Abstract
Two families of GTPases, the Toc34 and Toc159 GTPase families, take on the task of preprotein recognition at the translocon at the outer membrane of chloroplasts (TOC translocon). The major Toc159 family members have highly acidic N-terminal domains (A-domains) that are non-essential and so far have escaped functional characterization. But recently, interest in the role of the A-domain has strongly increased. The new data of three independent studies provide evidence that the Toc159 A-domain (I) participates in preprotein selectivity, (II) has typical features of intrinsically unfolded proteins and (III) is highly phosphorylated and possibly released from the rest of the protein by a proteolytic event. This hints at a complex regulation of A-domain function that is important for the maintenance of the preprotein selectivity at the TOC translocons.
Key words: chloroplast, import, Toc159, acidic domain, kinase, protease
Back in the Research Focus
Plastids are characteristic organelles of land plants and algae that evolved from a cyanobacterial endosymbiont.1 Today, they are fully integrated into the life cycle of the cell. The majority of plastid proteins are encoded in the nucleus and synthesized in the cytosol as preproteins with an N-terminal targeting signal, the transit peptide.2 Therefore the biogenesis of plastids and their differentiation into specialized forms e.g., photosynthetic chloroplasts depend on the coordination between gene-expression and the selective import of preproteins from the cytosol. For preprotein import plastids possess multi-protein import complexes in their dual-membrane envelope designated Toc (translocon at the outer envelope membrane of chloroplasts) and Tic (translocon at the inner envelope membrane of chloroplasts).3 The initial contact of preproteins with the chloroplast surface is mediated by a pair of homologous GTPases present in the Toc complex that act as receptor proteins and work together in the regulation of import.4,5 In higher plants the Toc GTPases are encoded by small gene families. In Arabidopsis thaliana the Toc34 family is encoded by two genes, TOC33 and TOC34, the Toc159 family by four genes, TOC159, TOC120, TOC132 and TOC90.6 The Toc34 family members are smaller in size (33 to 34 kDa) and consist of a GTPase domain (G-domain) that is anchored in the outer envelope membrane by a single stretch of hydrophobic amino acids at the C-terminus. The Toc159 family members are larger (90 to 159 kDa) and have a more complex domain structure. They contain a central G-domain flanked by a 52 kDa C-terminal membrane inserted domain (M-domain) and, with the exception of Toc90, a highly acidic N-terminal domain (A-domain). The N-terminal A-domains represent the most variable part of the Toc159-like proteins in length as well as in primary structure. They do not contain any conserved protein motifs but share properties with the growing class of intrinsically disordered proteins in higher eukaryotes.7,8 There is increasing evidence that the members of the Toc34/Toc159 gene families assemble into functionally and structurally distinct Toc complexes, each containing a small and a large Toc receptor GTPase9,10 and each representing a separate import pathway for a certain class of preproteins. It is likely that the selectivity of Toc complexes is mediated by receptor domains exposed to the cytosol. Indeed, binding of transit peptides to the cytosol-exposed G-domains of Toc34 and Toc159 has been demonstrated.11,12 However, it has now been shown that chloroplasts lacking the A-domain of Toc159 efficiently import preproteins, but in a manner no longer discriminating between different classes of preproteins.13 This suggests that the Toc159 A-domain confers selectivity to the Toc complexes and does so by a yet unknown mechanism. Neglected for a long time due to its non-essential function and high sensitivity to proteolysis, the A-domain is now back in the research focus.
Multi-Site Phosphorylation of the A-Domain
In our own recent work we made use of a N-terminal TAP-tag for purification of Arabidopsis thaliana Toc159.14 Using mass-spectrometry we discovered that a large proportion of purified Toc159 consisted only of a fragment of the full-length protein and was identical with the A-domain. This free A-domain was recovered from the soluble as well as the membrane fraction in cell fractionation experiments. By phosphospecific staining both Toc159 and the free A-domain were revealed as phosphoproteins. Furthermore phosphoproteomic surveys15–18 and our study experimentally identified a total of twelve in vivo phosphorylation sites in Toc159 that all map to the A-domain, most of them being predicted casein kinase 2 (CK2) phosphorylation sites. Many more phosphorylation sites are predicted and suggest an even higher phosphorylation rate of the A-domain. Some of the new findings were surprising and unexpected e.g., (i) the evident stability of the A-domain as a separate protein, (ii) its partial association with the membranes and (iii) its high degree of phosphorylation. In contrast earlier studies described the A-domain as highly sensitive to proteolysis and less stable than the rest of the protein consisting of the Toc159 G- and M-domains (referred to as Toc86).19 As such, the regulation of the A-domain by multi-site phosphorylation is not surprising, as it predominantly occurs in disordered proteins. However, which function of Toc159 is regulated by this post-translational modification was and still is an uncertain issue. At the date of publication of our study the function of the A-domain for Toc159 function was completely unclear. Only recently, a direct implication of the A-domain in the chloroplast import process was established.13 Chloroplasts lacking the A-domain exhibited a reduced selectivity for the import of the Toc159 model substrate preferredoxin, a representative preprotein of the “photosynthesis-associated” class. Conceivably, this function of the A-domain as determinant of Toc complex selectivity is regulated by phosphorylation.
Implication of the A-Domain in the Chloroplast Import Process
In our recent study we started to investigate the link of A-domain phosphorylation to the import mechanism into chloroplasts by modulating the phosphorylation status of isolated chloroplasts and studying their import capacity in in vitro experiments.14 Only heparin, a known casein kinase 2 inhibitor, inhibited in vitro phosphorylation of the A-domain as well as chloroplast import of the pre-small subunit of Rubisco (pSSU). The data were in accordance with a positive regulation of import by a heparin-sensitive kinase activity. However, the data we present here shows that heparin also inhibits import into chloroplasts that lack the A-domain of Toc159 (Fig. 1). This clearly demonstrates that the heparin effect on chloroplast import does not occur via the A-domain of Toc159. Considering this and the new data on the function of the A-domain in import selectivity13 future experiments on A-domain phosphorylation and chloroplast import will need to include a wider range of import substrates and competitors. A very interesting question is whether the A-domain exerts its determinant function on import selectivity in trans, that is as the free protein we discovered in our study. Possibly the selectivity of chloroplasts lacking the A-domain in the import assays could be reinstated by adding the purified A-domain. In addition such experiments would have the advantage that the phosphorylation status of recombinant A-domain is much easier to control than the one of the A-domain purified from plants.
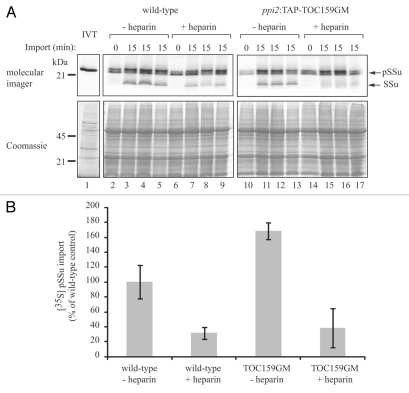
Figure 1.
The inhibitory effect of heparin on chloroplast import does not involve Toc159A. (A) In vitro import assays with chloroplasts of Arabidopsis thaliana wild-type plants and toc159 mutant plants (ppi2) complemented with TAP-tagged Toc159 lacking the acidic domain.26 Chloroplasts were preincubated for 20 min at 25°C in the dark in the presence of 15 Mg/ml heparin and 20 micromolar ATP. Then ATP was added to a final concentration of 5 mM and the in vitro translated precursor of the small subunit of Rubisco was added. After 0 min or 15 min of import the reactions were stopped and analyzed by SDS-PAGE and autoradiography. (B) Quantification of the import reactions. The amount of [35S] SSu imported after 15 min of import by untreated wild-type chloroplasts was set to 100%.
Toc159A Modifying Enzymes
Toc159 appears to be regulated by at least two mechanisms (Fig. 2), hyperphosphorylation and proteolytic processing. While there is no doubt that the A-domain is the target of at least two kinase activities,14 the evidence for the controlled proteolytic processing of the A-domain is less strong. Naturally, the concept of regulation by proteolysis raises more skepticism, as non-specific degradation is a frequent problem in biochemical experimentation. On the other hand proteases are known to act as key regulators of a variety of cellular and plastidal processes.20,21 Different from phosphorylation, regulation by proteases is irreversible. Therefore the release of the A-domain by a so far unidentified, possibly membrane located endopeptidase would definitively determine the fate of the receptor. So far only scarce information on outer envelope proteases is available. The first enzymes identified were SPaseI involved in the processing of the precursor of Toc75 and several thylakoid lumenal proteins (OE33, OE23, plastocyanin)22,23 and AraSP, an inner envelope protease, belonging to the S2P family of metalloproteases.24 Both enzymes are important for chloroplast biogenesis but due to their localization unlikely candidates for Toc159 processing enzymes. Recently, a study of the subplastidal proteome identified several other proteases, especially metalloproteases, in the chloroplast envelope (AT_CHLORO).25 It is conceivable that the Toc159 processing peptidase is among the list of envelope proteases of this study. The identification of Toc159 A-domain modifying enzymes (proteases, kinase[s] and phosphatases) is now the next step and will provide new and exciting insights into the chloroplast import process.
Figure 2.
Hypothetical model on the states of the Toc159 A-domain. The A-domain may exist in an un-phosphorylated state (bottom). Following phosphorylation by CK2 and another kinase at the chloroplast surface it may enter a hyperphosphorylated state (top). Proteolytic cleavage entails the release of the A-domain from the rest of the receptor (right). The signals that trigger the different states of the A-domain and the functional process that involves the A-domain are so far unknown.
Acknowledgements
Funding was provided by SNF grant 31003A_127380 and SystemsX PGCE to FK.
Footnotes
Previously published online: www.landesbioscience.com/journals/psb/article/13707
References
- 1.Margulis L. Origin of Eucaryotic Cells. New Haven, CT: Yale University Press; 1970. [Google Scholar]
- 2.Bruce BD. Chloroplast transit peptides: structure, function and evolution. Trends Cell Biol. 2000;10:440–447. doi: 10.1016/s0962-8924(00)01833-x. [DOI] [PubMed] [Google Scholar]
- 3.Schnell DJ, Blobel G, Keegstra K, Kessler F, Ko K, Soll J. A consensus nomenclature for the protein-import components of the chloroplast envelope. Trends Cell Biol. 1997;7:303–304. doi: 10.1016/S0962-8924(97)01111-2. [DOI] [PubMed] [Google Scholar]
- 4.Li HM, Kesavulu MM, Su PH, Yeh YH, Hsiao CD. Toc GTPases. J Biomed Sci. 2007;14:505–508. doi: 10.1007/s11373-007-9166-2. [DOI] [PubMed] [Google Scholar]
- 5.Kessler F, Schnell D. Chloroplast biogenesis: diversity and regulation of the protein import apparatus. Curr Opin Cell Biol. 2009;21:494–500. doi: 10.1016/j.ceb.2009.03.004. [DOI] [PubMed] [Google Scholar]
- 6.Jackson-Constan D, Keegstra K. Arabidopsis genes encoding components of the chloroplastic protein import apparatus. Plant Physiol. 2001;125:1567–1576. doi: 10.1104/pp.125.4.1567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hernandez Torres J, Maldonado MA, Chomilier J. Tandem duplications of a degenerated GTP-binding domain at the origin of GTPase receptors Toc159 and thylakoidal SRP. Biochem Biophys Res Commun. 2007;364:325–331. doi: 10.1016/j.bbrc.2007.10.006. [DOI] [PubMed] [Google Scholar]
- 8.Richardson LG, Jelokhani-Niaraki M, Smith MD. The acidic domains of the Toc159 chloroplast preprotein receptor family are intrinsically disordered protein domains. BMC Biochem. 2009;10:35. doi: 10.1186/1471-2091-10-35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kubis S, Patel R, Combe J, Bédard J, Kovacheva S, Lilley K, et al. Functional specialization amongst the Arabidopsis Toc159 family of chloroplast protein import receptors. Plant Cell. 2004;16:2059–2077. doi: 10.1105/tpc.104.023309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ivanova Y, Smith MD, Chen K, Schnell DJ. Members of the Toc159 import receptor family represent distinct pathways for protein targeting to plastids. Mol Biol Cell. 2004;15:3379–3392. doi: 10.1091/mbc.E03-12-0923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Smith MD, Rounds CM, Wang F, Chen K, Afitlhile M, Schnell DJ. atToc159 is a selective transit peptide receptor for the import of nucleus-encoded chloroplast proteins. J Cell Biol. 2004;165:323–334. doi: 10.1083/jcb.200311074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Sveshnikova N, Soll J, Schleiff E. Toc34 is a preprotein receptor regulated by GTP and phosphorylation. Proc Natl Acad Sci USA. 2000;97:4973–4978. doi: 10.1073/pnas.080491597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Inoue H, Rounds C, Schnell DJ. The molecular basis for distinct pathways for protein import into Arabidopsis chloroplasts. Plant Cell. 2010;22:1947–1960. doi: 10.1105/tpc.110.074328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Agne B, Andrès C, Montandon C, Christ B, Ertan A, Jung F, et al. The acidic A-domain of Arabidopsis TOC159 occurs as a hyperphosphorylated protein. Plant Physiol. 2010;153:1016–1030. doi: 10.1104/pp.110.158048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Reiland S, Messerli G, Baerenfaller K, Gerrits B, Endler A, Grossmann J, et al. Large-scale Arabidopsis phosphoproteome profiling reveals novel chloroplast kinase substrates and phosphorylation networks. Plant Physiol. 2009;150:889–903. doi: 10.1104/pp.109.138677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.de la Fuente van Bentem S, Anrather D, Dohnal I, Roitinger E, Csaszar E, Joore J, et al. Site-specific phosphorylation profiling of Arabidopsis proteins by mass spectrometry and peptide chip analysis. J Proteome Res. 2008;7:2458–2470. doi: 10.1021/pr8000173. [DOI] [PubMed] [Google Scholar]
- 17.Sugiyama N, Nakagami H, MOchida K, Daudi A, Tomita M, Shirasu K, et al. Large-scale phosphorylation mapping reveals the extent of tyrosine phosphorylation in Arabidopsis. Mol Syst Biol. 2008;4:193. doi: 10.1038/msb.2008.32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Whiteman SA, Serazetdinova L, Jones AM, Sanders D, Rathjen J, Peck SC, et al. Identification of novel proteins and phosphorylation sites in a tonoplast enriched membrane fraction of Arabidopsis thaliana. Proteomics. 2008;8:3536–3547. doi: 10.1002/pmic.200701104. [DOI] [PubMed] [Google Scholar]
- 19.Bolter B, May T, Soll J. A protein import receptor in pea chloroplasts, Toc86, is only a proteolytic fragment of a larger polypeptide. FEBS Lett. 1998;441:59–62. doi: 10.1016/s0014-5793(98)01525-7. [DOI] [PubMed] [Google Scholar]
- 20.Adam Z, Rudella A, van Wijk KJ. Recent advances in the study of Clp, FtsH and other proteases located in chloroplasts. Curr Opin Plant Biol. 2006;9:234–240. doi: 10.1016/j.pbi.2006.03.010. [DOI] [PubMed] [Google Scholar]
- 21.van der Hoorn RA. Plant proteases: from phenotypes to molecular mechanisms. Annu Rev Plant Biol. 2008;59:191–223. doi: 10.1146/annurev.arplant.59.032607.092835. [DOI] [PubMed] [Google Scholar]
- 22.Inoue K, Baldwin AJ, Shipman RL, Matsui K, Theg SM, Ohme-Takagi M. Complete maturation of the plastid protein translocation channel requires a type I signal peptidase. J Cell Biol. 2005;171:425–430. doi: 10.1083/jcb.200506171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Shipman-Roston RL, Ruppel NJ, Damoc C, Phinney BS, Inoue K. The significance of protein maturation by plastidic type I signal peptidase 1 for thylakoid development in Arabidopsis chloroplasts. Plant Physiol. 2010;152:1297–1308. doi: 10.1104/pp.109.151977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Bolter B, Nada A, Fulgosi H, Soll J. A chloroplastic inner envelope membrane protease is essential for plant development. FEBS Lett. 2006;580:789–794. doi: 10.1016/j.febslet.2005.12.098. [DOI] [PubMed] [Google Scholar]
- 25.Ferro M, Brugière S, Salvi D, Seigneuri-Berny D, Court M, Moyet L, et al. AT_CHLORO, a comprehensive chloroplast proteome database with subplastidial localization and curated information on envelope proteins. Mol Cell Proteomics. 2010;9:1063–1084. doi: 10.1074/mcp.M900325-MCP200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Agne B, Infanger S, Wang F, Hofstetter V, Rahim G, Martin M, et al. A toc159 import receptor mutant, defective in hydrolysis of GTP, supports preprotein import into chloroplasts. J Biol Chem. 2009;284:8670–8679. doi: 10.1074/jbc.M804235200. [DOI] [PMC free article] [PubMed] [Google Scholar]