Summary
During development, all cells make the decision to live or die. While the molecular mechanisms that execute the apoptotic program are well defined, less is known about how cells decide whether to live or die. In C. elegans, this decision is linked to how cells divide asymmetrically [1, 2]. Several classes of molecules are known to regulate asymmetric cell divisions in metazoans, yet these molecules do not appear to control C. elegans divisions that produce apoptotic cells [3]. We identified CNT-2, an Arf GAP protein of the AGAP family, as a novel regulator of this type of neuroblast division. Loss of CNT-2 altered daughter cell size and caused the apoptotic cell to adopt the fate of its sister cell, resulting in extra neurons. CNT-2’s Arf GAP activity was essential for its function in these divisions. The N-terminus of CNT-2, which contains a GTPase-like domain that defines the AGAP class of Arf GAPs, negatively regulates CNT-2’s function. We provide evidence that CNT-2 regulates receptor-mediated endocytosis and consider the implications of its role in asymmetric cell divisions.
Results and Discussion
Loss of CNT-2 results in the production of extra neurons
The two C. elegans Q.p neuroblasts divide during the first larval stage to produce a posterior daughter (Q.pp) that dies and an anterior daughter (Q.pa) that divides to produce a mechanosensory neuron (AVM on the right; PVM on the left) and an SDQ interneuron [1] (Figure 1A). The Q.p division requires PIG-1, the homolog of the stem cell regulator MELK [4]. pig-1 mutants produce extra A/PVM and SDQ neurons because Q.pp survives and adopts the fate of its sister (Figure 1B left). This phenotype differs from that caused by mutations in pro-apoptotic genes where Q.pp survives but does not divide (Figure 1B right) [4, 5].
Figure 1.
cnt-2 mutations disrupt the asymmetric divisions of the Q.p neuroblast. (A) The Q.p lineage. (B) The Q.p lineage in cell fate and cell death mutant animals. (C) Frequency of extra A/PVM neurons in cnt-2 mutants. All of the animalss carried the Pmec-4::gfp transgene zdIs5. cnt-2 mutant lineages either produced a single A/PVM or two A/PVMs (% Q.p lineages with extra A/PVMs). Animals were mutant for both maternal and zygotic cnt-2 (m−z−) or only for zygotic cnt-2 (m+z−). For this and subsequent graphs, the number of Q.p lineages scored is above each genotype. (D, E) Alleles analyzed were cnt-2(gm377), ced-4(n1162) and arf-1(ok796). Animals contained the Pegl-17::gfp transgene ayIs9. (D) Ratio of the Q.p daughter cell sizes. Above the bars are representative photomicrographs and drawings of Q.p daughter cells. (E) Fluorescence photomicrographs from time-lapse confocal recordings of wild-type, cnt-2 and arf-1 Q.p divisions. Also see supplemental movies. Numbers at the bottom left corner of each frame represents the time in seconds. Arrows indicate the position of the cleavage furrow. Normally, the QL.a cell migrates over the top of Q.p towards the tail and then divides. QL.ap, the PQR neuron, continues to migrate toward the tail, but QL.aa dies. The arrowhead indicates a QL.aa cell that did not die. Scale bar represents 2 m.
In screens for mutants with extra A/PVMs, we identified the gene cnt-2, which encodes an Arf GAP (GTPase activating protein) of the AGAP family (Arf GAP with G protein-like domain, Ankyrin repeat and PH domain) [6]. cnt-2 encodes three isoforms (A–C) that contain a G protein-like domain (GLD), a split pleckstrin homology (PH) domain and two ankyrin repeats (Figure 2A, B) [7]. Four mutant alleles exist (Figure 2A, B). The zd168 and gm390 nonsense mutations were identified in forward genetic screens. The gm377 and tm2328 deletions were isolated by reverse genetic approaches.
Figure 2.
CNT-2 and the role of its GAP domain in the Q.p division. (A) Structure of CNT-2B. GLD: GTPase-Like Domain; PH and DM: split Pleckstrin Homolog Domain; GAP: Arf GTPase Activating Protein domain; A: Ankyrin repeats. (B) Structure of the A, B and C mRNAs. Boxes represent exons, lines introns. Shaded boxes represent coding sequences; the open box represents the 5′ UTR of isoform A. The slashes in intron 1 of the B isoform indicate that intron length is larger than drawn. The tm2328 and gm377 deletions are predicted to shift the reading frame. (C and D) Effects of cnt-2 transgenes on the Q.p lineage. See Figure 1C for presentation. The mutation that changes a conserved arginine to lysine at position 709 (CNT-2B(RK)::GFP) abrogated the GAP activity of AGAP1 while still allowing the cognate Arf to bind [9]. The cysteine to serine changes at positions 681 and 684 within the zinc-finger motif (CNT-2-2B(CCSS)::GFP) are predicted to render the Arf GAP incapable of binding its cognate Arf [9, 10]. We generated one transgenic line for cnt-2B, five for cnt-2B::gfp, two each for the mutated versions of cnt-2B::gfp and ΔNcnt-2B::gfp, and three for ΔNcnt-2B::gfp. Each of the lines for a particular construct was tested and gave similar results. Data for only one line of each type is presented. GFP levels were similar for all of the transgenes. * P < 0.0001 (two-sample proportion test).
All cnt-2 mutants displayed a similar recessive extra A/PVM phenotype (Figure 1C). If cnt-2 mutations transform the cell fated to die into its sister, these mutations should also produce extra SDQ neurons. Indeed, 52% (n=150) of gm377 and 48% (n=290) of zd168 mutant lineages produced extra SDQs. Loss of cnt-2 also altered other asymmetric divisions that produce apoptotic cells (Table S1; Supplemental Information). Analysis of genetic interactions between cnt-2 and either pig-1 or ham-1, which is necessary for some of the asymmetric divisions that require pig-1 and cnt-2, suggest that they could act in the same pathway (Table S2; Supplemental Information).
Loss of CNT-2 affects Q.p neuroblast’s daughter size
Q.p daughters differ in size, with Q.pp fated to die being smaller than its mitotic sister Q.pa [4, 8]. The wild-type precursor is approximately four times the size of its sister, whereas cnt-2 Q.p daughters are equivalent in size (Figure 1D, 1E, Supplemental movies 1 and 2). We considered the possibility that apoptosis caused Q.pp to contract, resulting in the size asymmetry and ruled out this concern in two ways. First, we analyzed the sizes of the Q.p daughters in a ced-4 mutant where Q.pp survives and found that Q.pa was still more than three times the size of its sister (Figure 1D). Second, we followed the Q.p division using time-lapse confocal microscopy. The wild-type cleavage furrow was displaced toward the posterior, resulting in different sized daughter cells; the cnt-2 mutant furrow formed in a central position (Figure 1E, Supplemental movies 1 and 2).
CNT-2 Arf GAP activity is essential for its function
Of the three CNT-2 isoforms, B isoform expression with (CNT-2B::GFP) or without a GFP tag (CNT-2B) showed the most effective rescue of the extra A/PVM defect (Figure 2C and data not shown). To ask if CNT-2 activity depends on a functional GAP domain, we changed a conserved arginine to lysine at position 709 (CNT-2B(RK)::GFP), which abrogates AGAP1’s GAP activity while still allowing the cognate Arf to bind [9]. We also changed cysteine to serine at positions 681 and 684 (CNT-2B (CCSS)::GFP), which should render the Arf GAP incapable of binding its cognate Arf [9, 10]. Both changes abolished CNT-2B’s rescuing activity (Figure 2C).
We also asked whether excess CNT-2 caused an A/PVM phenotype. Expression of full-length wild-type or mutant CNT-2 did not (Figure 2C), but expression of the N-terminal deletion mutant ΔNCNT-2::GFP, which lacks the GLD domain, produced animals with extra A/PVMs (Figure 2D) and Q.p daughters that were more equivalent in size (Figure 3A). Mutations in pro-apoptotic genes enhance the A/PVM defect of pig-1 mutants or of animals treated with cnt-2 RNAi [4, 11, 12] (data not shown), and a ced-4 mutation enhanced the A/PVM phenotype caused by ΔNCNT-2::GFP (Figure 2D). The requirement for Arf GAP activity (Figure 2D), however, suggests that ΔNCNT-2::GFP does not act as a dominant negative. Moreover, the ability of ΔNCNT-2::GFP expression to partially rescue the A/PVM phenotype of cnt-2 mutants shows that the transgenic protein has normal CNT-2 activity (Figure 2D). Experiments presented below support the hypothesis that ΔNCNT-2::GFP possesses deregulated ArfGAP activity.
Figure 3.
Cell autonomous roles and genetic interactions for arf-1 and cnt-2. Alleles used were arf-1(ok796), arf-6(tm1447), ced-3(n717), ced-4(n1162) and cnt-2(gm377). (A) Ratio of the Q.p daughter cell sizes. See Figure 1D for presentation of results. (B, C) See Figure 1C for presentation of results. (B) Expression of a cnt-2 or an arf-1 cDNA from the mab-5 promoter rescued the QL (PVM) but not the QR (AVM) defects of cnt-2 or arf-1; ced-3 mutant animals, respectively. The frequency of extra AVMs (open bars) and PVMs (solid bars) are presented separately. (C) The roles of arf-1 and arf-6 in the Q.p division. (A, C) Because arf-1 and cnt-2 are linked on LG III, unc-32(e189) and dpy-18(e364) are visible markers used to construct the arf-1 cnt-2 double mutant. ^The arf-1 cnt-2 chromosome also contained an unc-32 mutation. #The cnt-2 control in C contained the unc-32 and dpy-18 mutations. * P < 0.0001, ** P <0.3, ***P <0.01.
CNT-2 functions cell autonomously
CNT-2 could regulate either secretion of a signal to Q.p or a membrane trafficking event in Q.p. To determine where CNT-2 acts, we expressed cnt-2B from the mab-5 promoter. mab-5 encodes a transcription factor expressed in cells near the tail [13, 14]. With the exception of Q descendents, bilaterally symmetric cells express mab-5 on both the left and right. The left Q.p, which generates PVM, but not the right Q.p, which generates AVM, expresses mab-5 [15]. If cnt-2 acts in the Q lineage, its expression from the mab-5 promoter should rescue the PVM but not the AVM defect of cnt-2 mutants. If cnt-2 acts in cells that signal to Q.p, its expression from the mab-5 promoter should either rescue or fail to rescue both the PVM and AVM defects, depending on whether CNT-2 acts in mab-5 expressing cells. Consistent with CNT-2 acting in the Q lineage, cnt-2B expression from the mab-5 promoter rescued the PVM but not the AVM defects (Figure 3B). We also placed the Pmab-5::cnt-2B transgene into a cnt-2; egl-20 mutant background. Transcription of mab-5 in the QL lineage but not in other cells requires the Wnt EGL-20 [16]. Loss of egl-20 abolished the ability of the Pmab-5::cnt-2 transgene to rescue the PVM defect of the cnt-2 mutant, consistent with CNT-2 acting in the Q lineage (data not shown).
CNT-2 functions in endocytosis
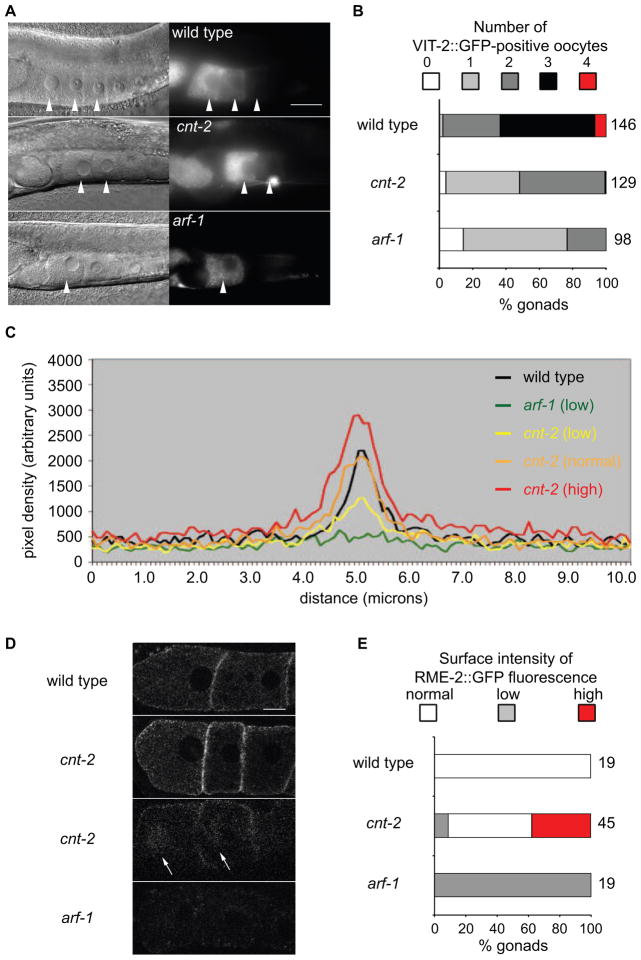
To address how CNT-2 regulates trafficking, we asked whether cnt-2 mutants exhibited defects in receptor-mediated endocytosis [17]. The intestine secretes vitellogenins, which are internalized by the RME-2 receptor on mature oocytes. Endocytosis-defective oocytes fail to internalize vitellogenins, resulting in their accumulation in the pseudocoelom and in their absence from mature oocytes. Tagged vitellogenin VIT-2::GFP accumulated in the pseudocoelom of cnt-2 mutants (not shown). Compared to wild-type oocytes, fewer cnt-2 oocytes contained VIT-2::GFP (Figure 4A, B).
Figure 4.
CNT-2 regulates receptor-mediated endocytosis. arf-1(ok796) and cnt-2(gm377) were used. (A) Nomarski (left panels) and fluorescence photomicrographs (right panels) of wild-type, cnt-2, and arf-1 animals containing [vit-2::gfp]. Arrowheads indicate the positions of individual oocytes. The scale bar represents 30 microns. (B) Quantification of the number of VIT-2::GFP-positive oocytes in a gonad arm. To the right of the bars are the number of gonad arms scored. P < 0.0001 for both arf-1 and cnt-2 compared to wild-type using a two-sample test for proportions. (C) Plots of line scans through the first two oocytes of wild-type, arf-1 and cnt-2 mutants that contain the rme-2::gfp transgene. ImageJ was used to draw a line perpendicular to, and centered on, the boundary between the first and the second oocyte on 12bit depth confocal images. The plot profile for this line was recorded, and five independent profiles were averaged for each gonad. Based on the maximum intensity of fluorescence (Imax) measured in arbitrary units (AU), gonads were grouped into low (Imax<1000 AU), normal (1000AU<Imax<2500AU) or high (Imax>2500) fluorescence classes. The three classes of RME-2::GFP surface intensity quantified in E are depicted here. The range of the normal class is illustrated by the wild-type and cnt-2 scans. (D) Confocal images RME-2::GFP in wild-type and mutant oocytes. Arrows indicate juxtanuclear dots. The scale bar represents 10 microns. The number of oocytes scored is to the right of the bar. (E) Quantification of RME-2::GFP levels at the oocyte cell surface. P < 0.0001 for both arf-1 and cnt-2 compared to wild-type using a two-sample test for proportions. Figure S2 describes the effects of interactions between arf-1 and cnt-2 on RME-2::GFP distribution.
Aberrant RME-2 trafficking can alter VIT-2::GFP distribution [18]. More RME-2::GFP accumulated at the cell surface of cnt-2 than wild-type oocytes (Figure 4C–E), and RME-2::GFP localized occasionally to juxta-nuclear dots that were observed in endocytosis mutants (Figure 4D) [18]. Taken together, the VIT-2::GFP and RME-2::GFP phenotypes suggest that CNT-2 regulates endocytosis. To test the hypothesis that compromised endocytosis contributes to the cnt-2 asymmetric division phenotypes, we asked whether reducing the levels of endocytosis molecules by RNAi affected the Q.p division. We used an rrf-3 mutant background to sensitize the animals to the effects of RNAi [19], and a weak ced-3(n2436) mutation. RNAi of endocytosis genes resulted in lethality, but escapers of the dynamin homolog dyn-1(RNAi) and rab-5(RNAi) resulted in significant numbers of extra A/PVMs (p < 0.001): 21% (n=66) for dyn-1 and 20% (n=133) for rab-5. The negative control produced 7% extra A/PVMs (n=197), and cnt-2(RNAi) produced 49% extra A/PVMs (n=218). Rab5 homologs specifically mediate endocytic trafficking [20], supporting the hypothesis that CNT-2 regulates endocytic events that control the Q.p division.
The small GTPase genes arf-1 and arf-6 regulate the Q.p division
The essential role for the GAP domain of CNT-2 indicates that arfs or arf-like (arl) genes regulate the Q.p division. Sequence analysis defines three Arf classes: Class I (Arf1–3), Class II (Arf4–5), and the more divergent Class III (Arf6) [21]. C. elegans has a single representative of each class [22]. The C. elegans genome also encodes 10 Arls.
Knocking down the function of each of the arf or arl homologs by RNAi in the rrf-3; ced-3(n2436) mutant background produced extra A/PVMs with arf-1, arf-3 and arf-6 (S. Cordes and J. Teuliere, unpublished observations). A single deletion allele of each arf gene exists. The arf-3 mutants arrest as larvae and do not have an A/PVM phenotype either alone or in a ced-3 mutant background (data not shown). Off-target effects could explain the discrepancy between the RNAi and mutant phenotypes since arf-3 is closely related to arf-1. Alternatively, maternal arf-3 provided by the heterozygous mothers could mask a role for arf-3 in the Q.p division. By contrast, arf-1 and arf-6 mutants displayed a weak extra A/PVM phenotype and enhanced the extra A/PVM phenotype of ced-3 or ced-4 mutants (Figure 3C). Because arf-1 interacted genetically with cnt-2 (see below), we addressed its role in the Q.p division: arf-1 regulates Q.p daughter cell size (Figures 1E, 3A and Supplemental movie 3) and acts autonomously in the Q lineage (Figure 3B).
arf-1 and cnt-2 genetic interactions
The arf-6 mutation failed to interact with either the cnt-2 mutant or ΔNCNT-2::GFP (Figure 3C). Loss of arf-1, by contrast, suppressed the extra A/PVM and size asymmetry defects of cnt-2 mutants (Figure 3B, C). Suppression of the extra A/PVMs defect resulted from an A/PVM differentiation defect (Figure S1; Supplemental Information).
To address the possibility that CNT-2 is an ARF-1 GAP, we asked whether arf-1 and cnt-2 mutants have similar oocyte phenotypes. RNAi of arf-1 disrupts VIT-2::GFP uptake into oocytes [17]. VIT-2::GFP accumulated in the pseudocoelom of the arf-1 mutant, and fewer mutant oocytes contained VIT-2::GFP (Figure 4A, B, data not shown). We rarely detected RME-2::GFP in arf-1 mutants, suggesting that it may not be trafficked to the plasma membrane (Figure 4C–E).
Distinct RME-2::GFP phenotypes support a model where ARF-1 and CNT-2 act in antagonistic cycles. This model predicts that an arf-1 cnt-2 double mutant should alter the RME-2::GFP phenotypes of the single mutants. As the model predicts, knocking down cnt-2 resulted in arf-1 oocytes with more RME-2::GFP at the cell surface (Figure S2; Supplemental Information).
The arf-1 mutation also interacted genetically with ΔNCNT-2::GFP, enhancing the daughter cell size and extra A/PVM phenotypes (Figure 3A, C). The opposite effects of arf-1 loss on cnt-2 mutants and transgenic animals confirm that ΔNCNT-2::GFP does not simply interfere with cnt-2 function. We propose that the GLD of CNT-2 negatively regulates CNT-2 function. Deregulated GAP activity of ΔNCNT-2::GFP might enhance endocytosis, leading to an asymmetric cell division defect. Excessive or decreased endocytosis might disrupt Q.p asymmetry. Alternatively, the N-terminus of CNT-2 might restrict its activity to a particular membrane compartment, the ΔNCNT-2::GFP phenotype resulting from CNT-2 acting in an inappropriate trafficking event.
The requirement for CNT-2 GAP function argues that Arfs or Arls regulate the Q.p division. Why then were we unable to identify a cognate GTPase? Eliminating CNT-2 or its cognate GTPase should disrupt a specific membrane trafficking step and result in a similar phenotype. If correct, one explanation for the discrepancy is CNT-2 regulates ARF-3, but the contribution of maternal arf-3 masked its role in mutant animals. Alternatively, CNT-2 could regulate more than one GTPase. Elimination of two or more GTPases would generate a phenotype similar to that of cnt-2 mutants. The GTPases could provide overlapping functions in the same trafficking event or could mediate distinct events. Consistent with the latter possibility, the two mammalian CNT-2 homologs interact with different adaptors that mediate distinct trafficking events [23]. As the sole C. elegans AGAP, CNT-2 could mediate both events. This model is consistent with the mixed oocyte phenotype of cnt-2 mutants: some oocytes accumulate excess cell surface RME-2::GFP, revealing an endocytosis defect, whereas other oocytes have low RME-2::GFP levels, revealing a distinct defect.
Membrane trafficking and development
Trafficking of membrane receptors plays important roles in developmental decisions. Defects in Notch trafficking can lead to inappropriate activation of the receptor [24]. Components of the trafficking machinery can also exhibit specificity for particular cargo. Neural tube closure requires specific interactions between the Van Gogh homolog Vangl2 and the sec24b subunit of the COPII complex to transport Vangl2 from the ER to the Golgi [25]. Whether CNT-2 exhibits specificity for particular cargo is unclear. Notch and Wnt signaling regulate asymmetric cell divisions in Drosophila and C. elegans, respectively [26, 27]. The involvement of these molecules in asymmetric cell divisions raise the interesting possibility that CNT-2 regulates Notch or Wnt pathways.
Experimental Procedures
C. elegans genetics and DNA manipulations are described in the Supplemental Experimental Procedures section.
Lineage analysis
Animals were maintained at 22°C. L1 larvae were anesthetized with 0.1% levamisole in M9 buffer and mounted on 2% agar pads. Images were acquired on a Zeiss Axiovert 200M microscope with a 100x 1.45 NA oil objective and an EM charge-coupled C9100-13 device camera (Hamamatsu Photonics) using the 488nm line of a CSU10 argon laser attached to a spinning-disk confocal scan head (Yokogawa; obtained from Solamere, Inc.). Images were acquired every 90s using μManager software (www.micro-manager.org).
Oocyte endocytosis
L4 larvae were collected and observed as young adults 24 hours later. Fluorescence images of bIs1[vit-2::GFP] gonads were obtained using an Axioskop 2 microscope (Carl Zeiss, Inc.). Images were collected using an ORCA-ER CCD camera (Hamamatsu Photonics) and Openlab imaging software (Improvision). Confocal sections of pwIs116[rme-2::RME-2-GFP] oocytes were obtained using a laser scanning 510 Meta confocal microscope (Carl Zeiss, Inc.) equipped with a 100x 1.3 NA oil objective and the 488-nm line of a LGK 7812 ML4 argon laser (Lasos Lasertechnik). 1024 × 1024 single monodirectional line scans were performed at medial positions of gonads to obtain 12bit images for light intensity measurements.
For Q.p daughter cell sizes, QL.pa and QL.pp cell areas were measured in L1 larvae that had their left sides up. Areas were measured in triplicate using ImageJ. The size ratio was calculated using average area values. QL.pa and QL.pp were imaged only when the Q.pp did not appear apoptotic, was not rounded and was still attached to Q.pa.
Supplementary Material
Acknowledgments
We thank Yuji Kohara for providing cnt-2 cDNAs, Shohei Mitani and the National Biosource Project for providing the cnt-2(tm2328) mutant, and Nancy Hawkins for providing unpublished observations. Some nematode strains used in this work were provided by the Caenorhabditis Genetics Center, which is funded by the NIH National Center for Research Resources (NCRR). This work was supported by National Institutes of Health grants NS39397 to S.C. and NS42213 to G.G. R.V. was supported by the Howard Hughes Medical Institute and NIH grant R37GM38499 G.O. was supported by a fellowship from the Damon Runyun Cancer Research Foundation, and J.T. by a fellowship from the Association pour la Recherche sur le Cancer.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Sulston JE, Horvitz HR. Post-embryonic cell lineages of the nematode, Caenorhabditis elegans. Dev Biol. 1977;56:110–156. doi: 10.1016/0012-1606(77)90158-0. [DOI] [PubMed] [Google Scholar]
- 2.Sulston JE, Schierenberg E, White JG, Thomson JN. The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev Biol. 1983;100:64–119. doi: 10.1016/0012-1606(83)90201-4. [DOI] [PubMed] [Google Scholar]
- 3.Singhvi A, Garriga G. Asymmetric divisions, aggresomes and apoptosis. Trends Cell Biol. 2009;19:1–7. doi: 10.1016/j.tcb.2008.10.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Cordes S, Frank CA, Garriga G. The C. elegans MELK ortholog PIG-1 regulates cell size asymmetry and daughter cell fate in asymmetric neuroblast divisions. Development. 2006;133:2747–2756. doi: 10.1242/dev.02447. [DOI] [PubMed] [Google Scholar]
- 5.Ellis HM, Horvitz HR. Genetic control of programmed cell death in the nematode C. elegans. Cell. 1986;44:817–829. doi: 10.1016/0092-8674(86)90004-8. [DOI] [PubMed] [Google Scholar]
- 6.Kahn RA, Bruford E, Inoue H, Logsdon JM, Jr, Nie Z, Premont RT, Randazzo PA, Satake M, Theibert AB, Zapp ML, et al. Consensus nomenclature for the human ArfGAP domain-containing proteins. J Cell Biol. 2008;182:1039–1044. doi: 10.1083/jcb.200806041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Randazzo PA, Hirsch DS. Arf GAPs: multifunctional proteins that regulate membrane traffic and actin remodelling. Cell Signal. 2004;16:401–413. doi: 10.1016/j.cellsig.2003.09.012. [DOI] [PubMed] [Google Scholar]
- 8.Ou G, Stuurman N, D’Ambrosio M, Vale RD. Polarized myosin produces unequal-size daughters during asymmetric cell division. Science. 2010;330:677–680. doi: 10.1126/science.1196112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Nie Z, Stanley KT, Stauffer S, Jacques KM, Hirsch DS, Takei J, Randazzo PA. AGAP1, an endosome-associated, phosphoinositide-dependent ADP-ribosylation factor GTPase-activating protein that affects actin cytoskeleton. J Biol Chem. 2002;277:48965–48975. doi: 10.1074/jbc.M202969200. [DOI] [PubMed] [Google Scholar]
- 10.Cukierman E, Huber I, Rotman M, Cassel D. The ARF1 GTPase-activating protein: zinc finger motif and Golgi complex localization. Science. 1995;270:1999–2002. doi: 10.1126/science.270.5244.1999. [DOI] [PubMed] [Google Scholar]
- 11.Frank CA, Hawkins NC, Guenther C, Horvitz HR, Garriga G. C. elegans HAM-1 positions the cleavage plane and regulates apoptosis in asymmetric neuroblast divisions. Dev Biol. 2005;284:301–310. doi: 10.1016/j.ydbio.2005.05.026. [DOI] [PubMed] [Google Scholar]
- 12.Guenther C, Garriga G. Asymmetric distribution of the C. elegans HAM-1 protein in neuroblasts enables daughter cells to adopt distinct fates. Development. 1996;122:3509–3518. doi: 10.1242/dev.122.11.3509. [DOI] [PubMed] [Google Scholar]
- 13.Costa M, Weir M, Coulson A, Sulston J, Kenyon C. Posterior pattern formation in C. elegans involves position-specific expression of a gene containing a homeobox. Cell. 1988;55:747–756. doi: 10.1016/0092-8674(88)90131-6. [DOI] [PubMed] [Google Scholar]
- 14.Cowing DW, Kenyon C. Expression of the homeotic gene mab-5 during Caenorhabditis elegans embryogenesis. Development. 1992;116:481–490. doi: 10.1242/dev.116.2.481. [DOI] [PubMed] [Google Scholar]
- 15.Salser SJ, Kenyon C. Activation of a C. elegans Antennapedia homologue in migrating cells controls their direction of migration. Nature. 1992;355:255–258. doi: 10.1038/355255a0. [DOI] [PubMed] [Google Scholar]
- 16.Maloof JN, Whangbo J, Harris JM, Jongeward GD, Kenyon C. A Wnt signaling pathway controls hox gene expression and neuroblast migration in C. elegans. Development. 1999;126:37–49. doi: 10.1242/dev.126.1.37. [DOI] [PubMed] [Google Scholar]
- 17.Grant B, Hirsh D. Receptor-mediated endocytosis in the Caenorhabditis elegans oocyte. Mol Biol Cell. 1999;10:4311–4326. doi: 10.1091/mbc.10.12.4311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Balklava Z, Pant S, Fares H, Grant BD. Genome-wide analysis identifies a general requirement for polarity proteins in endocytic traffic. Nat Cell Biol. 2007;9:1066–1073. doi: 10.1038/ncb1627. [DOI] [PubMed] [Google Scholar]
- 19.Simmer F, Tijsterman M, Parrish S, Koushika SP, Nonet ML, Fire A, Ahringer J, Plasterk RH. Loss of the putative RNA-directed RNA polymerase RRF-3 makes C. elegans hypersensitive to RNAi. Curr Biol. 2002;12:1317–1319. doi: 10.1016/s0960-9822(02)01041-2. [DOI] [PubMed] [Google Scholar]
- 20.Zerial M, McBride H. Rab proteins as membrane organizers. Nat Rev Mol Cell Biol. 2001;2:107–117. doi: 10.1038/35052055. [DOI] [PubMed] [Google Scholar]
- 21.Kahn RA, Cherfils J, Elias M, Lovering RC, Munro S, Schurmann A. Nomenclature for the human Arf family of GTP-binding proteins: ARF, ARL, and SAR proteins. J Cell Biol. 2006;172:645–650. doi: 10.1083/jcb.200512057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Li Y, Kelly WG, Logsdon JM, Jr, Schurko AM, Harfe BD, Hill-Harfe KL, Kahn RA. Functional genomic analysis of the ADP-ribosylation factor family of GTPases: phylogeny among diverse eukaryotes and function in C. elegans. FASEB J. 2004;18:1834–1850. doi: 10.1096/fj.04-2273com. [DOI] [PubMed] [Google Scholar]
- 23.Nie Z, Fei J, Premont RT, Randazzo PA. The Arf GAPs AGAP1 and AGAP2 distinguish between the adaptor protein complexes AP-1 and AP-3. J Cell Sci. 2005;118:3555–3566. doi: 10.1242/jcs.02486. [DOI] [PubMed] [Google Scholar]
- 24.Fortini ME, Bilder D. Endocytic regulation of Notch signaling. Curr Opin Genet Dev. 2009;19:323–328. doi: 10.1016/j.gde.2009.04.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Merte J, Jensen D, Wright K, Sarsfield S, Wang Y, Schekman R, Ginty DD. Sec24b selectively sorts Vangl2 to regulate planar cell polarity during neural tube closure. Nat Cell Biol. 2010;12:41–46. doi: 10.1038/ncb2002. sup pp 41–48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Doe CQ, Spana EP. A collection of cortical crescents: asymmetric protein localization in CNS precursor cells. Neuron. 1995;15:991–995. doi: 10.1016/0896-6273(95)90088-8. [DOI] [PubMed] [Google Scholar]
- 27.Mizumoto K, Sawa H. Two betas or not two betas: regulation of asymmetric division by beta-catenin. Trends Cell Biol. 2007;17:465–473. doi: 10.1016/j.tcb.2007.08.004. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.