Abstract
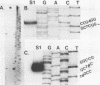
A combination of a single stranded plasmid vector and ordered deletions were used to determine the complete nucleotide sequence of a rDNA spacer region isolated from the DNA of maize Black Mexican Sweet suspension cells. The sequence reveals the presence of ten "200 base" subrepeats within the maize rDNA spacer region. By S1 protection experiments we have tentatively determined that the start of maize rRNA transcription is 144 bases 3' of the end of the last spacer subrepeat. We propose that the spacer subrepeats may have an important role in regulating maize rRNA transcription.
Full text
PDF















Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bakken A., Morgan G., Sollner-Webb B., Roan J., Busby S., Reeder R. H. Mapping of transcription initiation and termination signals on Xenopus laevis ribosomal DNA. Proc Natl Acad Sci U S A. 1982 Jan;79(1):56–60. doi: 10.1073/pnas.79.1.56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boseley P., Moss T., Mächler M., Portmann R., Birnstiel M. Sequence organization of the spacer DNA in a ribosomal gene unit of Xenopus laevis. Cell. 1979 May;17(1):19–31. doi: 10.1016/0092-8674(79)90291-5. [DOI] [PubMed] [Google Scholar]
- Coen E. S., Dover G. A. Multiple Pol I initiation sequences in rDNA spacers of Drosophila melanogaster. Nucleic Acids Res. 1982 Nov 11;10(21):7017–7026. doi: 10.1093/nar/10.21.7017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dale R. M., McClure B. A., Houchins J. P. A rapid single-stranded cloning strategy for producing a sequential series of overlapping clones for use in DNA sequencing: application to sequencing the corn mitochondrial 18 S rDNA. Plasmid. 1985 Jan;13(1):31–40. doi: 10.1016/0147-619x(85)90053-8. [DOI] [PubMed] [Google Scholar]
- Dunaway M., Reeder R. H. DNase I footprinting shows three protected regions in the promoter of the rRNA genes of Xenopus laevis. Mol Cell Biol. 1985 Feb;5(2):313–319. doi: 10.1128/mcb.5.2.313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fromm M., Taylor L. P., Walbot V. Expression of genes transferred into monocot and dicot plant cells by electroporation. Proc Natl Acad Sci U S A. 1985 Sep;82(17):5824–5828. doi: 10.1073/pnas.82.17.5824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldsbrough P. B., Cullis C. A. Characterisation of the genes for ribosomal RNA in flax. Nucleic Acids Res. 1981 Mar 25;9(6):1301–1309. doi: 10.1093/nar/9.6.1301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohorn B. D., Rae P. M. Nontranscribed spacer sequences promote in vitro transcription of Drosophila ribosomal DNA. Nucleic Acids Res. 1982 Nov 11;10(21):6879–6886. doi: 10.1093/nar/10.21.6879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Labhart P., Reeder R. H. Enhancer-like properties of the 60/81 bp elements in the ribosomal gene spacer of Xenopus laevis. Cell. 1984 May;37(1):285–289. doi: 10.1016/0092-8674(84)90324-6. [DOI] [PubMed] [Google Scholar]
- Long E. O., Dawid I. B. Repeated genes in eukaryotes. Annu Rev Biochem. 1980;49:727–764. doi: 10.1146/annurev.bi.49.070180.003455. [DOI] [PubMed] [Google Scholar]
- Long E. O., Dawid I. B. Restriction analysis of spacers in ribosomal DNA of Drosophila melanogaster. Nucleic Acids Res. 1979 Sep 11;7(1):205–215. doi: 10.1093/nar/7.1.205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mandal R. K., Dawid I. B. The nucleotide sequence at the transcription termination site of ribosomal RNA in Drosophila melanogaster. Nucleic Acids Res. 1981 Apr 24;9(8):1801–1811. doi: 10.1093/nar/9.8.1801. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mandal R. K. The organization and transcription of eukaryotic ribosomal RNA genes. Prog Nucleic Acid Res Mol Biol. 1984;31:115–160. doi: 10.1016/s0079-6603(08)60376-1. [DOI] [PubMed] [Google Scholar]
- Messing J., Carlson J., Hagen G., Rubenstein I., Oleson A. Cloning and sequencing of the ribosomal RNA genes in maize: the 17S region. DNA. 1984;3(1):31–40. doi: 10.1089/dna.1.1984.3.31. [DOI] [PubMed] [Google Scholar]
- Messing J. New M13 vectors for cloning. Methods Enzymol. 1983;101:20–78. doi: 10.1016/0076-6879(83)01005-8. [DOI] [PubMed] [Google Scholar]
- Miller J. R., Hayward D. C., Glover D. M. Transcription of the 'non-transcribed' spacer of Drosophila melanogaster rDNA. Nucleic Acids Res. 1983 Jan 11;11(1):11–19. doi: 10.1093/nar/11.1.11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller O. L., Jr, Beatty B. R. Visualization of nucleolar genes. Science. 1969 May 23;164(3882):955–957. doi: 10.1126/science.164.3882.955. [DOI] [PubMed] [Google Scholar]
- Moss T. A transcriptional function for the repetitive ribosomal spacer in Xenopus laevis. Nature. 1983 Mar 17;302(5905):223–228. doi: 10.1038/302223a0. [DOI] [PubMed] [Google Scholar]
- Murtif V. L., Rae P. M. In vivo transcription of rDNA spacers in Drosophila. Nucleic Acids Res. 1985 May 10;13(9):3221–3239. doi: 10.1093/nar/13.9.3221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perry R. P. Processing of RNA. Annu Rev Biochem. 1976;45:605–629. doi: 10.1146/annurev.bi.45.070176.003133. [DOI] [PubMed] [Google Scholar]
- Reeder R. H., Roan J. G., Dunaway M. Spacer regulation of Xenopus ribosomal gene transcription: competition in oocytes. Cell. 1983 Dec;35(2 Pt 1):449–456. doi: 10.1016/0092-8674(83)90178-2. [DOI] [PubMed] [Google Scholar]
- Saedler H., Nevers P. Transposition in plants: a molecular model. EMBO J. 1985 Mar;4(3):585–590. doi: 10.1002/j.1460-2075.1985.tb03670.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saghai-Maroof M. A., Soliman K. M., Jorgensen R. A., Allard R. W. Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci U S A. 1984 Dec;81(24):8014–8018. doi: 10.1073/pnas.81.24.8014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siegel A., Kolacz K. Heterogeneity of pumpkin ribosomal DNA. Plant Physiol. 1983 May;72(1):166–171. doi: 10.1104/pp.72.1.166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simeone A., La Volpe A., Boncinelli E. Nucleotide sequence of a complete ribosomal spacer of D. melanogaster. Nucleic Acids Res. 1985 Feb 25;13(4):1089–1101. doi: 10.1093/nar/13.4.1089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sollner-Webb B., Reeder R. H. The nucleotide sequence of the initiation and termination sites for ribosomal RNA transcription in X. laevis. Cell. 1979 Oct;18(2):485–499. doi: 10.1016/0092-8674(79)90066-7. [DOI] [PubMed] [Google Scholar]
- Varsanyi-Breiner A., Gusella J. F., Keys C., Housman D. E., Sullivan D., Brisson N., Verma D. P. The organization of a nuclear DNA sequence from a higher plant: molecular cloning and characterization of soybean ribosomal DNA. Gene. 1979 Nov;7(3-4):317–334. doi: 10.1016/0378-1119(79)90051-9. [DOI] [PubMed] [Google Scholar]
- Veldman G. M., Klootwijk J., de Jonge P., Leer R. J., Planta R. J. The transcription termination site of the ribosomal RNA operon in yeast. Nucleic Acids Res. 1980 Nov 25;8(22):5179–5192. doi: 10.1093/nar/8.22.5179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wellauer P. K., Reeder R. H., Carroll D., Brown D. D., Deutch A., Higashinakagawa T., Dawid I. B. Amplified ribosomal DNA from Xenopus laevis has heterogeneous spacer lengths. Proc Natl Acad Sci U S A. 1974 Jul;71(7):2823–2827. doi: 10.1073/pnas.71.7.2823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang R., Lis J., Wu R. Elution of DNA from agarose gels after electrophoresis. Methods Enzymol. 1979;68:176–182. doi: 10.1016/0076-6879(79)68012-6. [DOI] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]