Abstract
Circadian clocks are endogenous timekeeping mechanisms that allow organisms to anticipate rhythmic, daily environmental changes. Temporal coordination of transcription results in a set of gene expression patterns with peak levels occurring at precise times of the day. An intriguing question is how a single clock can generate different oscillatory rhythms, and it has been proposed that hormone signaling might act in plants as a relay mechanism to modulate the amplitude and the phase of output rhythms. Here we show that the circadian clock gates gibberellin (GA) signaling through transcriptional regulation of the GA receptors, resulting in higher stability of DELLA proteins during daytime and higher GA sensitivity at night. Oscillation of GA signaling appears to be particularly critical for rhythmic growth, given that constitutive expression of the GA receptor expands the daily growth period in seedlings, and complete loss of DELLA function causes continuous, arrhythmic hypocotyl growth. Moreover, transcriptomic analysis of a pentuple della KO mutant indicates that the GA pathway mediates the rhythmic expression of many clock-regulated genes related to biotic and abiotic stress responses and cell wall modification. Thus, gating of GA sensitivity by the circadian clock represents an additional layer of regulation that might provide extra robustness to the diurnal growth rhythm and constitute a regulatory module that coordinates the circadian clock with additional endogenous and environmental signals.
Keywords: crosstalk, plasticity, light, photoperiod
The pervasive role of the circadian clock driving plant physiology is reflected by the extensive regulation it exerts on gene expression, as more than one third of Arabidopsis genes are under circadian control (1). Remarkably, the expression of almost every single gene of Arabidopsis cycles when plants are grown under more realistic situations, for example combinations of thermo- and photocycles (2). This suggests that the entrainment of the circadian clock by light and temperature signals might allow plants to adapt to the daily changes in the environment by timing every physiological pathway to the specific time of day when it is more advantageous. For instance, the concerted action of the circadian clock and phyB-mediated light signaling allows the expression of a set of hormone-related genes toward dawn (3), which may provide robustness to the rhythmic patterns of growth of the seedling under diurnal conditions (4). Accordingly, a correlation exists between the oscillation of auxin-related genes and changes in the hypocotyl growth rate of seedlings grown under free-running conditions (5, 6), although the physiological significance of this correlation remains to be explored. In addition to light-mediated growth, the circadian clock controls the time of day that other environmental response pathways can be activated, often by triggering the oscillation of key signaling genes involved in these pathways (7). This type of regulation is known as gating because the clock can be thought of as opening or closing a gate to control the flow of information through a signaling pathway. Through such gating the circadian clock regulates many physiological responses including the photoperiodic induction of flowering and stress responses (8).
The phytohormone gibberellin (GA) has a prominent role in the regulation of several developmental programs also affected by light and the circadian clock, including the establishment of photomorphogenesis (9–11) and cell expansion (12), and the question arises of whether GA activity might mediate circadian regulation of clock targets. Given that such a regulatory mechanism would impact the robustness and flexibility of circadian regulation of development, we decided to explore this possibility and its physiological relevance.
Results and Discussion
Expression of GA Receptors Is Controlled by the Circadian Clock.
To investigate whether the circadian clock regulates GA signaling in Arabidopsis, we examined the daily expression pattern of all known GA signaling elements in the DIURNAL database (http://diurnal.cgrb.oregonstate.edu/) (2, 13). Although a weak oscillation could be detected in some cases, mostly linked to temperature rhythms, only the GID1 receptor genes displayed robust cycling under short days (Fig. S1A). The cycling of GID1a and GID1b was validated by real-time quantitative RT-PCR in independent time-course experiments, whereas we were not able detect oscillation for GID1c transcript (Fig. 1A). The anticipation of changes in transcript levels to the light-to-dark and dark-to-light transitions and the oscillation under continuous light in entrained seedlings (Fig. S1B) suggested circadian rather than diurnal regulation. This was confirmed by analyzing mRNA levels in mutants defective for clock function, toc1-1 (14) and lhy (15). The waveform of the oscillation in toc1-1 seedlings was different from the WT, and the peak was narrower and was phased earlier (Fig. 1F). The phase advance is typical of toc1-1 mutants, as a result of the deviation between their endogenous period (21 h) and the length of the day (24 h) (16). Transcript levels of both genes were altered also in the arrhythmic mutant lhy (Fig. 1K).
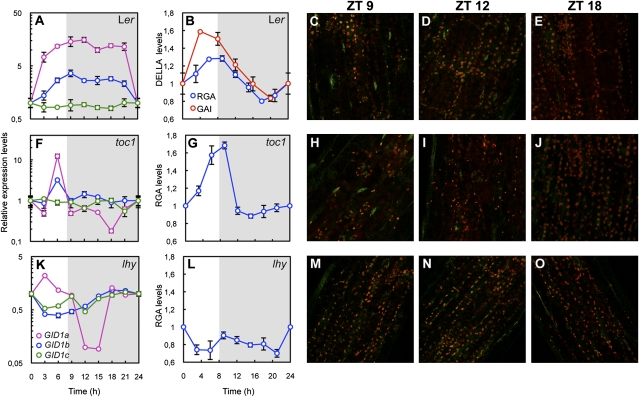
Fig. 1.
The circadian clock controls the diurnal oscillation of DELLA proteins in the cell expansion zone of hypocotyls. Expression of GID1a, GID1b, and GID1c in 5-d-old Ler WT seedlings (A), in toc1-1 (F), and in lhy (K) mutants grown under short-day photocycles (8 h light/16 h dark). Values are expressed relative to PP2a expression. In B–E, G–J, and L–O, seedlings carrying the 35S::TAP-GAI or RGA::GFP-RGA constructs were grown for 5 d under short-day photocycles (8 h light/16 h dark). DELLA protein levels in the Ler WT (B) and in the toc1-1 (G) and lhy (J) mutants were determined by Western blot analysis. TAP-GAI and GFP-RGA proteins were detected with commercial antibodies against the myc tag and GFP, respectively. DELLA levels were normalized against levels of DET3, which was used as loading control. Data are average of three independent experiments and plotted as mean ± SEM. Protein level at ZT0 was set to 1 and used as reference for all other time points. White and gray areas represent day and night, respectively. Fluorescence of GFP-RGA oscillates in the upper part of hypocotyls of Ler WT (C–E) and toc1-1 mutant seedlings (H–J), but not in the lhy mutant (M–O). Fluorescence was detected by confocal microscopy. Images are representative of three independent biological repeats including 12 to 15 seedlings per time point and per genotype.
The expression of GA receptor genes is known to respond to endogenous GA levels through a DELLA-mediated feedback mechanism, i.e., their expression increases when GA levels are low and decreases when hormone levels are high (17). Hence, oscillation of GID1 transcript levels might be a direct consequence of the circadian clock activity or, alternatively, it might respond to a putative oscillation of GA levels. However, GID1a expression was not altered in seedlings of the quadruple della mutant (rga-t2 gai-t6 rgl1-1 rgl2-1) (18, 19) (Fig. S2), indicating that the circadian clock controls the expression of GID1 genes independently of the status of the GA pathway.
Levels of DELLA Proteins Oscillate with a Daily Rhythm.
GID1 receptors are known to interact with DELLA proteins in a GA-dependent way and promote their degradation (17). Thus, if the oscillation of GID1 expression is physiologically relevant, it should cause coherent changes in DELLA accumulation with a daily rhythm. We focused our attention in two DELLA proteins, GA INSENSITIVE (GAI) and REPRESSOR OF ga1-3 (RGA), which are the most abundant DELLAs in young seedlings and shoots (20). To monitor the level of these proteins, we used lines that express the RGA::GFP-RGA (21) or 35S::TAP-GAI (22) transgenes. Consistent with the clock regulation of GID1 genes, both GFP-RGA and TAP-GAI protein levels oscillated in a diurnal manner, showing peak levels at the end of the light period (Fig. 1B). Strikingly, GFP-RGA oscillation was detected in the growing region of the hypocotyls (Fig. 1 C–E and H–J). Fluorescence from the fusion protein accumulated in nuclei of the uppermost part of hypocotyls at ZT9, whereas it was below the detection limit late in the night, at ZT18 (Fig. 1 C–E), coinciding with periods of minimum and maximum growth rates, respectively (4). The periodicity of RGA accumulation must largely be caused by the activity of GID1 receptors, as the RGA transcript did not show significant oscillation (Fig. S1C). Thus, given (i) the major role of RGA and GAI in controlling growth (23, 24) and (ii) that seedling growth under diurnal conditions is gated by the circadian clock (4), this result suggests that DELLA proteins are regulatory components for the control of the clock output, such as daily growth rhythm in young seedlings (4).
The oscillation of GFP-RGA levels was affected also in clock mutants. The waveform of GFP-RGA oscillation in toc1-1 seedlings was slightly different from the WT, the amplitude was higher and the peak narrower because of an advance in the phase of the trough (Fig. 1 G–J), according to the phase advance observed in the expression of GA receptor genes (Fig. 1F). On the contrary, GFP-RGA protein levels were constant and low in the lhy mutant (Fig. 1 L–O), which correlates with the long hypocotyl phenotype observed in this mutant when grown in short days (3).
Circadian Clock Gates GA Signaling Activity.
The observation that lower DELLA protein levels coincide with higher growth rates at the end of the night suggests that they participate in the core mechanism that controls rhythmic growth of hypocotyls. To test this hypothesis, we examined the impact upon growth of an alteration of the normal rhythm of GA signaling with two complementary approaches. First, we used a transgenic line that expresses a dominant version of GAI under the control of a heat-shock inducible promoter, HS::gai-1D (9). This line allowed us to block GA signaling by applying a 10-min heat shock at 33 °C at two different times of the day: ZT5, when the growth rate is low and DELLA levels high; and ZT17, which coincides with the beginning of the growing phase and with the trough of DELLA levels (Fig. 2A). The effectiveness of the treatments was confirmed by expression analysis of gai-1D and one of its known direct targets, AtGA20ox2 (Fig. 2B). Interestingly, blocking GA activity at ZT5 during four consecutive days did not have any effect upon hypocotyl growth (Fig. 2C). On the contrary, the heat treatment had a strong inhibitory effect on the hypocotyl growth of HS::gai-1D seedlings when applied at ZT17 (Fig. 2C).
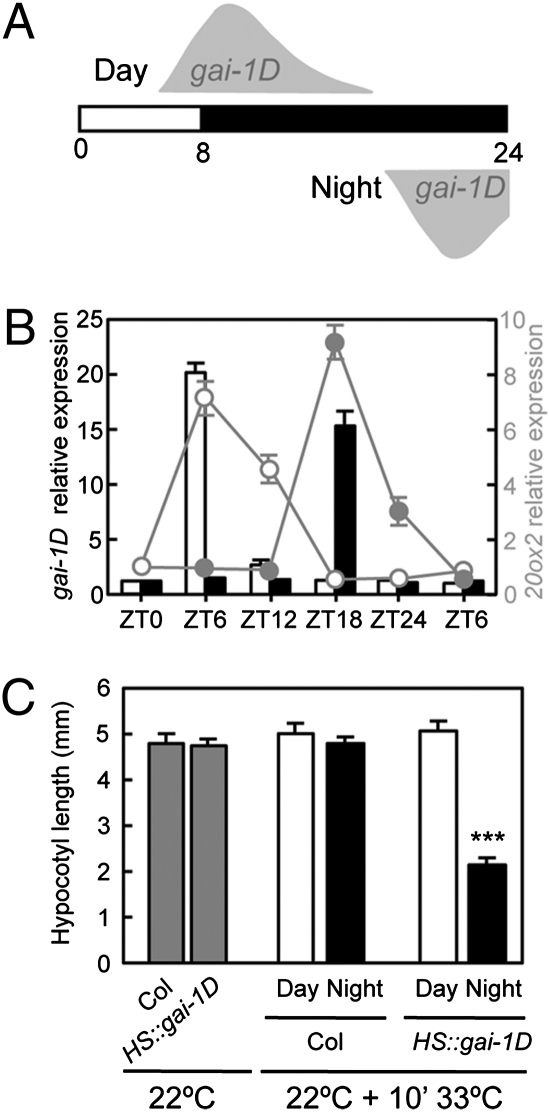
Fig. 2.
Blocking GA signaling at night affects hypocotyl growth. Seedlings of the HS::gai-1D line were grown under short-day photocycles (8 h light/16 h dark) and received heat treatments of 33 °C for 10 min at ZT5 or ZT17 (Materials and Methods). In A, shaded areas mark the period of the day during which gai-1D accumulates. (B) Expression of gai-1D (bars) and its target gene GA20ox2 (circles, scale on right) after heat treatments at ZT5 (white symbols) and ZT17 (dark symbols). (C) Hypocotyl length of Col-0 WT and HS::gai-1D seedlings that did not receive heat treatments (gray bars) or that received treatments at ZT5 (white bars) or ZT17 (black bars). The experiment was repeated three times with similar results. Data represent the mean ± SD (n ≥ 15 seedlings), and asterisks indicate P < 0.0001.
Second, we examined how transient application of GA4 at two different times of the day would rescue the dwarfism caused by continuous incubation with paclobutrazol (PAC), a compound that causes accumulation of DELLA proteins (Fig. 3A and B). GA application provoked rapid degradation of DELLA proteins that lasted for the next 10 h: between ZT1 and ZT10 when the GA treatment was applied at dawn, and between ZT13 and ZT22 when applied at ZT12 (Fig. 3B). Importantly, GA treatment applied at ZT12 during two consecutive days (fourth and fifth) significantly alleviated the growth-repressing effects of PAC, whereas it had no effect when applied at dawn (Fig. 3C). In summary, these results confirm that there is a DELLA-sensitive period that overlaps the growing phase of the night, and that under short-day conditions the circadian clock might allow growth by preventing accumulation of DELLA proteins during that particular period.
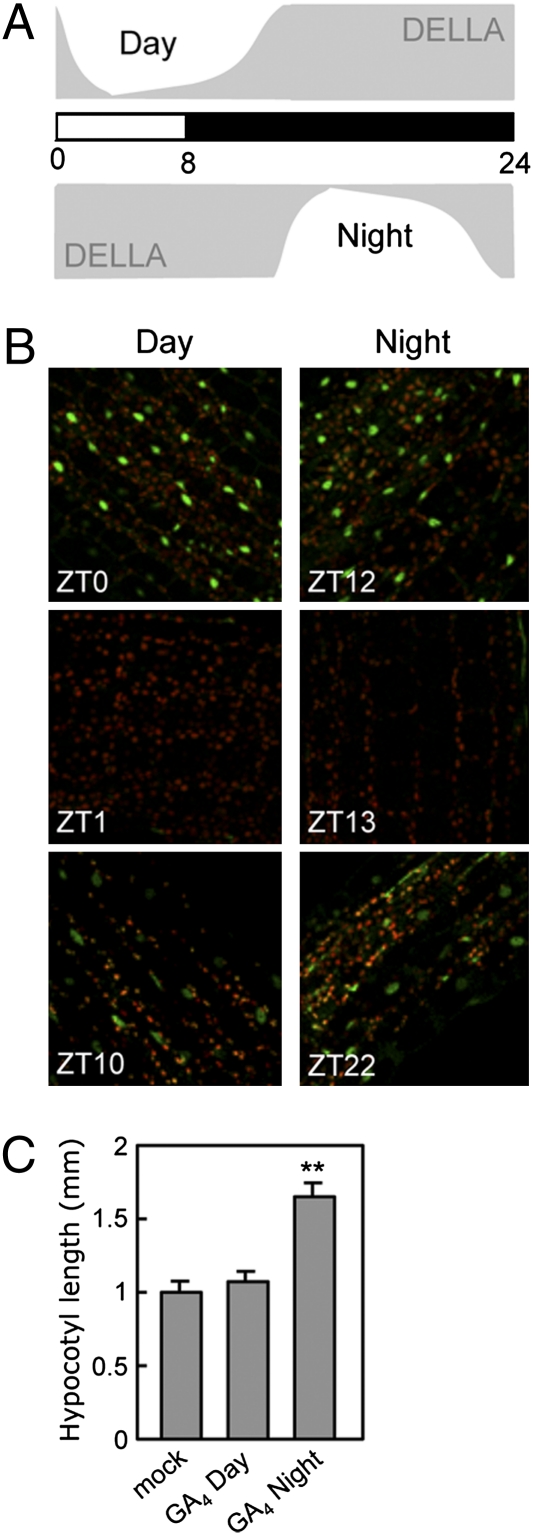
Fig. 3.
GA application at night releases the growth restrain imposed by DELLAs. RGA::GFP-RGA seedlings grown under short-day photocycles (8 h light/16 h dark) in the presence of 0.2 μM PAC were treated with 1 μM GA4 at ZT0 or ZT12 or untreated (Materials and Methods). (A) Scheme of DELLA accumulation after GA4 treatments, deduced from the GFP-RGA fluorescence of seedlings grown under the same conditions (B). Confocal images taken at the time of GA4 treatment (ZT0 and ZT12) and 1 h (ZT1 and ZT13) and 10 h later (ZT10 and ZT22) show that the maximum period with low DELLA levels spans less than 10 h. Images are representative of three independent biological repeats including eight to 10 seedlings per time point. (C) Hypocotyl length of WT (Ler) seedlings grown in the presence of 0.2 μM PAC that did not receive any additional treatment (mock) or that were treated with 1 μM GA4 at ZT0 (day) or ZT12 (night). The WT seedlings contain the RGA::GFP-RGA transgene. Data represent mean ± SD (n ≥ 15 seedlings). Asterisks indicate P < 0.001.
Oscillation of GA Signaling Refines Rhythmic Growth.
If the oscillation of GA signaling constitutes part of the mechanism that ensures rhythmic growth, a prediction of this model is that GA signaling mutants should display not only a defect in the final size of the hypocotyl (12), but also an altered rhythmic growth pattern. In agreement with this, transgenic plants that expressed GID1a under the control of the 35S promoter showed an expanded growth phase that started at the same time as in the WT and extended well into daytime, almost spanning the whole light period (Fig. 4A). In agreement with this, plants overexpressing GID1a, GID1b, or GID1c displayed longer hypocotyls compared with the WT (P < 0.05, Student t test; Fig. S3A), and seedlings of gid1a-1, gid1b-1, and gid1c-1 loss-of-function mutants, and of the different double mutant combinations (17), had shorter hypocotyls than WT (P < 0.05, Student t test; Fig. S3B). Taken together, these results indicate that GID1 expression is limiting for promotion of hypocotyl elongation under diurnal conditions and that oscillation of GID1 is necessary for the establishment of proper patterns of rhythmic growth.
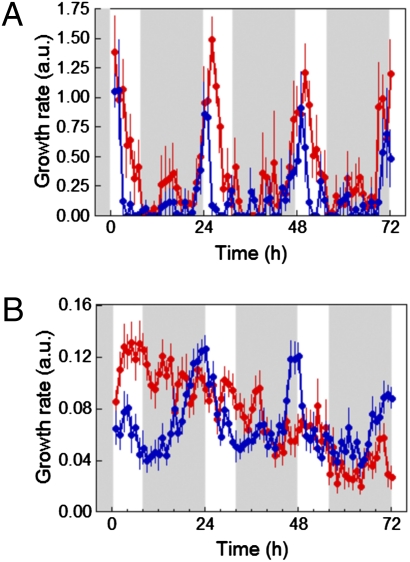
Fig. 4.
GA activity regulates diurnal rhythms of hypocotyl elongation. Col 0 and 35S::GID1a seedlings (A) and Ler and quintuple della mutant seedlings (B) were grown under short-day photocycles (8 h light/16 h dark) for 3 d before they were imaged under the same conditions for three additional days. Blue and red symbols and lines denote the WT and mutant/transgenic seedlings, respectively. Seedlings’ growth rates were measured as described in Materials and Methods. White and gray areas represent day and night, respectively. Data represent mean ± SD (n ≥ 10 seedlings). a.u., arbitrary units.
Nonetheless, seedlings overexpressing GID1a still exhibited robust rhythmic growth. Given that this behavior cannot be attributed to circadian regulation of GID1 protein stability (Fig. S4), it could be taken as an indication that GA levels oscillate in a diurnal or in a clock-controlled manner. In fact, expression of several genes encoding GA metabolic enzymes oscillate diurnally in Arabidopsis (25, 26), in a manner that would likely result in lower GA levels at dusk; and diurnal changes in GA levels have been described in sorghum (27).
More clear evidence for an integral role of GA signaling in the establishment of rhythmic growth was provided by the quintuple della KO mutant (22). Growth of this mutant was completely arrhythmic under short days, showing a relatively high growth rate during the first day examined that progressively decreased over the next few days (Fig. 4B). This arrhythmic phenotype is not likely caused by a dysfunction of the core clock mechanism, as the expression of the clock genes TOC1 and CCA1 was not significantly affected in the mutant (Fig. S5). DELLAs have been proposed to regulate cell expansion through the inhibition of PIF's activity (22, 28). The observation that pif4 pif5 double mutants are impaired in rhythmic growth (4) suggests that the DELLA-PIF interaction might thus constitute the main output pathway that controls rhythmic growth. However, given that DELLAs also exert part of their action through the HY5 transcription factor (9) and hy5 mutants display arrhythmic growth (4), it is also possible that other components different from PIF might also be part of this regulatory module.
DELLA Proteins Mediate Daily Rhythms of Gene Expression.
GA signaling participates in the regulation of multiple developmental and physiological processes other than cell expansion (29). It is therefore possible that the role of DELLA proteins as regulatory components of the output of the circadian clock extends beyond the control of rhythmic growth and affects other processes. To test this hypothesis, we performed a genome-wide search for genes that would oscillate in a circadian manner in WT seedlings, and whose oscillation would be affected in a della KO mutant. For this purpose, we examined global gene expression in short-day–grown WT and della mutant seedlings at ZT9 (high DELLA levels) and ZT21 (low DELLA levels). In this search, we found 5,087 genes whose expression varied between both time points in the WT. We found that this list was significantly enriched (66%; P < 2.2 × 10−16 by Fisher exact test) for genes differentially expressed across time points in a previously published full short-day data set (2, 13), indicating that our approximation to identify genes that oscillated with a specific diurnal phase was acceptable. To ask what role DELLA genes play in diurnal regulation of gene expression, we looked for genes that were differentially expressed when della mutants were compared with WT. We found 58 genes differentially expressed between della and WT; 37 of these were only found at ZT9 and four were found at ZT21, consistent with DELLAs being most important at ZT9 (Fig. 5A and Dataset S1). Furthermore, the 37 genes affected by the della KO at ZT9 were significantly enriched for diurnally regulated genes (70%; P = 0.001 by Fisher exact test), whereas the remaining differentially expressed genes were not enriched.
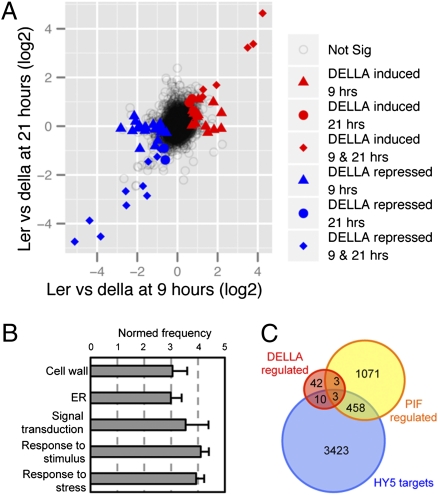
Fig. 5.
DELLAs mediate circadian regulation of transcription. (A) Scatter plot of genes differentially regulated at ZT21 versus ZT9 in WT Ler and della mutants. Genes showing statistically significant (FDR < 0.1) differential expression between Ler and della are displayed for each time point in blue and red. (B) Enrichment of gene ontology categories among genes regulated by DELLAs at ZT9 (P < 0.0001 in all cases). (C) Venn diagram showing overlap between genes regulated by DELLAs at ZT9; genes directly bound by HY5, as detected by ChIP-chip experiments in light-grown seedlings (30); and genes regulated by PIF transcription factors, as genes differentially expressed in the quadruple pif1 pif3 pif4 pif5 mutant in darkness and light, with respect to WT (31).
Importantly, Gene Ontology analysis showed that, among the genes whose oscillation was altered in the della mutant, there was an enrichment in functional categories related to the response to stress and environmental signals, as well as in genes encoding proteins located in the cell wall and the endoplasmic reticulum (Fig. 5B). Although part of the genes represented in the “cell wall” category could include those with a function in growth and cell expansion, the enrichment of other growth-unrelated categories suggests that DELLAs mediate the regulation of a larger array of circadian clock-controlled processes. These results attribute a more general role to DELLA proteins in the modulation of the output of the clock. Moreover, meta-analysis of the DELLA targets at ZT9 (Fig. 5C) indicated that only 45% of these genes were either direct targets for HY5 (30) or genes also regulated by the PIF transcription factors (31, 32). This implies that DELLA proteins control gene expression through the interaction with additional transcription factors, which is in agreement with the observation that DELLA proteins can interact with several members of the bHLH family of transcription factors other than PIFs (33, 34).
Concluding Remarks.
The enormous plasticity of plant growth and development is based on a web of interacting signaling pathways, which provides the plant with multiple entry points to adjust their physiology in response to frequent, unpredicted environmental changes (35). The circadian system, on the contrary, provides the plant with the ability to anticipate predictable daily and seasonal environmental changes (36) and buffers plant responses against casual environmental variability (37). The circadian clock therefore provides stability to plants’ life. Then, can responses regulated by the circadian clock be plastic? We suggest that the regulation of GA activity by the circadian clock might provide such ability, acting as a link between two properties critical for plant growth and development, robustness, and plasticity (38). Thus, the concurrency of clock and GA regulation of certain processes guarantees a precise and robust response to unpredicted, transitory, and above-noise changes in the environment that have an impact on the GA pathway, such as nutrient availability (39), salt stress (18), ambient temperature (40), or flooding (41). The observation that the functioning of the circadian clock does not seem to be affected significantly by GA (ref. 42 and the present study) supports the suggested role of GA as a regulatory output module that fine-tunes clock-regulated gene expression in response to environmental signals.
Materials and Methods
Plant Material.
Arabidopsis thaliana accessions Col-0 and Ler were used as WT. Seeds of gid1a-1, gid1b-1, gid1c-1, gid1a-1 gid1b-1, gid1b-1 gid1c-1, gid1a-1 gid1c-1, toc1-1, lhy, rga-t2 gai-t6 rgl1-1 rgl2-1, rga-t2 gai-t6 rgl1-1 rgl2-1 rgl3-1, RGA::GFP-RGA, 35S::TAP-GAI, and HS::gai-1D have been previously described (9, 15, 17, 18, 21, 22, 43). RGA::GFP-RGA toc1-1 and RGA::GFP-RGA lhy lines were obtained by genetic crosses and isolated from an F3 population.
Plasmid Constructs and Transgenic Plants.
The preparation of transgenic lines expressing 35S::GID1a-YFP-HA, 35S::GID1b-YFP-HA, or 35S::GID1c-YFP-HA was as follows. Coding sequences of GID1a-1c, excluding the stop codon, were PCR-amplified with Pwo polymerase (Roche) from cDNA obtained from 7-d-old, light-grown WT Col-0 seedlings. Oligonucleotides used as primers for PCR (Dataset S2) included the attB sites needed for Gateway-mediated cloning and were designed to allow expression of a C-terminal fusion. PCR products were first cloned into vector pDONR-221 (Invitrogen) by BP recombination, and then transferred to the binary vector pEarleyGate-101 (44) by LR recombination to create a C-terminal fusion with YFP and HA-tag. The final constructs were transferred to WT Col-0 plants by Agrobacterium-mediated transformation. Primary transformants were selected in MS plates containing 50 mM glufosinate ammonium (Fluka). Transgenic lines with a 3:1 (resistant:sensitive) segregation ratio were selected, and several homozygous lines were identified in the T3 generation for each construction.
Seedling Growth Assays.
All seeds were surface-sterilized with 70% (vol/vol) ethanol and 0.01% (vol/vol) Triton X-100 for 5 min, followed by 96% (vol/vol) ethanol for 5 min. Seeds were sown on plates of 0.5 MS medium (Duchefa) and 0.8% (wt/vol) agar without sucrose, and stratified at 4 °C in darkness for 5 d. Germination was induced by placing the plates under white fluorescent light (190–200 μmol m−2 s−1) at 22 °C for 8 h. Seedlings were grown at 22 °C under short-day photocycles, 8 h light (70–200 μmol m−2s−1, depending on the experiment)/16 h dark in an E-30B growth cabinet (Percival).
For heat-shock experiments, WT Col-0 and HS::gai-1D (9) seedlings grown in the same plate under short days (70 μmol m−2 s−1) received a heat treatment (10 min at 33 °C in darkness) at either ZT5 or ZT17. Control seedlings of both genotypes were kept at 22 °C. Heat treatments were applied at days 3, 4, 5, and 6. Hypocotyl length was measured on day 7.
For GA sensitivity assays, RGA::GFP-RGA seeds were sown on sterile filter papers placed on 0.5 MS, 0.8% (wt/vol) agar plates without sucrose, stratified, and induced to germinate as described earlier. After induction of germination, filter papers harboring seeds were transferred to treatment plates containing 0.2 μM PAC (Duchefa) and grown under short days (190–200 μmol m−2 s−1) at 22 °C for 3 d. Filter papers containing 4-d-old seedlings were transferred at either ZT0 or ZT12 for 1 h to Petri dishes with 5 mL of 0.5 MS liquid media containing 0.1 μM GA4 (Sigma) plus 0.2 μM PAC, or just 0.2 μM PAC. Filter papers containing seedlings were then rinsed three times for 20 min in Petri dishes containing 0.5 MS with PAC 0.2 μM. After washes, seedlings were transferred into a new sterile filter paper, placed on a fresh 0.2-μM PAC treatment plate, and returned to short-day conditions. GA treatments were given during two consecutive days (fourth and fifth), and hypocotyl length was measured on day 6. Handling of seedlings during the dark period was performed under a safe green light.
To measure hypocotyl length, seedlings were placed on an acetate sheet and scanned at a resolution of 600 dpi, and the length was measured with ImageJ software.
The time-lapse photography and image analysis to determine hypocotyl growth rate was performed as previously described (4), except that the growth medium contained 0.5 MS and 1% sucrose.
RNA Extraction and Gene Expression Analysis by Quantitative RT-PCR.
Total RNA was isolated from whole seedlings grown as described earlier (190–200 μmol m−2 s−1) by using the E.Z.N.A. Plant RNA Mini Kit (Omega Bio-tek) according to the manufacturer´s instructions. cDNA synthesis and quantitative PCR conditions were as described (9). Primers used are listed in Table S2.
Microarray Analysis.
WT Ler and rga-t2 gai-t6 rgl1-1 rgl2-1 rgl3-1 pentuple della mutant seedlings were grown under short days (190–200 μmolm−2s−1) at 22 °C, and seedlings were sampled at ZT9 and ZT12 of day 5. Total RNA from whole seedlings was extracted with RNeasy Plant Mini kit (Qiagen). RNA labeling and hybridization to Affymetrix ATH1 arrays were performed by the Nottingham Arabidopsis Stock Centre. Analysis was performed in R (45) and Bioconductor (46). Microarrays were normalized with the RMA procedure as implemented in the affy package (47), and differential expression was determined using limma (48, 49) with a false discovery rate lower than 0.05. To determine genes whose expression varies in diurnal short-day conditions, the previously published data set (13) was downloaded from Array Express (http://www.ebi.ac.uk/arrayexpress/; accession E-MEXP-1304), Robust Multichip Average-normalized, and analyzed in limma by using a one-way ANOVA model with time as the grouping variable. Samples from each of the 2 d of collection were used as replicates (so, in total, there were two replicates for each of six time points). Gene annotations were based on the TAIR9 version of the Arabidopsis Web site (http://arabidopsis.org). Gene Ontology analysis was performed with the Classification SuperViewer tool at the Bio-Array Resource of the University of Toronto.
Protein Extraction and Western Blot Analysis.
Protein extraction and Western blot analysis from whole 5-d-old seedlings grown under short days (190–200 μmol m−2 s−1) were performed as described. The GFP, TAP, and HA fusion proteins were detected using anti-GFP (JL8; Clontech), anti–c-myc (9E10; Roche), and anti-HA (3F10; Roche) antibodies, respectively. Antibodies against DET3 were used to check protein loading (50). Signal from bound antibodies was revealed using ECL Advance Western Blotting detection Kit (GE Healthcare) and visualized and quantified by using the Luminescence Image Analyzer LAS-3000 (Fujifilm) and Image Gauge (version 4.0; Fujifilm), respectively.
Confocal Imaging.
Fluorescence from the GFP-RGA fusion protein was detected by using a confocal microscope (TCS SL; Leica) as previously described (40).
Supplementary Material
Acknowledgments
We are indebted to the Nottingham Arabidopsis Stock Centre, Stephen G. Thomas, Tai-ping Sun, and Nicholas P. Harberd for providing us with seeds. M.V.A. was the recipient of a postdoctoral contract from the Spanish Ministry of Science and Innovation for the mobility of young researchers into Spanish universities. N.A.M. holds a Consejo Superior de Investigaciones Científicas Fellowship of the JAE-Pre program. This study was supported by Spanish Ministry of Science and Innovation Grants BIO2007-60923, BIO2010-15071, and CSD2007-00057; Generalitat Valenciana Grants ACOMP/2010/190; and National Science Foundation Grants DBI0820854 and IOS0923752.
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
Data deposition: The microarrays reported in this paper have been deposited in Nottingham Arabidopsis Stock Centre (code: NASCArray 587).
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1101050108/-/DCSupplemental.
References
- 1.Pruneda-Paz JL, Kay SA. An expanding universe of circadian networks in higher plants. Trends Plant Sci. 2010;15:259–265. doi: 10.1016/j.tplants.2010.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Michael TP, et al. Network discovery pipeline elucidates conserved time-of-day-specific cis-regulatory modules. PLoS Genet. 2008;4:e14. doi: 10.1371/journal.pgen.0040014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Michael TP, et al. A morning-specific phytohormone gene expression program underlying rhythmic plant growth. PLoS Biol. 2008;6:e225. doi: 10.1371/journal.pbio.0060225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Nozue K, et al. Rhythmic growth explained by coincidence between internal and external cues. Nature. 2007;448:358–361. doi: 10.1038/nature05946. [DOI] [PubMed] [Google Scholar]
- 5.Covington MF, Harmer SL. The circadian clock regulates auxin signaling and responses in Arabidopsis. PLoS Biol. 2007;5:e222. doi: 10.1371/journal.pbio.0050222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Dowson-Day MJ, Millar AJ. Circadian dysfunction causes aberrant hypocotyl elongation patterns in Arabidopsis. Plant J. 1999;17:63–71. doi: 10.1046/j.1365-313x.1999.00353.x. [DOI] [PubMed] [Google Scholar]
- 7.Harmer SL, et al. Orchestrated transcription of key pathways in Arabidopsis by the circadian clock. Science. 2000;290:2110–2113. doi: 10.1126/science.290.5499.2110. [DOI] [PubMed] [Google Scholar]
- 8.de Montaigu A, Tóth R, Coupland G. Plant development goes like clockwork. Trends Genet. 2010;26:296–306. doi: 10.1016/j.tig.2010.04.003. [DOI] [PubMed] [Google Scholar]
- 9.Alabadí D, et al. Gibberellins modulate light signaling pathways to prevent Arabidopsis seedling de-etiolation in darkness. Plant J. 2008;53:324–335. doi: 10.1111/j.1365-313X.2007.03346.x. [DOI] [PubMed] [Google Scholar]
- 10.Alabadí D, Gil J, Blázquez MA, García-Martínez JL. Gibberellins repress photomorphogenesis in darkness. Plant Physiol. 2004;134:1050–1057. doi: 10.1104/pp.103.035451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Achard P, et al. DELLAs contribute to plant photomorphogenesis. Plant Physiol. 2007;143:1163–1172. doi: 10.1104/pp.106.092254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cowling RJ, Harberd NP. Gibberellins control Arabidopsis hypocotyl growth via regulation of cellular elongation. J Exp Bot. 1999;50:1351–1357. [Google Scholar]
- 13.Mockler TC, et al. The DIURNAL project: DIURNAL and circadian expression profiling, model-based pattern matching, and promoter analysis. Cold Spring Harb Symp Quant Biol. 2007;72:353–363. doi: 10.1101/sqb.2007.72.006. [DOI] [PubMed] [Google Scholar]
- 14.Millar AJ, Straume M, Chory J, Chua NH, Kay SA. The regulation of circadian period by phototransduction pathways in Arabidopsis. Science. 1995;267:1163–1166. doi: 10.1126/science.7855596. [DOI] [PubMed] [Google Scholar]
- 15.Schaffer R, et al. The late elongated hypocotyl mutation of Arabidopsis disrupts circadian rhythms and the photoperiodic control of flowering. Cell. 1998;93:1219–1229. doi: 10.1016/s0092-8674(00)81465-8. [DOI] [PubMed] [Google Scholar]
- 16.Strayer C, et al. Cloning of the Arabidopsis clock gene TOC1, an autoregulatory response regulator homolog. Science. 2000;289:768–771. doi: 10.1126/science.289.5480.768. [DOI] [PubMed] [Google Scholar]
- 17.Griffiths J, et al. Genetic characterization and functional analysis of the GID1 gibberellin receptors in Arabidopsis. Plant Cell. 2006;18:3399–3414. doi: 10.1105/tpc.106.047415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Achard P, et al. Integration of plant responses to environmentally activated phytohormonal signals. Science. 2006;311:91–94. doi: 10.1126/science.1118642. [DOI] [PubMed] [Google Scholar]
- 19.Cheng H, et al. Gibberellin regulates Arabidopsis floral development via suppression of DELLA protein function. Development. 2004;131:1055–1064. doi: 10.1242/dev.00992. [DOI] [PubMed] [Google Scholar]
- 20.Tyler L, et al. Della proteins and gibberellin-regulated seed germination and floral development in Arabidopsis. Plant Physiol. 2004;135:1008–1019. doi: 10.1104/pp.104.039578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Silverstone AL, et al. Repressing a repressor: Gibberellin-induced rapid reduction of the RGA protein in Arabidopsis. Plant Cell. 2001;13:1555–1566. doi: 10.1105/TPC.010047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Feng S, et al. Coordinated regulation of Arabidopsis thaliana development by light and gibberellins. Nature. 2008;451:475–479. doi: 10.1038/nature06448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Dill A, Sun T. Synergistic derepression of gibberellin signaling by removing RGA and GAI function in Arabidopsis thaliana. Genetics. 2001;159:777–785. doi: 10.1093/genetics/159.2.777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.King KE, Moritz T, Harberd NP. Gibberellins are not required for normal stem growth in Arabidopsis thaliana in the absence of GAI and RGA. Genetics. 2001;159:767–776. doi: 10.1093/genetics/159.2.767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Hisamatsu T, King RW, Helliwell CA, Koshioka M. The involvement of gibberellin 20-oxidase genes in phytochrome-regulated petiole elongation of Arabidopsis. Plant Physiol. 2005;138:1106–1116. doi: 10.1104/pp.104.059055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhao X, et al. A study of gibberellin homeostasis and cryptochrome-mediated blue light inhibition of hypocotyl elongation. Plant Physiol. 2007;145:106–118. doi: 10.1104/pp.107.099838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lee IJ, Foster KR, Morgan PW. Photoperiod control of gibberellin levels and flowering in sorghum. Plant Physiol. 1998;116:1003–1011. doi: 10.1104/pp.116.3.1003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.de Lucas M, et al. A molecular framework for light and gibberellin control of cell elongation. Nature. 2008;451:480–484. doi: 10.1038/nature06520. [DOI] [PubMed] [Google Scholar]
- 29.Sun TP. Gibberellin-GID1-DELLA: a pivotal regulatory module for plant growth and development. Plant Physiol. 2010;154:567–570. doi: 10.1104/pp.110.161554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Lee J, et al. Analysis of transcription factor HY5 genomic binding sites revealed its hierarchical role in light regulation of development. Plant Cell. 2007;19:731–749. doi: 10.1105/tpc.106.047688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Leivar P, et al. Definition of early transcriptional circuitry involved in light-induced reversal of PIF-imposed repression of photomorphogenesis in young Arabidopsis seedlings. Plant Cell. 2009;21:3535–3553. doi: 10.1105/tpc.109.070672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Shin J, et al. Phytochromes promote seedling light responses by inhibiting four negatively-acting phytochrome-interacting factors. Proc Natl Acad Sci USA. 2009;106:7660–7665. doi: 10.1073/pnas.0812219106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Arnaud N, et al. Gibberellins control fruit patterning in Arabidopsis thaliana. Genes Dev. 2010;24:2127–2132. doi: 10.1101/gad.593410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gallego-Bartolomé J, et al. Transcriptional diversification and functional conservation between DELLA proteins in Arabidopsis. Mol Biol Evol. 2010;27:1247–1256. doi: 10.1093/molbev/msq012. [DOI] [PubMed] [Google Scholar]
- 35.Casal JJ, Fankhauser C, Coupland G, Blázquez MA. Signalling for developmental plasticity. Trends Plant Sci. 2004;9:309–314. doi: 10.1016/j.tplants.2004.04.007. [DOI] [PubMed] [Google Scholar]
- 36.Harmer SL. The circadian system in higher plants. Annu Rev Plant Biol. 2009;60:357–377. doi: 10.1146/annurev.arplant.043008.092054. [DOI] [PubMed] [Google Scholar]
- 37.Troein C, Locke JC, Turner MS, Millar AJ. Weather and seasons together demand complex biological clocks. Curr Biol. 2009;19:1961–1964. doi: 10.1016/j.cub.2009.09.024. [DOI] [PubMed] [Google Scholar]
- 38.Alabadí D, Blázquez MA. Molecular interactions between light and hormone signaling to control plant growth. Plant Mol Biol. 2009;69:409–417. doi: 10.1007/s11103-008-9400-y. [DOI] [PubMed] [Google Scholar]
- 39.Jiang C, Gao X, Liao L, Harberd NP, Fu X. Phosphate starvation root architecture and anthocyanin accumulation responses are modulated by the gibberellin-DELLA signaling pathway in Arabidopsis. Plant Physiol. 2007;145:1460–1470. doi: 10.1104/pp.107.103788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Stavang JA, et al. Hormonal regulation of temperature-induced growth in Arabidopsis. Plant J. 2009;60:589–601. doi: 10.1111/j.1365-313X.2009.03983.x. [DOI] [PubMed] [Google Scholar]
- 41.Hattori Y, et al. The ethylene response factors SNORKEL1 and SNORKEL2 allow rice to adapt to deep water. Nature. 2009;460:1026–1030. doi: 10.1038/nature08258. [DOI] [PubMed] [Google Scholar]
- 42.Hanano S, Domagalska MA, Nagy F, Davis SJ. Multiple phytohormones influence distinct parameters of the plant circadian clock. Genes Cells. 2006;11:1381–1392. doi: 10.1111/j.1365-2443.2006.01026.x. [DOI] [PubMed] [Google Scholar]
- 43.Millar AJ, Carré IA, Strayer CA, Chua NH, Kay SA. Circadian clock mutants in Arabidopsis identified by luciferase imaging. Science. 1995;267:1161–1163. doi: 10.1126/science.7855595. [DOI] [PubMed] [Google Scholar]
- 44.Earley KW, et al. Gateway-compatible vectors for plant functional genomics and proteomics. Plant J. 2006;45:616–629. doi: 10.1111/j.1365-313X.2005.02617.x. [DOI] [PubMed] [Google Scholar]
- 45.R Development Core Team . R: A Language and Environment for Statistical Computing. Vienna: R Foundation for Statistical Computing; 2010. [Google Scholar]
- 46.Reimers M, Carey VJ. Bioconductor: An open source framework for bioinformatics and computational biology. Methods Enzymol. 2006;411:119–134. doi: 10.1016/S0076-6879(06)11008-3. [DOI] [PubMed] [Google Scholar]
- 47.Gautier L, Cope L, Bolstad BM, Irizarry RA. affy—analysis of Affymetrix GeneChip data at the probe level. Bioinformatics. 2004;20:307–315. doi: 10.1093/bioinformatics/btg405. [DOI] [PubMed] [Google Scholar]
- 48.Smyth GK. Limma: linear models for microarray data. In: Gentleman R, Carey V, Dudoit S, Irizarry R, Huber W, editors. Bioinformatics and Computational Biology Solutions Using R and Bioconductor. New York: Springer; 2005. pp. 397–420. [Google Scholar]
- 49.Smyth GK. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol. 2004;3:Article 3. doi: 10.2202/1544-6115.1027. [DOI] [PubMed] [Google Scholar]
- 50.Duek PD, Elmer MV, van Oosten VR, Fankhauser C. The degradation of HFR1, a putative bHLH class transcription factor involved in light signaling, is regulated by phosphorylation and requires COP1. Curr Biol. 2004;14:2296–2301. doi: 10.1016/j.cub.2004.12.026. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.