Abstract
Macromolecular assemblies that regulate chromatin structure using the energy of ATP hydrolysis have critical roles in development, cancer, and stem cell biology. The ATPases of this family are encoded by 27 human genes and are usually associated with several other proteins that are stable, non-exchangeable subunits. One fundamental mechanism used by these complexes is thought to be the movement or exchange of nucleosomes to regulate transcription. However, recent genetic studies indicate that chromatin remodelers may also be involved in regulating other aspects of chromatin structure during many cellular processes. The SWI/SNF family in particular appears to have undergone a substantial change in subunit composition and mechanism coincident with the evolutionary advent of multicellularity and the appearance of linking histones. The differential usage of this greater diversity of mammalian BAF subunits is essential for the development of specific cell fates, including the progression from pluripotency to multipotency to committed neurons. Recent human genetic screens have revealed that BRG1, ARID1A, BAF155, and hSNF5 are frequently mutated in tumors, indicating that BAF complexes also play a critical role in the initiation or progression of cancer. The mechanistic bases underlying the genetic requirements for BAF and other chromatin remodelers in development and cancer are relatively unexplored and will be a focus of this review.
Keywords: chromatin, genetics, biochemistry, BAF, SWI/SNF, development
Introduction
In eukaryotic cells, genetic information encoded in over 1 meter of DNA is packaged into chromatin and compartmentalized in the nucleus. The basic unit of chromatin is the nucleosome 1, which consists of 146 basepairs of duplex DNA wrapped around a histone octamer comprised of two of each of the conventional histone proteins: H2A, H2B, H3 and H4. Highly related histone variants are also incorporated throughout the genome for regulatory purposes 2. A fifth histone protein, H1, promotes higher order chromatin structures by encouraging compaction of neighboring nucleosomes from “beads on a string” to the 30 nm fiber. H1 is more abundant in heterochromatin than in euchromatin, leading to the greater compaction and more condensed appearance of heterochromatin. As the 30 nm fiber accounts for only ∼25-fold of a 5 000-fold DNA-to-nucleus compaction ratio, several other as yet unknown mechanisms must contribute to higher order compaction and nuclear organization.
Despite this intricate packaging, DNA must be accessible for critical cellular processes such as transcription, replication, recombination, and repair. DNA accessibility is facilitated by two classes of enzymes, ATP-dependent nucleosome remodelers and histone modifying enzymes. Histone modifying enzymes post-translationally modify the N-terminal tails of histone proteins to alter the structure of chromatin and provide binding sites for regulatory proteins. Many chromatin-associated proteins contain protein domains that bind these moieties such as the bromodomain that recognizes acetylated residues. Through direct interactions with histone tails, these proteins are targeted to specific sites on chromatin, such as transcriptionally active regions abundant in H3K4me3, or repressed regions marked with H3K27me3.
Chromatin remodeling complexes (CRCs) in contrast utilize the energy of ATP to disrupt nucleosome DNA contacts, move nucleosomes along DNA, and remove or exchange nucleosomes. They thus make DNA /chromatin available to proteins that need to access DNA or histones directly during cellular processes. The essential role of these enzymes is reflected in the fact that many of them are required for diverse but specific aspects of embryonic development including pluripotency, cardiac development, dendritic morphogenesis and self-renewal of neural stem cells. In the adult, deletion or mutation of these proteins often leads to apoptosis or tumorigenesis as a consequence of dysregulated cell cycle control. Here we review the four major families of CRCs, highlighting their specialized functions.
Early genetics
The first chromatin remodeling proteins were identified in two independent screens for mutants affecting mating-type switching and growth on sucrose, and were named SWI or SNF for switching defective or sucrose nonfermenting 3, 4. Along with several cytoplasmic signaling proteins, these screens returned what were subsequently found to be regulators of chromatin structure. The swi2 and snf2 mutations could be suppressed by mutations in H2A and H2B, and the mutants exhibit altered nuclease sensitivity at the suc2 gene, indicating that SWI/SNF proteins alter chromatin structure to enable transcription 5.
The yeast proteins SWI1/ADR6, SWI2/SNF2, SWI3, SNF5, and SNF6 are co-associated in a large multi-subunit complex called SWI/SNF 6, 7. Inactivation of any single subunit similarly affects transcription due to dissolution of the complex, although these proteins are not essential for yeast viability 4, 8, 9. SWI/SNF proteins are required for transcription by sequence-specific transcription factors, including yeast GAL4 and the glucocorticoid receptor expressed in yeast 4, 10, 11. In addition, LexA fusions of SNF2, SNF5, and SNF6 can activate transcription when bound to DNA 6, 10. The SWI/SNF complex was thus proposed to be a general activator of transcription, working in coordination with sequence-specific transactivators and the histone acetylase GCN5 12.
The SWI/SNF family is evolutionarily conserved, and homologous proteins were subsequently identified in flies, plants, and mammals. The Drosophila homologue of SWI2/SNF2, Brahma (BRM), was identified in a screen for genes that suppress the body segment defects caused by mutations in Polycomb 13. BRM mutants cause homeotic transformations consistent with the reduced transcription of homeotic genes 14. The demonstration of antagonism between BRM and Polycomb, a known chromatin regulator, provided complementary support in flies for the role of SWI/SNF as a regulator of chromatin structure 15. It was subsequently found that BRM localizes to a fraction of actively transcribed genes on polytene chromosomes and is required for Pol II localization to these loci 16, 17. However, in contrast to the genetic studies in yeast, mutants for the fly homologue of SNF5 do not phenocopy the BRM mutant 18, 19.
The ATPase homology
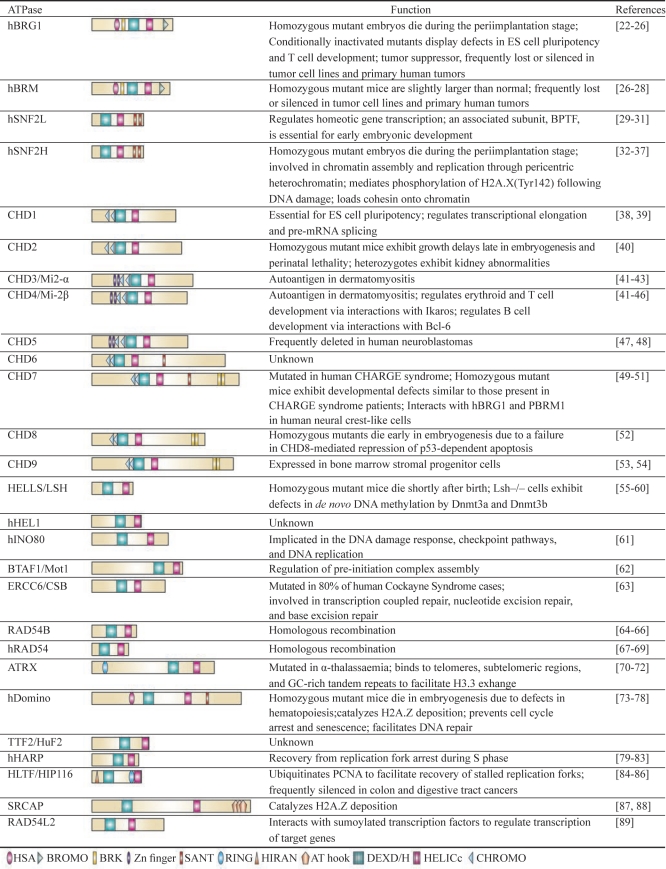
The yeast SWI2/SNF2 gene is homologous to a number of other ATP-binding helicases of the DEAD/H family 20. The sequence similarity includes the catalytic ATPase domain and seven characteristic protein motifs 21. Performing a homology search in the human genome against the human homologue of SWI2/SNF2, BRG1, reveals a high degree of homology with 26 other ATPase domain-containing proteins (Table 1). These include proteins involved in transcriptional regulation (BTAF1), DNA methylation/gene silencing (LSH), DNA repair (CSB, HARP, HIP116), DNA recombination (RAD54, RAD54b), chromosome stability (ATRX), proteins of unknown function (HuF2, HEL1), as well as other chromatin remodelers. Despite the similarity in their core enzymatic domains, these proteins are genetically nonredundant in vivo indicating that they have specialized functions. This nonredundancy is not simply due to exclusive expression patterns as coexpressed ATPases exhibit very different phenotypes upon deletion in the same cell type.
Table 1. ATPases of the DEAD/H helicase family.
The chromatin remodelers can be further subdivided into four families: BRG1 and hBRM are subunits of the SWI/SNF family, hINO80, hDomino, and SRCAP are ATPases of the INO80/SWR1 family, hSNF2H and hSNF2L belong to the ISWI family, and CHD1-9 serve the CHD family. These ATPases share affinity for the nucleosome and display DNA- and nucleosome-dependent ATPase activity in vitro90. Most chromatin remodelers form large multi-subunit complexes, which most likely alters the activity of the core ATPase in vivo. The accessory subunits commonly contain interaction domains that may directly regulate the enzymatic activity of the complex, facilitate binding to transcription factors and other chromatin modifying enzymes, and target the complex to DNA and/or modified histones. Among others, Actin-related proteins (Arps) are frequently associated with chromatin remodelers and genetic studies in yeast and mice have demonstrated an unequivocal requirement for these proteins 91. β-actin is also an intrinsic subunit of the INO80, SWR1, and mammalian SWI/SNF or BAF complexes; however, its function has been difficult to discern because actin is essential for viability. It is most likely the contribution of these additional subunits that lends functional specificity to highly similar ATPases.
Mechanistic features of DNA-dependent ATPases
Until recently, the mechanisms of action of most of these complexes were thought to be relatively well understood. However, recent genomic and genetic data in mammals have raised questions about the mechanisms by which these complexes produce their biologic activities. Early studies done primarily on nucleosomal templates suggested that these complexes might mediate their biologic actions by simply moving or exchanging nucleosomes. Indeed for the SWR1 complex, genetic studies have nicely documented the importance of the nucleosome exchange reaction first observed in vitro92. However, many investigators including ourselves have failed to find substantial changes in nucleosome positioning after acute conditional mutation of proteins such as Brg1, despite rapid and extensive changes in the regulation of target genes. These studies have suggested that the biologic actions of CRCs may be achieved not only by nucleosome movement but also additional mechanisms. Genome-wide studies of Brg1 have shown that BAF complexes generally do not bind at promoters, which are less compact than other genomic regions. Hence, these complexes may primarily occupy nucleosomal assemblies that are more complex than a simple “beads on a string” template where nucleosomes are not free to slide on DNA as observed in vitro. In addition, despite the definitive requirement for complex subunits in vivo, they are not required for the remodeling activity of the ATPase in vitro, further cautioning that in vitro mechanistic studies may be only partially informative. Thus, there is an urgent need to develop new and better mechanistic approaches to study the complexes upon chromatin templates with all of the histone modifications and complexity of the actual genomic sites where they carry out their functions. With these caveats, we will review what is known of the mechanism underlying the biologic actions of ATP-dependent chromatin remodelers.
Structural studies have focused on understanding how the hydrolysis of ATP catalyzes the movement of DNA around nucleosomes. Although the electrostatic interactions governing the DNA-nucleosome association are energetically unfavorable to unwrapping, it has been shown that CRCs can disrupt these contacts in order to promote nucleosome sliding, DNA exposure, and nucleosome exchange on DNA templates in vitro. An early hypothesis supported a mechanism of twist diffusion, whereby DNA twists around the nucleosome to accommodate the gain of a base pair from the linker DNA. The twist is propagated through the rest of the DNA:histone contacts, resulting in the advance of the nucleosome along DNA. However, this model was rejected on the basis that large impediments to DNA twisting, such as DNA hairpins or biotin crosslinks, produced no defect in nucleosome sliding 93, 94. A more favored model is the 'loop recapture' model, which argues for the generation of a loop of DNA created by new histone contacts with neighboring linker DNA 94. As in the twist diffusion model, this loop is propagated around the nucleosome to advance the nucleosome along DNA. Although energetically more costly, the loop recapture model explains how DNA might move around the nucleosome without changes to rotational phasing.
The initial creation of the DNA loop or bulge may be the result of the translocase activity that has been ascribed to the SWI/SNF, ISWI, and ACF ATPases 95, 96, 97, 98. In the case of SWI/SNF, Saha and colleagues postulate that the ATPase binds to a specific location on the nucleosome, from which it utilizes its 3′ → 5′ translocase activity to draw DNA from one entry/exit and pump it to the other in a directional wave 99. DNA footprinting and crosslinking experiments have placed the ATPase at a site of weak DNA:histone contact, where torsional strain might be tolerated for the propagation of the loop 96, 99, 100, 101. In addition, electron micrographic reconstructions of the larger RSC and SWI/SNF complexes indicate that these CRCs form multi-lobed C-shaped structures that cradle the nucleosome in a central cavity with its entry and exit points exposed 101, 102, 103. Binding of the complex to the nucleosome creates significant rearrangement of the DNA with respect to the histone octamer even in the absence of ATP hydrolysis, which may facilitate the creation of a DNA bulge required for ATP-dependent translocation 104.
In contrast to SWI/SNF, the smaller ISWI CRCs make limited contacts with the nucleosome and the extranucleosomal DNA 100, 105, 106, 107. These complexes bind their substrate as a dimeric motor to facilitate the bidirectional and processive translocation of DNA over the nucleosome 95, 108, 109. This is consistent with the role of these remodelers in nucleosome spacing, and their ability to sample DNA linker lengths to position nucleosomes equidistant from either end 94, 107, 110, 111. In this case, ATP hydrolysis by one of the two ATPases results in loosened DNA-histone contacts that may act similarly to the ATP-independent conformational change upon SWI/SNF binding to promote DNA translocation following ATP hydrolysis by the second ATPase 112.
Although the in vitro translocase activity of the SWI/SNF and ISWI ATPases is remarkably similar, their associated subunits have a profound impact on the resulting function of the remodeling complex. For example, subunit composition determines the stoichiometry of the complex to the nucleosome, 1:1 for SWI/SNF versus 2:1 for ISWI. In addition, the Dbp4 subunit of ISWI makes significant contacts with extranucleosomal DNA, thus rendering ISWI activity dependent on linker length 105. These biochemical differences correspond to functional specialization in vivo. Namely, nucleosome sliding and perturbation by SWI/SNF promotes access to DNA, while nucleosome spacing by ISWI facilitates chromatin formation and gene silencing.
Structural studies may also shed light on template selection in vivo. For example, it has been determined by mutational analysis that the catalytic activity of ISWI is uniquely affected by a basic patch of residues (K16R17H18R19) on the H4 tail 113, 114. Similarly, acetylation of H4K16 reduces remodeling by the ISWI remodeler ACF 115. This suggests that ISWI is targeted away from transcriptionally active, H4K16 acetylated chromatin and areas where even spacing of nucleosomes occludes either the H4 tail or linker DNA. On a related note, the extensive contact between SWI/SNF and RSC and their nucleosome substrate would seem to preclude the binding of these complexes to condensed chromatin. This is in agreement with reports suggesting that H1 incorporation renders chromatin resistant to remodeling 116, 117, 118.
Bioinformatic analysis of chromatin remodelers suggests that the core ATPase domain is remarkably similar to that of the DEAD/H helicases, indicating that they most likely utilize ATP to facilitate movement of DNA by similar mechanisms. Indeed, these enzymes and the related RAD54 protein share an ability to translocate DNA similar to the DEAD/H helicases, despite no evidence for helicase activity of SWI/SNF proteins 119. Despite these similarities, these proteins are genetically nonredundant, demonstrating functional specificity 120. Extensive structural analyses of the SWI/SNF and ISWI complexes have demonstrated differences in the reaction product, substrate selection, enzymatic activity, and targeting of these complexes. However, as alluded to above, our understanding of the relative ability of CRCs to remodel chromatinized templates may be misguided due to the complexity of reproducing physiological templates in vitro. It is possible that nucleosome movement is only one aspect of CRC function and that studies utilizing native chromatin templates will reveal other activities.
Chromatin regulators as macromolecular machines
Given the importance of associated subunits, it is appropriate to consider what defines a macromolecular complex and why nature would choose to partner proteins in this manner. Perhaps the most well defined macromolecular complex is the ribosome, which we can use as a standard for what constitutes a subunit of a complex as opposed to an associated protein. By this criteria, a subunit can be defined as a protein that forms a stable, interlocking association with the complex which is resistant to exchange with free subunits and can only be disrupted by denaturation. Subunits exhibit biochemical dedication to the complex as assessed by glycerol sedimentation, which is often 50% or more of the total protein in case of CRCs. Certain subunits can be found in more than one complex. For example, virtually all BAF53b and BAF45 migrate with the ATPase Brg1 in mammalian BAF complexes. However, Polybromo migrates with Brg1 and with a large unidentified complex, yet Polybromo is a stable subunit of BAF complexes 121. Lastly, genetic deletion of subunits often results in similar phenotypes. However, different phenotypes can emerge if a subunit is employed in another complex or is only required for a subset of complex functions.
Why would a DNA-dependent ATPase be driven evolutionarily to function with other subunits, rather than simply use the activities of these other proteins in solution? One answer seems to lie in the rapid coupling of reactions. The probability of an effective collision between two molecules is a 3rd order function of proximity. Hence complexes are probably assembled from active subunits during the course of evolution to provide proximity for coupled reactions. This means that if a chromatin regulatory complex carries out steps 1, 2, 3, 4, 5, 6 and 7 on its surface, then the genetic lack of a subunit might result in loss of reactions 5-7 while another mutation would result in the loss of reactions 1-7 thereby giving a different phenotype to two genes that are entirely dedicated, for example BRM and OSA in flies 18. Unfortunately, we have very little understanding of the other coupled reactions that almost certainly are serially and stereo-specifically organized on the surface of chromatin regulators. The development of small molecule inhibitors to parse the sequence of steps is likely to be invaluable to understanding the nature of these coupled mechanisms.
A second evolutionary force driving the formation of macromolecular complexes is the opportunity for diversification of function afforded through combinatorial assembly of the complexes. Combinatorial assembly seems to be a feature of mammalian complexes and arises from the fact that subunits are encoded by gene families. These family members provide mechanistic variation leading to the functional specialization of a specific complex assembly. As we will discuss later, this latter point is an intriguing property of the mammalian BAF complex and probably most other chromatin remodeling complexes, which has dramatically broadened its role from yeast to mammals. Here again, small molecule inhibitors interfering with the function of specific subunits or emerging from unique interfaces between subunits could be vital to understanding how the subunits work as a complex. Of note, the development of small molecule inhibitors targeting the bromodomain of BRD4 was recently reported, which could be modified to target any of the several bromodomains present in subunits of the BAF complex 122, 123.
Actin and actin related proteins in chromatin remodeling
The long debate over the existence of actin in the nucleus centered on the fact that actin filaments could never be detected in the nucleus despite concerted efforts. However, the discovery that monomeric actin is an intrinsic subunit of the BAF complexes resolved this argument and also explained why filaments were not visible 124. β-actin is present at what appears to be a 1:1 molar ratio with Brg1 and binds directly to the Brg1 ATPase 125. Actin is not exchangeable in vitro and remains associated with BAF complexes even in 3 M urea. How monomeric actin might be contributing to chromatin remodeling by the BAF complex is unknown, but it is intriguing to speculate that the ATPase activity of β-actin is involved analogous to its regulation of myosin. In association with mysosin, actin acts as an exchange factor to remove ADP from the active site of myosin, which initiates the myosin power stroke and thereby increases the rate of ATP hydrolysis by myosin 126. Thus one possible role of actin in the BAF complex is to act as an exchange factor to promote a similar conformational twist of the Brg/Brm ATPase. Actin appears to be necessary for DNA-dependent ATPase activity of BRG1 as removal of actin from its binding site on BRG1 by treatment of BAF complexes with 5 M urea followed by renaturation leads to impairment of ATPase activity 124. However, it is not clear that the BRG1 protein was fully renatured in these experiments. Actin was also found to be a component of the SWR1 complex and recent evidence indicates that it also is required for optimum ATPase activity of the SWR1 complex 92.
Arps are highly similar to actin, particularly over the ATP/ADP-binding pocket of the actin ATPase domain known as the 'actin fold'. However, unlike actin, Arps cannot polymerize and do not have ATPase activity, with the possible exception of yeast Arp4. Arps have recently been classified into 11 subfamilies with Arp4-Arp9 being predominantly localized in the nucleus 127. These proteins are dedicated, conserved subunits of the SWI/SNF and INO80 CRCs that associate directly with the ATPase via the conserved N-terminal HSA domain 128. Arp7 and Arp9 appear to be specific to fungi and may be functionally similar to Arp4 and actin, which are present in the orthologous CRCs of higher eukaryotes 127. Due to a unique insertion, Arp4 has the ability to bind histones, modified histones, and nucleosomes, which is critical to the function of the yeast NuA4, INO80, and SWR1 complexes in DNA repair 129. In addition, Arp4 is the only known Arp with ATP-binding activity, which contributes to NuA4 function through Arp4 association and disassociation from the complex 130. Arp6 is a unique subunit of SWR1, which regulates binding to H2AZ prior to H2AZ exchange by the SWR1 complex 131. Finally, Arp5 and Arp8 have been shown to be essential subunits of yINO80 as deletion of Arp5 or Arp8 phenocopies Ino80 deletion in yeast 132.
In the mammalian BAF complexes, the two Arp4 homologues, BAF53a and BAF53b, are used sequentially in the development of the nervous system. BAF53a is present in neural stem cells lining the ventricles and is rapidly replaced by BAF53b at cell cycle exit 121. BAF53b is a dedicated subunit of the neuron-specific nBAF complex and its deletion leads to death shortly after birth due to a failure of dendritic morphogenesis in post-mitotic neurons 133. Similarly, knockdown of the Drosophila homologue of BAF53a/b, BAP55, in class I dendrite arborization neurons during embryogenesis results in aberrantly oriented dendrites and reduced arborization while deletion of BAP55 leads to a highly specific mistargeting of olfactory projection neurons 134, 135. Because BAF53a and b do not possess ATPase activity and are not required for the enzymatic activity of Brg in vitro 124, 133, they must contribute to complex function by other means, perhaps through complex targeting to histones or nucleosomes. Alternatively, they might function like Arp2/3 in generation of dimers that control actin dynamics 136.
Chromatin remodeling complexes by family
SWI/SNF family
SWI/SNF Yeast SWI2/SNF2 is incorporated into a 1.14 MDa multi-subunit complex of 8-11 subunits. When we purified a family of related mammalian complexes in HeLa cells we called them mSWI/SNF because we found that four subunits were clearly homologous to ones present in the yeast SWI/SNF complex 137, 138, 139. However, we and other laboratories have characterized the complexes from non-transformed cells and find that the mammalian complexes are less similar to ySWI/SNF than we initially thought 120, 121, 124. Recent studies have shown that many BAF subunits are bona fide tumor suppressors, which are generally inactivated in malignant cell lines and can be deleted with no apparent phenotype 140. Hence, we and other investigators who took the same tack were most likely studying partially assembled complexes. Hence we have used BAF for Brg/Brm-associated factors for the mammalian complexes to discourage extrapolation, which has often led to incorrect conclusions. Because SWI, SNF, and BAF were not acceptable to the HUGO nomenclature committee, we suggested an alternate nomenclature of SMARC (SWI/SNF related, actin containing, regulators of chromatin). Unfortunately, this nomenclature has been extended by HUGO to proteins that do not have these characteristics. At present all of these nomenclatures are in use and Table 2 provides a translation for the reader.
Table 2. The SWI/SNF family.
| Subfamily |
SWI/SNF |
||||||
|---|---|---|---|---|---|---|---|
| Species | Yeast | Yeast | Fly | Human | Mouse | Mouse | Mouse |
| Complex | SWI/SNF | RSC | BAP | BAF | esBAF | npBAF | nBAF |
| Swi2/Snf2 | Sth1 | BRM | BRG1/SMARCA4 or hBRM/SMARCA2 | BRG1/SMARCA4 | BRG1/SMARCA4 | BRG1/SMARCA4 | |
| OSA/eyelid or BAP170 | BAF250a/ARID1A or BAF250b/ARID1B or BAF200/ARID2* | BAF250a/ARID1A | BAF250a/ARID1A or BAF250b/ARID1B or BAF200/ARID2* | BAF250a/ARID1A or BAF250b/ARID1B or BAF200/ARID2* | |||
| Rsc1, Rsc2, Rsc4 | Polybromo | BAF180* | |||||
| Swi3 | Rsc8 | MOR/BAP155 | BAF155/SMARCC1 and/or BAF170/SMARCC2 | BAF155/SMARCC1 | BAF155/SMARCC1 and/or BAF170/SMARCC2 | BAF155/SMARCC1 and/or BAF170/SMARCC2 | |
| Swp73 | Rsc6 | BAP60 | BAF60a/SMARCD1 or BAF60b/SMARCD2 or BAF60c/SMARCD3 | BAF60a/SMARCD1 | BAF60a/SMARCD1 | BAF60a/SMARCD1 | |
| Arp7, Arp9 | Arp7, Arp9 | BAF55 or BAP47 | BAF53a or BAF53b | BAF53a | BAF53a | BAF53b | |
| Snf5 | Sfh1 | SNR1/BAP45 | BAF47/hSNF5/SMARCB1 | BAF47/SNF5/SMARCB1 | BAF47/SNF5/SMARCB1 | BAF47/SNF5/SMARCB1 | |
| SAYP | BAF45a or BAF45b or BAF45c or BAF45d | BAF45d | BAF45a or BAF45d | BAF45b or BAF45c or BAF45d | |||
| BAP111/dalao | BAF57/SMARCE1 | BAF57/SMARCE1 | BAF57/SMARCE1 | BAF57/SMARCE1 | |||
| β-actin | β-actin | β-actin | β-actin | β-actin | |||
| Unique Subunits | Swi1/Adr6, Swp82, Taf14, Snf6, Snf11 | Rsc3, 5, 7, 9, 10, 30, Htl1, Lbd7, Rtt102 | Brd7*, Brd9 |
*ARID2, BAF180, and Brd7 are proposed subunits of the PBAF complex. Despite coassociation with BRG1, these proteins are not dedicated subunits of the BAF complex as assessed by glycerol gradient centrifugation (121, Hargreaves DC and Crabtree GR, unpublished observations).
Mammalian BAF and fly BAP (for Brahma-associated factors) complexes have lost, gained, and poached SWI/SNF subunits in response to the changes in epigenetic regulation that accompanied the evolution of multicellularity. The BAF complex is composed of five ySWI/SNF orthologues (BRG1/hBRM, BAF155/170, BAF60, BAF53a/b, and BAF47) and several additional unique subunits (BAF250a/BAF250b, BAF200, BAF45a/b/c/d, Brd9, and Brd7), including two subunits, BAF57 and actin, which are related to Nhp10 and actin found in the yeast INO80 and SWR1 complexes. Five yeast subunits have been discarded, resulting in a mammalian complex of about 2 MDa, which is larger than the calculated molecular weight of the known subunits, indicating that several subunits have yet to be identified. In addition to the amino terminal DEXDc and HELICc subdomains shared with other DEAD/H helicases, Brg1 and Brm have a C-terminal bromodomain. The bromodomain was first identified in BRM, the Drosophila homologue of ySWI2/SNF2, and binds acetylated residues on histone tails 14. Additional domains present in the accessory subunits presumably facilitate interactions with proteins (LXXLL, BAH, SANT, SWIRM, SWIB), DNA (ARID, HMG, Zn finger, Leucine zipper), and modified histones (Bromodomain, Chromodomain, PHD domain), although this has yet to be confirmed genetically. The bromodomain of Brg1 is not required for its in vivo function, as a mutant lacking the bromodomain fully rescued the knockout phenotype in both flies and mice 90, 141. However, additional bromodomains in Brd7 and Brd9 might function redundantly in recruiting the complexes to specific genetic loci. Finally, while ySWI/SNF lacks actin, the mammalian BAF complex has approximately one actin molecule per complex, which may enhance the ATPase activity of BRG1/hBRM, as in the case of SWR1 124.
Perhaps even more important than gain or loss of subunits in the evolution of SWI/SNF is the expansion of gene families that encode homologous BAF subunits. Whereas ySWI/SNF is monomorphic, BAF complexes are arrayed from several possible options in each of the following gene families: Brg1/Brm, BAF250a/b, BAF155/BAF170, BAF60a/b/c, BAF45a/b/c/d, BAF57, BAF53a/b, BAF47, and actin. Genetic studies indicate that subunit exchange helps drive the transition from pluripotency to multipotency to the committed post-mitotic neuron. Pluripotent embryonic stem (ES) cells express BAF155 but not BAF170, Brg1 but not Brm, and BAF53a but not BAF53b 23, 24, 142, 143. Consistent with a requirement for this particular assembly for pluripotency, deletion of Brg1 results in early embryonic lethality during implantation 22, while Brm-deficient mice are viable, though a little larger than normal 28. Differentiation of ES cells into neural progenitors leads to the activation of Brm and BAF170 and repression of BAF60b 121. The final step to post-mitotic neurons is accompanied by the repression of BAF53a and the activation of BAF53b, BAF45b and BAF45c, which assemble in nBAF complexes, apparently unique to the nervous system 121. The mechanism underlying the switch in subunits during the development of the vertebrate nervous system appears to be a triple negative genetic circuit in which REST represses miR-9/9* and miR-124, which in turn repress BAF53a leading to cell cycle exit, the activation of the alternative BAF53b subunit, and neural differentiation 144. This npBAF to nBAF switch is essential for the development of the vertebrate nervous system, since mutations affecting either state are lethal. Another example of selective assembly occurs in the developing heart, where BAF60c designates the region of the embryo with cardiogenic potential 145. Complexes containing BAF60c are uniquely required for heart development, and can directly facilitate the formation of heart tissue from mesoderm in the presence of tissue-specific factors 145, 146. In each of these cases, the expression of homologous subunits could not compensate for the loss of the correct subunit, arguing for the exquisite specificity of each composite in instructing cell fate decisions.
The unique composition of BAF at each developmental stage correlates with a specific gene expression program that is required for maintaining cell state, although the mechanistic basis for this maintenance is unknown. In ES cells, Brg1 binding was mapped by chromatin immunoprecipitation in combination with high resolution sequencing 147. Brg1 binds to about 6 000 sites in large islands of several hundred bases. About 300 000 molecules are present on average in each ES cell, indicating that the peaks are likely the result of the binding of several complexes, rather than an individual complex. In contrast to the expectation that Brg1 would function primarily at promoters, only 12% of peaks were within 500 bp of the transcription initiation site 147. Brg1 occupancy is more highly correlated with H3K4me1, a modification enriched at enhancers and regulatory elements, than H3K4me3, a mark of active transcription 147, 148. Consistent with this, pou5f1 and nanog show peaks of Brg1 binding at each of the mapped enhancers critical to the positive feedback loop of pluripotency 149, 150. Rapid deletion of Brg1 in ES cells followed by analysis of RNA by transcript arrays led to the unexpected discovery that Brg1 represses most of its direct targets. As Brg1 binding is non-overlapping with Suz12, a Polycomb group protein, BAF must mediate repression via a distinct mechanism(s). Interestingly, Brg1-dependent repression occurs in functional coordination with pluripotency genes such as Oct4 and Sox2, to which it binds in solution and on chromatin. Genome-wide, Brg1 binding sites in ES cells physically overlap with the binding sites of Oct4, Sox2, Nanog, Smad1, and STAT3, all critical regulators of pluripotency, at far higher levels than predicted 147. Proteomic studies in ES cells also revealed that the esBAF complex binds to many of the pluripotency factors suggesting that the specialized subunit composition of esBAF complexes is tailored to the needs of a pluripotent cell.
The possibility that BAF complexes mediate repression at a distance originated from studies of the role of Brg1 and BAF complexes in lymphocytes indicating that these complexes act on a distal silencer to repress transcription of the Cd4 gene 151. Deletion of Brg1 or the silencer gives similar phenotypes of premature derepression of CD4 expression 151. These studies and the fact that Brg1 most commonly binds to intragenic regions suggest that Brg1 acts from a distance to regulate transcription, perhaps by creating long-range interactions. Indeed, Brg1-dependent looping has been observed at both the α- and β-globin loci and is required for developmentally regulated transcription 152, 153. Thus, long−range interactions from distal regulatory sites, and not nucleosome sliding, may more accurately account for BAF's observed phenotypes. In the example of the CD4 silencer in T cells, BAF is thought to prevent accumulation of H1 linker histones, allowing the corepressor Runx1 to bind and repress CD4 154.
Finally, the antagonism between BRM and Polycomb first observed in Drosophila also warrants revisiting. Brahma was identified in a screen for mutants that suppress the body segment phenotypes resulting from mutations in Polycomb. In vitro, Polycomb can inhibit remodeling by Brm. However, as Brahma is required for homeotic gene transcription and, by some accounts, transcription of all Drosophila genes, it is unclear how direct this antagonism is. In mouse ES cells, Brg1 binding was anti-correlated with Polycomb and H3K27me3, indicating that co-occupancy of Brg1 and Polycomb is actively avoided 147. Recently, a more direct antagonism was demonstrated at the p16ink4a/p19Arf locus where expression of BAF47 in a BAF47-deficient tumor cell line resulted in Polycomb eviction from the locus, re-expression of the gene, and growth arrest 155. It has subsequently been shown that Polycomb represses many genes upon loss of BAF47 in tumorigenesis and that the double mutant with Ezh2, a subunit of the Polycomb Repressive Complex 2 (PRC2), expresses wild-type levels of these genes and survives tumor-free 156. Interestingly, many of the genes affected by this antagonism are part of a stem cell-associated gene set, suggesting that this antagonism may also be critical during normal development.
As mammalian genetic studies come to light, several longstanding tenants concerning the biologic role of SWI/SNF are being challenged. Clearly, extrapolation of findings from the ySWI/SNF complex to the mammalian BAF complex is risky and direct genetic and biochemical analyses of the mammalian complex have led to surprising conclusions. Yeast SWI/SNF binds almost exclusively to promoters and activates its direct targets concomitant with nucleosome displacement. In contrast, BAF complexes are most often found in intergenic regions where they both activate and repress genes and the role of nucleosome movement seems less clear. Furthermore, BAF complexes are polymorphic and hence possess greater functional diversity through combinatorial assembly. By exchanging subunits, BAF can reinterpret the genome, in effect increasing the total number of potential BAF targets in an organism. To date, this mechanism has only been observed in vertebrates, possibly as a means of increasing effective gene number or patterns of gene regulation during development. Although a similar exchange of subunits has not been described for other CRCs, it seems likely that they also take advantage of the diversification of function afforded by combinatorial assembly.
RSC A complex related to ySWI/SNF was identified called RSC for Remodel the Structure of Chromatin based on shared homology between the yeast ATPases STH1 and SWI2/SNF2 157. RSC shares two identical subunits with ySWI/SNF, Arp7 and Arp9, and several homologous subunits 157 (Table 2). There is also evidence of an additional biochemically distinct SWI/SNF-related complex in Drosophila and mammals that nevertheless shares many subunits with the BAF and BAP complexes. These are the PBAP and PBAF complexes, named after their signature subunit, Polybromo (BAF180), a large protein containing several bromodomains, two BAH protein interaction domains, an HMG domain, and a C2H2 zinc finger domain. BAF180 is structurally related to the yeast Rsc1, Rsc2, and Rsc4 proteins, which are unique to the RSC complex, and thus, PBAP and PBAF are presumed to be evolutionary counterparts of RSC 158. BAF200 is also present only in PBAF, while hBRM and BAF250 are unique to BAF 159. It is worth noting that similarly to the case with ySWI/SNF and BAF, there are significant biochemical differences between RSC and PBAF that suggest these complexes may not be functional orthologues.
Genetic studies in yeast indicate that ySWI/SNF and RSC are nonredundant and functionally specialized. Both complexes appear to regulate gene expression, but target non-overlapping sets of genes 8, 9, 160. STH1, SFH1, RSC3, and RSC9, all components of the RSC complex, are required for cell cycle progression through G2/M, while ySWI/SNF is not essential for viability 161, 162, 163. Related to this, RSC has been implicated in sister chromatid cohesion, chromosome segregation, and ploidy maintenance 164, 165, 166, 167. In vitro, PBAF was implicated in ligand-dependent transactivation by nuclear hormone receptors 168. Consistent with this, deletion of BAF180 in mice resulted in severe hypoplastic ventricle development and trophoblast placental defects due to reduced retinoic acid-induced gene expression 169. However, flies deficient in Polybromo and/or BAP170 have no defects in ecdysone-induced transcription suggesting that the role of PBAF in nuclear receptor-dependent activation is not evolutionarily conserved 170. Whether PBAP or PBAF acts more broadly to influence cell cycle is not known, although there is evidence that BAF180 localizes to kinetochores during mitosis 158. Of note, BAF180 predominantly co-migrates on glycerol gradients with a very large 4 MDa complex distinct from the other BAF subunits, which could account for some of its activities, for example at the kinetochore 121.
INO80 family
Continuing in the theme of unexpected roles for chromatin remodelers, we next discuss the INO80 family. The INO80 family includes the yeast INO80 complex and its orthologues Pho-dINO80 (Drosophila) and INO80 (human), the yeast SWR1 complex and its orthologue SRCAP (human), and the yeast NuA4 complex and its orthologues Tip60 (Drosophila) and TRAAP/Tip60 (humans) (Table 3). The core ATPase subunits of the INO80 family are characterized by a split ATPase domain. This unique module retains ATPase activity, and acts as a scaffold for the association of two RuvB-like proteins, Rvb1 and Rvb2. RuvB is a bacterial ATP-dependent helicase that forms a double hexamer around Holliday junctions to promote their migration during homologous recombination 171. Rvb1 and Rvb2 are present at 6:1 stoichiometry, suggesting that they form hexamer structures similar to the bacterial RuvB protein. Unlike CRCs of other families, the INO80 complex exhibits DNA helicase activity in vitro and binds specialized DNA structures in vitro that resemble Holliday junctions and replication forks 172, 173. However, it should be noted that the INO80 family does have affinity for nucleosomes, in particular the H2A variants H2AZ and H2AX. INO80 and SWR1 can slide nucleosomes in vitro on a reconstituted chromatin template and even evict histones from DNA 132, 174, 175.
Table 3. The INO80 family.
| Subfamily |
INO80 |
SWR1 |
||||||
|---|---|---|---|---|---|---|---|---|
| Species | Yeast | Fly | Human | Yeast | Human | Fly | Human | Yeast |
| Complex | INO80 | Pho-dINO80 | INO80 | SWR1 | SRCAP | Tip60 | TRAAP/Tip60 | NuA4 |
| Ino80 | dIno80 | hIno80 | Swr1 | SRCAP | Domino | p400 | ||
| Rvb1, Rvb2 | Reptin, Pontin | Tip49a, Tip49b | Rvb1, Rvb2 | Tip49a, Tip49b | Reptin, Pontin | Tip49a, Tip49b | ||
| Arp4, Arp5, Arp8, Act1 | dArp5, dArp8, dActin | Baf53a, Arp5, Arp8 | Arp4, Arp6, Act1 | BAF53a, Arp6 | BAP55, Act87E | BAF53a, Actin | Arp4, Act1 | |
| Taf14 | Yaf9 | GAS41 | dGAS41 | GAS41 | Yaf9 | |||
| Ies2 | hIes2 | |||||||
| Ies6 | hIes6 | |||||||
| Swc4/Eaf2 | DMAP1 | dMAP1 | DMAP1 | Swc4/Eaf2 | ||||
| Swc2/Vps72 | YL-1 | dYL-1 | YL-1 | |||||
| Bdf1 | dBrd8 | BRD8/TRCp120 | ||||||
| H2AZ, H2B | H2AZ, H2B | H2Av, H2B | ||||||
| Swc6/Vps71 | Znf-HIT1 | |||||||
| dTra1 | Tra1 | |||||||
| dTIP60 | Tip60 | Esa1 | ||||||
| dMRG15 | MRG15, MRGX | Eaf3 | ||||||
| dEaf6 | FLJ11730 | Eaf6 | ||||||
| dMRGBP | MRGBP | Eaf7 | ||||||
| E(pc) | EPC1,EPC-like | Epl1 | ||||||
| dING3 | ING3 | Yng2 | ||||||
| Unique Subunits | Ies1, Ies3-5, Nhp10 | Pho | Amida, NFRKB, MCRS1, FLJ90652, FLJ20309 | Swc3,5,7 | Eaf5, Eaf1/Vid21 | |||
The INO80 complexes contain β-actin and several Arps, which are shared with the mammalian BAF and fly BAP complexes. The yINO80 complex contains Arp4, Arp5, Arp8, and actin, of which Arp5, Arp8, and actin are conserved in flies and mammals. Arp4 and actin are also present in the yeast SWR1 and NuA4 complexes. The Arp4 mammalian homologue, BAF53a, is present in the INO80, SRCAP, and Tip60 complexes of the INO80 family, and as mentioned previously, the BAF and PBAF complexes. In flies, the SWR1 complex contains BAP55, an Arp that is also present in the BAP and PBAP complexes. This conservation argues for a critical actin- or Arp4-dependent role shared by INO80 and BAF complexes, possibly in targeting these complexes to structural elements. The fact that ySWI/SNF has neither Arp4 nor actin implies that the BAP and BAF complexes may have acquired mechanistic features of the INO80 family that are not operational in ySWI/SNF.
INO80 and SWR1 Yeast Ino80, the Swi2/Snf2-related ATPase of the INO80 complex, was identified in a screen for regulators of phospholipid biosynthesis 176. It was subsequently found that yIno80 associates with 14 other proteins to form a complex of 1.2 MDa 172, 176. As outlined above, Rvb1, Rvb2, Arp4, Arp5, Arp8, and actin are among these subunits, as well as Taf14, Ies1-6, and Nhp10. As evidenced by its role in phospholipid biosynthesis, yINO80 regulates gene expression downstream of several promoters, both positively and negatively 177.
Swr1 was identified based on its homology to Ino80 and was subsequently found to catalyze the exchange of H2A for Htz1 92, 178. Htz1 and H2B copurify with Swr1, and genome-wide transcription profiles reveal a ∼40% overlap in genes regulated by Htz1 and Swr1 92. Rvb1, Rvb2, Arp4, and actin are shared subunits with INO80. The remaining subunits are Arp6, Yaf9, Swc2-7, and Bdf1, Swc2 being a potential histone chaperone-like subunit based on its affinity for Htz1 179. The role of Bdf1, a bromodomain-containing protein, is not known, but this may suggest that H2AZ exchange is regulated by Bdf1-dependent binding to acetylated residues on histone tails. The role of SWR1 in H2A variant exchange is conserved throughout evolution: the Drosophila Tip60 complex catalyzes the exchange of H2A for H2Av (a variant with similarities to both H2AZ and H2AX) 180 and mammalian SRCAP complex catalyzes the exchange of H2A for H2AZ 87, 88.
Yeast ino80 and swr1 mutants are hypersensitive to DNA damage-inducing agents, but the transcriptional response induced by damage is unaffected 181, 182. This implies a direct role for these complexes in DNA repair, perhaps at the site of damage. Upon DNA damage, H2AX is phosphorylated at double-stranded breaks (DSBs) by the Mec1/Tel1 kinases (ATM/ATR in mammals), leading to the recruitment of several DNA repair proteins. The lesion can then be repaired by homologous recombination (HR) or non-homologous end joining (NHEJ) depending on which downstream pathway is initiated. ino80 mutants have defects in both the HR and NHEJ pathways, while swr1 mutants have defects in NHEJ alone 175. Both the INO80 and SWR1 complexes are recruited to DSBs via direct binding of complex subunits to phosphorylated H2AX, or γ−H2AX. Arp4, a shared subunit of INO80 and SWR1, mediates SWR1 recruitment to DSBs, while Nph10 (and Ies3) is required for INO80 recruitment, perhaps in association with Arp4 129, 182. Based on its ability to exchange histones, it was initially proposed that SWR1 is recruited to DSBs to exchange γ−H2AX for an unmodified histone. However, it is INO80 that facilitates nucleosome eviction at DSBs, allowing the recruitment of repair factors. Specifically, ino80 and arp8 mutants are defective in histone eviction, which leads to impaired recruitment of Mec1 and Rad51 and defects in ssDNA resection by the nuclease Mre11, both prerequisites for repair by HR 174. INO80 and an associated protein YY1 also promote HR repair in human cells, indicating that the role of INO80 in repair is conserved 173. Although SWR1 does not appear to exchange histones at the site of damage, it is required for efficient recruitment of Mec1 and Ku80 to DNA ends during NHEJ 175.
Following the initiation of repair at DSBs, checkpoint pathways are activated downstream of Mec1/Tel1 to coordinate the cell cycle with DNA repair. Interestingly, ino80, ies4, arp5 and arp8 mutants have defects in checkpoint recovery and reduced viability in the face of replicative stress. It was shown that Mec1/Tel1 phosphorylate Ies4 of the INO80 complex, which acts redundantly with checkpoint factor Tof1 to regulate the cell cycle following release from replicative stress 183. In parallel, INO80 was shown to promote recovery of stalled replication forks during the S-phase checkpoint response 184. Specifically, INO80 binds replication forks during S phase and recruits Rad18 and Rad51 to stalled replication forks in order that they might be processed 184, 185. In contrast, SWR1 is not present at stalled replicative forks and the corresponding mutant does not exhibit decreased viability in response to replicative stress 186, 187.
Finally, INO80 and SWR1 have been identified in genetic screens for proteins that regulate telomere length 187, 188. Ino80, Nph10, Ies3, unique subunits of the INO80 complex, and Arp4, a common subunit of both INO80 and SWR1, localize to telomeres. Mechanistically, it is possible that INO80 is recruited to telomeres because they share several features with DSBs, including the incorporation of γ-H2AX and the recruitment of Mec1/Tel1. These features appear to regulate telomere length during normal replication and intensify during telomere shortening to “repair” the DNA end. Ies3, which is not involved in DSB repair, may mediate the specialized function of the INO80 complex at telomeres following γ-H2AX-dependent recruitment. Indeed, Ies3 associates directly with the telomerase complex and deletion of Ies3 specifically results in impaired growth and telomeric instability 188.
NuA4/Tip60 As mentioned above, the Drosophila Tip60 complex catalyzes the exchange of H2A for H2Av, similar to the yeast SWR1 and human SRCAP complexes 180. However, the dTip60 complex harbors an additional seven subunits, including the eponymous subunit Tip60, a histone acetyltransferase (HAT). In fact, based on a number of shared subunits, the Tip60 complex seems to be the evolutionary product of the combined SWR1 and NuA4 yeast complexes. The related Tip60/TRRAP complex in humans also has both chromatin remodeling and HAT activity, the key enzymatic subunits being the Swr1-related ATPase, p400, and Tip60 189, 190, 191, 192. Intriguingly, Arp4 and/or actin are common to the entire INO80 family in all organisms.
Tip60 has been shown to regulate the DNA damage response through the acetylation of histones and other regulatory proteins. In all organisms, Tip60 is recruited to DSBs, possibly through an Arp4-dependent mechanism similar to that shown for SWR1. In yeast, NuA4 acetylates H4 at breaks following recruitment by Arp4 129. Drosophila Tip60 acetylates phosphorylated H2Av at DSBs and facilitates its exchange with unmodified H2Av 180. Likwise, the acetylation of γ-H2AX by human TIP60 after ionizing radiation is required for its subsequent ubiquitination and removal 193. Tip60 also acetylates ATM upon recruitment to DSBs, which activates this kinase to initiate the repair process. Interestingly, Tip60 enzymatic activity is dependent on an interaction between the Tip60 chromodomain and H3K9me3 at DSBs and disruption of this interaction or depletion of H3K9me3 results in increased sensitivity to ionizing radiation and genomic instability 194. In an unrelated fashion, Tip60 was found to acetylate lysine 120 on p53 upon increasing levels of DNA damage 195, 196. p53 acetylation by Tip60 leads to p53's selective activation of pro-apoptotic genes, shifting the balance from cell cycle arrest to apoptosis. Finally, Tip60 appears to play a regulatory role in oncogene-induced DNA damage response as haplo-insufficiency of Tip60 acclerates Myc-induced tumor progression 197.
In addition to its role in the DNA damage response, Tip60 appears to regulate several other pathways critical for cell viability. Along with Brg1, Tip60 was identified in a screen for factors affecting ES cell self-renewal and morphology 198. Using promoter ChIP-on-chip analysis, Fazzio and colleagues found that the p400-Tip60 complex is bound to about 55% of all promoter regions, including active and silent genes, where it acetylates H4. Knockdown of Tip60 resulted in the upregulation of many developmental genes coregulated by Nanog and dysregulation of the ES cell cycle. Perhaps as a result, Tip60 ablation causes embryonic lethality in mice and flies 199, 200.
In summary, the INO80 family of CRCs function in a diverse array of cellular processes, including DNA repair, cell cycle checkpoint, and telomeric stability. Some intriguing features of these complexes include the special affinity for the histone variants H2AZ and H2AX and the recruitment of these complexes to specialized DNA structures, such as DSBs, Holliday junctions, telomeric ends, and replication forks. The unique split ATPase domain of Ino80 and Swr1 and their association with the RuvB-like proteins, Rvb1 and Rvb2, may account for the specialized function of these complexes. Of note, RAD54, a related ATPase, can bind Holliday junction-like structures and promote bidirectional branch migration in an ATP-dependent manner 201. Alternatively, their recruitment may reflect the incorporation of H2A variants at critical sites. It is clear that the associated Arps are absolutely required for the function of these complexes, which may reflect a critical role for Arps and actin in targeting and/or regulation of these complexes.
ISWI family
The CRCs of the imitation SWI (ISWI) family were first identified in in vitro assays for nucleosome remodeling activities in Drosophila embryo extracts 202. These complexes are characterized by the DEAD/H-related ATPase ISWI, which in addition to the conserved ATPase domain contains a SANT and a SLIDE domain. ISWI is incorporated into several distinct complexes in Drosophila, including the ACF, CHRAC, and NURF complexes (Table 4). ACF and CHRAC are highly similar as they both contain the large Acf1 subunit in addition to ISWI 203, 204. Acf1 has a bromodomain, 2 PHD fingers, and a putative heterochromatin targeting domain. NURF is characterized by the NURF301 subunit, which is structurally related to Acf1 apart from four LXXLL motifs, which allow interactions with nuclear hormone receptors. Homologous complexes have been identified in yeast and humans. The related human ATPases hSNF2H and hSNF2L are accompanied by hACF and BPTF in ACF/CHRAC and NURF complexes, respectively (Table 4).
Table 4. The ISWI family.
| Subfamily |
ACF/CHRAC |
NURF |
||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Species | Yeast | Yeast | Yeast | Fly | Fly | Human | Human | Human | Fly | Human |
| Complex | ISWIa | ISWIb | ISW2 | ACF | CHRAC | ACF | CHRAC | WICH | NURF | NURF |
| Isw1 | Isw1 | Isw2 | ISWI | ISWI | hSNF2H | hSNF2H | hSNF2H | ISWI | hSNF2L | |
| Itc1 | ACF1 | ACF1 | WCRF180/hACF1 | WCRF180/hACF1 | WSTF | NURF301 | BPTF | |||
| Dpb4 | CHRAC14 | hCHRAC17 | NURF55/p55 | RbAp46 | ||||||
| Dls1 | CHRAC16 | hCHRAC15 | RbAp48 | |||||||
| Unique Subunits | Ioc3 | Ioc2, Ioc4 | NURF38 | |||||||
Initial identification of these complexes indicated a role in transcriptional activation as they promote transcription from in vitro templates 202, 205, 206. In vivo, NURF301-deficient flies and BPTF morpholino frogs exhibit developmental defects consistent with reduced expression of homeotic genes 207, 208. NURF301 is required for transcription of the GAGA gene targets ultrabithorax, engrailed, hsp70, and hsp26, and the wingless target senseless through direct association with cognate transcription factors 205, 208. BPTF is similarly required for homeotic gene transcription in Xenopus via an interaction between the PHD domain of BPTF and H3K4me3 at HOX genes 31. In mice, deletion of SNF2h or BPTF is embryonic lethal. ES cell lines could not be derived from SNF2h-deficient mice due to the requirement for SNF2h in survival of the inner cell mass ex vivo 32. BPTF is required at a later stage in the differentiation to the endoderm, mesoderm, and ectoderm lineages, possibly in conjunction with the Smad pathway 30.
The abovementioned studies highlight the somewhat predicted role of ISWI in regulating gene expression in euchromatic regions. However, ISWI is also enriched in heterochromatic regions, suggesting a role for ISWI in initiation or maintenance of heterochromatin formation 209. ISWI is required for fly embryogenesis, so flies expressing an ATPase-dead dominant negative mutant of ISWI, ISWIK159R, were analyzed. Expression of ISWIK159R results in decondensation of the male X chromosome, presumably reflecting a loss of chromatin compaction. Interestingly, mutations in the dosage compensation complex that acetylates H4K16, cause the male X to appear more condensed, consistent with increased ISWI activity in the absence of H4 acetylation. NURF301 mutant flies are viable and recapitulate the male X phenotype, while ACF mutant flies are semi-embryonic lethal, providing a clear example of functional specialization of the ISWI complexes by associated subunits 210, 211. In the 30% of ACF-deficient flies that do survive, normal diversification of euchromatin versus heterochromatin during blastoderm development is perturbed 210. In addition, ACF mutant flies display defects in heterochromatic silencing as assessed by defects in pericentric position effect variegation and Polycomb-mediated repression 35. The phenotypes observed upon loss of ISWI, ACF, and NURF301 may correlate with the unique ability of ISWI to catalyze the formation of regularly spaced nucleosomal arrays in vitro, as regular spacing of nucleosomes in native chromatin may result in greater compaction. However, closer examination revealed that nucleosome spacing is normal ISWI mutant flies despite global perturbations in chromatin compaction 212, 213. The authors instead ascribed the defect to a loss of H1 on chromatin as H1 mutant flies have a similar defect in compaction.
A specific role for the mammalian WICH complex in DNA replication of condensed regions has also been observed. hSNF2h, ACF1, and the ACF1 homologue WSTF are particularly enriched in replicating pericentric heterochromatin, suggesting that they might have a role in the replication of condensed chromatin 33, 214. Indeed, knockdown of these proteins led to slower S phase progression that could be ameliorated by treatment with 5-aza-2-deoxycytidine, a DNA methylase inhibitor that results in chromatin decondensation 33. Further exploration revealed that WSTF is directly recruited to replication foci by PCNA and that knockdown of WSTF results in greater compaction and increased HP1 association of newly replicated DNA 34. Finally, WSTF binds to γ-H2AX following DNA damage and stabilizes γ-H2AX by phosphorylating Tyr142 36.
The ISWI family has been implicated in a variety of cellular functions, including transcriptional repression, heterochromatin formation, DNA replication, and ES cell pluripotency. Like Brg1 and Brm, ISWI can assemble with different homologous subunits such as ACF1 or NURF301 to form distinct functional complexes. It is tempting to speculate that combinatorial assembly is an important regulatory mechanism that governs the function of ISWI in developmental or cellular transitions as it does for the BAF complex. Consistent with this hypothesis, ACF1 expression appears to be developmentally regulated in Drosophila and constitutive ACF1 expression is lethal 210. Careful genetic analysis of this type may reveal that subunit switch is a common theme among chromatin remodelers. Finally, it will be important to determine how the unique ability of the ISWI ATPase to order nucleosomes in vitro relates to the phenotypes observed in vivo.
CHD family
The ATPases of the CHD family are characterized by N-terminal tandem chromodomains in addition to the conserved DEAD/H-related ATPase domain. There are nine known family members that have been further categorized into three families based on similarities in domain structure. CHD1 and CHD2 have a C-terminal DNA binding domain, CHD3 and CHD4 lack the DNA binding domain but have N-terminal paired PHD fingers, and CHD5-9 have additional functional domains in their C termini. Of these nine members, CHD1 is the only family member in yeast. Here, we will review the known functions of a few of the well-characterized family members, beginning with CHD1.
CHD1/2 Biochemical characterization of CHD1 demonstrated that CHD1 exists as a monomer or dimer, unlike the enzymatic subunits of other CRCs 215, 216. yChd1 has also been shown to interact with the multi-subunit SAGA and SLIK complexes, two highly related yeast HAT complexes that preferentially acetylate H3 and H2B 216. CHD1 is targeted to sites of active transcription through PHD-mediated recognition of H3K4me3 217, 218 and associates with other preinitiation factors to facilitate transcriptional elongation and splicing 39. Consistent with these findings, genome-wide binding of Chd1 correlates globally with H3K4me3 and Pol II in ES cells 38. Knockdown of Chd1 in ES cells results in the loss of endoderm lineage potential and the appearance of heterochromatic foci, suggesting that Chd1 is required to maintain an open chromatin state for pluripotency 38. Interestingly, Drosophila CHD1 is required for the deposition of histone variant H3.3 into decondensing sperm chromatin during embryogenesis 219. Although it is not known whether the related CHD2 protein also functions in transcription or nucleosome assembly, Chd2-deficient mice exhibit decreased neonatal viability and organ failure 40.
CHD3/4 CHD3 and CHD4, or Mi-2α and Mi-2β, are incorporated into a large protein complex with both histone deacetylase and remodeling activities called NURD (Table 5). The other subunits of NURD include HDAC1, HDAC2, RbAp48, RbAp46, MTA1, MTA2, MTA3, MBD2, and MBD3. The deacetylase activity of HDAC1/2 on nucleosomal templates is stimulated by ATP, suggesting that the chromatin remodeling activity of NURD facilitates access to acetylated histones 220, 221. It is thus tempting to assume that NURD represses transcription by binding to methylated DNA in an MBD2/3-dependent manner, remodeling the surrounding chromatin, and removing active histone marks. However, it is unclear how these accessory proteins contribute to the considerably more complex NURD-dependent processes observed in vivo.
Table 5. The NURD complex.
| Subfamily |
NURD |
|
|---|---|---|
| Species | Fly | Human |
| Complex | Mi-2/NuRD | NuRD |
| dMi-2 | CHD3/Mi-2α CHD4/Mi-2β | |
| dMBD2/3 | MBD2,MBD3 | |
| dMTA dRPD3 | MTA1,2,3 HDAC1,2 | |
| p55 | RbAp46,48 | |
| p66/68 | p66α,β |
Our limited genetic knowledge suggests that NURD associates with transcription factors to regulate key developmental transitions. The role of Mi2-β in T lymphocytes was extensively analyzed after it was found to associate with Ikaros, a transcriptional repressor important in the development of T cells 222. Like Ikaros, Mi-2β is required for thymocyte development and for proliferative expansion of mature peripheral T cells. Interestingly, Mi-2β also promotes CD4 expression during T cell development by antagonizing Ikaros 46, 223. This is in contrast to Brg, which represses CD4 expression in developing T lymphocytes 151. Through association with another repressor, Bcl-6, MTA-3/NURD also regulates cell fate decisions in the B lymphocyte lineage 44. The role of NURD in embryonic development has not been reported. MBD3 was shown to be critical for maintaining pluripotency in ES cells through suppression of the trophoectoderm lineage, but it is unclear whether this transcriptional repression is mediated by Mi-2/NURD 224, 225. Of note, dMi-2 is essential for the development of germ cells in flies 226.
Similarly to INO80, SWR1 and TIP60, NURD is recruited to sites of DNA damage, although this recruitment appears to be mediated by the DNA repair protein PARP rather than γ-H2AX itself 227, 228. Following recruitment, Mi-2β is phosphorylated by ATM and facilitates the RNF8/RNF168-dependent ubiquitination of γ-H2AX and subsequent RNF8 and BRCA1 accumulation at DSBs 229, 230. Loss of Mi-2β, in turn, results in aberrant DSB repair and G2/M checkpoint activation, leading to increased sensitivity to DNA damage. Interestingly, loss of Mi-2β in the absence of induced DNA damage led to spontaneous DNA damage, increased levels of p53 and p21, reduced cell proliferation, and apoptosis 230, 231. Consistent with these findings, it was shown that NURD subunits are lost during premature and normal aging, leading to changes in higher order chromatin structure and spontaneous DNA damage 232. These data indicate that NURD acts as a gatekeeper of genomic stability.
CHD5 CHD5 was recently found to be the tumor suppressor gene located in 1p36, a region that is commonly deleted in malignancies of epithelial, neural, and hematopoietic origin 233. CHD5 is required for p16Ink4a/p19Arf expression and CHD5 knockdown results in dysregulation of the p19Arf/p53- and p16Ink4a/Rb-mediated tumor suppressive pathways. Interestingly, loss of heterozygosity (LOH) was not observed in most tumor cell lines, indicating that heterozygosity predisposes to tumor formation. The frequency of this deletion in a diversity of cancers indicates that the requirement for CHD5 in transactivation of Ink4a/Arf may be relatively universal.
CHD7 The final Chd family member of note is CHD7. Recently, haploinsufficiency of CHD7 was found to account for 10 of 17 cases of the human CHARGE syndrome, features of which include coloboma of the eye, heart defects, atresia of chonae, renal anomalies, genital anomalies, and ear abnormalities 49. Mice deficient in CHD7 recapitulate many of the human defects as a result of dysregulated transcription of key developmental genes 50, 234, 235. Interestingly, CHD7 was found to collaborate with BRG1 in neural crest-like cells at distal enhancer elements to regulate the neural crest transcriptional program 51. Although this regulation was initially ascribed to the PBAF complex based on the association of CHD7 with BAF180, it is more likely mediated by nBAF that is present in post-mitotic neurons and neural crest cells 121. Genome-wide binding studies further demonstrated that CHD7 colocalizes with Brg1 and H3K4me1 at distal enhancer sites in mouse ES cells, although SWI/SNF clearly has a more extensive role in defining pluripotency as CHD7-deficient mice survive as late as E10.5, when they presumably die due to defects in cardiac development 51, 236, 237.
Chromatin remodeling complexes and cancer
Several CRCs, most prominently the BAF complex, have been implicated in cancer initiation or progression. Early studies demonstrated that many cell lines have lost both BRG1 and hBRM expression and that introduction of BRG1 or hBRM results in slower or arrested growth 238. Other cell lines such as HeLa have partially assembled complexes whose deletion often does not affect the cells and hence these partial complexes seem to be inactive. However, the first definitive evidence that BAF complexes are tumor suppressive came from Versteege and colleagues who found that BAF47 (SNF5) underwent LOH in nearly all cases of pediatric rhabdoid sarcoma, in which the other allele was mutated or silenced by methylation 239. These are extraordinarily aggressive tumors of young children that appear to have a short interval between loss of the second allele and malignancy. This disease can be recapitulated in Snf5+/− mice, which give rise to tumors that are histologically similar to human rhabdoid tumors following LOH 240. Furthermore, conditional inactivation of Baf47 in mice leads to lymphomas with 100% penetrance in a median onset time of 11 weeks 241. Thus BAF47 is a bona fide tumor suppressor that when lost gives rise to tumors more quickly than in any other tumorigenic mouse model. Biallelic loss of BRG1 also occurs in prostate, lung, breast and pancreatic cancer cell lines, and has been shown to result in pediatric rhabdoid sarcomas 27, 242, 243. hBRM and BRG1 were lost in 26% and 23% of small-cell lung cancer cell lines and 76% and 77% of non-small-cell lung cancer cell lines, respectively 244. Consistent with these data, about 10% of Brg1+/− mice spontaneously acquire tumors within one year 22, 245. Among the other BAF subunits, ARID1A (BAF250a) was found to be deleted in 10% of breast carcinomas, 30% of renal carcinomas, and 57% of primary ovarian clear cell carcinomas 246, 247. PBRM1 was deleted in 48% of breast cancer cell lines tested 248, and BRD7 was frequently deleted in a subset of p53 intact human breast tumors 249. Collectively, these data argue that BAF and possibly PBAF complexes play a critical role in suppressing tumorigenesis.
Efforts are now focused on how loss of BRG1 might promote cell growth or dysregulation of the cell cycle. It was shown, for example, that Rb is unable to induce G1 arrest in BRG1/BRM-deficient cell lines, while expression of BRG1 or BRM in these cells restores growth inhibition. BRG1 and Rb physically interact and regulate cell cycle in conjunction with cyclin proteins 238, 250. During G1, Rb-SWI/SNF associates with HDACs to repress the cyclin A and E genes. Phosphorylation of Rb by cyclin D/CDK disrupts the interaction between Rb-SWI/SNF and HDACs, resulting in cyclin E upregulation and progression into S phase. The Rb-SWI/SNF complex remains intact and repressive for the cyclin A gene until cyclin E/CDK2 accumulates and phosphorylates BRG1/BRM and BAF155, leading to disruption of the Rb-SWI/SNF complex and exit from S phase 251. In addition, BRG1 promotes hypophosphorylation of Rb through enhancing the expression of p21 252. In other work, it was shown that SWI/SNF co-associates with p53 and BRCA1, and that SWI/SNF is required for p53- and BRCA1-mediated transcription 253, 254, 255. Similarly, BAF180 and BRD7 regulate transcription of a subset of p53-dependent targets induced during oncogene-induced and replicative senescence 249, 256.
Despite considerable progress on the mechanism of tumorigenesis, a comprehensive understanding of the BAF complex in cancer is lacking. Why deletion of dedicated subunits does not result in similar tumorigenic phenotypes is unknown. One might imagine that loss of BRG1, BAF250a and BAF47 would result in tumorigenesis with similar kinetics and penetrance, but their phenotypes are distinct and tissue specific. The answer may lie in the combinatorial diversity of the BAF complex, which underlies important developmental transitions and appears to have an instructive role in certain cell fates 146. Differential subunit usage could contribute to tissue- or gene-specific effects, which might result in distinct tumorigenic phenotypes upon loss of a given subunit. For example, incorporation of BAF250a or BAF250b is anti- and pro-proliferative, respectively, and it is BAF250a that is lost in human carcinomas 247, 257. If oncogenesis does indeed arise from differential subunit usage, the unique complex interface of malignancy could be specifically targeted using small molecule inhibitors. This theory does not address the specific role of BAF47, which is monomorphic. In this case, it is possible that BAF47 is not required for all of the functions of the BAF complex or that it assembles into specific complexes selectively. Alternatively, loss of BAF47 may result in formation of partial complexes with detrimental and nonphysiological activities. At the molecular level, loss of BAF47 results in upregulation of the Polycomb protein Ezh2 and subsequent repression of p16Ink4a/p19Arf 155, 156. It remains to be seen whether BRG1/hBRM and other subunits of the SWI/SNF complex similarly antagonize Polycomb to suppress tumorigenesis.
The loss or inactivation of CRC-associated proteins is frequently observed in cancer, and many such as BAF47, BAF250, BRG1, and CHD5 are bona fide tumor suppressors. Others appear to play a more supporting role, and are among the class often referred to as passengers in the process of oncogenesis. Mechanistically, CRCs may directly repress growth by interacting with other key tumor suppressors, or they may act independently to maintain genomic integrity. Certainly, members of the BAF complex have been implicated in growth restriction and cell cycle arrest mediated by Rb, p53, BRCA1, and p16ink4a/p19Arf. However, given the unprecedented rapid onset and penetrance of tumors following loss of BAF47, it is likely that interactions with known tumor suppressors will not fully explain its tumor suppressive function. The extensive role of related CRCs in DNA repair, DNA replication, and genomic integrity suggests the intriguing possibility that BAF may also contribute to these surveillance mechanisms. Further mechanistic studies are imperative to understand whether the loss of chromatin remodeling activity is related to a breakdown in these basic cellular processes.
Outlook
The first CRC, ySWI/SNF, was isolated for its role in transcriptional activation of signal-dependent gene targets. Subsequent work in yeast and Drosophila contributed to the strong perception that SWI/SNF and related CRCs are recruited by signal-dependent transcription factors to disrupt nucleosome:DNA contacts to enable transcription. Certainly, it has been well documented that ATP-dependent remodelers are required for transcription and can catalyze nucleosome sliding and exchange on DNA templates in vitro. However, these machines are absolutely critical for many other cellular processes such as DNA replication, recombination, and repair. It is not clear that nucleosomal movement can account for all of the biologic activities of CRCs observed in vivo despite clear affinity of the complexes for nucleosomes. CRCs may bind nucleosomal assemblies where they conduct a range of activities, including, but not limited to, nucleosome remodeling. There is a clear need for experimental model systems that faithfully recapitulate the native state of chromatin with its higher order structure and local histone modifications in order to define and characterize these as yet undefined mechanisms.
This unique class of machines is both omnipresent and hard to define. Despite the similarity between their core enzymatic subunits, their common affinity for nucleosomes, and their common ability to disrupt nucleosomal templates in vitro, there is little functional similarity between members in vivo and as a result, very little predictive power for members of unknown function. In many cases, additional associated subunits radically alter the activity, mechanism, or targeting of the complex to lend it specific function, as in the case of BAF53b in post-mitotic neurons. The utilization of genetic models and genome-wide binding studies to parse out the role of each subunit may reveal that subunit usage is a common theme among CRCs to generate specificity. Capitalizing on this knowledge, we may reveal new therapeutic targets for the treatment of cancer and other CRC-associated diseases.
References
- Kornberg RD. Chromatin structure: a repeating unit of histones and DNA. Science. 1974;184:868–871. doi: 10.1126/science.184.4139.868. [DOI] [PubMed] [Google Scholar]
- Talbert PB, Henikoff S. Histone variants--ancient wrap artists of the epigenome. Nat Rev Mol Cell Biol. 11:264–275. doi: 10.1038/nrm2861. [DOI] [PubMed] [Google Scholar]
- Neigeborn L, Carlson M. Genes affecting the regulation of SUC2 gene expression by glucose repression in Saccharomyces cerevisiae. Genetics. 1984;108:845–858. doi: 10.1093/genetics/108.4.845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peterson CL, Herskowitz I. Characterization of the yeast SWI1, SWI2, and SWI3 genes, which encode a global activator of transcription. Cell. 1992;68:573–583. doi: 10.1016/0092-8674(92)90192-f. [DOI] [PubMed] [Google Scholar]
- Hirschhorn JN, Brown SA, Clark CD, Winston F. Evidence that SNF2/SWI2 and SNF5 activate transcription in yeast by altering chromatin structure. Genes Dev. 1992;6:2288–2298. doi: 10.1101/gad.6.12a.2288. [DOI] [PubMed] [Google Scholar]
- Laurent BC, Treitel MA, Carlson M. Functional interdependence of the yeast SNF2, SNF5, and SNF6 proteins in transcriptional activation. Proc Natl Acad Sci USA. 1991;88:2687–2691. doi: 10.1073/pnas.88.7.2687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cairns BR, Kim YJ, Sayre MH, Laurent BC, Kornberg RD. A multisubunit complex containing the SWI1/ADR6, SWI2/SNF2, SWI3, SNF5, and SNF6 gene products isolated from yeast. Proc Natl Acad Sci USA. 1994;91:1950–1954. doi: 10.1073/pnas.91.5.1950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sudarsanam P, Iyer VR, Brown PO, Winston F. Whole-genome expression analysis of snf/swi mutants of Saccharomyces cerevisiae. Proc Natl Acad Sci USA. 2000;97:3364–3369. doi: 10.1073/pnas.050407197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holstege FC, Jennings EG, Wyrick JJ, et al. Dissecting the regulatory circuitry of a eukaryotic genome. Cell. 1998;95:717–728. doi: 10.1016/s0092-8674(00)81641-4. [DOI] [PubMed] [Google Scholar]
- Laurent BC, Carlson M. Yeast SNF2/SWI2, SNF5, and SNF6 proteins function coordinately with the gene-specific transcriptional activators GAL4 and Bicoid. Genes Dev. 1992;6:1707–1715. doi: 10.1101/gad.6.9.1707. [DOI] [PubMed] [Google Scholar]
- Yoshinaga SK, Peterson CL, Herskowitz I, Yamamoto KR. Roles of SWI1, SWI2, and SWI3 proteins for transcriptional enhancement by steroid receptors. Science. 1992;258:1598–1604. doi: 10.1126/science.1360703. [DOI] [PubMed] [Google Scholar]
- Biggar SR, Crabtree GR. Continuous and widespread roles for the Swi-Snf complex in transcription. EMBO J. 1999;18:2254–2264. doi: 10.1093/emboj/18.8.2254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kennison JA, Tamkun JW. Dosage-dependent modifiers of polycomb and antennapedia mutations in Drosophila. Proc Natl Acad Sci USA. 1988;85:8136–8140. doi: 10.1073/pnas.85.21.8136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamkun JW, Deuring R, Scott MP, et al. brahma: a regulator of Drosophila homeotic genes structurally related to the yeast transcriptional activator SNF2/SWI2. Cell. 1992;68:561–572. doi: 10.1016/0092-8674(92)90191-e. [DOI] [PubMed] [Google Scholar]
- Elfring LK, Deuring R, McCallum CM, Peterson CL, Tamkun JW. Identification and characterization of Drosophila relatives of the yeast transcriptional activator SNF2/SWI2. Mol Cell Biol. 1994;14:2225–2234. doi: 10.1128/mcb.14.4.2225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Armstrong JA, Papoulas O, Daubresse G, et al. The Drosophila BRM complex facilitates global transcription by RNA polymerase II. EMBO J. 2002;21:5245–5254. doi: 10.1093/emboj/cdf517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mohrmann L, Langenberg K, Krijgsveld J, et al. Differential targeting of two distinct SWI/SNF-related Drosophila chromatin-remodeling complexes. Mol Cell Biol. 2004;24:3077–3088. doi: 10.1128/MCB.24.8.3077-3088.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vazquez M, Moore L, Kennison JA. The trithorax group gene osa encodes an ARID-domain protein that genetically interacts with the brahma chromatin-remodeling factor to regulate transcription. Development. 1999;126:733–742. doi: 10.1242/dev.126.4.733. [DOI] [PubMed] [Google Scholar]
- Zraly CB, Marenda DR, Nanchal R, et al. SNR1 is an essential subunit in a subset of Drosophila brm complexes, targeting specific functions during development. Dev Biol. 2003;253:291–308. doi: 10.1016/s0012-1606(02)00011-8. [DOI] [PubMed] [Google Scholar]
- Laurent BC, Yang X, Carlson M. An essential Saccharomyces cerevisiae gene homologous to SNF2 encodes a helicase-related protein in a new family. Mol Cell Biol. 1992;12:1893–1902. doi: 10.1128/mcb.12.4.1893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eisen JA, Sweder KS, Hanawalt PC. Evolution of the SNF2 family of proteins: subfamilies with distinct sequences and functions. Nucleic Acids Res. 1995;23:2715–2723. doi: 10.1093/nar/23.14.2715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bultman S, Gebuhr T, Yee D, et al. A Brg1 null mutation in the mouse reveals functional differences among mammalian SWI/SNF complexes. Mol Cell. 2000;6:1287–1295. doi: 10.1016/s1097-2765(00)00127-1. [DOI] [PubMed] [Google Scholar]
- Ho L, Ronan JL, Wu J, et al. An embryonic stem cell chromatin remodeling complex, esBAF, is essential for embryonic stem cell self-renewal and pluripotency. Proc Natl Acad Sci USA. 2009;106:5181–5186. doi: 10.1073/pnas.0812889106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kidder BL, Palmer S, Knott JG. SWI/SNF-Brg1 regulates self-renewal and occupies core pluripotency-related genes in embryonic stem cells. Stem Cells. 2009;27:317–328. doi: 10.1634/stemcells.2008-0710. [DOI] [PubMed] [Google Scholar]
- Chi TH, Wan M, Lee PP, et al. Sequential roles of Brg, the ATPase subunit of BAF chromatin remodeling complexes, in thymocyte development. Immunity. 2003;19:169–182. doi: 10.1016/s1074-7613(03)00199-7. [DOI] [PubMed] [Google Scholar]
- Reisman DN, Sciarrotta J, Wang W, Funkhouser WK, Weissman BE. Loss of BRG1/BRM in human lung cancer cell lines and primary lung cancers: correlation with poor prognosis. Cancer Res. 2003;63:560–566. [PubMed] [Google Scholar]
- Glaros S, Cirrincione GM, Muchardt C, et al. The reversible epigenetic silencing of BRM: implications for clinical targeted therapy. Oncogene. 2007;26:7058–7066. doi: 10.1038/sj.onc.1210514. [DOI] [PubMed] [Google Scholar]
- Reyes JC, Barra J, Muchardt C, et al. Altered control of cellular proliferation in the absence of mammalian brahma (SNF2alpha) EMBO J. 1998;17:6979–6991. doi: 10.1093/emboj/17.23.6979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barak O, Lazzaro MA, Lane WS, et al. Isolation of human NURF: a regulator of Engrailed gene expression. EMBO J. 2003;22:6089–6100. doi: 10.1093/emboj/cdg582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Landry J, Sharov AA, Piao Y, et al. Essential role of chromatin remodeling protein Bptf in early mouse embryos and embryonic stem cells. PLoS Genet. 2008;4:e1000241. doi: 10.1371/journal.pgen.1000241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wysocka J, Swigut T, Xiao H, et al. A PHD finger of NURF couples histone H3 lysine 4 trimethylation with chromatin remodelling. Nature. 2006;442:86–90. doi: 10.1038/nature04815. [DOI] [PubMed] [Google Scholar]
- Stopka T, Skoultchi AI. The ISWI ATPase Snf2h is required for early mouse development. Proc Natl Acad Sci USA. 2003;100:14097–14102. doi: 10.1073/pnas.2336105100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Collins N, Poot RA, Kukimoto I, et al. An ACF1-ISWI chromatin-remodeling complex is required for DNA replication through heterochromatin. Nat Genet. 2002;32:627–632. doi: 10.1038/ng1046. [DOI] [PubMed] [Google Scholar]
- Poot RA, Bozhenok L, van den Berg DL, et al. The Williams syndrome transcription factor interacts with PCNA to target chromatin remodelling by ISWI to replication foci. Nat Cell Biol. 2004;6:1236–1244. doi: 10.1038/ncb1196. [DOI] [PubMed] [Google Scholar]
- Fyodorov DV, Blower MD, Karpen GH, Kadonaga JT. Acf1 confers unique activities to ACF/CHRAC and promotes the formation rather than disruption of chromatin in vivo. Genes Dev. 2004;18:170–183. doi: 10.1101/gad.1139604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiao A, Li H, Shechter D, et al. WSTF regulates the H2A.X DNA damage response via a novel tyrosine kinase activity. Nature. 2009;457:57–62. doi: 10.1038/nature07668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hakimi MA, Bochar DA, Schmiesing JA, et al. A chromatin remodelling complex that loads cohesin onto human chromosomes. Nature. 2002;418:994–998. doi: 10.1038/nature01024. [DOI] [PubMed] [Google Scholar]
- Gaspar-Maia A, Alajem A, Polesso F, et al. Chd1 regulates open chromatin and pluripotency of embryonic stem cells. Nature. 2009;460:863–868. doi: 10.1038/nature08212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sims RJ, 3rd, Millhouse S, Chen CF, et al. Recognition of trimethylated histone H3 lysine 4 facilitates the recruitment of transcription postinitiation factors and pre-mRNA splicing. Mol Cell. 2007;28:665–676. doi: 10.1016/j.molcel.2007.11.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marfella CG, Ohkawa Y, Coles AH, et al. Mutation of the SNF2 family member Chd2 affects mouse development and survival. J Cell Physiol. 2006;209:162–171. doi: 10.1002/jcp.20718. [DOI] [PubMed] [Google Scholar]
- Ge Q, Nilasena DS, O'Brien CA, Frank MB, Targoff IN. Molecular analysis of a major antigenic region of the 240-kD protein of Mi-2 autoantigen. J Clin Invest. 1995;96:1730–1737. doi: 10.1172/JCI118218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seelig HP, Moosbrugger I, Ehrfeld H, et al. The major dermatomyositis-specific Mi-2 autoantigen is a presumed helicase involved in transcriptional activation. Arthritis Rheum. 1995;38:1389–1399. doi: 10.1002/art.1780381006. [DOI] [PubMed] [Google Scholar]
- Seelig HP, Renz M, Targoff IN, Ge Q, Frank MB. Two forms of the major antigenic protein of the dermatomyositis-specific Mi-2 autoantigen. Arthritis Rheum. 1996;39:1769–1771. doi: 10.1002/art.1780391029. [DOI] [PubMed] [Google Scholar]
- Fujita N, Jaye DL, Geigerman C, et al. MTA3 and the Mi-2/NuRD complex regulate cell fate during B lymphocyte differentiation. Cell. 2004;119:75–86. doi: 10.1016/j.cell.2004.09.014. [DOI] [PubMed] [Google Scholar]
- O'Neill DW, Schoetz SS, Lopez RA, et al. An ikaros-containing chromatin-remodeling complex in adult-type erythroid cells. Mol Cell Biol. 2000;20:7572–7582. doi: 10.1128/mcb.20.20.7572-7582.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams CJ, Naito T, Arco PG, et al. The chromatin remodeler Mi-2beta is required for CD4 expression and T cell development. Immunity. 2004;20:719–733. doi: 10.1016/j.immuni.2004.05.005. [DOI] [PubMed] [Google Scholar]
- Law ME, Templeton KL, Kitange G, et al. Molecular cytogenetic analysis of chromosomes 1 and 19 in glioma cell lines. Cancer Genet Cytogenet. 2005;160:1–14. doi: 10.1016/j.cancergencyto.2004.11.012. [DOI] [PubMed] [Google Scholar]
- White PS, Thompson PM, Gotoh T, et al. Definition and characterization of a region of 1p36.3 consistently deleted in neuroblastoma. Oncogene. 2005;24:2684–2694. doi: 10.1038/sj.onc.1208306. [DOI] [PubMed] [Google Scholar]
- Vissers LE, van Ravenswaaij CM, Admiraal R, et al. Mutations in a new member of the chromodomain gene family cause CHARGE syndrome. Nat Genet. 2004;36:955–957. doi: 10.1038/ng1407. [DOI] [PubMed] [Google Scholar]
- Layman WS, McEwen DP, Beyer LA, et al. Defects in neural stem cell proliferation and olfaction in Chd7 deficient mice indicate a mechanism for hyposmia in human CHARGE syndrome. Hum Mol Genet. 2009;18:1909–1923. doi: 10.1093/hmg/ddp112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bajpai R, Chen DA, Rada-Iglesias A, et al. CHD7 cooperates with PBAF to control multipotent neural crest formation. Nature. 2010;463:958–962. doi: 10.1038/nature08733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishiyama M, Oshikawa K, Tsukada Y, et al. CHD8 suppresses p53-mediated apoptosis through histone H1 recruitment during early embryogenesis. Nat Cell Biol. 2009;11:172–182. doi: 10.1038/ncb1831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marom R, Shur I, Hager GL, Benayahu D. Expression and regulation of CReMM, a chromodomain helicase-DNA-binding (CHD), in marrow stroma derived osteoprogenitors. J Cell Physiol. 2006;207:628–635. doi: 10.1002/jcp.20611. [DOI] [PubMed] [Google Scholar]
- Shur I, Socher R, Benayahu D. In vivo association of CReMM/CHD9 with promoters in osteogenic cells. J Cell Physiol. 2006;207:374–378. doi: 10.1002/jcp.20586. [DOI] [PubMed] [Google Scholar]
- De La Fuente R, Baumann C, Fan T, et al. Lsh is required for meiotic chromosome synapsis and retrotransposon silencing in female germ cells. Nat Cell Biol. 2006;8:1448–1454. doi: 10.1038/ncb1513. [DOI] [PubMed] [Google Scholar]
- Dennis K, Fan T, Geiman T, Yan Q, Muegge K. Lsh, a member of the SNF2 family, is required for genome-wide methylation. Genes Dev. 2001;15:2940–2944. doi: 10.1101/gad.929101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geiman TM, Tessarollo L, Anver MR, et al. Lsh, a SNF2 family member, is required for normal murine development. Biochim Biophys Acta. 2001;1526:211–220. doi: 10.1016/s0304-4165(01)00129-5. [DOI] [PubMed] [Google Scholar]
- Xi S, Geiman TM, Briones V, et al. Lsh participates in DNA methylation and silencing of stem cell genes. Stem Cells. 2009;27:2691–2702. doi: 10.1002/stem.183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xi S, Zhu H, Xu H, et al. Lsh controls Hox gene silencing during development. Proc Natl Acad Sci USA. 2007;104:14366–14371. doi: 10.1073/pnas.0703669104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu H, Geiman TM, Xi S, et al. Lsh is involved in de novo methylation of DNA. Embo J. 2006;25:335–345. doi: 10.1038/sj.emboj.7600925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morrison AJ, Shen X. Chromatin remodelling beyond transcription: the INO80 and SWR1 complexes. Nat Rev Mol Cell Biol. 2009;10:373–384. doi: 10.1038/nrm2693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sikorski TW, Buratowski S. The basal initiation machinery: beyond the general transcription factors. Curr Opin Cell Biol. 2009;21:344–351. doi: 10.1016/j.ceb.2009.03.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stevnsner T, Muftuoglu M, Aamann MD, Bohr VA. The role of Cockayne Syndrome group B (CSB) protein in base excision repair and aging. Mech Ageing Dev. 2008;129:441–448. doi: 10.1016/j.mad.2008.04.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyagawa K, Tsuruga T, Kinomura A, et al. A role for RAD54B in homologous recombination in human cells. EMBO J. 2002;21:175–180. doi: 10.1093/emboj/21.1.175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanaka K, Hiramoto T, Fukuda T, Miyagawa K. A novel human rad54 homologue, Rad54B, associates with Rad51. J Biol Chem. 2000;275:26316–26321. doi: 10.1074/jbc.M910306199. [DOI] [PubMed] [Google Scholar]
- Tanaka K, Kagawa W, Kinebuchi T, Kurumizaka H, Miyagawa K. Human Rad54B is a double-stranded DNA-dependent ATPase and has biochemical properties different from its structural homolog in yeast, Tid1/Rdh54. Nucleic Acids Res. 2002;30:1346–1353. doi: 10.1093/nar/30.6.1346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Essers J, Hendriks RW, Swagemakers SM, et al. Disruption of mouse RAD54 reduces ionizing radiation resistance and homologous recombination. Cell. 1997;89:195–204. doi: 10.1016/s0092-8674(00)80199-3. [DOI] [PubMed] [Google Scholar]
- Essers J, van Steeg H, de Wit J, et al. Homologous and non-homologous recombination differentially affect DNA damage repair in mice. EMBO J. 2000;19:1703–1710. doi: 10.1093/emboj/19.7.1703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mazin AV, Mazina OM, Bugreev DV, Rossi MJ. Rad54, the motor of homologous recombination. DNA Repair (Amst) 2010;9:286–302. doi: 10.1016/j.dnarep.2009.12.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldberg AD, Banaszynski LA, Noh KM, et al. Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell. 2010;140:678–691. doi: 10.1016/j.cell.2010.01.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Law MJ, Lower KM, Voon HP, et al. ATR-X syndrome protein targets tandem repeats and influences allele-specific expression in a size-dependent manner. Cell. 2010;143:367–378. doi: 10.1016/j.cell.2010.09.023. [DOI] [PubMed] [Google Scholar]
- Lewis PW, Elsaesser SJ, Noh KM, Stadler SC, Allis CD. Daxx is an H3.3-specific histone chaperone and cooperates with ATRX in replication-independent chromatin assembly at telomeres. Proc Natl Acad Sci USA. 2010;107:14075–14080. doi: 10.1073/pnas.1008850107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mattera L, Courilleau C, Legube G, et al. The E1A-associated p400 protein modulates cell fate decisions by the regulation of ROS homeostasis. PLoS Genet. 2010;6:e1000983. doi: 10.1371/journal.pgen.1000983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mattera L, Escaffit F, Pillaire MJ, et al. The p400/Tip60 ratio is critical for colorectal cancer cell proliferation through DNA damage response pathways. Oncogene. 2009;28:1506–1517. doi: 10.1038/onc.2008.499. [DOI] [PubMed] [Google Scholar]
- Ueda T, Watanabe-Fukunaga R, Ogawa H, et al. Critical role of the p400/mDomino chromatin-remodeling ATPase in embryonic hematopoiesis. Genes Cells. 2007;12:581–592. doi: 10.1111/j.1365-2443.2007.01080.x. [DOI] [PubMed] [Google Scholar]
- Xu Y, Sun Y, Jiang X, et al. The p400 ATPase regulates nucleosome stability and chromatin ubiquitination during DNA repair. J Cell Biol. 2010;191:31–43. doi: 10.1083/jcb.201001160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujii T, Ueda T, Nagata S, Fukunaga R. Essential role of p400/mDomino chromatin-remodeling ATPase in bone marrow hematopoiesis and cell-cycle progression. J Biol Chem. 2010;285:30214–30223. doi: 10.1074/jbc.M110.104513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gevry N, Chan HM, Laflamme L, Livingston DM, Gaudreau L. p21 transcription is regulated by differential localization of histone H2A.Z. Genes Dev. 2007;21:1869–1881. doi: 10.1101/gad.1545707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bansbach CE, Betous R, Lovejoy CA, Glick GG, Cortez D. The annealing helicase SMARCAL1 maintains genome integrity at stalled replication forks. Genes Dev. 2009;23:2405–2414. doi: 10.1101/gad.1839909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ciccia A, Bredemeyer AL, Sowa ME, et al. The SIOD disorder protein SMARCAL1 is an RPA-interacting protein involved in replication fork restart. Genes Dev. 2009;23:2415–2425. doi: 10.1101/gad.1832309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Postow L, Woo EM, Chait BT, Funabiki H. Identification of SMARCAL1 as a component of the DNA damage response. J Biol Chem. 2009;284:35951–35961. doi: 10.1074/jbc.M109.048330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yusufzai T, Kong X, Yokomori K, Kadonaga JT. The annealing helicase HARP is recruited to DNA repair sites via an interaction with RPA. Genes Dev. 2009;23:2400–2404. doi: 10.1101/gad.1831509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuan J, Ghosal G, Chen J. The annealing helicase HARP protects stalled replication forks. Genes Dev. 2009;23:2394–2399. doi: 10.1101/gad.1836409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moinova HR, Chen WD, Shen L, et al. HLTF gene silencing in human colon cancer. Proc Natl Acad Sci USA. 2002;99:4562–4567. doi: 10.1073/pnas.062459899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Motegi A, Liaw HJ, Lee KY, et al. Polyubiquitination of proliferating cell nuclear antigen by HLTF and SHPRH prevents genomic instability from stalled replication forks. Proc Natl Acad Sci USA. 2008;105:12411–12416. doi: 10.1073/pnas.0805685105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Unk I, Hajdu I, Fatyol K, et al. Human HLTF functions as a ubiquitin ligase for proliferating cell nuclear antigen polyubiquitination. Proc Natl Acad Sci USA. 2008;105:3768–3773. doi: 10.1073/pnas.0800563105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruhl DD, Jin J, Cai Y, et al. Purification of a human SRCAP complex that remodels chromatin by incorporating the histone variant H2A.Z into nucleosomes. Biochemistry. 2006;45:5671–5677. doi: 10.1021/bi060043d. [DOI] [PubMed] [Google Scholar]
- Wong MM, Cox LK, Chrivia JC. The chromatin remodeling protein, SRCAP, is critical for deposition of the histone variant H2A.Z at promoters. J Biol Chem. 2007;282:26132–26139. doi: 10.1074/jbc.M703418200. [DOI] [PubMed] [Google Scholar]
- Ogawa H, Komatsu T, Hiraoka Y, Morohashi K. Transcriptional Suppression by Transient Recruitment of ARIP4 to Sumoylated nuclear receptor Ad4BP/SF-1. Mol Biol Cell. 2009;20:4235–4245. doi: 10.1091/mbc.E08-12-1247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laurent BC, Treich I, Carlson M. The yeast SNF2/SWI2 protein has DNA-stimulated ATPase activity required for transcriptional activation. Genes Dev. 1993;7:583–591. doi: 10.1101/gad.7.4.583. [DOI] [PubMed] [Google Scholar]
- Olave IA, Reck-Peterson SL, Crabtree GR. Nuclear actin and actin-related proteins in chromatin remodeling. Annu Rev Biochem. 2002;71:755–781. doi: 10.1146/annurev.biochem.71.110601.135507. [DOI] [PubMed] [Google Scholar]
- Mizuguchi G, Shen X, Landry J, et al. ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex. Science. 2004;303:343–348. doi: 10.1126/science.1090701. [DOI] [PubMed] [Google Scholar]
- Aoyagi S, Hayes JJ. hSWI/SNF-catalyzed nucleosome sliding does not occur solely via a twist-diffusion mechanism. Mol Cell Biol. 2002;22:7484–7490. doi: 10.1128/MCB.22.21.7484-7490.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strohner R, Wachsmuth M, Dachauer K, et al. A 'loop recapture' mechanism for ACF-dependent nucleosome remodeling. Nat Struct Mol Biol. 2005;12:683–690. doi: 10.1038/nsmb966. [DOI] [PubMed] [Google Scholar]
- Blosser TR, Yang JG, Stone MD, Narlikar GJ, Zhuang X. Dynamics of nucleosome remodelling by individual ACF complexes. Nature. 2009;462:1022–1027. doi: 10.1038/nature08627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zofall M, Persinger J, Kassabov SR, Bartholomew B. Chromatin remodeling by ISW2 and SWI/SNF requires DNA translocation inside the nucleosome. Nat Struct Mol Biol. 2006;13:339–346. doi: 10.1038/nsmb1071. [DOI] [PubMed] [Google Scholar]
- Whitehouse I, Stockdale C, Flaus A, Szczelkun MD, Owen-Hughes T. Evidence for DNA translocation by the ISWI chromatin-remodeling enzyme. Mol Cell Biol. 2003;23:1935–1945. doi: 10.1128/MCB.23.6.1935-1945.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saha A, Wittmeyer J, Cairns BR. Chromatin remodeling by RSC involves ATP-dependent DNA translocation. Genes Dev. 2002;16:2120–2134. doi: 10.1101/gad.995002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saha A, Wittmeyer J, Cairns BR. Chromatin remodeling through directional DNA translocation from an internal nucleosomal site. Nat Struct Mol Biol. 2005;12:747–755. doi: 10.1038/nsmb973. [DOI] [PubMed] [Google Scholar]
- Schwanbeck R, Xiao H, Wu C. Spatial contacts and nucleosome step movements induced by the NURF chromatin remodeling complex. J Biol Chem. 2004;279:39933–39941. doi: 10.1074/jbc.M406060200. [DOI] [PubMed] [Google Scholar]
- Dechassa ML, Zhang B, Horowitz-Scherer R, et al. Architecture of the SWI/SNF-nucleosome complex. Mol Cell Biol. 2008;28:6010–6021. doi: 10.1128/MCB.00693-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chaban Y, Ezeokonkwo C, Chung WH, et al. Structure of a RSC-nucleosome complex and insights into chromatin remodeling. Nat Struct Mol Biol. 2008;15:1272–1277. doi: 10.1038/nsmb.1524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leschziner AE, Lemon B, Tjian R, Nogales E. Structural studies of the human PBAF chromatin-remodeling complex. Structure. 2005;13:267–275. doi: 10.1016/j.str.2004.12.008. [DOI] [PubMed] [Google Scholar]
- Lorch Y, Maier-Davis B, Kornberg RD. Mechanism of chromatin remodeling. Proc Natl Acad Sci USA. 2010;107:3458–3462. doi: 10.1073/pnas.1000398107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dang W, Kagalwala MN, Bartholomew B. The Dpb4 subunit of ISW2 is anchored to extranucleosomal DNA. J Biol Chem. 2007;282:19418–19425. doi: 10.1074/jbc.M700640200. [DOI] [PubMed] [Google Scholar]
- Kagalwala MN, Glaus BJ, Dang W, Zofall M, Bartholomew B. Topography of the ISW2-nucleosome complex: insights into nucleosome spacing and chromatin remodeling. EMBO J. 2004;23:2092–2104. doi: 10.1038/sj.emboj.7600220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gangaraju VK, Bartholomew B. Dependency of ISW1a chromatin remodeling on extranucleosomal DNA. Mol Cell Biol. 2007;27:3217–3225. doi: 10.1128/MCB.01731-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Racki LR, Yang JG, Naber N, et al. The chromatin remodeller ACF acts as a dimeric motor to space nucleosomes. Nature. 2009;462:1016–1021. doi: 10.1038/nature08621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fyodorov DV, Kadonaga JT. Dynamics of ATP-dependent chromatin assembly by ACF. Nature. 2002;418:897–900. doi: 10.1038/nature00929. [DOI] [PubMed] [Google Scholar]
- Zofall M, Persinger J, Bartholomew B. Functional role of extranucleosomal DNA and the entry site of the nucleosome in chromatin remodeling by ISW2. Mol Cell Biol. 2004;24:10047–10057. doi: 10.1128/MCB.24.22.10047-10057.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang JG, Madrid TS, Sevastopoulos E, Narlikar GJ. The chromatin-remodeling enzyme ACF is an ATP-dependent DNA length sensor that regulates nucleosome spacing. Nat Struct Mol Biol. 2006;13:1078–1083. doi: 10.1038/nsmb1170. [DOI] [PubMed] [Google Scholar]
- Gangaraju VK, Prasad P, Srour A, Kagalwala MN, Bartholomew B. Conformational changes associated with template commitment in ATP-dependent chromatin remodeling by ISW2. Mol Cell. 2009;35:58–69. doi: 10.1016/j.molcel.2009.05.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clapier CR, Nightingale KP, Becker PB. A critical epitope for substrate recognition by the nucleosome remodeling ATPase ISWI. Nucleic Acids Res. 2002;30:649–655. doi: 10.1093/nar/30.3.649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamiche A, Kang JG, Dennis C, Xiao H, Wu C. Histone tails modulate nucleosome mobility and regulate ATP-dependent nucleosome sliding by NURF. Proc Natl Acad Sci USA. 2001;98:14316–14321. doi: 10.1073/pnas.251421398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shogren-Knaak M, Ishii H, Sun JM, et al. Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science. 2006;311:844–847. doi: 10.1126/science.1124000. [DOI] [PubMed] [Google Scholar]
- Horn PJ, Carruthers LM, Logie C, et al. Phosphorylation of linker histones regulates ATP-dependent chromatin remodeling enzymes. Nat Struct Biol. 2002;9:263–267. doi: 10.1038/nsb776. [DOI] [PubMed] [Google Scholar]
- Saeki H, Ohsumi K, Aihara H, et al. Linker histone variants control chromatin dynamics during early embryogenesis. Proc Natl Acad Sci USA. 2005;102:5697–5702. doi: 10.1073/pnas.0409824102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramachandran A, Omar M, Cheslock P, Schnitzler GR. Linker histone H1 modulates nucleosome remodeling by human SWI/SNF. J Biol Chem. 2003;278:48590–48601. doi: 10.1074/jbc.M309033200. [DOI] [PubMed] [Google Scholar]
- Thoma NH, Czyzewski BK, Alexeev AA, et al. Structure of the SWI2/SNF2 chromatin-remodeling domain of eukaryotic Rad54. Nat Struct Mol Biol. 2005;12:350–356. doi: 10.1038/nsmb919. [DOI] [PubMed] [Google Scholar]
- Ho L, Crabtree GR. Chromatin remodelling during development. Nature. 463:474–484. doi: 10.1038/nature08911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lessard J, Wu JI, Ranish JA, et al. An essential switch in subunit composition of a chromatin remodeling complex during neural development. Neuron. 2007;55:201–215. doi: 10.1016/j.neuron.2007.06.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicodeme E, Jeffrey KL, Schaefer U, et al. Suppression of inflammation by a synthetic histone mimic. Nature. 2010;468:1119–1123. doi: 10.1038/nature09589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Filippakopoulos P, Qi J, Picaud S, et al. Selective inhibition of BET bromodomains. Nature. 2010;468:1067–1073. doi: 10.1038/nature09504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao K, Wang W, Rando OJ, et al. Rapid and phosphoinositol-dependent binding of the SWI/SNF-like BAF complex to chromatin after T lymphocyte receptor signaling. Cell. 1998;95:625–636. doi: 10.1016/s0092-8674(00)81633-5. [DOI] [PubMed] [Google Scholar]
- Rando OJ, Zhao K, Janmey P, Crabtree GR. Phosphatidylinositol-dependent actin filament binding by the SWI/SNF-like BAF chromatin remodeling complex. Proc Natl Acad Sci USA. 2002;99:2824–2829. doi: 10.1073/pnas.032662899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lymn RW, Taylor EW. Mechanism of adenosine triphosphate hydrolysis by actomyosin. Biochemistry. 1971;10:4617–4624. doi: 10.1021/bi00801a004. [DOI] [PubMed] [Google Scholar]
- Muller J, Oma Y, Vallar L, et al. Sequence and comparative genomic analysis of actin-related proteins. Mol Biol Cell. 2005;16:5736–5748. doi: 10.1091/mbc.E05-06-0508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szerlong H, Hinata K, Viswanathan R, et al. The HSA domain binds nuclear actin-related proteins to regulate chromatin-remodeling ATPases. Nat Struct Mol Biol. 2008;15:469–476. doi: 10.1038/nsmb.1403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Downs JA, Allard S, Jobin-Robitaille O, et al. Binding of chromatin-modifying activities to phosphorylated histone H2A at DNA damage sites. Mol Cell. 2004;16:979–990. doi: 10.1016/j.molcel.2004.12.003. [DOI] [PubMed] [Google Scholar]
- Sunada R, Gorzer I, Oma Y, et al. The nuclear actin-related protein Act3p/Arp4p is involved in the dynamics of chromatin-modulating complexes. Yeast. 2005;22:753–768. doi: 10.1002/yea.1239. [DOI] [PubMed] [Google Scholar]
- Wu WH, Alami S, Luk E, et al. Swc2 is a widely conserved H2AZ-binding module essential for ATP-dependent histone exchange. Nat Struct Mol Biol. 2005;12:1064–1071. doi: 10.1038/nsmb1023. [DOI] [PubMed] [Google Scholar]
- Shen X, Ranallo R, Choi E, Wu C. Involvement of actin-related proteins in ATP-dependent chromatin remodeling. Mol Cell. 2003;12:147–155. doi: 10.1016/s1097-2765(03)00264-8. [DOI] [PubMed] [Google Scholar]
- Wu JI, Lessard J, Olave IA, et al. Regulation of dendritic development by neuron-specific chromatin remodeling complexes. Neuron. 2007;56:94–108. doi: 10.1016/j.neuron.2007.08.021. [DOI] [PubMed] [Google Scholar]
- Parrish JZ, Kim MD, Jan LY, Jan YN. Genome-wide analyses identify transcription factors required for proper morphogenesis of Drosophila sensory neuron dendrites. Genes Dev. 2006;20:820–835. doi: 10.1101/gad.1391006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tea JS, Luo L. The chromatin remodeling factor Bap55 functions through the TIP60 complex to regulate olfactory projection neuron dendrite targeting. Neural Dev. 2011;6:5. doi: 10.1186/1749-8104-6-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Theriot JA, Rosenblatt J, Portnoy DA, Goldschmidt-Clermont PJ, Mitchison TJ. Involvement of profilin in the actin-based motility of L. monocytogenes in cells and in cell-free extracts. Cell. 1994;76:505–517. doi: 10.1016/0092-8674(94)90114-7. [DOI] [PubMed] [Google Scholar]
- Khavari PA, Peterson CL, Tamkun JW, Mendel DB, Crabtree GR. BRG1 contains a conserved domain of the SWI2/SNF2 family necessary for normal mitotic growth and transcription. Nature. 1993;366:170–174. doi: 10.1038/366170a0. [DOI] [PubMed] [Google Scholar]
- Wang W, Cote J, Xue Y, et al. Purification and biochemical heterogeneity of the mammalian SWI-SNF complex. EMBO J. 1996;15:5370–5382. [PMC free article] [PubMed] [Google Scholar]
- Wang W, Xue Y, Zhou S, et al. Diversity and specialization of mammalian SWI/SNF complexes. Genes Dev. 1996;10:2117–2130. doi: 10.1101/gad.10.17.2117. [DOI] [PubMed] [Google Scholar]
- Roberts CW, Orkin SH. The SWI/SNF complex--chromatin and cancer. Nat Rev Cancer. 2004;4:133–142. doi: 10.1038/nrc1273. [DOI] [PubMed] [Google Scholar]
- Elfring LK, Daniel C, Papoulas O, et al. Genetic analysis of brahma: the Drosophila homolog of the yeast chromatin remodeling factor SWI2/SNF2. Genetics. 1998;148:251–265. doi: 10.1093/genetics/148.1.251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaeser MD, Aslanian A, Dong MQ, Yates JR, 3rd, Emerson BM. BRD7, a novel PBAF-specific SWI/SNF subunit, is required for target gene activation and repression in embryonic stem cells. J Biol Chem. 2008;283:32254–32263. doi: 10.1074/jbc.M806061200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao X, Tate P, Hu P, et al. ES cell pluripotency and germ-layer formation require the SWI/SNF chromatin remodeling component BAF250a. Proc Natl Acad Sci USA. 2008;105:6656–6661. doi: 10.1073/pnas.0801802105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoo AS, Staahl BT, Chen L, Crabtree GR. MicroRNA-mediated switching of chromatin-remodelling complexes in neural development. Nature. 2009;460:642–646. doi: 10.1038/nature08139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lickert H, Takeuchi JK, Von Both I, et al. Baf60c is essential for function of BAF chromatin remodelling complexes in heart development. Nature. 2004;432:107–112. doi: 10.1038/nature03071. [DOI] [PubMed] [Google Scholar]
- Ieda M, Fu JD, Delgado-Olguin P, et al. Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell. 142:375–386. doi: 10.1016/j.cell.2010.07.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ho L, Jothi R, Ronan JL, et al. An embryonic stem cell chromatin remodeling complex, esBAF, is an essential component of the core pluripotency transcriptional network. Proc Natl Acad Sci USA. 2009;106:5187–5191. doi: 10.1073/pnas.0812888106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heintzman ND, Stuart RK, Hon G, et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat Genet. 2007;39:311–318. doi: 10.1038/ng1966. [DOI] [PubMed] [Google Scholar]
- Yeom YI, Fuhrmann G, Ovitt CE, et al. Germline regulatory element of Oct-4 specific for the totipotent cycle of embryonal cells. Development. 1996;122:881–894. doi: 10.1242/dev.122.3.881. [DOI] [PubMed] [Google Scholar]
- Boyer LA, Lee TI, Cole MF, et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell. 2005;122:947–956. doi: 10.1016/j.cell.2005.08.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chi TH, Wan M, Zhao K, et al. Reciprocal regulation of CD4/CD8 expression by SWI/SNF-like BAF complexes. Nature. 2002;418:195–199. doi: 10.1038/nature00876. [DOI] [PubMed] [Google Scholar]
- Kim SI, Bresnick EH, Bultman SJ. BRG1 directly regulates nucleosome structure and chromatin looping of the alpha globin locus to activate transcription. Nucleic Acids Res. 2009;37:6019–6027. doi: 10.1093/nar/gkp677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim SI, Bultman SJ, Kiefer CM, Dean A, Bresnick EH. BRG1 requirement for long-range interaction of a locus control region with a downstream promoter. Proc Natl Acad Sci USA. 2009;106:2259–2264. doi: 10.1073/pnas.0806420106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wan M, Zhang J, Lai D, et al. Molecular basis of CD4 repression by the Swi/Snf-like BAF chromatin remodeling complex. Eur J Immunol. 2009;39:580–588. doi: 10.1002/eji.200838909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kia SK, Gorski MM, Giannakopoulos S, Verrijzer CP. SWI/SNF mediates polycomb eviction and epigenetic reprogramming of the INK4b-ARF-INK4a locus. Mol Cell Biol. 2008;28:3457–3464. doi: 10.1128/MCB.02019-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson BG, Wang X, Shen X, et al. Epigenetic antagonism between polycomb and SWI/SNF complexes during oncogenic transformation. Cancer Cell. 2010;18:316–328. doi: 10.1016/j.ccr.2010.09.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cairns BR, Lorch Y, Li Y, et al. RSC, an essential, abundant chromatin-remodeling complex. Cell. 1996;87:1249–1260. doi: 10.1016/s0092-8674(00)81820-6. [DOI] [PubMed] [Google Scholar]
- Xue Y, Canman JC, Lee CS, et al. The human SWI/SNF-B chromatin-remodeling complex is related to yeast rsc and localizes at kinetochores of mitotic chromosomes. Proc Natl Acad Sci USA. 2000;97:13015–13020. doi: 10.1073/pnas.240208597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yan Z, Cui K, Murray DM, et al. PBAF chromatin-remodeling complex requires a novel specificity subunit, BAF200, to regulate expression of selective interferon-responsive genes. Genes Dev. 2005;19:1662–1667. doi: 10.1101/gad.1323805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Damelin M, Simon I, Moy TI, et al. The genome-wide localization of Rsc9, a component of the RSC chromatin-remodeling complex, changes in response to stress. Mol Cell. 2002;9:563–573. doi: 10.1016/s1097-2765(02)00475-6. [DOI] [PubMed] [Google Scholar]
- Angus-Hill ML, Schlichter A, Roberts D, et al. A Rsc3/Rsc30 zinc cluster dimer reveals novel roles for the chromatin remodeler RSC in gene expression and cell cycle control. Mol Cell. 2001;7:741–751. doi: 10.1016/s1097-2765(01)00219-2. [DOI] [PubMed] [Google Scholar]
- Cao Y, Cairns BR, Kornberg RD, Laurent BC. Sfh1p, a component of a novel chromatin-remodeling complex, is required for cell cycle progression. Mol Cell Biol. 1997;17:3323–3334. doi: 10.1128/mcb.17.6.3323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du J, Nasir I, Benton BK, Kladde MP, Laurent BC. Sth1p, a Saccharomyces cerevisiae Snf2p/Swi2p homolog, is an essential ATPase in RSC and differs from Snf/Swi in its interactions with histones and chromatin-associated proteins. Genetics. 1998;150:987–1005. doi: 10.1093/genetics/150.3.987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baetz KK, Krogan NJ, Emili A, Greenblatt J, Hieter P. The ctf13-30/CTF13 genomic haploinsufficiency modifier screen identifies the yeast chromatin remodeling complex RSC, which is required for the establishment of sister chromatid cohesion. Mol Cell Biol. 2004;24:1232–1244. doi: 10.1128/MCB.24.3.1232-1244.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang J, Hsu JM, Laurent BC. The RSC nucleosome-remodeling complex is required for Cohesin's association with chromosome arms. Mol Cell. 2004;13:739–750. doi: 10.1016/s1097-2765(04)00103-0. [DOI] [PubMed] [Google Scholar]
- Hsu JM, Huang J, Meluh PB, Laurent BC. The yeast RSC chromatin-remodeling complex is required for kinetochore function in chromosome segregation. Mol Cell Biol. 2003;23:3202–3215. doi: 10.1128/MCB.23.9.3202-3215.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Campsteijn C, Wijnands-Collin AM, Logie C. Reverse genetic analysis of the yeast RSC chromatin remodeler reveals a role for RSC3 and SNF5 homolog 1 in ploidy maintenance. PLoS Genet. 2007;3:e92. doi: 10.1371/journal.pgen.0030092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lemon B, Inouye C, King DS, Tjian R. Selectivity of chromatin-remodelling cofactors for ligand-activated transcription. Nature. 2001;414:924–928. doi: 10.1038/414924a. [DOI] [PubMed] [Google Scholar]
- Wang Z, Zhai W, Richardson JA, et al. Polybromo protein BAF180 functions in mammalian cardiac chamber maturation. Genes Dev. 2004;18:3106–3116. doi: 10.1101/gad.1238104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carrera I, Zavadil J, Treisman JE. Two subunits specific to the PBAP chromatin remodeling complex have distinct and redundant functions during Drosophila development. Mol Cell Biol. 2008;28:5238–5250. doi: 10.1128/MCB.00747-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- West SC. Processing of recombination intermediates by the RuvABC proteins. Annu Rev Genet. 1997;31:213–244. doi: 10.1146/annurev.genet.31.1.213. [DOI] [PubMed] [Google Scholar]
- Shen X, Mizuguchi G, Hamiche A, Wu C. A chromatin remodelling complex involved in transcription and DNA processing. Nature. 2000;406:541–544. doi: 10.1038/35020123. [DOI] [PubMed] [Google Scholar]
- Wu S, Shi Y, Mulligan P, et al. A YY1-INO80 complex regulates genomic stability through homologous recombination-based repair. Nat Struct Mol Biol. 2007;14:1165–1172. doi: 10.1038/nsmb1332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsukuda T, Fleming AB, Nickoloff JA, Osley MA. Chromatin remodelling at a DNA double-strand break site in Saccharomyces cerevisiae. Nature. 2005;438:379–383. doi: 10.1038/nature04148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Attikum H, Fritsch O, Gasser SM. Distinct roles for SWR1 and INO80 chromatin remodeling complexes at chromosomal double-strand breaks. EMBO J. 2007;26:4113–4125. doi: 10.1038/sj.emboj.7601835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ebbert R, Birkmann A, Schuller HJ. The product of the SNF2/SWI2 paralogue INO80 of Saccharomyces cerevisiae required for efficient expression of various yeast structural genes is part of a high-molecular-weight protein complex. Mol Microbiol. 1999;32:741–751. doi: 10.1046/j.1365-2958.1999.01390.x. [DOI] [PubMed] [Google Scholar]
- Jonsson ZO, Jha S, Wohlschlegel JA, Dutta A. Rvb1p/Rvb2p recruit Arp5p and assemble a functional Ino80 chromatin remodeling complex. Mol Cell. 2004;16:465–477. doi: 10.1016/j.molcel.2004.09.033. [DOI] [PubMed] [Google Scholar]
- Krogan NJ, Keogh MC, Datta N, et al. A Snf2 family ATPase complex required for recruitment of the histone H2A variant Htz1. Mol Cell. 2003;12:1565–1576. doi: 10.1016/s1097-2765(03)00497-0. [DOI] [PubMed] [Google Scholar]
- Wu WH, Wu CH, Ladurner A, et al. N terminus of Swr1 binds to histone H2AZ and provides a platform for subunit assembly in the chromatin remodeling complex. J Biol Chem. 2009;284:6200–6207. doi: 10.1074/jbc.M808830200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kusch T, Florens L, Macdonald WH, et al. Acetylation by Tip60 is required for selective histone variant exchange at DNA lesions. Science. 2004;306:2084–2087. doi: 10.1126/science.1103455. [DOI] [PubMed] [Google Scholar]
- van Attikum H, Fritsch O, Hohn B, Gasser SM. Recruitment of the INO80 complex by H2A phosphorylation links ATP-dependent chromatin remodeling with DNA double-strand break repair. Cell. 2004;119:777–788. doi: 10.1016/j.cell.2004.11.033. [DOI] [PubMed] [Google Scholar]
- Morrison AJ, Highland J, Krogan NJ, et al. INO80 and gamma-H2AX interaction links ATP-dependent chromatin remodeling to DNA damage repair. Cell. 2004;119:767–775. doi: 10.1016/j.cell.2004.11.037. [DOI] [PubMed] [Google Scholar]
- Morrison AJ, Kim JA, Person MD, et al. Mec1/Tel1 phosphorylation of the INO80 chromatin remodeling complex influences DNA damage checkpoint responses. Cell. 2007;130:499–511. doi: 10.1016/j.cell.2007.06.010. [DOI] [PubMed] [Google Scholar]
- Papamichos-Chronakis M, Peterson CL. The Ino80 chromatin-remodeling enzyme regulates replisome function and stability. Nat Struct Mol Biol. 2008;15:338–345. doi: 10.1038/nsmb.1413. [DOI] [PubMed] [Google Scholar]
- Falbo KB, Alabert C, Katou Y, et al. Involvement of a chromatin remodeling complex in damage tolerance during DNA replication. Nat Struct Mol Biol. 2009;16:1167–1172. doi: 10.1038/nsmb.1686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pan X, Ye P, Yuan DS, et al. A DNA integrity network in the yeast Saccharomyces cerevisiae. Cell. 2006;124:1069–1081. doi: 10.1016/j.cell.2005.12.036. [DOI] [PubMed] [Google Scholar]
- Collins SR, Miller KM, Maas NL, et al. Functional dissection of protein complexes involved in yeast chromosome biology using a genetic interaction map. Nature. 2007;446:806–810. doi: 10.1038/nature05649. [DOI] [PubMed] [Google Scholar]
- Yu EY, Steinberg-Neifach O, Dandjinou AT, et al. Regulation of telomere structure and functions by subunits of the INO80 chromatin remodeling complex. Mol Cell Biol. 2007;27:5639–5649. doi: 10.1128/MCB.00418-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doyon Y, Selleck W, Lane WS, Tan S, Cote J. Structural and functional conservation of the NuA4 histone acetyltransferase complex from yeast to humans. Mol Cell Biol. 2004;24:1884–1896. doi: 10.1128/MCB.24.5.1884-1896.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai Y, Jin J, Tomomori-Sato C, et al. Identification of new subunits of the multiprotein mammalian TRRAP/TIP60-containing histone acetyltransferase complex. J Biol Chem. 2003;278:42733–42736. doi: 10.1074/jbc.C300389200. [DOI] [PubMed] [Google Scholar]
- Ikura T, Ogryzko VV, Grigoriev M, et al. Involvement of the TIP60 histone acetylase complex in DNA repair and apoptosis. Cell. 2000;102:463–473. doi: 10.1016/s0092-8674(00)00051-9. [DOI] [PubMed] [Google Scholar]
- Fuchs M, Gerber J, Drapkin R, et al. The p400 complex is an essential E1A transformation target. Cell. 2001;106:297–307. doi: 10.1016/s0092-8674(01)00450-0. [DOI] [PubMed] [Google Scholar]
- Ikura T, Tashiro S, Kakino A, et al. DNA damage-dependent acetylation and ubiquitination of H2AX enhances chromatin dynamics. Mol Cell Biol. 2007;27:7028–7040. doi: 10.1128/MCB.00579-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun Y, Jiang X, Xu Y, et al. Histone H3 methylation links DNA damage detection to activation of the tumour suppressor Tip60. Nat Cell Biol. 2009;11:1376–1382. doi: 10.1038/ncb1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sykes SM, Mellert HS, Holbert MA, et al. Acetylation of the p53 DNA-binding domain regulates apoptosis induction. Mol Cell. 2006;24:841–851. doi: 10.1016/j.molcel.2006.11.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang Y, Luo J, Zhang W, Gu W. Tip60-dependent acetylation of p53 modulates the decision between cell-cycle arrest and apoptosis. Mol Cell. 2006;24:827–839. doi: 10.1016/j.molcel.2006.11.021. [DOI] [PubMed] [Google Scholar]
- Gorrini C, Squatrito M, Luise C, et al. Tip60 is a haplo-insufficient tumour suppressor required for an oncogene-induced DNA damage response. Nature. 2007;448:1063–1067. doi: 10.1038/nature06055. [DOI] [PubMed] [Google Scholar]
- Fazzio TG, Huff JT, Panning B. An RNAi screen of chromatin proteins identifies Tip60-p400 as a regulator of embryonic stem cell identity. Cell. 2008;134:162–174. doi: 10.1016/j.cell.2008.05.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu Y, Fisher JB, Koprowski S, et al. Homozygous disruption of the Tip60 gene causes early embryonic lethality. Dev Dyn. 2009;238:2912–2921. doi: 10.1002/dvdy.22110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu X, Singh N, Donnelly C, Boimel P, Elefant F. The cloning and characterization of the histone acetyltransferase human homolog Dmel\\TIP60 in Drosophila melanogaster: Dmel\\TIP60 is essential for multicellular development. Genetics. 2007;175:1229–1240. doi: 10.1534/genetics.106.063685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bugreev DV, Mazina OM, Mazin AV. Rad54 protein promotes branch migration of Holliday junctions. Nature. 2006;442:590–593. doi: 10.1038/nature04889. [DOI] [PubMed] [Google Scholar]
- Tsukiyama T, Daniel C, Tamkun J, Wu C. ISWI, a member of the SWI2/SNF2 ATPase family, encodes the 140 kDa subunit of the nucleosome remodeling factor. Cell. 1995;83:1021–1026. doi: 10.1016/0092-8674(95)90217-1. [DOI] [PubMed] [Google Scholar]
- Ito T, Levenstein ME, Fyodorov DV, et al. ACF consists of two subunits, Acf1 and ISWI, that function cooperatively in the ATP-dependent catalysis of chromatin assembly. Genes Dev. 1999;13:1529–1539. doi: 10.1101/gad.13.12.1529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varga-Weisz PD, Wilm M, Bonte E, et al. Chromatin-remodelling factor CHRAC contains the ATPases ISWI and topoisomerase II. Nature. 1997;388:598–602. doi: 10.1038/41587. [DOI] [PubMed] [Google Scholar]
- Xiao H, Sandaltzopoulos R, Wang HM, et al. Dual functions of largest NURF subunit NURF301 in nucleosome sliding and transcription factor interactions. Mol Cell. 2001;8:531–543. doi: 10.1016/s1097-2765(01)00345-8. [DOI] [PubMed] [Google Scholar]
- Mizuguchi G, Tsukiyama T, Wisniewski J, Wu C. Role of nucleosome remodeling factor NURF in transcriptional activation of chromatin. Mol Cell. 1997;1:141–150. doi: 10.1016/s1097-2765(00)80015-5. [DOI] [PubMed] [Google Scholar]
- Badenhorst P, Voas M, Rebay I, Wu C. Biological functions of the ISWI chromatin remodeling complex NURF. Genes Dev. 2002;16:3186–3198. doi: 10.1101/gad.1032202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song H, Spichiger-Haeusermann C, Basler K. The ISWI-containing NURF complex regulates the output of the canonical Wingless pathway. EMBO Rep. 2009;10:1140–1146. doi: 10.1038/embor.2009.157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deuring R, Fanti L, Armstrong JA, et al. The ISWI chromatin-remodeling protein is required for gene expression and the maintenance of higher order chromatin structure in vivo. Mol Cell. 2000;5:355–365. doi: 10.1016/s1097-2765(00)80430-x. [DOI] [PubMed] [Google Scholar]
- Chioda M, Vengadasalam S, Kremmer E, Eberharter A, Becker PB. Developmental role for ACF1-containing nucleosome remodellers in chromatin organisation. Development. 137:3513–3522. doi: 10.1242/dev.048405. [DOI] [PubMed] [Google Scholar]
- Kwon SY, Xiao H, Wu C, Badenhorst P. Alternative splicing of NURF301 generates distinct NURF chromatin remodeling complexes with altered modified histone binding specificities. PLoS Genet. 2009;5:e1000574. doi: 10.1371/journal.pgen.1000574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corona DF, Siriaco G, Armstrong JA, et al. ISWI regulates higher-order chromatin structure and histone H1 assembly in vivo. PLoS Biol. 2007;5:e232. doi: 10.1371/journal.pbio.0050232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siriaco G, Deuring R, Chioda M, Becker PB, Tamkun JW. Drosophila ISWI regulates the association of histone H1 with interphase chromosomes in vivo. Genetics. 2009;182:661–669. doi: 10.1534/genetics.109.102053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bozhenok L, Wade PA, Varga-Weisz P. WSTF-ISWI chromatin remodeling complex targets heterochromatic replication foci. EMBO J. 2002;21:2231–2241. doi: 10.1093/emboj/21.9.2231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tran HG, Steger DJ, Iyer VR, Johnson AD. The chromo domain protein chd1p from budding yeast is an ATP-dependent chromatin-modifying factor. EMBO J. 2000;19:2323–2331. doi: 10.1093/emboj/19.10.2323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pray-Grant MG, Daniel JA, Schieltz D, Yates JR, Grant PA. Chd1 chromodomain links histone H3 methylation with SAGA- and SLIK-dependent acetylation. Nature. 2005;433:434–438. doi: 10.1038/nature03242. [DOI] [PubMed] [Google Scholar]
- Sims RJ, Chen CF, Santos-Rosa H, et al. Human but not yeast CHD1 binds directly and selectively to histone H3 methylated at lysine 4 via its tandem chromodomains. J Biol Chem. 2005;280:41789–41792. doi: 10.1074/jbc.C500395200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flanagan JF, Mi LZ, Chruszcz M, et al. Double chromodomains cooperate to recognize the methylated histone H3 tail. Nature. 2005;438:1181–1185. doi: 10.1038/nature04290. [DOI] [PubMed] [Google Scholar]
- Konev AY, Tribus M, Park SY, et al. CHD1 motor protein is required for deposition of histone variant H3.3 into chromatin in vivo. Science. 2007;317:1087–1090. doi: 10.1126/science.1145339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tong JK, Hassig CA, Schnitzler GR, Kingston RE, Schreiber SL. Chromatin deacetylation by an ATP-dependent nucleosome remodelling complex. Nature. 1998;395:917–921. doi: 10.1038/27699. [DOI] [PubMed] [Google Scholar]
- Xue Y, Wong J, Moreno GT, et al. NURD, a novel complex with both ATP-dependent chromatin-remodeling and histone deacetylase activities. Mol Cell. 1998;2:851–861. doi: 10.1016/s1097-2765(00)80299-3. [DOI] [PubMed] [Google Scholar]
- Kim J, Sif S, Jones B, et al. Ikaros DNA-binding proteins direct formation of chromatin remodeling complexes in lymphocytes. Immunity. 1999;10:345–355. doi: 10.1016/s1074-7613(00)80034-5. [DOI] [PubMed] [Google Scholar]
- Naito T, Gomez-Del Arco P, Williams CJ, Georgopoulos K. Antagonistic interactions between Ikaros and the chromatin remodeler Mi-2beta determine silencer activity and Cd4 gene expression. Immunity. 2007;27:723–734. doi: 10.1016/j.immuni.2007.09.008. [DOI] [PubMed] [Google Scholar]
- Kaji K, Caballero IM, MacLeod R, et al. The NuRD component Mbd3 is required for pluripotency of embryonic stem cells. Nat Cell Biol. 2006;8:285–292. doi: 10.1038/ncb1372. [DOI] [PubMed] [Google Scholar]
- Zhu D, Fang J, Li Y, Zhang J. Mbd3, a component of NuRD/Mi-2 complex, helps maintain pluripotency of mouse embryonic stem cells by repressing trophectoderm differentiation. PLoS One. 2009;4:e7684. doi: 10.1371/journal.pone.0007684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kehle J, Beuchle D, Treuheit S, et al. dMi-2, a hunchback-interacting protein that functions in polycomb repression. Science. 1998;282:1897–1900. doi: 10.1126/science.282.5395.1897. [DOI] [PubMed] [Google Scholar]
- Chou DM, Adamson B, Dephoure NE, et al. A chromatin localization screen reveals poly (ADP ribose)-regulated recruitment of the repressive polycomb and NuRD complexes to sites of DNA damage. Proc Natl Acad Sci USA. 2010;107:18475–18480. doi: 10.1073/pnas.1012946107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Polo SE, Kaidi A, Baskcomb L, Galanty Y, Jackson SP. Regulation of DNA-damage responses and cell-cycle progression by the chromatin remodelling factor CHD4. EMBO J. 2010;29:3130–3139. doi: 10.1038/emboj.2010.188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larsen DH, Poinsignon C, Gudjonsson T, et al. The chromatin-remodeling factor CHD4 coordinates signaling and repair after DNA damage. J Cell Biol. 190:731–740. doi: 10.1083/jcb.200912135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smeenk G, Wiegant WW, Vrolijk H, et al. The NuRD chromatin-remodeling complex regulates signaling and repair of DNA damage. J Cell Biol. 190:741–749. doi: 10.1083/jcb.201001048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo J, Su F, Chen D, Shiloh A, Gu W. Deacetylation of p53 modulates its effect on cell growth and apoptosis. Nature. 2000;408:377–381. doi: 10.1038/35042612. [DOI] [PubMed] [Google Scholar]
- Pegoraro G, Kubben N, Wickert U, et al. Ageing-related chromatin defects through loss of the NURD complex. Nat Cell Biol. 2009;11:1261–1267. doi: 10.1038/ncb1971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bagchi A, Papazoglu C, Wu Y, et al. CHD5 is a tumor suppressor at human 1p36. Cell. 2007;128:459–475. doi: 10.1016/j.cell.2006.11.052. [DOI] [PubMed] [Google Scholar]
- Hurd EA, Poucher HK, Cheng K, Raphael Y, Martin DM. The ATP-dependent chromatin remodeling enzyme CHD7 regulates pro-neural gene expression and neurogenesis in the inner ear. Development. 2010;137:3139–3150. doi: 10.1242/dev.047894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bosman EA, Penn AC, Ambrose JC, et al. Multiple mutations in mouse Chd7 provide models for CHARGE syndrome. Hum Mol Genet. 2005;14:3463–3476. doi: 10.1093/hmg/ddi375. [DOI] [PubMed] [Google Scholar]
- Schnetz MP, Handoko L, Akhtar-Zaidi B, et al. CHD7 targets active gene enhancer elements to modulate ES cell-specific gene expression. PLoS Genet. 6:e1001023. doi: 10.1371/journal.pgen.1001023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hurd EA, Capers PL, Blauwkamp MN, et al. Loss of Chd7 function in gene-trapped reporter mice is embryonic lethal and associated with severe defects in multiple developing tissues. Mamm Genome. 2007;18:94–104. doi: 10.1007/s00335-006-0107-6. [DOI] [PubMed] [Google Scholar]
- Dunaief JL, Strober BE, Guha S, et al. The retinoblastoma protein and BRG1 form a complex and cooperate to induce cell cycle arrest. Cell. 1994;79:119–130. doi: 10.1016/0092-8674(94)90405-7. [DOI] [PubMed] [Google Scholar]
- Versteege I, Sevenet N, Lange J, et al. Truncating mutations of hSNF5/INI1 in aggressive paediatric cancer. Nature. 1998;394:203–206. doi: 10.1038/28212. [DOI] [PubMed] [Google Scholar]
- Roberts CW, Galusha SA, McMenamin ME, Fletcher CD, Orkin SH. Haploinsufficiency of Snf5 (integrase interactor 1) predisposes to malignant rhabdoid tumors in mice. Proc Natl Acad Sci USA. 2000;97:13796–13800. doi: 10.1073/pnas.250492697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts CW, Leroux MM, Fleming MD, Orkin SH. Highly penetrant, rapid tumorigenesis through conditional inversion of the tumor suppressor gene Snf5. Cancer Cell. 2002;2:415–425. doi: 10.1016/s1535-6108(02)00185-x. [DOI] [PubMed] [Google Scholar]
- Decristofaro MF, Betz BL, Rorie CJ, et al. Characterization of SWI/SNF protein expression in human breast cancer cell lines and other malignancies. J Cell Physiol. 2001;186:136–145. doi: 10.1002/1097-4652(200101)186:1<136::AID-JCP1010>3.0.CO;2-4. [DOI] [PubMed] [Google Scholar]
- Schneppenheim R, Fruhwald MC, Gesk S, et al. Germline nonsense mutation and somatic inactivation of SMARCA4/BRG1 in a family with rhabdoid tumor predisposition syndrome. Am J Hum Genet. 86:279–284. doi: 10.1016/j.ajhg.2010.01.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Girard L, Zochbauer-Muller S, Virmani AK, Gazdar AF, Minna JD. Genome-wide allelotyping of lung cancer identifies new regions of allelic loss, differences between small cell lung cancer and non-small cell lung cancer, and loci clustering. Cancer Res. 2000;60:4894–4906. [PubMed] [Google Scholar]
- Bultman SJ, Herschkowitz JI, Godfrey V, et al. Characterization of mammary tumors from Brg1 heterozygous mice. Oncogene. 2008;27:460–468. doi: 10.1038/sj.onc.1210664. [DOI] [PubMed] [Google Scholar]
- Wang X, Nagl NG, Flowers S, et al. Expression of p270 (ARID1A), a component of human SWI/SNF complexes, in human tumors. Int J Cancer. 2004;112:636. doi: 10.1002/ijc.20450. [DOI] [PubMed] [Google Scholar]
- Jones S, Wang TL, Shih IeM, et al. Frequent mutations of chromatin remodeling gene ARID1A in ovarian clear cell carcinoma. Science. 2010;330:228–231. doi: 10.1126/science.1196333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xia W, Nagase S, Montia AG, et al. BAF180 is a critical regulator of p21 induction and a tumor suppressor mutated in breast cancer. Cancer Res. 2008;68:1667–1674. doi: 10.1158/0008-5472.CAN-07-5276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drost J, Mantovani F, Tocco F, et al. BRD7 is a candidate tumour suppressor gene required for p53 function. Nat Cell Biol. 2010;12:380–389. doi: 10.1038/ncb2038. [DOI] [PubMed] [Google Scholar]
- Zhang HS, Gavin M, Dahiya A, et al. Exit from G1 and S phase of the cell cycle is regulated by repressor complexes containing HDAC-Rb-hSWI/SNF and Rb-hSWI/SNF. Cell. 2000;101:79–89. doi: 10.1016/S0092-8674(00)80625-X. [DOI] [PubMed] [Google Scholar]
- Muchardt C, Reyes JC, Bourachot B, Leguoy E, Yaniv M. The hbrm and BRG-1 proteins, components of the human SNF/SWI complex, are phosphorylated and excluded from the condensed chromosomes during mitosis. EMBO J. 1996;15:3394–3402. [PMC free article] [PubMed] [Google Scholar]
- Kang H, Cui K, Zhao K. BRG1 controls the activity of the retinoblastoma protein via regulation of p21CIP1/WAF1/SDI. Mol Cell Biol. 2004;24:1188–1199. doi: 10.1128/MCB.24.3.1188-1199.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bochar DA, Wang L, Beniya H, et al. BRCA1 is associated with a human SWI/SNF-related complex: linking chromatin remodeling to breast cancer. Cell. 2000;102:257–265. doi: 10.1016/s0092-8674(00)00030-1. [DOI] [PubMed] [Google Scholar]
- Lee D, Kim JW, Seo T, et al. SWI/SNF complex interacts with tumor suppressor p53 and is necessary for the activation of p53-mediated transcription. J Biol Chem. 2002;277:22330–22337. doi: 10.1074/jbc.M111987200. [DOI] [PubMed] [Google Scholar]
- Wang M, Gu C, Qi T, et al. BAF53 interacts with p53 and functions in p53-mediated p21-gene transcription. J Biochem. 2007;142:613–620. doi: 10.1093/jb/mvm176. [DOI] [PubMed] [Google Scholar]
- Burrows AE, Smogorzewska A, Elledge SJ. Polybromo-associated BRG1-associated factor components BRD7 and BAF180 are critical regulators of p53 required for induction of replicative senescence. Proc Natl Acad Sci USA. 2010;107:14280–14285. doi: 10.1073/pnas.1009559107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagl NG Jr, Wang X, Patsialou A, Van Scoy M, Moran E. Distinct mammalian SWI/SNF chromatin remodeling complexes with opposing roles in cell-cycle control. EMBO J. 2007;26:752–763. doi: 10.1038/sj.emboj.7601541. [DOI] [PMC free article] [PubMed] [Google Scholar]