Abstract
Plant growth and development are sustained by continuous cell division in the meristems, which is perturbed by various environmental stresses. For the maintenance of meristematic functions, it is essential that cell division be coordinated with cell differentiation. However, it is unknown how the proliferative activities of the meristems and the coordination between cell division and differentiation are maintained under stressful conditions. Here we show that a rice protein, RSS1, whose stability is controlled by cell cycle phases, contributes to the vigour of meristematic cells and viability under salinity conditions. These effects of RSS1 are exerted by regulating the G1–S transition, possibly through an interaction of RSS1 with protein phosphatase 1, and are mediated by the phytohormone, cytokinin. RSS1 is conserved widely in plant lineages, except eudicots, suggesting that RSS1-dependent mechanisms might have been adopted in specific lineages during the evolutionary radiation of angiosperms.
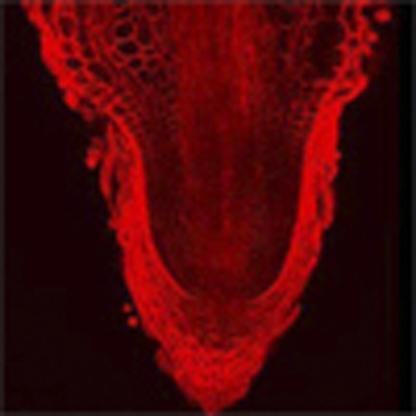 Cell proliferation in plant meristems is often altered during conditions of stress. In this study, the authors identify a plant protein, RSS1, that is regulated in a cell-cycle dependent manner and is required to maintain shoot and root meristems in the presence of abiotic stress.
Cell proliferation in plant meristems is often altered during conditions of stress. In this study, the authors identify a plant protein, RSS1, that is regulated in a cell-cycle dependent manner and is required to maintain shoot and root meristems in the presence of abiotic stress.
In accordance with their sessile lifestyle, land plants have acquired various systems to adapt to environmental conditions. To maintain growth under stress conditions, the activity of meristems needs to be maintained even while mature tissues undergo accelerated senescence. This is because post-embryonic growth of plants largely depends on the activities of the apical and lateral meristems. Particularly in the monocot cereals, proliferating cells in the shoot apex give rise to most of the shoot and root tissues, as the majority of the roots generate adventitiously from shoots, independently of the seminal root.
The meristems are composed of undifferentiated stem cells with a low dividing activity and actively dividing cells that provide new cells to undergo differentiation. In addition to the cell cycle phase-dependent control by cyclins and cyclin-dependent protein kinases (CDKs), cell proliferation in meristems is regulated in coordination with the differentiation of neighbouring cells1,2. In fact, the aberrant expression of a B-type CDK has been shown to cause abnormal meristem organization in Arabidopsis3. Moreover, the downregulation or induced overexpression of Arabidopsis retinoblastoma-related protein, which acts downstream of cyclin D and regulates G1–S transition, also affects stem cell maintenance and meristem organization4,5. Under stress conditions, cell division is slowed by checkpoint systems to ensure accurate cell cycle progression in many organisms2,6. In plants, inhibitors of CDK, such as Arabidopsis ICK1 and rice EL2, are reported to be induced by abiotic stresses7,8. However, it is not understood how cell proliferation in meristems is maintained in coordination with cell differentiation under environmental stress conditions.
Rice and other monocot cereals are the principal foods for the world's population. To achieve increased food production, it is necessary to identify the genetic determinants and the underlying mechanisms that control tolerance to environmental stress, especially soil salinity and drought, in these crops9. However, in monocots, genetic studies aimed at elucidating the mechanisms underlying salt tolerance are scarce, as compared with those in the model dicot, Arabidopsis; studies in the latter have revealed that a number of genes are required for salt tolerance, including the SOS genes involved in the Na+ translocation system10. In rice, one of the quantitative trait loci controlling salt tolerance was recently cloned: it encodes a plasma membrane Na+ transporter, which regulates Na+ unloading from the xylem and, thereby, the K+/Na+ ratio in the shoot11. In contrast to the accumulation of knowledge for the genetic mechanisms of salt tolerance related to the ionic status, the mechanisms that ensure cell-proliferative activity under saline conditions are largely unknown in both monocots and dicots.
We report here that a previously uncharacterized protein, RSS1, has a critical role in tolerance to multiple abiotic stresses in rice plants. RSS1 functions preferentially in dividing cells, and its stability is regulated by cell cycle phase-dependent protein degradation through the anaphase-promoting complex/cyclosome (APC/C)–26S proteasome pathway. Moreover, RSS1 interacts with a type 1 protein phosphatase (PP1), which regulates many cellular processes, including the cell division cycle12,13,14,15,16. We found that the activity of RSS1 was required for the balance between cell division and differentiation, and thus the proper organization of meristems. On the basis of these results, we propose that RSS1 acts as a key factor in the maintenance of meristematic activity by ensuring cell division under stressful environmental conditions.
Results
rss1 is sensitive to salinity stress
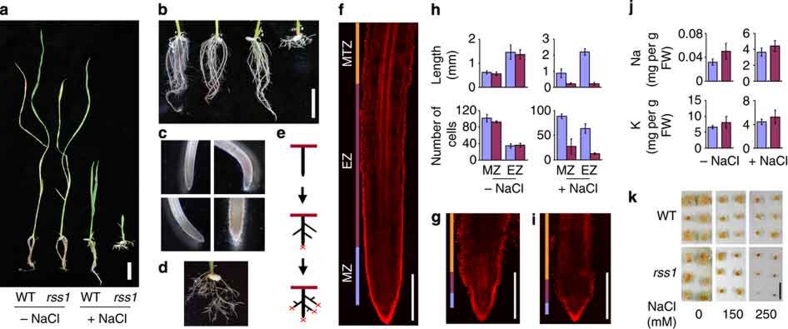
To explore the mechanisms underlying salt tolerance in monocots, we performed a genetic screen of rice populations mutagenized with the retrotransposon, Tos17 (ref. 17), and we identified a gene whose defect caused an extreme dwarf and short-root phenotype under high-salt, but not normal growth conditions (Fig. 1a). The obtained loss-of-function mutant, designated rss1 (rice salt sensitive 1), exhibited irreversible growth inhibition after exposure to 150 mM NaCl for 2 weeks. The rss1 seedlings exhibited highly branched roots, possibly reflecting impaired root apex activity (Fig. 1b–e). Such a branching pattern was not observed in wild-type (WT) plants treated with various concentrations of salt, including those higher than 150 mM. In the root apex of young rss1 seedlings, the size of the meristematic zone (MZ) and the elongation zone (EZ) decreased under salinity, concomitant with a reduction in the number of cells in both zones (Fig. 1f–h and Supplementary Fig. S1). Interestingly, the defects in rss1 were mimicked in WT roots by the application of the DNA synthesis inhibitor, aphidicolin, which induces the cell cycle arrest at the G1–S boundary (Fig. 1i). The dramatic reduction in the sizes of both the MZ and EZ in the rss1 root under high salinities probably indicates that the number of cells supplied from the MZ to EZ was considerably decreased, while the rate of differentiation was less affected. A decreased rate of cell division and a constant rate of differentiation will result in the reduction of the MZ size. The salt hypersensitivity of rss1 was not due to the overloading of Na+ because the shoot K+/Na+ ratio in the mutant was nearly the same as that in WT under high-salt conditions (Fig. 1j). Moreover, rss1 calli were hypersensitive to salt, suggesting that RSS1 confers salt tolerance primarily at the cellular level (Fig. 1k and Supplementary Fig. S2). Salinity is known to cause detrimental effects through not only ionic toxicity but also a water deficit that results from hyper-osmotic stress10,18. Notably, rss1 was hypersensitive to both ionic and hyper-osmotic stress caused by low concentrations of LiCl and high concentrations of sorbitol, respectively (Supplementary Fig. S3).
Figure 1. rss1 is hypersensitive to salt.
(a, b) The phenotype of rss1. WT and rss1-1 seedlings were grown in the absence (−) or presence of 150 mM NaCl (+) for 15 days. (a) Seedlings. (b) The roots in (a) were visualized by floating in water. Left, WT (−NaCl); middle-left, rss1-1 (−NaCl); middle-right, WT (+NaCl); right, rss1-1 (+NaCl). (c) Root tips of WT (top) and rss1-1 (bottom) seedlings grown in the absence (left panels) or presence of 150 mM NaCl (right panels). (d) The root branching pattern of rss1-1 grown in the presence of 150 mM NaCl. (e) A schematic illustration of rss1-1 root branching under high-salt conditions. (f, g) Propidium iodide (PI) staining of the root cells of WT (f) and rss1-2 (g) plants grown under high-salt conditions (150 mM NaCl, 5 days). (h) The length of (top) and the number of cells in (bottom) the MZ and EZ in the root of WT (blue) and rss1-1 (magenta) plants grown under normal (−NaCl, 4 days) or high-salt conditions (+150 mM NaCl, 5 days). Mean±s.d., n=4. (i) PI staining of root cells of WT grown in the presence of 150 mM NaCl and 10 mg/l aphidicolin. (j) The amount of Na+ (top) and K+ (bottom) in the shoot of WT (blue) and rss1 (magenta) seedlings grown in the absence (−) or presence (+) of 150 mM NaCl for 14 days. The data represent the mean±s.d., n=5. The K+/Na+ ratios were 211.1±46.2 in WT versus 164.4±32.2 in rss1-1 under normal conditions and 1.23±0.17 in WT versus 1.19±0.24 in rss1-1 under high-salt conditions (mean±s.d., n=5). (k) rss1 calli are hypersensitive to salt. WT or rss1-1 calli derived from seeds were grown in MS medium containing 2 mg/l 2,4-dichlorophenoxy acetic acid. Fresh callus pieces were then transferred to fresh medium of the indicated concentrations of NaCl and cultured at 25 °C for 3 weeks. Bars: 2 cm (a, b), 0.5 mm (f, g, i), 1 cm (k). EZ, elongation zone; MTZ, maturation zone; MZ, meristematic zone.
RSS1 is controlled in a cell cycle phase-dependent manner
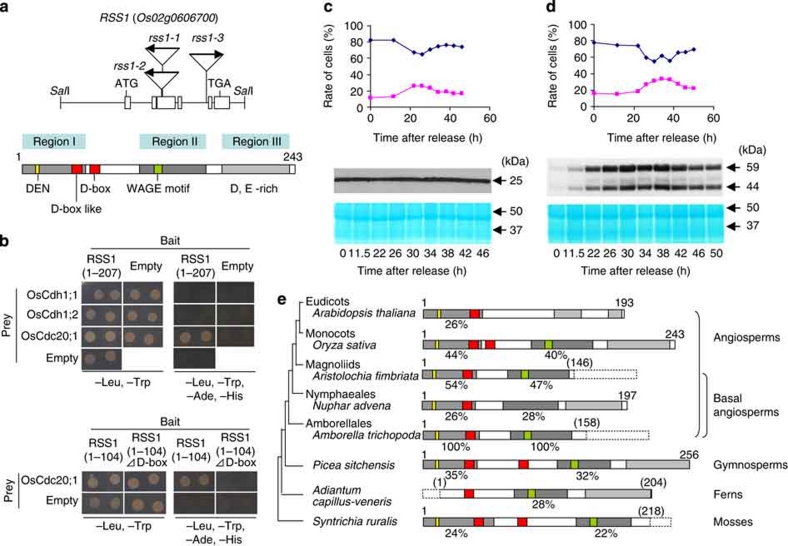
We molecularly identified RSS1 by transposon tagging and the subsequent isolation and characterization of two additional mutant alleles (Fig. 2a). Tos17 (4.1 kb) was inserted into the third exon of RSS1 in rss1-1 and rss1-2 and into the fourth intron in rss1-3. A full-length RSS1 cDNA driven by the rice actin or RSS1 promoter was able to complement the rss1 mutation (Supplementary Fig. S4). RSS1 encodes an uncharacterized protein consisting of 243 amino acids that was localized in both the nucleus and cytosol when fused to GFP (Supplementary Fig. S5). RSS1 showed no significant sequence similarity to any proteins with known function in the publicly accessible databases; however, it has a canonical motif, a 'destruction box' (D-box) and a D-box-like motif in its amino-terminal region (Supplementary Fig. S6). The D-box acts as a target signal for ubiquitin-26S proteasome-dependent degradation from prometaphase to the late G1 phase, which is regulated by the APC/C E3 ubiquitin ligase19,20,21. This targeting is often specified in conjunction with another cis-sequence, or 'KEN-box'20. RSS1 also possesses a KEN-related sequence, the 'DEN-box,' and RSS1 was stabilized by treatment with a specific inhibitor of the 26S proteasome, MG132 (Supplementary Fig. S4). The N-terminal 104-amino-acid sequence containing both the D- and DEN-boxes was required and was sufficient for the 26S proteasome-dependent degradation. The deletion or amino-acid substitution of the D-boxes in RSS1 partially disrupted the effect of MG132. These results suggest that RSS1 is regulated by APC/C.
Figure 2. RSS1 encodes an uncharacterized protein containing a D-box.
(a) (top) The schematic structure of RSS1. Rectangles indicate the exons. The first ATG, the stop codon, and the insertion sites of the Tos17 retrotransposon (arrows) in the three mutant alleles are shown. (bottom) The RSS1 protein including its conserved motifs. The three regions containing amino acid (a.a.) sequences that are conserved among the RSS1 homologues, the DEN and double D-Box, the WAGE motif, and the C-terminal acidic amino-acid-rich region are shown. (b) Assay for the interaction of RSS1 with the subunits of APC/C by the yeast two-hybrid system. (top) The GAL4 DNA-binding domain fused to the truncated RSS1 (a.a. 1–207) and GAL4 activation domain fused to OsCdh1;1 (Os01g0972900), OsCdh1;2 (Os03g0123300) or OsCdc20;1 (Os04g0599800) were co-expressed in the yeast cells carrying the reporter genes, ADE2 and HIS3. The yeast cells were grown on the appropriate medium (which did not contain the indicated amino acids). (bottom) The two-hybrid assay using truncated RSS1 (a.a. 1–104) with or without the D-boxes as the bait. Empty indicates the vector control. (c, d) The cell cycle-dependent accumulation of GFP (c) and RSS1-GFP (d). Rice Oc cells carrying the respective transgenes were synchronized and used for ploidy analysis (top) and immunoblot analysis using anti-GFP (middle) at the indicated time points after release from G1 arrest by starvation. (top) The rates of 2C cells (blue) and 4C cells (magenta) are plotted. The 59-kDa band corresponds to the full-length RSS1-GFP protein. (bottom) Protein staining for the loading control of the immunoblot. (e) A simplified phylogeny of plants and the schematic structure of RSS1-related proteins representative of the respective lineages are illustrated. The percent amino-acid identity among sequences homologous to regions I and II is indicated. The C-terminal regions containing acidic amino acids are not necessarily homologous to region III. The dashed line indicates truncated sequence information. GenBank accession and code numbers are as follows: Arabidopsis AT3G14910 (NM_112353), Oryza sativa Os02g0606700 (RSS1) (NC_008395), Aristolochia fimbriata (FD758252), Nuphar advena (PUT-165a-Nuphar_advena-6361, Plant GDB (http://www.plantgdb.org/), Amborella trichopoda (CK757545), Picea sitchensis (ABK26784), Adiantum capillus-veneris (DK961685), and Syntrichia ruralis (CN201203).
It is known that a substrate of APC/C is specified by the binding of Cdh1 or Cdc20 to the D-box20,21. Therefore, we examined whether RSS1 was recognized by these proteins. As shown in Figure 2b and Supplementary Figure S7, RSS1 specifically interacted with a rice homologue of Cdc20, but not a homologue of Cdh1, in a D-box-dependent manner. We further tested whether the stability of RSS1 was controlled in a cell cycle phase-dependent manner, using rice cultured cells expressing an RSS1–GFP fusion protein or GFP alone (Fig. 2c,d). When expressed under the control of the constitutive actin gene promoter, the levels of GFP were not affected according to the changes of the cell cycle phases. In contrast, RSS1–GFP began to be detectable nearly at the onset of DNA replication (after 11.5 h), and increased its levels during S-phase progression, and was then destabilized concomitant with progression of mitosis (after 42 h). This result is consistent with the fact that protein degradation by APC/C occurs during M–G1 phase20,21. Taken together, the stability of RSS1 is regulated in a cell cycle phase-dependent manner, which is mediated by APC/C.
Loss and diversification of RSS1 homologues in angiosperms
RSS1-related sequences are not only highly conserved among monocots, but can also be identified in basal angiosperms, including Amborella trichopoda, which belongs to the most ancient lineage of extant angiosperms (Fig. 2e and Supplementary Figs. S8 and S9)22. RSS1-related genes also exist in gymnosperms, ferns and mosses. In addition to the N-terminal region containing the DEN-box and D-box/D-box-like motifs (region I), these RSS1 homologues have a well-conserved region in the middle region (region II), which includes a sequence (W–A–K/R–D/E/G–G–V/I–E) designated the 'WAGE' motif and a moderately conserved carboxy-terminal acidic region (region III) (Fig. 2a,e and Supplementary Fig. S8). Notably, no proteins with overall similarity to RSS1 exist in eudicots, although sequences related to region I were found. Moreover, region I of RSS1 was not necessary for salt tolerance (Supplementary Fig. S4). These results suggest a lineage-specific loss of RSS1 counterparts in eudicots. Interestingly, an RSS1-related gene in Nuphar advena is highly diverged and lacks the WAGE motif (Fig. 2e and Supplementary Fig. S8). The loss and/or pronounced diversification of RSS1-related genes in angiosperms may reflect evolutionary radiation22,23 or lineage-specific adaptations to environmental conditions (Supplementary Figs S8 and S9 and Supplementary Discussion).
RSS1 is expressed preferentially in dividing cells
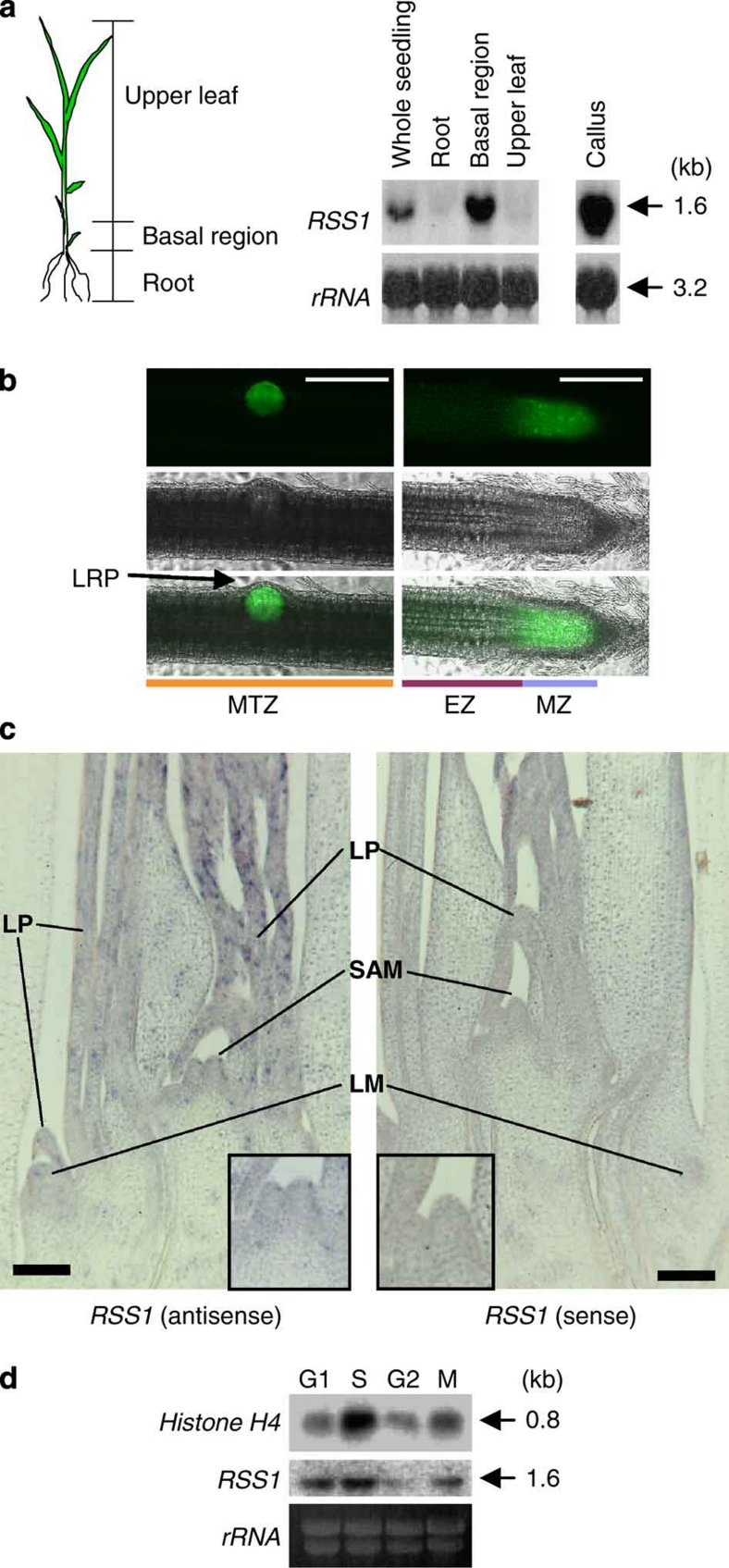
RSS1 mRNA is expressed abundantly in proliferating tissue, such as the basal region of the shoot, which contains apical and lateral shoot meristems and leaf primordia, and, to a lesser extent, in the upper region of the shoot (Fig. 3a). RSS1 expression was upregulated by low temperatures, but not by high-salt conditions (Supplementary Fig. S10). In the root, RSS1 promoter activity was detected predominantly in the MZ or lateral root primordia (Fig. 3b). At the reproductive stages, RSS1 was expressed preferentially in immature inflorescences, as compared with mature florets or leaves (Supplementary Fig. S10). In situ hybridization analysis revealed patchy signals for RSS1 mRNA in the leaf primordia and in the meristems of the shoot and root apices, which are the major sites of cell cycling (Fig. 3c and Supplementary Figs. S10 and S11). In the shoot apical meristem (SAM), RSS1 mRNA was detected only weakly in the central zone, where the cells are less active in terms of division and are completely undifferentiated. This is in sharp contrast to the expression pattern of OSH1, a homeobox gene orthologous to Arabidopsis STM, which is expressed in indeterminate cells, but not in leaf primordia, and is suggested to function in the maintenance of undifferentiated states24. The spatial expression pattern of RSS1 is reminiscent of those of genes encoding histones, cyclins or CDKs24,25,26. The expression of these cell cycle-related genes is often associated with specific phases of the cell cycle. RSS1 was expressed most abundantly during the G1 and S phases (Fig. 3d and Supplementary Fig. S10), similar to cyclin D27. These results, together with the finding of cell cycle phase-dependent proteolysis, suggest that RSS1 functions in cell division.
Figure 3. RSS1 is expressed preferentially in dividing cells.
(a) The expression of RSS1 in seedlings and calli. Two-week-old seedlings of rice were dissected into the three parts as in the left panel and used for northern blot analysis with an RSS1-specific probe (right). The basal region of the shoot contains the shoot apical meristem (SAM) and leaf primordia (LP). rRNA, loading control. (b) GFP signals detected in the root of transgenic rice carrying GFP driven by the RSS1 promoter. Dark-field GFP images (top) and bright-field images (middle) are merged in the panels on the bottom. (left panels) The MTZ of the root with an emerging lateral root primordium (LRP). (right panels) The apex of an adventitious root. Bars, 200 μm. (c) In situ localization of RSS1 mRNA in the shoot. Longitudinal sections were hybridized with anti-sense and sense DIG-labelled RNA probes specific for RSS1, respectively. LM, lateral meristem. Insets Increased magnification of the SAM region. Bars, 100 μm. (d) Expression of RSS1 during the cell cycle. The synchronized rice Oc cells were harvested at the indicated cell cycle phases and analysed by northern blotting using histone H4- or RSS1-specific probes. rRNA, loading control.
RSS1 ensures cell division activity under stress conditions
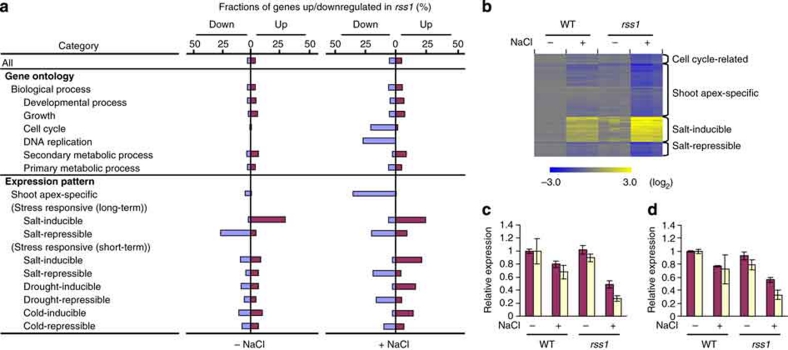
To obtain further insight into the function of RSS1, we characterized the transcriptomic profiles in the basal proliferative region of the shoot between wild type and rss1 grown under normal or high-salt conditions. In agreement with the proposed function of RSS1, genes involved in the cell cycle and DNA replication were preferentially downregulated in rss1 under high-salt conditions (Fig. 4a,b), as exemplified by an S-phase-specific gene, PCNA, and an M-phase cyclin, CycB2;1 (Fig. 4c,d). Moreover, ploidy analysis demonstrated that the relative number of cells in G2-M phase (4C cells) compared with G1 phase (2C cells) was decreased in rss1 under salinity (Fig. 5a,b). These results suggest that RSS1 is required for the G1–S transition and subsequent cell cycle progression under stressful conditions. Conspicuously, more than 30% of the genes that were expressed specifically in the shoot apex were coordinately downregulated in rss1 under high-salt conditions (Fig. 4a,b), supporting the proposed function of RSS1 in maintaining proliferative tissue activity.
Figure 4. Altered expression of genes in rss1.
(a) The classification of the genes whose expression was affected by the defects in rss1. RNA levels in the shoot basal proliferative tissues of 1-week-old WT and rss1-2 seedlings grown in the absence (−) or presence of 150 mM NaCl (+) were examined by microarray with three biological replicates. Differentially expressed genes were statistically determined by the rank product (P<0.01) and two-way ANOVA (P<0.01). The ratio of the number of genes, whose expression was significantly upregulated (magenta) or downregulated (blue) in rss1, to the number of genes in the indicated gene category is shown. Among the total number of 29,859 genes examined, 1,129 genes (3.8%) and 792 genes (2.7%) were up- and downregulated in rss1, respectively, under the normal growth condition, whereas 1,607 genes (5.4%) and 1,581 genes (5.3%) were up- and downregulated in rss1, respectively, under conditions of salinity. Categories included in 'Biological process' were selected from the Gene Ontology (GO) database (http://www.gramene.org/plant_ontology/index.html). Categories of 'Shoot apex-specific,' stress responsive (long-term) and stress responsive (short-term) were selected from our microarray data and public array databases (see Methods). (b) A heat map view of the expression pattern of down- or upregulated genes selected from the indicated categories. 'Salt-inducible' (or 'salt-repressible') genes are a mixture of genes that were commonly up- (or down)regulated under the both long- and short-term salinity conditions. The levels of expression are shown with log2-transformed values (bottom) based on the microarray data, as in (a). The patterns of expression with the three biological replicates are shown. (c, d) The expression of PCNA (c) and CycB2;1 (d) in the basal shoot tissues of one-week-old WT and rss1-2 seedlings, as determined by microarray analysis (magenta), or of WT and rss1-1, as determined by quantitative RT-PCR (yellow). Seedlings were grown in the absence (−) or presence of 150 mM NaCl (+). Mean±s.d., n=3. The relative expression levels in the real-time RT-PCR analysis were normalized by the expression level of ubiquitin E2.
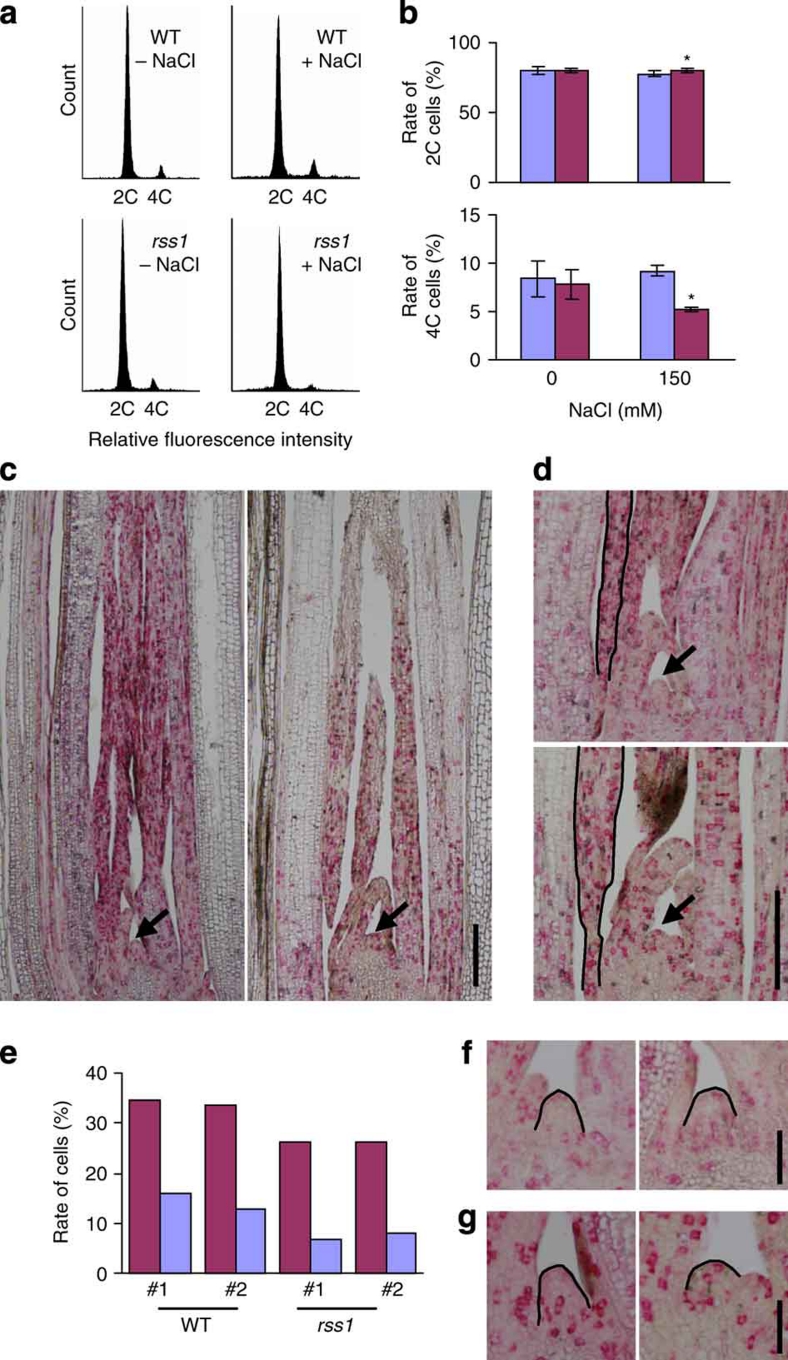
Figure 5. RSS1 ensures cell division activity at the G1–S transition.
(a, b) Flow cytometry measurements of nuclei from the basal shoot tissues in 1-week-old WT and rss1-2 plants grown under non-stressed (−NaCl) and salt-stressed (+150 mM NaCl) conditions. (a) The representative data of the 2C and 4C cells. The number of nuclei is plotted according to their fluorescence intensity. (b) The ratio of the 2C (top) and 4C (bottom) cells in WT (blue) and rss1-2 (magenta). Mean±s.d., n=5–9. Asterisks, a significant difference between WT and rss1-2 (P=0.0052 (top panel), P=0.0002 (bottom panel); one-tailed t-test). (c, d) In situ analysis of the expression of histone H4 (red) and cyclin B2 (purple). Antisense probes for histone H4 and cyclin B2 were labelled differentially and hybridized with longitudinal sections of the basal shoot in WT and rss1-2 seedlings grown in the presence of 150 mM NaCl. (c) Overviews of the longitudinal sections, including the shoot apical meristem (SAM) and leaf primordia of WT (left) and rss1-2 (right). (d) Magnified views of the in situ detection of transcripts for histone H4 (red) and cyclin B2 (purple) in the shoot basal tissue. (top) WT. (bottom) rss1-2. Black lines trace the leaf margin of primordium 3 (P3). (e) Rate of cells that express transcripts for histone H4 (magenta) and cyclin B2 (blue) in the leaf margin of P3 in (d). Two seedlings (nos. 1 and 2) were analysed for WT and rss1-2, respectively. The average scores based on the observation of two to three serial sections are shown. (f, g) In situ detection of histone H4 mRNA (red signals) in the SAM region in WT (left each) and rss1-2 (right each) plants grown under non-stressed (f) and salt-stressed conditions (g). The regions of layer 1 (L1) in the SAM used for counting the cell number are traced by black lines. Arrows indicate the SAM in (c, d). Bars indicate 200 μm in (c, d), and 50 μm in (f, g).
To confirm the reduction of cell division activity under salinity in the basal shoot region in rss1, we investigated the in situ expression patterns of histone H4 and cyclin B2 as representative markers for S phase and G2/M phase cells, respectively. In rss1, the size of the cell division zone, marked with both transcripts, was decreased under the salt-stress condition (Fig. 5c). In addition, the ratio of cells expressing histone H4 and cyclin B2 was decreased (Fig. 5d,e; in the leaf margin, P3). These results indicated that RSS1 was required for the maintenance of the number of proliferative cells, and support the hypothesis that cell cycle progression is inhibited primarily at the G1–S transition in rss1 under salinity.
rss1 affects SAM maintenance and abiotic stress responses
In rss1, the size of the SAM was decreased under salinity, but not normal growth conditions (Fig. 5f,g). The rss1 SAM was flattened, as compared with the dome-shaped SAM in WT. Concomitant with this, the number of cells in layer 1 (L1) of the SAM was decreased in rss1 under the salinity condition (19 and 20 in WT, 10 and 10 in rss1; n=2), but not under normal conditions (18 and 18 in WT, 18 and 16 in rss1; n=2). In WT, the central dome region consists of stem cells that have a low cell division activity. However, transcripts for histone H4 were often detected in the central region of rss1 under salinity (Fig. 5g). The activation of cell cycle-related genes or cell divisions in the central region of the SAM is often associated with defects in the proper organization of the SAM and results in the loss of indeterminate cells24. Thus, RSS1 was required for the maintenance of the SAM under stress conditions.
We investigated the transcriptional profiles of abiotic stress-responsive genes. Consistent with the observed salt hypersensitivity, responses to salt, in terms of gene expression, were considerably enhanced in rss1 (Fig. 4a,b). The genes that were induced by salt and further upregulated in rss1 under salinity included genes encoding proteins for osmolyte accumulation, free radical elimination and transcriptional regulation, which have been implicated in salt-stress tolerance (Supplementary Fig. S12). The expression of long-term salinity-induced genes was often, though slightly, increased in rss1, even under normal growth conditions, possibly reflecting a weak provocation of the stress response. Despite the overall tendency towards the enhancement of stress responses, several genes involved in salt tolerance were downregulated in rss1, as exemplified by OsNHX1 and OsDREB1A, which encode a vacuolar Na+/H+ antiporter and transcription factor, respectively (Supplementary Fig. S12)28,29. Thus, the rss1 mutation perturbs some responses required for the adaptation to salinity. In plants, the mechanisms and regulatory networks mediating tolerance to distinct abiotic stresses overlap30,31. Indeed, in rss1, the expression of a large number of drought- and cold-inducible genes was upregulated under high-salt conditions (Fig. 4a). Moreover, rss1 displayed hypersensitivity to both cold and heat (Supplementary Fig. S12). Therefore, RSS1 is involved in a common mechanism required for the tolerance to several abiotic stresses.
RSS1 contributes to maintenance of active cytokinin levels
Cytokinin is a phytohormone with central roles in cell division32,33,34 and in the maintenance of stem cells in the SAM35,36, and it negatively regulates abiotic stress-induced senescence37. Hence, the function of RSS1 in the regulation of the cell cycle under conditions of high salinity and the effects of rss1 on stress-regulated gene expression led us to examine the possible mediation of cytokinins. The exogenous application of cytokinins alleviated the lethality of the rss1 mutation in the shoot under high-salt conditions (Fig. 6a). In the basal region of the shoot, the expression of OsCKX2, which encodes a cytokinin oxidase that inactivates cytokinins38, was upregulated in rss1 (Fig. 6b). Moreover, the levels of an active cytokinin, trans-zeatin (tZ), were increased under high-salt conditions in WT plants, but not in rss1 (Fig. 6c,d), although the level of another cytokinin, N6-(Δ2-isopentenyl) adenine (iP), was decreased in both WT and rss1 plants (Supplementary Fig. S13). The reduced level of tZ in rss1 was accompanied by a decline in the level of its precursor and an increase in the inactive tZ-O-glucoside (tZOG) content (Fig. 6c,d). Decreased cytokinin activity in rss1 was also evident from the downregulation of the cytokinin-inducible response regulator gene, OsRR139 (Fig. 6b). These results suggest that the function of RSS1 is somehow connected to the sustainment of proliferating shoot tissues by maintaining the active cytokinin level in the face of salt stress.
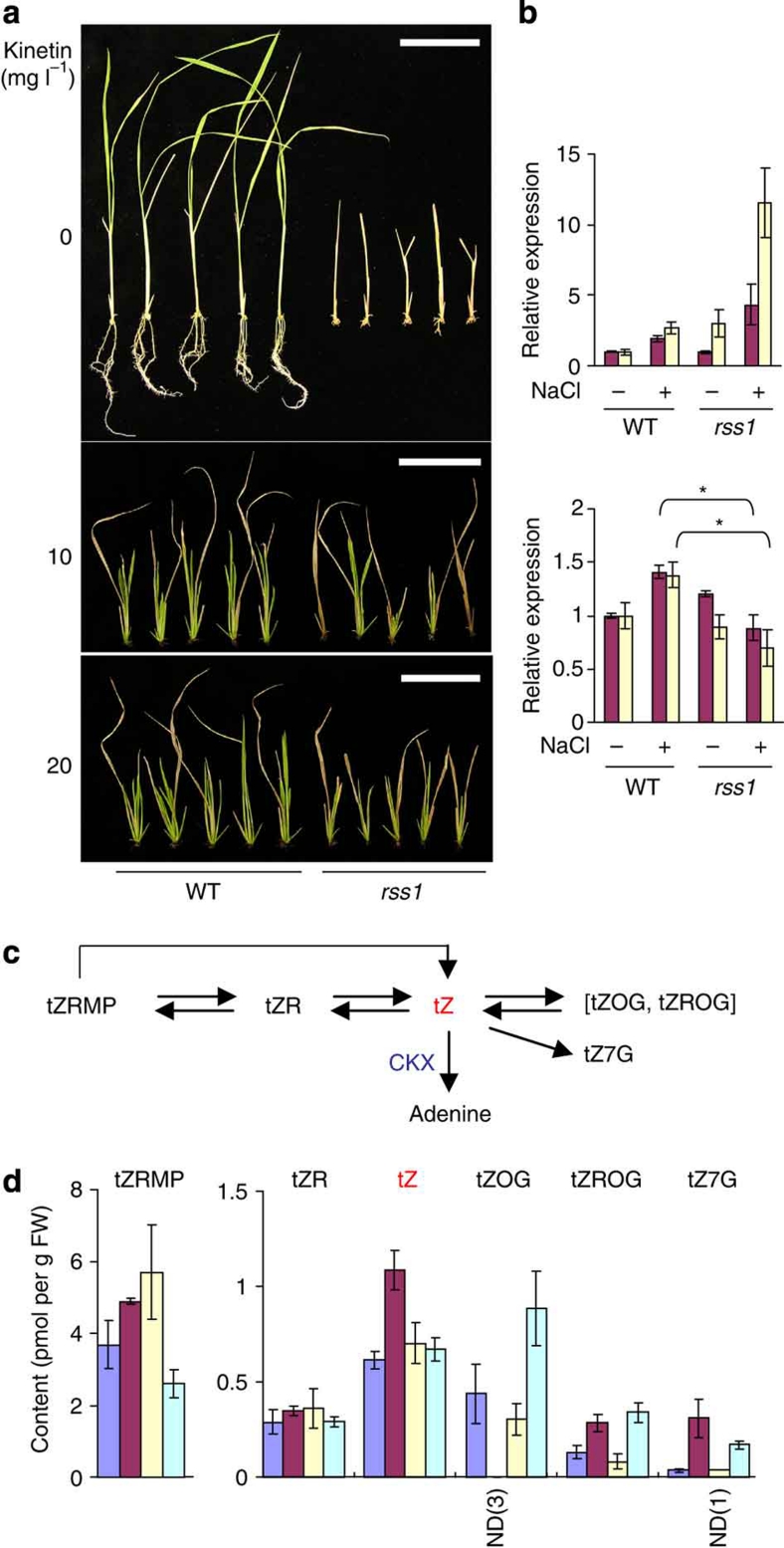
Figure 6. rss1 affects cytokinin action.
(a) The exogenous application of cytokinins alleviated the lethality of rss1 under high-salt conditions. The basal shoot segments without roots (2.5 cm long) were excised from 1-week-old WT and rss1-2 plants, transferred to the medium containing 150 mM NaCl with or without the indicated concentrations of kinetin, and grown for 19 days. Bars, 5 cm. (b) The expression of CKX2 (top) and RR1 (bottom) in the basal shoot tissues of 1-week-old WT and rss1-2 seedlings, as determined by microarray analysis (magenta), or of WT and rss1-1 plants, as determined by quantitative RT-PCR (yellow). Seedlings grown under the normal (−NaCl) and saline (+NaCl) conditions were tested. The expression levels of the tested genes quantified by RT-PCR were normalized with that of ubiquitin E2. Mean±s.d., n=3. Asterisks, a significant difference between WT and rss1-2 under the saline condition. (P=0.002 (by real-time RT-PCR), P=0.001 (by microarray); one-tailed t-test). (c) A schematic illustration of the metabolic pathway of trans-zeatin (tZ) adapted from previous reports33,34. tZR, tZ riboside; tZRMP, tZR 5′-monophosphate; tZOG, tZ-O-glucoside; tZROG, tZ O-glucoside riboside; tZ7G, tZ-N-7-glucoside; CKX, cytokinin oxidase. (d) The levels of tZ and related metabolites in the 1-cm-long shoot basal tissues from 1-week-old seedlings of WT (blue and magenta) and rss1-2 seedlings (yellow and light blue) grown in the absence (blue and yellow) or presence of 150 mM NaCl (magenta and light blue) (mean±s.d., n=3). ND(1), the average of two independent samples due to one value being below the detection limit. ND(3), all values were below the detection limit in triplicate samples.
RSS1 interacts with PP1
To gain further insight into the molecular function of RSS1, we searched for RSS1-interacting proteins by a yeast two-hybrid screen. Notably, we found that RSS1 binds to a catalytic subunit of PP1 (Fig. 7a,b). PP1 is a major serine/threonine protein phosphatase that regulates physiological processes in eukaryotes, including cellular signalling that leads to gene expression, cell cycle and cellular metabolism12,13,14,15,16. We designated this RSS1-interacting PP1 OsPP1. Our data indicated that an N-terminal region of OsPP1 was sufficient for the interaction with RSS1 (Supplementary Fig. S14). When expressed in onion epidermal cells, OsPP1-fused GFP was localized both in the nucleus and in the cytosol, as was RSS1-GFP and YFP-RSS1 (Supplementary Fig. S5). The in vivo interaction between RSS1 and OsPP1 was confirmed by the bimolecular fluorescence complementation (BiFC) assay (Fig. 7c).
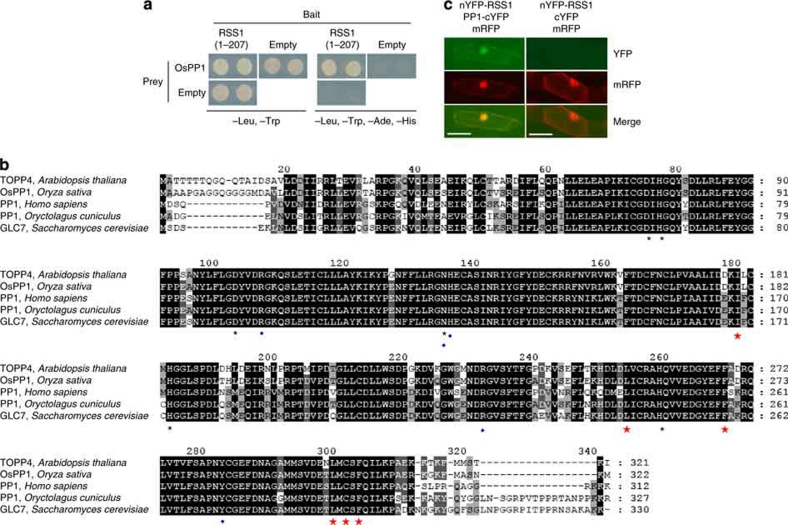
Figure 7. RSS1 interacts with a catalytic subunit of PP1.
(a) The interaction of RSS1 with OsPP1 by yeast two-hybrid systems. The GAL4 DNA-binding domain fused to the truncated RSS1 (a.a. 1–207) and the GAL4 activation domain fused to OsPP1 (Os03g0268000) were co-expressed in yeast cells carrying the reporter genes, ADE2 and HIS3. The yeast cells were grown on the appropriate medium (which did not contain the indicated amino acids). Empty indicates the vector control. (b) An alignment of the amino-acid sequences of PP1 homologues (Arabidopsis thaliana TOPP4 (AT2G39840), Oryza sativa OsPP1, Homo sapiens PP1 (NP_002699.1), Oryctolagus cuniculus, PP1 (CAA43820), Saccharomyces cerevisiae GLC7 (NP_011059.1)). Black asterisks and blue diamonds indicate the residues that contribute to metal coordination and phosphate binding, respectively12. Red stars indicate the conserved residues involved in Rb binding16. (c) The bimolecular fluorescent complementation (BiFC) assay for interactions between RSS1 and OsPP1 in onion epidermal cells. The plasmids for expression of the indicated split YFP variants with the reference gene encoding mRFP were introduced by particle bombardment. Colour images for fluorescence of YFP (green, top) and mRFP (red, middle) are merged (bottom). Bars indicate 0.1 mm.
The overall amino-acid sequence of OsPP1 is similar to PP1s of other eukaryotes (Fig. 7b). This includes a conservation of amino acids not only for phosphate and metal binding but also for the interaction with the retinoblastoma protein (Rb), a substrate of PP1 in animals16,40,41. Rb is a tumour suppressor and prevents G1–S transition when dephosphorylated by PP1. Although most of the substrates of PP1 are specified by the regulatory subunits of PP1, Rb is recognized directly by the catalytic subunit alone, through the RVxF-like motif (KLRF) located at the C-terminal region16. The RVxF motif is also conserved in rice Rb homologues, in addition to the overall similarity among Rb homologues in animals and plants (Supplementary Fig. S15). Thus, Rb is possibly a substrate of the RSS1-interacting OsPP1 phosphatase.
Discussion
In this study, we demonstrated that RSS1 functions in the regulation of the cell cycle and is required for the maintenance of meristems in both the shoot and root under stressful environmental conditions. We propose that RSS1 functions to antagonize the G1–S checkpoint of the cell in response to stress, such that the cell cycle is slowed through the activity of cyclin-dependent kinase inhibitors2,6,7,8 (Fig. 8). This function is probably required to balance cell division and differentiation under undesirable growth conditions. In rice, cell differentiation is presumably not arrested immediately upon exposure to stress. In such a situation, the population of meristematic cells would be depleted unless cell division was maintained. Supporting this idea, the size and cell number of the MZ in rss1 roots were decreased under high-salt conditions, eventually leading to the loss of the MZ.
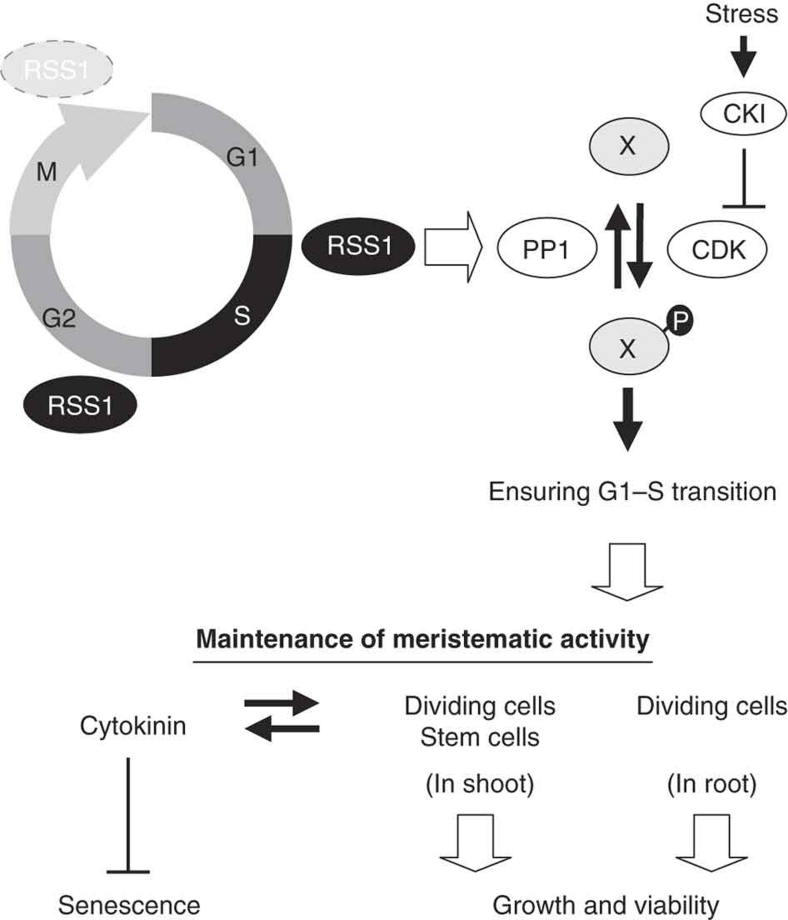
Figure 8. A possible model of the stress tolerance mechanism conferred by RSS1.
RSS1 ensures cell cycle progression, and thereby contributes to the maintenance of meristematic cells under undesirable environmental conditions. RSS1 stability is controlled by the APC/C-26S proteasome pathway, leading to RSS1 accumulation from G1/S to M phase. RSS1 functions primarily at the G1–S transition, possibly through the regulation of protein phosphatase 1 (PP1), which antagonizes the cyclin-dependent protein kinase (CDK) activity. The phosphorylation states of the substrate X are the determinants of G1–S transition (and subsequent cell cycle progression). Under stressful conditions, the G1/S checkpoint is conducted by cyclin-dependent kinase inhibitor (CKI) to inhibit the CDK activity that leads to a restrained transition to S phase, whereas RSS1 sustains cell cycle progression to balance cell division with cell differentiation. Defects in the cell cycle progression in rss1 plants affect, at least, the maintenance of proliferative tissues in the roots and the number of dividing cells and the organization of the shoot apical meristems. The latter interferes with cytokinin metabolism, which in turn influences cell division control and anti-senescence activity. Taken together, the role of RSS1 is essential for growth and viability under stressful conditions.
In the shoot, RSS1 contributes to the maintenance of the SAM size under stress conditions. This seems to be achieved not only by the maintenance of the G1–S transition, but also by the cytokinin activity in the shoots, considering the lower levels of tZ in the rss1 mutant under salinity. In agreement with this, several lines of evidence have suggested that the function of the SAM is regulated by cytokinins. Accordingly, it has been shown that the SAM cannot be maintained in the triple mutants of cytokinin receptor genes42,43. Moreover, the size of the SAM is reduced by the enhanced expression of cytokinin oxidases that degrade active cytokinins35 and by loss-of-function mutations of cytokinin biosynthetic genes44. Conversely, the size of the SAM is increased by exogenous cytokinin and by the loss of a negative regulator of cytokinin signalling36.
Under normal conditions, the cell division activity, the amount of stem cells and cytokinin metabolism are controlled interdependently32,45,46,47 (Fig. 8). Although these controls may be balanced under conditions of stress in WT, they seem to be collapsed in the rss1 mutant. Consistent with this, perturbation of the cell cycle phase-dependent expression of Arabidopsis CDKB2 has been shown to cause defects in the SAM organization concomitantly with a decrease in the levels of active cytokinins3. Moreover, the reported changes in the profiles of cytokinin metabolites in the Arabidopsis line are strikingly similar to those observed in the rss1 mutant under salinity, as represented by the decrease in the levels of tZRMP, tZR and tZ and the increase in the levels of tZOG3.
Our results suggest that RSS1 functions through an interaction with PP1, in a cell cycle-dependent manner (Fig. 8). We speculate that RSS1 interferes with the activity of cell cycle regulators controlling the G1–S transition (or the proliferation competence of the cell). In human cells, one of the best-characterized regulators of the G1–S transition is Rb, and PP1 is a known regulator of Rb. PP1 dephosphorylates and activates Rb16,40,41, which prevents the G1–S transition by the inhibitory binding to the E2F transcription factors and a DP protein48. When a cell enters into S phase, Rb is inactivated through phosphorylation by CDK, leading to the release of the E2F-DP, which, in turn, activate the expression of genes that are required for DNA replication1,2,16,48,49. In plants, Rb-related proteins have also been proposed to function at the G1–S transition, which requires the phosphorylation by CDK1,4,49,50. It will be interesting to determine whether RSS1 function is linked to the control of Rb activity.
Importantly, many PP1-binding proteins have been characterized as regulators of PP1, some of which have evolved late in their evolutionary lineages15. Considering the role of RSS1 in ensuring the G1–S transition and subsequent cell cycle progression, RSS1 may function by modulating the PP1 activity that antagonizes the CDK activity (Fig. 8). This is in line with the fact that human inhibitor-5 of PP1 promotes the G1–S transition51. Conversely, the introduction of a constitutively active PP1 (Thr320Ala) into human cancer cells was shown to cause G1 arrest13. It is also noteworthy that PP1 is inactivated by phosphorylation at the C-terminal Thr320 by CDK at the onset of S-phase52. This conserved CDK phosphorylation site is also found in a rice PP1 protein (Os06g0164100), but not in OsPP1 (Os3g0268000). Therefore, the activity of OsPP1 might be regulated not by CDK, but possibly through RSS1.
Whereas the function of RSS1 is required for G1–S transition, RSS1 is degraded during M–G1 phase by the APC/C-26S proteasome pathway. In the course of the deletion analyses, we could not obtain any transgenic plants that expressed an RSS1 variant lacking only the N-terminal 104 amino acids, which are required for destabilization. This may reflect a situation where the destabilization of RSS1 is crucial to restrict the function of RSS1 to the specific cell cycle phases. It has been reported that PP1 activity is required for mitosis53,54 and for dephosphorylation of Rb during M–G1 phase40,55. If RSS1 negatively regulated PP1 during M–G1 phase, normal cell cycle progression would be prevented.
In addition to the function that ensures cell cycle progression, RSS1 seems to have more general roles in controlling stress responses, as a weak constitutive activation of stress-responsive genes was observed in the rss1 mutant. This might also be mediated by the interaction with PP1 and by changing the phosphorylation status of intracellular signalling molecules. Conceivably, RSS1 has broader roles in protecting proliferative tissues from cellular damage by stress, on the basis of the fact that the stress-responsive expression of several genes was enhanced in rss1; this may also be explained, in part, by the antisenescence activity of cytokinins37. Alternatively, RSS1 might have another biochemical function, as has been reported for protein phosphatase inhibitor-1, which has an extra activity in the protection of cellular proteins from the denaturation induced by abiotic stresses56.
In conclusion, RSS1 functions to maintain the self-supplying activity of proliferative cells in meristems under salinity stress. RSS1, therefore, is required for continuous growth and survival under stressful environmental conditions. The loss of RSS1 homologues during the evolution of eudicots advocates the adoption of RSS1-mediated stress tolerance mechanism by specific lineages (see Supplementary Discussion). Further studies of RSS1 will provide new clues concerning the history of angiosperms22 in the context of their environmental adaptations, and will also contribute to the improvement of stress tolerance in extant plant species for agriculture.
Methods
Plant materials
Tos17-mutagenized M2 rice seeds (Oryza sativa, cv. Nipponbare; ∼2,500 lines) were surface sterilized and germinated on a sterile MS-based medium (Murashige and Skoog basal medium, 1% sucrose, 0.25% gellan gum, and 0.05% MES-KOH, pH 5.8) in the presence of 150 mM NaCl in 13-cm-high capped bottles. The seedlings were then grown at 25 °C under a photoperiod of 14 h light (4,000 lux, white light) and 10 h dark for 2–4 weeks. For comparative phenotyping and seed amplification, the same set of lines was grown in the field.
Cell cycle synchronization
Synchronized rice Oc cells for RNA analysis were prepared by treatment with 0.1 mM aphidicolin for 24 h. After removal of the drug, the cells were harvested at the S (0.5 h), G2 (10 h), M (12.5 h) and G1 phase (15.5 h)57. Untreated cells were harvested as a control. Alternatively, cells were synchronized by starvation with an 8-day incubation in the same medium. For release from G1 arrest by starvation, cells were transferred into new R2S medium (30 g/l sucrose, 4 g/l KNO3, 0.355 g/l (NH4)2SO4, 0.25 g/l MgSO4 (7H2O), 0.15 g/l CaCl2 (2H2O), 0.273 g/l NaH2PO4 (H2O), 1.6 mg/l MnSO4 (4H2O), 2.2 mg/l ZnSO4 (7H2O), 0.125 mg/l CuSO4 (5H2O), 3 mg/l H3BO3, 0.125 mg/l Na2MoO4 (2H2O), 5.5 mg/l FeSO4 (7H2O), 7.5 mg/l Na2EDTA, 0.1 g/l myo-inositol, 0.5 mg/l nicotine acid, 0.5 mg/l pyridoxine acid, 0.5 mg/l thiamine acid, 2 mg/l glycine and 2 mg/l 2.4-D, pH5.6) and cultured at 28 °C with gentle shaking at 80 r.p.m.
Microscopic observation of the root cells
Root samples were fixed, stained with propidium iodide, and treated with chloral hydrate. The tissues were then observed under a microscope (Axio Imager.M1; Carl Zeiss) or under a confocal laser-scanning microscope (FV1000D IX81; Olympus). The meristematic, elongation and maturation zones were defined as follows: the meristematic zone included small, actively dividing cells in the root apex; the elongation zone consisted of elongated cells between the meristematic and maturation zones; and the maturation zone cells were marked by root hairs proximal to the shoot.
Cloning of RSS1 and the isolation of allelic mutants
The genomic DNA fragment for RSS1 flanking the transposed Tos17 copy was isolated by PCR using XbaI-digested, self-ligated DNA as a template and the Tos17-specific primers, Tos17-3911F (5′-GAGAGCATCATCGGTTACATCTTCTC-3′) and Tos17-XbaI-R (5′-CATGAAATAGATCCATGTATATCT-3′). The amplified fragment was then subcloned into pCR2.1 (Invitrogen). Cosegregation of the Tos17 insertion with the mutant phenotype was confirmed by DNA gel blot analysis. RSS1 cDNA was isolated from a rice seedling cDNA library prepared using pADGAL4-2.1 and the HybriZAP-2.1 cDNA cloning kit (Stratagene). Nucleotide sequences were determined using a 310 or 3100 Genetic Analyzer (Applied Biosystems). rss1-2 was identified through PCR by the screening of three-dimensional pools of genomic DNA mixtures from Tos17-insertional mutant lines using RSS1- (5′-GTCTGGCCAGTCGTGCAATG-3′, 5′-CTGCTGGCAAGAGGGCTGAT-3′, 5′-TGGGTTCTTGGTGGCCTCAT-3′ and 5′-GCCAAGCCACTGAAGCCATT-3′) and Tos17-specific primers. rss1-3 (derived from Oryza sativa cv. Hitomebore) was identified by a database search for Tos17 flanking sequences (http://tos.nias.affrc.go.jp/)17. RSS1-related genes were identified by a mutual BLAST search. Multiple amino-acid sequence alignments were performed using the MAFFT program (version 6; http://mafft.cbrc.jp/alignment/server/) followed by visualization using GeneDoc (http://www.nrbsc.org/gfx/genedoc/).
Immunoblot analysis
The protein (30 μg) was purified by acetone precipitation, separated by 10% SDS-PAGE, blotted onto a nitrocellulose membrane (Trans-BlotN transfer medium; Bio-Rad Laboratories) and analysed using anti-GFP polyclonal antibodies (Code-598; Medical & Biological Laboratories).
In situ hybridization
DIG-labelled RNA probes for RSS1 and histone H4 and fluorescein (FLU)-labelled RNA probes for cyclin B2 were synthesized by in vitro transcription. Tissues were fixed in 50 mM sodium phosphate buffer (pH 7.2) containing 4% paraformaldehyde and 0.25% glutaraldehyde at 4 °C for 1 h (for roots) or overnight (for shoots). The preparation of paraffin-embedded sections (8 or 10 μm) and the in situ hybridization were carried out essentially as described26, but with some modification. For the deparaffinization, Histo-clear (National Diagnostics) was used instead of xylene. For double-colour staining, DIG-labelled RNA probes were detected by anti-DIG conjugated with alkaline phosphatase (AP) and SIGMAFAST Fast Red TR/naphthol AS-MX Tablets (Sigma-Aldrich). For the deactivation of alkaline phosphatase, sections were incubated in 2× SSC at 70 °C for 8 h. FLU-labelled RNA probes were detected by anti-FLU conjugated with AP (Roche) and NBT/BCIP.
Yeast two-hybrid experiments
Fragments of cDNAs encoding RSS1 and the activator subunits of APC/C that recognize the D-box were amplified by PCR with specific primers (RSS1: 5′-CCAACATATGGCTGCCCCAACTGCAACA-3′ and 5′-CAACGAATTCAGCCAGCTTCATTTTATCAC-3′, OsCdh1;1: 5′-CAATCATATGGCGGGGGGGCTCCGCCTC-3′ and 5′-TGATGGATCCGTATGTGGCTTCTCGAGAA-3′, OsCdh1;2: 5′-CACTCATATGGATCACCACCACCACCAC-3′ and 5′-GCTCGAGCTCGCCGGATGTAGCTCCTAACAAA-3′, and OsCdc20: 5′-CAAGGAATTCATGGATGCAGGTTCGCACTC-3′ and 5′-CTTTCTCGAGCACGGATGTGAATGTGGTTGG-3′). The amplified fragments were digested by NdeI and EcoRI (for RSS1), NdeI and BamHI (for OsCdh1;1), NdeI and SacI (for OsCdh1;2), and EcoRI and XhoI (for OsCdc20), respectively. The fragments were then introduced into the bait vector, pGBKT7 or the prey vector, pGADT7 AD. For screening of RSS1 interactors, rice cDNA libraries derived from shoot basal tissues were amplified and cloned into the prey vector (pGADT7-Rec) in yeast using Matchmaker Library Construction and Screening kits (Clontech). The interaction assay was performed according to the manufacturer's instruction manual. For selection, 1.92×106 independent yeast transformants were incubated on solid SCD media without leucine, tryptophan and histidine and with 0.1 or 0.2 mM of 3-amino-1,2,4-triazole at 30 °C for 2 to 3 days. The genes encoding the candidates for RSS1-interacting proteins were retested with different reporter genes.
Microarray and statistical data analysis
The transcriptomic profiles of rss1 were investigated using Agilent-015241 Rice Gene Expression 4×44 K Microarray (Agilent Technology). Labelling and hybridization were performed as described58. Raw microarray data were normalized by the variance stabilization normalization method using R software (http://www.r-project.org/). Data are available at Gene Expression Omnibus, accession code: GSE27884. The differentially expressed gene list is provided as Supplementary Data 1 and gene ontology analysis is presented in Supplementary Data 2. Significant differences were examined using both the Rank product (P<0.01) with the R software and two-way ANOVA (P<0.01) with the TIGR Multiexperiment Viewer (MeV; http://www.tm4.org/mev/). Affymetrix microarray data from various tissues (GSE6893) and seedlings exposed to stress (GSE6901) were downloaded from GEO. These raw data were normalized with the GCRMA (GC Robust Multi-Array) method and analysed statistically using R software. The shoot apex-specific genes were classified as those genes that were expressed at significantly higher levels in the shoot apical and rachis meristems than in young leaves, mature leaves and roots (P<0.01, rank product). Stress-responsive genes were categorized in the same manner with data obtained in this study (for 'long term') or with public microarray data for seedlings treated with or without salt, drought or cold for 3 h, respectively (for 'short-term'). The salt-inducible (or salt-repressible) genes shown in Figure 4b represent the genes commonly up- (or down)regulated under long- and short-term salt stress conditions. See Supplementary Information for the gene lists.
Ploidy analysis
Ploidy analysis was performed with Ploidy Analyser PA (Partec), following the procedures and with the buffers provided by the manufacturer. For nuclear staining, 0.5-cm-long shoot basal tissues from 7-day-old rice seedlings were excised and chopped into pieces for 60 s with a razor blade in nuclei extraction buffer and then filtered with a 30-μm CellTrics filter.
Quantification of cytokinins
The concentrations of cytokinins and their conjugates in 1-cm-long shoot basal tissues from 1-week-old seedlings were measured essentially as described59. In brief, the extract was passed through an Oasis HLB column (Waters), and the cytokinin-containing fraction was obtained by solid-phase extraction with an Oasis MCX column (Waters). Cytokinins were measured with a liquid chromatography-tandem mass chromatography system (AQUITY UPLC™ System/Quattro Ultima Pt; Waters) with an ODS column (AQUITY UPLC BEH C18, 1.7 μm, 2.1×100 mm, Waters). Data were processed by MassLynx software with QuanLynx (version 4.0, Waters).
Fluorescence microscopy and BiFC
For the construction of nYFP-RSS1, cYFP-PP1 and YFP-PP1 genes driven by the CaMV 35S promoter, cDNAs for RSS1 and PP1 were amplified by PCR using primers (RSS1: 5′-CACCATGGCTGCCCCAACTGCAAC-3′ and 5′-TCAGTTCATTCCAGGCTCCA-3′ and PP1: 5′-CACCATGGCGGCGGCACCGGGGGC-3′ and 5′-CATTTTGTTTGATGCCATAA-3′), cloned into the pENTR/D-TOPO vector, and transferred to Gateway expression vectors (nYFP/pUGW0 and cYFP/pUGW2) by LR recombination (Gateway Technology; Invitrogen)60. The pDH51-mRFP1 plasmid carrying the mRFP1 gene driven by the CaMV 35S promoter was used as a control for BiFC. Transient expression assays to assess the subcellular localization of GFP or YFP-fused proteins and mRFP in onion epidermal cells were carried out using the particle bombardment technique. For BiFC, plasmids encoding nYFP-fused proteins, cYFP-fused proteins and mRFP were mixed at a 5:5:1 (w/w) ratio and were adsorbed onto gold particles 1.6 μm in diameter, according to the manufacturer's instructions (Bio-Rad Laboratories). For each bombardment, 25 μl of DNA-coated gold particles was placed on the macrocarrier. Onion epidermal cells were bombarded using the PDS-1000/He Biolistic Particle Delivery System (Bio-Rad) at 7,584 kPa under a vacuum of 94.82 kPa. Following the bombardment, the onion tissues were incubated in the same Petri dish for 12 h at 30 °C in the dark. Onion epidermal cells were then observed under a fluorescent microscope (Axio Imager.M1; Carl Zeiss) with a set of filters capable of distinguishing between yellow (YFP) and red (RFP) fluorescence.
Further details for the above methods and other experimental procedures are described in the Supplementary Methods.
Author contributions
S.T., K.A., A.M and H.H. isolated, cloned and genetically characterized RSS1, and D.O., M.M., H.M., Y.T., T. Hobo, Y.S., T.H. and S.T. contributed to further functional analysis of RSS1. M.K. and H.S. measured cytokinin contents. D.O., T.H. and S.T. wrote the manuscript with contributions from all co-authors.
Additional information
Accession codes: Microarray data are available at the Gene Expression Omnibus, under accession code GSE27884.
How to cite this article: Ogawa, D. et al. RSS1 regulates the cell cycle and maintains meristematic activity under stress conditions in rice. Nat. Commun. 2:278 doi: 10.1038/ncomms1279 (2011).
Supplementary Material
Supplementary Discussion, Supplementary Figures S1-S15, Supplementary Table S1, Supplementary Methods and Supplementary References
Transcriptomic analysis of WT and the rss1 mutant under normal and salt-stress conditions using the Rice 44K (Agilent) microarray
Gene Ontology classification of genes up- or down-regulated in rss1 mutants
Acknowledgments
We thank K. Sugimoto, M. Yamazaki, G.K. Agrawal, T. Watanabe, K. Murata, S. Komatsu and M. Yoshii for experimental suggestions, C. Ueguchi, A. Morikami, E. Katoh and H. Tsukagoshi for helpful discussion, and A. Baba, H. Takatsuji, J. Hashimoto, T. Furukawa, S. Endo, M. Hattori, R. Motoyama, Y. Nagamura, S. Mitsuya and Y. Kitom for technical assistance. We thank T. Nakagawa and S. Mano for providing nYFP or cYFP/pUGW0 or pUGW2 vectors, and Nottingham Arabidopsis Stock Centre for providing pDH51-mRFP1 plasmid. This work was supported in part by grants from the Ministry of Agriculture, Forestry, and Fisheries of Japan (Integrated research project for plant, insect, and animal using genome technology MP-2146 and IP-5002; Genomics for Agricultural Innovation AMR0002) and from Bio-oriented Technology Research Advancement Institute of Japan, Grants-in-Aid for Scientific Research on Priority Areas (18075007), and the Center of Excellence from the Japanese Ministry of Education, Culture, Sports, Science and Technology. D.O. is supported by the fellowship of the Japan Society for the Promotion of Sciences.
References
- Inze D. & De Veylder L. Cell cycle regulation in plant development. Annu. Rev. Genet. 40, 77–105 (2006). [DOI] [PubMed] [Google Scholar]
- De Veylder L., Beeckman T. & Inze D. The ins and outs of the plant cell cycle. Nat. Rev. Mol. Cell Biol. 8, 655–665 (2007). [DOI] [PubMed] [Google Scholar]
- Andersen S. U. et al. Requirement of B2-type cyclin-dependent kinases for meristem integrity in Arabidopsis thaliana. Plant Cell 20, 88–100 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wildwater M. et al. The RETINOBLASTOMA-RELATED gene regulates stem cell maintenance in Arabidopsis roots. Cell 123, 1337–1349 (2005). [DOI] [PubMed] [Google Scholar]
- Borghi L. et al. Arabidopsis RETINOBLASTOMA-RELATED is required for stem cell maintenance, cell differentiation, and lateral organ production. Plant Cell 22, 1792–1811 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clotet J. & Posas F. Control of cell cycle in response to osmostress: lessons from yeast. Methods Enzymol. 428, 63–76 (2007). [DOI] [PubMed] [Google Scholar]
- Ruggiero B. et al. Uncoupling the effects of abscisic acid on plant growth and water relations. Analysis of sto1/nced3, an abscisic acid-deficient but salt stress-tolerant mutant in Arabidopsis. Plant Physiol. 136, 3134–3147 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peres A. et al. Novel plant-specific cyclin-dependent kinase inhibitors induced by biotic and abiotic stresses. J. Biol. Chem. 282, 25588–25596 (2007). [DOI] [PubMed] [Google Scholar]
- Takeda S. & Matsuoka M. Genetic approaches to crop improvement: responding to environmental and population changes. Nat. Rev. Genet. 9, 444–457 (2008). [DOI] [PubMed] [Google Scholar]
- Zhu J. K. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 53, 247–273 (2002). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ren Z. H. et al. A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat. Genet. 37, 1141–1146 (2005). [DOI] [PubMed] [Google Scholar]
- Goldberg J. et al. Three-dimensional structure of the catalytic subunit of protein serine/threonine phosphatase-1. Nature 376, 745–753 (1995). [DOI] [PubMed] [Google Scholar]
- Berndt N., Dohadwala M. & Liu C. W. Constitutively active protein phosphatase 1alpha causes Rb-dependent G1 arrest in human cancer cells. Curr. Biol. 7, 375–386 (1997). [DOI] [PubMed] [Google Scholar]
- Cohen P. T. Protein phosphatase 1—targeted in many directions. J. Cell Sci. 115, 241–256 (2002). [DOI] [PubMed] [Google Scholar]
- Ceulemans H. & Bollen M. Functional diversity of protein phosphatase-1, a cellular economizer and reset button. Physiol. Rev. 84, 1–39 (2004). [DOI] [PubMed] [Google Scholar]
- Hirschi A. et al. An overlapping kinase and phosphatase docking site regulates activity of the retinoblastoma protein. Nat. Struct. Mol. Biol. 17, 1051–1057 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyao A. et al. Target site specificity of the Tos17 retrotransposon shows a preference for insertion within genes and against insertion in retrotransposon-rich regions of the genome. Plant Cell 15, 1771–1780 (2003). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Munns R. & Tester M. Mechanisms of salinity tolerance. Annu. Rev. Plant Biol. 59, 651–681 (2008). [DOI] [PubMed] [Google Scholar]
- Glotzer M., Murray A. W. & Kirschner M. W. Cyclin is degraded by the ubiquitin pathway. Nature 349, 132–138 (1991). [DOI] [PubMed] [Google Scholar]
- Vodermaier H. C. APC/C and SCF: controlling each other and the cell cycle. Curr. Biol. 14, R787–R796 (2004). [DOI] [PubMed] [Google Scholar]
- Castro A., Bernis C., Vigneron S., Labbe J. C. & Lorca T. The anaphase-promoting complex: a key factor in the regulation of cell cycle. Oncogene 24, 314–325 (2005). [DOI] [PubMed] [Google Scholar]
- Frohlich M. W. & Chase M. W. After a dozen years of progress the origin of angiosperms is still a great mystery. Nature 450, 1184–1189 (2007). [DOI] [PubMed] [Google Scholar]
- Soltis D. E., Bell C. D., Kim S. & Soltis P. S. Origin and early evolution of angiosperms. Ann. N Y Acad. Sci. 1133, 3–25 (2008). [DOI] [PubMed] [Google Scholar]
- Itoh J. I., Kitano H., Matsuoka M. & Nagato Y. Shoot organization genes regulate shoot apical meristem organization and the pattern of leaf primordium initiation in rice. Plant Cell 12, 2161–2174 (2000). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fobert P. R., Coen E. S., Murphy G. J. & Doonan J. H. Patterns of cell division revealed by transcriptional regulation of genes during the cell cycle in plants. EMBO J. 13, 616–624 (1994). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kouchi H., Sekine M. & Hata S. Distinct classes of mitotic cyclins are differentially expressed in the soybean shoot apex during the cell cycle. Plant Cell 7, 1143–1155 (1995). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Menges M., de Jager S. M., Gruissem W. & Murray J. A. Global analysis of the core cell cycle regulators of Arabidopsis identifies novel genes, reveals multiple and highly specific profiles of expression and provides a coherent model for plant cell cycle control. Plant J. 41, 546–566 (2005). [DOI] [PubMed] [Google Scholar]
- Dubouzet J. G. et al. OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J. 33, 751–763 (2003). [DOI] [PubMed] [Google Scholar]
- Fukuda A. et al. Function, intracellular localization and the importance in salt tolerance of a vacuolar Na(+)/H(+) antiporter from rice. Plant Cell Physiol. 45, 146–159 (2004). [DOI] [PubMed] [Google Scholar]
- Chinnusamy V., Schumaker K. & Zhu J. K. Molecular genetic perspectives on cross-talk and specificity in abiotic stress signalling in plants. J. Exp. Bot. 55, 225–236 (2004). [DOI] [PubMed] [Google Scholar]
- Yamaguchi-Shinozaki K. & Shinozaki K. Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu. Rev. Plant Biol. 57, 781–803 (2006). [DOI] [PubMed] [Google Scholar]
- Riou-Khamlichi C., Huntley R., Jacqmard A. & Murray J. A. Cytokinin activation of Arabidopsis cell division through a D-type cyclin. Science 283, 1541–1544 (1999). [DOI] [PubMed] [Google Scholar]
- Sakakibara H. Cytokinins: activity, biosynthesis, and translocation. Annu. Rev. Plant Biol. 57, 431–449 (2006). [DOI] [PubMed] [Google Scholar]
- Kyozuka J. Control of shoot and root meristem function by cytokinin. Curr. Opin. Plant Biol. 10, 442–446 (2007). [DOI] [PubMed] [Google Scholar]
- Werner T. et al. Cytokinin-deficient transgenic Arabidopsis plants show multiple developmental alterations indicating opposite functions of cytokinins in the regulation of shoot and root meristem activity. Plant Cell 15, 2532–2550 (2003). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giulini A., Wang J. & Jackson D. Control of phyllotaxy by the cytokinin-inducible response regulator homologue ABPHYL1. Nature 430, 1031–1034 (2004). [DOI] [PubMed] [Google Scholar]
- Rivero R. M. et al. Delayed leaf senescence induces extreme drought tolerance in a flowering plant. Proc. Natl Acad. Sci. USA 104, 19631–19636 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ashikari M. et al. Cytokinin oxidase regulates rice grain production. Science 309, 741–745 (2005). [DOI] [PubMed] [Google Scholar]
- Kurakawa T. et al. Direct control of shoot meristem activity by a cytokinin-activating enzyme. Nature 445, 652–655 (2007). [DOI] [PubMed] [Google Scholar]
- Ludlow J. W., Glendening C. L., Livingston D. M. & DeCarprio J. A. Specific enzymatic dephosphorylation of the retinoblastoma protein. Mol. Cell Biol. 13, 367–372 (1993). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nelson D. A. & Ludlow J. W. Characterization of the mitotic phase pRb-directed protein phosphatase activity. Oncogene 14, 2407–2415 (1997). [DOI] [PubMed] [Google Scholar]
- Higuchi M. et al. In planta functions of the Arabidopsis cytokinin receptor family. Proc. Natl Acad. Sci. USA 101, 8821–8826 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishimura C. et al. Histidine kinase homologs that act as cytokinin receptors possess overlapping functions in the regulation of shoot and root growth in Arabidopsis. Plant Cell 16, 1365–1377 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyawaki K. et al. Roles of Arabidopsis ATP/ADP isopentenyltransferases and tRNA isopentenyltransferases in cytokinin biosynthesis. Proc. Natl Acad. Sci. USA 103, 16598–16603 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boucheron E. et al. Ectopic expression of Arabidopsis CYCD2 and CYCD3 in tobacco has distinct effects on the structural organization of the shoot apical meristem. J. Exp. Bot. 56, 123–134 (2005). [DOI] [PubMed] [Google Scholar]
- Yanai O. et al. Arabidopsis KNOXI proteins activate cytokinin biosynthesis. Curr. Biol. 15, 1566–1571 (2005). [DOI] [PubMed] [Google Scholar]
- Dewitte W. et al. Arabidopsis CYCD3 D-type cyclins link cell proliferation and endocycles and are rate-limiting for cytokinin responses. Proc. Natl Acad. Sci. USA 104, 14537–14542 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubin S. M., Gall A. L., Zheng N. & Pavletich N. P. Structure of the Rb C-terminal domain bound to E2F1-DP1: a mechanism for phosphorylation-induced E2F release. Cell 123, 1093–1106 (2005). [DOI] [PubMed] [Google Scholar]
- Boniotti M. B. & Gutierrez C. A cell-cycle-regulated kinase activity phosphorylates plant retinoblastoma protein and contains, in Arabidopsis, a CDKA/cyclin D complex. Plant J. 28, 341–350 (2001). [DOI] [PubMed] [Google Scholar]
- Shimizu-Sato S., Ike Y. & Mori H. PsRBR1 encodes a pea retinoblastoma-related protein that is phosphorylated in axillary buds during dormancy-to-growth transition. Plant Mol. Biol. 66, 125–135 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X. et al. IPP5, a novel protein inhibitor of protein phosphatase 1, promotes G1/S progression in a Thr-40-dependent manner. J. Biol. Chem. 283, 12076–12084 (2008). [DOI] [PubMed] [Google Scholar]
- Liu C. W. et al. Inhibitory phosphorylation of PP1alpha catalytic subunit during the G(1)/S transition. J. Biol. Chem. 274, 29470–29475 (1999). [DOI] [PubMed] [Google Scholar]
- Sassoon I. et al. Regulation of Saccharomyces cerevisiae kinetochores by the type 1 phosphatase Glc7p. Genes Dev. 13, 545–555 (1999). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu D. et al. Regulated targeting of protein phosphatase 1 to the outer kinetochore by KNL1 opposes Aurora B kinase. J. Cell Biol. 188, 809–820 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buchkovich K., Duffy L. A. & Harlow E. The retinoblastoma protein is phosphorylated during specific phases of the cell cycle. Cell 58, 1097–1105 (1989). [DOI] [PubMed] [Google Scholar]
- Kim T. D. Protein phosphatase inhibitor-1 (PPI-1) has protective activities in stress conditions in E. coli. Int. J. Biol. Macromol. 38, 70–76 (2006). [DOI] [PubMed] [Google Scholar]
- Takase T., Yanagawa Y., Komatsu S., Nakagawa H. & Hashimoto J. Cell-cycle-related variation in proteins in suspension-cultured rice cells. J. Plant Res. 116, 469–475 (2003). [DOI] [PubMed] [Google Scholar]
- Shimono M. et al. Rice WRKY45 plays a crucial role in benzothiadiazole-inducible blast resistance. Plant Cell 19, 2064–2076 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kojima M. et al. Highly sensitive and high-throughput analysis of plant hormones using MS-probe modification and liquid chromatography-tandem mass spectrometry: an application for hormone profiling in Oryza sativa. Plant Cell Physiol. 50, 1201–1214 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakagawa T. et al. Development of series of gateway binary vectors, pGWBs, for realizing efficient construction of fusion genes for plant transformation. J. Biosci. Bioeng. 104, 34–41 (2007). [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Discussion, Supplementary Figures S1-S15, Supplementary Table S1, Supplementary Methods and Supplementary References
Transcriptomic analysis of WT and the rss1 mutant under normal and salt-stress conditions using the Rice 44K (Agilent) microarray
Gene Ontology classification of genes up- or down-regulated in rss1 mutants