys S., Koch W. J. (2009) Proc. Natl. Acad. Sci. U.S.A. 106, 5825–5830 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Kendall R. T., Luttrell L. M. (2009) Cell Mol. Life Sci. 66, 2953–2973 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Luttrell L. M., Gesty-Palmer D. (2010) Pharmacol. Rev. 62, 305–330 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Lee M. H., El-Shewy H. M., Luttrell D. K., Luttrell L. M. (2008) J. Biol. Chem. 283, 2088–2097 [DOI] [PubMed] [Google Scholar]
- 12. Morinelli T. A., Kendall R. T., Luttrell L. M., Walker L. P., Ullian M. E. (2009) J. Pharmacol. Exp. Ther. 330, 118–124 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Barak L. S., Ferguson S. S., Zhang J., Caron M. G. (1997) J. Biol. Chem. 272, 27497–27500 [DOI] [PubMed] [Google Scholar]
- 14. Yu L. G., Packman L. C., Weldon M., Hamlett J., Rhodes J. M. (2004) J. Biol. Chem. 279, 41377–41383 [DOI] [PubMed] [Google Scholar]
- 15. ten Klooster J. P., Leeuwen I., Scheres N., Anthony E. C., Hordijk P. L. (2007) EMBO J. 26, 336–345 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Tanioka T., Nakatani Y., Semmyo N., Murakami M., Kudo I. (2000) J. Biol. Chem. 275, 32775–32782 [DOI] [PubMed] [Google Scholar]
- 17. Kobayashi T., Nakatani Y., Tanioka T., Tsujimoto M., Nakajo S., Nakaya K., Murakami M., Kudo I. (2004) Biochem. J. 381, 59–69 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Beaulieu J. M., Sotnikova T. D., Marion S., Lefkowitz R. J., Gainetdinov R. R., Caron M. G. (2005) Cell 122, 261–273 [DOI] [PubMed] [Google Scholar]
- 19. Simon V., Guidry J., Gettys T. W., Tobin A. B., Lanier S. M. (2006) J. Biol. Chem. 281, 40310–40320 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Ishihara H., Martin B. L., Brautigan D. L., Karaki H., Ozaki H., Kato Y., Fusetani N., Watabe S., Hashimoto K., Uemura D., et al. (1989) Biochem. Biophys. Res. Commun. 159, 871–877 [DOI] [PubMed] [Google Scholar]
- 21. Brami-Cherrier K., Valjent E., Garcia M., Pagès C., Hipskind R. A., Caboche J. (2002) J. Neurosci. 22, 8911–8921 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Liao Y., Hung M. C. (2010) Am. J. Transl. Res. 2, 19–42 [PMC free article] [PubMed] [Google Scholar]
- 23. Christensen G. L., Kelstrup C. D., Lyngsø C., Sarwar U., Bøgebo R., Sheikh S. P., Gammeltoft S., Olsen J. V., Hansen J. L. (2010) Mol. Cell Proteomics. 9, 1540–1553 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Schilling M., Knapp D. R. (2008) J. Proteome Res. 7, 4164–4172 [DOI] [PubMed] [Google Scholar]
- 25. Chicz R. M., Regnier F. E. (1990) Methods Enzymol. 182, 392–421 [DOI] [PubMed] [Google Scholar]
- 26. Xiao K., Sun J., Kim J., Rajagopal S., Zhai B., Villén J., Haas W., Kovacs J. J., Shukla A. K., Hara M. R., Hernandez M., Lachmann A., Zhao S., Lin Y., Cheng Y., Mizuno K., Ma'ayan A., Gygi S. P., Lefkowitz R. J. (2010) Proc. Natl. Acad. Sci. U.S.A. 107, 15299–15304 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Beaulieu J. M., Marion S., Rodriguiz R. M., Medvedev I. O., Sotnikova T. D., Ghisi V., Wetsel W. C., Lefkowitz R. J., Gainetdinov R. R., Caron M. G. (2008) Cell 132, 125–136 [DOI] [PubMed] [Google Scholar]
- 28. Kutney S. N., Hong R., Macfarlan T., Chakravarti D. (2004) J. Biol. Chem. 279, 30850–30855 [DOI] [PubMed] [Google Scholar]
- 29. Fan Z., Beresford P. J., Oh D. Y., Zhang D., Lieberman J. (2003) Cell 112, 659–672 [DOI] [PubMed] [Google Scholar]
- 30. Johnson J. L., Beito T. G., Krco C. J., Toft D. O. (1994) Mol. Cell. Biol. 14, 1956–1963 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Walters R. W., Shukla A. K., Kovacs J. J., Violin J. D., DeWire S. M., Lam C. M., Chen J. R., Muehlbauer M. J., Whalen E. J., Lefkowitz R. J. (2009) J. Clin. Invest. 119, 1312–1321 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Gimbrone M. A., Jr., Alexander R. W. (1975) Science 189, 219–220 [DOI] [PubMed] [Google Scholar]
- 33. Alexander R. W., Gimbrone M. A., Jr. (1976) Proc. Natl. Acad. Sci. U.S.A. 73, 1617–1620 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Qi Z., Hao C. M., Langenbach R. I., Breyer R. M., Redha R., Morrow J. D., Breyer M. D. (2002) J. Clin. Invest. 110, 61–69 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Luttrell L. M., Ferguson S. S., Daaka Y., Miller W. E., Maudsley S., Della Rocca G. J., Lin F., Kawakatsu H., Owada K., Luttrell D. K., Caron M. G., Lefkowitz R. J. (1999) Science 283, 655–661 [DOI] [PubMed] [Google Scholar]