Abstract
In view of the likely role of H1-H1 interactions in the stabilization of chromatin higher order structure, we have asked whether interactions can occur between the globular domains of the histone molecules. We have studied the properties of the isolated globular domains of H1 and the variant H5 (GH1 and GH5) and we have shown (by sedimentation analysis, electron microscopy, chemical cross-linking and nucleoprotein gel electrophoresis) that although GH1 shows no, and GH5 little if any, tendency to self-associate in dilute solution, they bind highly cooperatively to DNA. The resulting complexes appear to contain essentially continuous arrays of globular domains bridging 'tramlines' of DNA, similar to those formed with intact H1, presumably reflecting the ability of the globular domain to bind more than one DNA segment, as it is likely to do in the nucleosome. Additional (thicker) complexes are also formed with GH5, probably resulting from association of the primary complexes, possibly with binding of additional GH5. The highly cooperative nature of the binding, in close apposition, of GH1 and GH5 to DNA is fully compatible with the involvement of interactions between the globular domains of H1 and its variants in chromatin folding.
Full text
PDF






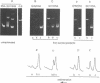
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Allan J., Hartman P. G., Crane-Robinson C., Aviles F. X. The structure of histone H1 and its location in chromatin. Nature. 1980 Dec 25;288(5792):675–679. doi: 10.1038/288675a0. [DOI] [PubMed] [Google Scholar]
- Aviles F. J., Chapman G. E., Kneale G. G., Crane-Robinson C., Bradbury E. M. The conformation of histone H5. Isolation and characterisation of the globular segment. Eur J Biochem. 1978 Aug 1;88(2):363–371. doi: 10.1111/j.1432-1033.1978.tb12457.x. [DOI] [PubMed] [Google Scholar]
- Bates D. L., Butler P. J., Pearson E. C., Thomas J. O. Stability of the higher-order structure of chicken-erythrocyte chromatin in solution. Eur J Biochem. 1981 Oct;119(3):469–476. doi: 10.1111/j.1432-1033.1981.tb05631.x. [DOI] [PubMed] [Google Scholar]
- Briand G., Kmiecik D., Sautiere P., Wouters D., Borie-Loy O., Biserte G., Mazen A., Champagne M. Chicken erythrocyte histone H5. IV. Sequence of the carboxy-termined half of the molecule (96 residues) and complete sequence. FEBS Lett. 1980 Apr 7;112(2):147–151. doi: 10.1016/0014-5793(80)80167-0. [DOI] [PubMed] [Google Scholar]
- Butler P. J., Thomas J. O. Changes in chromatin folding in solution. J Mol Biol. 1980 Jul 15;140(4):505–529. doi: 10.1016/0022-2836(80)90268-5. [DOI] [PubMed] [Google Scholar]
- Clark D. J., Thomas J. O. Differences in the binding of H1 variants to DNA. Cooperativity and linker-length related distribution. Eur J Biochem. 1988 Dec 1;178(1):225–233. doi: 10.1111/j.1432-1033.1988.tb14447.x. [DOI] [PubMed] [Google Scholar]
- Clark D. J., Thomas J. O. Salt-dependent co-operative interaction of histone H1 with linear DNA. J Mol Biol. 1986 Feb 20;187(4):569–580. doi: 10.1016/0022-2836(86)90335-9. [DOI] [PubMed] [Google Scholar]
- Clore G. M., Gronenborn A. M., Nilges M., Sukumaran D. K., Zarbock J. The polypeptide fold of the globular domain of histone H5 in solution. A study using nuclear magnetic resonance, distance geometry and restrained molecular dynamics. EMBO J. 1987 Jun;6(6):1833–1842. doi: 10.1002/j.1460-2075.1987.tb02438.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coles L. S., Robins A. J., Madley L. K., Wells J. R. Characterization of the chicken histone H1 gene complement. Generation of a complete set of vertebrate H1 protein sequences. J Biol Chem. 1987 Jul 15;262(20):9656–9663. [PubMed] [Google Scholar]
- Crane-Robinson C., Ptitsyn O. B. Binding of the globular domain of linker histones H5/H1 to the nucleosome: a hypothesis. Protein Eng. 1989 Aug;2(8):577–582. doi: 10.1093/protein/2.8.577. [DOI] [PubMed] [Google Scholar]
- De Petrocellis L., Quagliarotti G., Tomei L., Geraci G. Structuring of H1 histone. Evidence of high-affinity binding sites for phosphate ions. Eur J Biochem. 1986 Apr 1;156(1):143–148. doi: 10.1111/j.1432-1033.1986.tb09559.x. [DOI] [PubMed] [Google Scholar]
- Dimitrov S. I., Russanova V. R., Pashev I. G. The globular domain of histone H5 is internally located in the 30 nm chromatin fiber: an immunochemical study. EMBO J. 1987 Aug;6(8):2387–2392. doi: 10.1002/j.1460-2075.1987.tb02516.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ericsson C., Grossbach U., Björkroth B., Daneholt B. Presence of histone H1 on an active Balbiani ring gene. Cell. 1990 Jan 12;60(1):73–83. doi: 10.1016/0092-8674(90)90717-s. [DOI] [PubMed] [Google Scholar]
- Fenn J. B., Mann M., Meng C. K., Wong S. F., Whitehouse C. M. Electrospray ionization for mass spectrometry of large biomolecules. Science. 1989 Oct 6;246(4926):64–71. doi: 10.1126/science.2675315. [DOI] [PubMed] [Google Scholar]
- Fried M. G., Crothers D. M. Equilibrium studies of the cyclic AMP receptor protein-DNA interaction. J Mol Biol. 1984 Jan 25;172(3):241–262. doi: 10.1016/s0022-2836(84)80025-x. [DOI] [PubMed] [Google Scholar]
- Graziano V., Gerchman S. E., Wonacott A. J., Sweet R. M., Wells J. R., White S. W., Ramakrishnan V. Crystallization of the globular domain of histone H5. J Mol Biol. 1990 Mar 20;212(2):253–257. doi: 10.1016/0022-2836(90)90122-3. [DOI] [PubMed] [Google Scholar]
- Hartman P. G., Chapman G. E., Moss T., Bradbury E. M. Studies on the role and mode of operation of the very-lysine-rich histone H1 in eukaryote chromatin. The three structural regions of the histone H1 molecule. Eur J Biochem. 1977 Jul 1;77(1):45–51. doi: 10.1111/j.1432-1033.1977.tb11639.x. [DOI] [PubMed] [Google Scholar]
- Hill C. S., Rimmer J. M., Green B. N., Finch J. T., Thomas J. O. Histone-DNA interactions and their modulation by phosphorylation of -Ser-Pro-X-Lys/Arg- motifs. EMBO J. 1991 Jul;10(7):1939–1948. doi: 10.1002/j.1460-2075.1991.tb07720.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ichimura S., Mita K., Zama M. Essential role of arginine residues in the folding of deoxyribonucleic acid into nucleosome cores. Biochemistry. 1982 Oct 12;21(21):5329–5334. doi: 10.1021/bi00264a032. [DOI] [PubMed] [Google Scholar]
- Kamakaka R. T., Thomas J. O. Chromatin structure of transcriptionally competent and repressed genes. EMBO J. 1990 Dec;9(12):3997–4006. doi: 10.1002/j.1460-2075.1990.tb07621.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klug A., Rhodes D., Smith J., Finch J. T., Thomas J. O. A low resolution structure for the histone core of the nucleosome. Nature. 1980 Oct 9;287(5782):509–516. doi: 10.1038/287509a0. [DOI] [PubMed] [Google Scholar]
- Lambert S. F., Thomas J. O. Lysine-containing DNA-binding regions on the surface of the histone octamer in the nucleosome core particle. Eur J Biochem. 1986 Oct 1;160(1):191–201. doi: 10.1111/j.1432-1033.1986.tb09957.x. [DOI] [PubMed] [Google Scholar]
- Lennard A. C., Thomas J. O. The arrangement of H5 molecules in extended and condensed chicken erythrocyte chromatin. EMBO J. 1985 Dec 16;4(13A):3455–3462. doi: 10.1002/j.1460-2075.1985.tb04104.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Losa R., Thoma F., Koller T. Involvement of the globular domain of histone H1 in the higher order structures of chromatin. J Mol Biol. 1984 Jun 5;175(4):529–551. doi: 10.1016/0022-2836(84)90183-9. [DOI] [PubMed] [Google Scholar]
- Nacheva G. A., Guschin D. Y., Preobrazhenskaya O. V., Karpov V. L., Ebralidse K. K., Mirzabekov A. D. Change in the pattern of histone binding to DNA upon transcriptional activation. Cell. 1989 Jul 14;58(1):27–36. doi: 10.1016/0092-8674(89)90399-1. [DOI] [PubMed] [Google Scholar]
- Nikolaev L. G., Glotov B. O., Dashkevich V. K., Barbashov S. F., Severin E. S. Raspolozhenie gistona H1 v khromatine. Poperechnoe sshivanie tsentral'nykh globuliarnykh chastei molekul H1 bifunktsional'nym reagentom. Mol Biol (Mosk) 1983 Nov-Dec;17(6):1255–1261. [PubMed] [Google Scholar]
- Noll M., Kornberg R. D. Action of micrococcal nuclease on chromatin and the location of histone H1. J Mol Biol. 1977 Jan 25;109(3):393–404. doi: 10.1016/s0022-2836(77)80019-3. [DOI] [PubMed] [Google Scholar]
- Pearson E. C., Butler P. J., Thomas J. O. Higher-order structure of nucleosome oligomers from short-repeat chromatin. EMBO J. 1983;2(8):1367–1372. doi: 10.1002/j.1460-2075.1983.tb01593.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Russanova V. R., Dimitrov S. I., Makarov V. L., Pashev I. G. Accessibility of the globular domain of histones H1 and H5 to antibodies upon folding of chromatin. Eur J Biochem. 1987 Sep 1;167(2):321–326. doi: 10.1111/j.1432-1033.1987.tb13339.x. [DOI] [PubMed] [Google Scholar]
- Russo E., Giancotti V., Crane-Robinson C., Geraci G. Histone H1 and chromatin higher order structure. Does histone H1 exhibit specific self-association? Int J Biochem. 1983;15(4):487–493. doi: 10.1016/0020-711x(83)90121-0. [DOI] [PubMed] [Google Scholar]
- Simpson R. T. Structure of the chromatosome, a chromatin particle containing 160 base pairs of DNA and all the histones. Biochemistry. 1978 Dec 12;17(25):5524–5531. doi: 10.1021/bi00618a030. [DOI] [PubMed] [Google Scholar]
- Staynov D. Z., Crane-Robinson C. Footprinting of linker histones H5 and H1 on the nucleosome. EMBO J. 1988 Dec 1;7(12):3685–3691. doi: 10.1002/j.1460-2075.1988.tb03250.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sugarman B. J., Dodgson J. B., Engel J. D. Genomic organization, DNA sequence, and expression of chicken embryonic histone genes. J Biol Chem. 1983 Jul 25;258(14):9005–9016. [PubMed] [Google Scholar]
- Thibodeau A., Ruiz-Carrillo A. The globular region of histone H5 is equally accessible to antibodies in relaxed and condensed chromatin. J Biol Chem. 1988 Nov 5;263(31):16236–16241. [PubMed] [Google Scholar]
- Thoma F., Koller T., Klug A. Involvement of histone H1 in the organization of the nucleosome and of the salt-dependent superstructures of chromatin. J Cell Biol. 1979 Nov;83(2 Pt 1):403–427. doi: 10.1083/jcb.83.2.403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thoma F., Losa R., Koller T. Involvement of the domains of histones H1 and H5 in the structural organization of soluble chromatin. J Mol Biol. 1983 Jul 5;167(3):619–640. doi: 10.1016/s0022-2836(83)80102-8. [DOI] [PubMed] [Google Scholar]
- Thomas J. O. Chemical cross-linking of histones. Methods Enzymol. 1989;170:549–571. doi: 10.1016/0076-6879(89)70064-1. [DOI] [PubMed] [Google Scholar]
- Thomas J. O., Khabaza A. J. Cross-linking of histone H1 in chromatin. Eur J Biochem. 1980 Dec;112(3):501–511. doi: 10.1111/j.1432-1033.1980.tb06113.x. [DOI] [PubMed] [Google Scholar]
- Thomas J. O., Kornberg R. D. The study of histone--histone associations by chemical cross-linking. Methods Cell Biol. 1978;18:429–440. [PubMed] [Google Scholar]
- Thomas J. O., Rees C., Butler P. J. Salt-induced folding of sea urchin sperm chromatin. Eur J Biochem. 1986 Jan 15;154(2):343–348. doi: 10.1111/j.1432-1033.1986.tb09403.x. [DOI] [PubMed] [Google Scholar]
- Thomas J. O., Rees C. Exchange of histones H1 and H5 between chromatin fragments. A preference of H5 for higher-order structures. Eur J Biochem. 1983 Jul 15;134(1):109–115. doi: 10.1111/j.1432-1033.1983.tb07538.x. [DOI] [PubMed] [Google Scholar]
- Thomas J. O., Wilson C. M. Selective radiolabelling and identification of a strong nucleosome binding site on the globular domain of histone H5. EMBO J. 1986 Dec 20;5(13):3531–3537. doi: 10.1002/j.1460-2075.1986.tb04679.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turnell W. G., Satchwell S. C., Travers A. A. A decapeptide motif for binding to the minor groove of DNA. A proposal. FEBS Lett. 1988 May 23;232(2):263–268. doi: 10.1016/0014-5793(88)80750-6. [DOI] [PubMed] [Google Scholar]
- Weintraub H. Assembly and propagation of repressed and depressed chromosomal states. Cell. 1985 Oct;42(3):705–711. doi: 10.1016/0092-8674(85)90267-3. [DOI] [PubMed] [Google Scholar]
- Wells D., McBride C. A comprehensive compilation and alignment of histones and histone genes. Nucleic Acids Res. 1989;17 (Suppl):r311–r346. doi: 10.1093/nar/17.suppl.r311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Widom J. Toward a unified model of chromatin folding. Annu Rev Biophys Biophys Chem. 1989;18:365–395. doi: 10.1146/annurev.bb.18.060189.002053. [DOI] [PubMed] [Google Scholar]
- Wray W., Boulikas T., Wray V. P., Hancock R. Silver staining of proteins in polyacrylamide gels. Anal Biochem. 1981 Nov 15;118(1):197–203. doi: 10.1016/0003-2697(81)90179-2. [DOI] [PubMed] [Google Scholar]