Abstract
Phosphatidylinositol phosphate kinase (PIPK) catalyzes a key step controlling cellular contents of phosphatidylinositol-4,5-bisphosphate [PtdIns(4,5)P2], a critical intracellular messenger involved in vesicle trafficking and modulation of actin cytoskeleton and also a substrate of phospholipase C to produce the two intracellular messengers, diacylglycerol and inositol-1,4,5-trisphosphate. In addition to the conserved C-terminal PIPK catalytic domain, plant PIPKs contain a unique structural feature consisting of a repeat of membrane occupation and recognition nexus (MORN) motifs, called the MORN domain, in the N-terminal half. The MORN domain has previously been proposed to regulate plasma membrane localization and phosphatidic acid (PA)-inducible activation. Recently, the importance of the catalytic domain, but not the MORN domain, in these aspects was demonstrated. These conflicting data raise the question about the function of the MORN domain in plant PIPKs. We therefore performed analyses of PpPIPK1 from the moss Physcomitrella patens to elucidate the importance of the MORN domain in the control of enzymatic activity; however, we found no effect on either enzymatic activity or activation by PA. Taken together with our previous findings of lack of function in plasma membrane localization, there is no positive evidence indicating roles of the MORN domain in enzymatic and functional regulations of PpPIPK1. Therefore, further biochemical and reverse genetic analyses are necessary to understand the biological significance of the MORN domain in plant PIPKs.
Key words: membrane occupation and recognition nexus (MORN) domain, phosphatidylinositol phosphate kinase, phosphatidic acid, Physcomitrella patens
Phosphoinositides (PIs) are minor membrane phospholipds that play pivotal roles in various signal transduction cascades involved in development and stress response via the regulation of cytoskeletal organization, ion channel activation and vesicle trafficking.1,2 These are derivatives of phosphatidylinositol (PtdIns) produced by phosphorylation of the 3-, 4- and 5- positions of the inositol ring.2 To address the roles of PIs, enzymes involved in their production have been extensively studied using biochemical and molecular biological approaches. Of these enzymes, phosphatidylinositol monophosphate kinases (PIPKs) catalyze the reaction producing phosphatidylinositol-4,5-bisphosphate [PtdIns(4,5)P2] that is a substrate of phospholipase C and phosphatidylinositol 3-kinase, and also acts as an intracellular messenger involved in the regulation of F-actin organization and activity of ion channels.1–3 Although PtdIns(4,5)P2 is produced by sequential phosphorylation by phosphatidylinositol 4-kinase, producing phosphatidylinositol-4-phosphate [PtdIns(4)P], and then by PIPK,1,2 the cellular levels of PtdIns(4)P are much higher compared to PtdIns(4,5)P2.4–6 Thus, a restriction step controlling cellular PtdIns(4,5)P2 contents is mediated by PIPKs, indicating the importance of PIPK regulation in various kinds of physiological processes.
The roles of plant PIPKs have been established in growth regulation, such as polarized tip growth of root hairs and pollen tubes, via their localization at plasma membranes.7–12 It is worth to note that plant PIPKs contain a unique structure consisting of a repeat of a membrane occupation recognition nexus (MORN) motifs, called MORN domain, at the N-terminal region and a C-terminal PIPK catalytic domain, except for AtPIP5K10 and AtPIP5K11 from Arabidopsis thaliana, which lack the N-terminal MORN domain.13 The MORN domain was first identified as plasma membrane-binding module in junctophilin14 and the involvement of the MORN domain in plasma membrane localization was proposed for A. thaliana AtPIP5K1 and AtPIP5K3.9,15,16
Another remarkable feature of eukaryotic PIPKs is dependency of the enzymatic activity on phosphatidic acid (PA).17,18 Indeed, PA-dependent activation of PIPKs has been observed in A. thaliana and in the moss Physcomitrella patens,6,19,20 as with animal type I PIPKs.21 Although much less is known about how PA activates PIPKs in plants, biochemical analyses suggested the involvement of the MORN domain in PA-dependent activation of AtPIP5K1.15
Based on above findings, it was proposed that plasma membrane-localization and PA-dependent activation of plant PIPKs might be regulated by the MORN domain.9,15,16 In contrast, we recently demonstrated the critical involvement of the C-terminal half containing the catalytic domain of plant PIPKs in both plasma membrane-localization and PA-dependent activation.22 Thus, the function of the MORN domain remains elusive in plant PIPKs.
As shown earlier, the N-terminal half of P. patens PpPIPK1 containing the MORN domain enhances its catalytic activity.22 Thus, to identify the region required for the activation of PpPIPK1, we further dissected the N-terminal half into 3 regions; the N-terminal region (amino acid nos. 1–154), the MORN repeat (amino acid nos. 155–316) and the linker region (amino acid nos. 338–452), and made deletion mutants of PpPIPK1 as shown in Figure 1A. Using Pfu Turbo DNA polymerase (Stratagene, La Jolla, USA), DNA fragments corresponding to deletion mutants lacking the N-terminal and N-terminal plus the MORN repeat, designated PpPIPK1ΔN and PpPIPK1ΔN-MORN, respectively, were amplified with primer sets; one is M_PIPK1_fb (5′-GGC AAG CAC GTG TAT AAT GTC TGA AGG GCT T-3′) and XhoIPIPK1 (5′-TAA ACT CGA GTT AGC TGG GTA GGA GGA AA-3′) and the other is M_PIPK1_f7 (5′-AGA GAA CAC GTG TAT AAT GTC TGA CTT CTA CGT CGG T-3′) and XhoIPIPK1. For building an expression plasmid for a deletion mutant lacking the MORN repeat, designated PpPIPK1ΔMORN, the N-terminal region and PpPIPK1ΔN-MORN were amplified with primer sets, M_PIPK1_fb and M_PIPK1_r3 (5′-TTG TAA GTC TCG GGT GCC ATT TGA GAG CTC-3′) M_PIPK1_f6 (5′-GAG CTC TCA AAT GGC ACC CGA GAC TTA CAA-3′) and XhoIPIPK1, respectively, using Pfu Turbo DNA polymerase and resultant DNA fragments were fused by PCR with a primer set, M_PIPK1_fb and XhoIPIPK1 using the same enzyme. These PCR products were digested with Pml1 and XhoI and inserted into Pml1-XhoI digested pPICZB (Invitrogen) to construct expression plasmids, pPICZB-PpPIPK1ΔN, pPICZB-PpPIPK1ΔN-MORN and pPICZB-PpPIPK1ΔMORN. Transformation of P. pastoris X-33 cells with the above expression plasmids, colony PCR of transformants and following expression, purification and western blot analysis of His-tagged recombinant proteins were performed as described previously.6 The PIPK activity assay using purified His-tagged proteins was carried out as described previously23 with the modifications.6
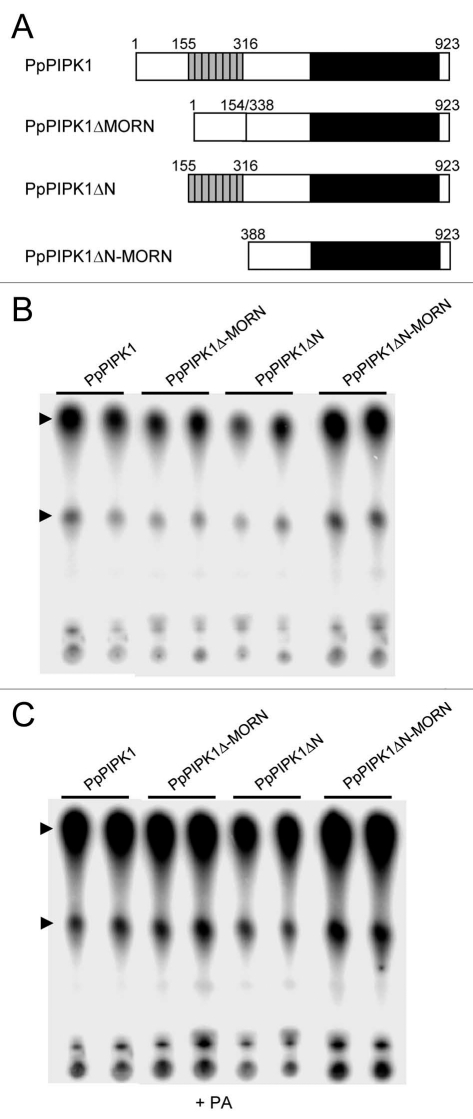
Figure 1.
Functional dissection of the N-terminal region of PpPIPK1 identifies positive regulatory regions. (A) His-tagged recombinant PpPIPK1 proteins. A repetition of eight MORN motifs (grey boxes) and the conserved catalytic domain (black box) are indicated in wild type and mutant PpPIPK1s. The MORN repeat and junction of internal deletion are indicated by amino acid position numbers. (B) In vitro lipid kinase activity of His-tagged recombinant proteins. The activities of recombinant proteins bound to Ni-NTA agarose beads were assayed with PtdIns4P. (C) In vitro PA-dependent lipid kinase activity of His-tagged proteins. The activities of recombinant proteins bound to Ni-NTA agarose beads were assayed with PtdIns4P with 143 µM PA. Top and bottom arrowheads represent reaction products PtdIns(4,5)P2 and lysoPtdIns(4,5)P2, respectively.
Biochemical analyses of these enzymes after expression in yeast P. pastoris X-33 cells followed by purification showed that deletion of the N-terminal region (PpPIPK1ΔN) reduced PpPIPK1 activity ca 40% compared to the full length enzyme, whereas loss of the MORN repeat (PpPIPK1ΔMORN) had no significant effect (Fig. 1B). In agreement, a mutant lacking four MORN repeats of the total eight repeats showed no difference in the activity compared the full length enzyme (data not shown). These results indicate a positive role of the N-terminal region, but not the MORN repeats, on PpPIPK1 activity. However, these findings differ from those obtained with AtPIP5K1, where the MORN domain represses enzymatic activity.15 Interestingly, PpPIPK1ΔN-MORN containing the linker and catalytic regions showed higher enzymatic activity of ca 23 % compared to the full length PpPIPK1 (Fig. 1B). The C-terminal half only containing the catalytic domain of PpPIPK1 and thus lacking the linker region showed a reduced activity.22 It is therefore proposed that the linker region carries a positive regulatory element. Although details are unknown, negligible effects of the N-terminal and MORN domains for the enzymatic activity has been indicated in AtPIP5K3 from A. thaliana.11 Moreover, it is noteworthy that PA-dependent activation was not affected by any deletion as shown in Figure 1C, confirming that the N-terminal half is not involved in PA dependency of the PpPIPK1 activity.22
Our results indicated that the MORN domain is not involved in the regulation of the catalytic activity in PpPIPK1. Similarly, the function of the MORN domain found in the accumulation and replication of chloroplasts 3 (ARC3) was not resolved. ARC3 is an FtsZ homologue involved in chloroplast division24 and the only protein containing the MORN repeats other than PIPKs in A. thaliana. It was shown that the ARC3 MORN domain did not interact with any stromal plastid division components.25 Moreover, there are reports representing functions of the MORN domain other than plasma membrane binding. Human amyotrophic lateral sclerosis 2 (ALS2), a guanine nucleotide exchange factor (GEF) specific to the small GTPase Rab5, contains the MORN domain at the central region that is essential for the GEF activity but not for interaction with Rab5.26 In contrast, specific interaction of the MORN domain with Rab-E GTPases and resultant enzymatic activation was recently demonstrated for AtPIP5K2.12 It is interesting that these results are inconsistent with each other in terms of interaction of the MORN domain with small GTPases.
Taken together, with no function of the MORN domain in plasma membrane localization of PpPIPK1 and AtPIP5K1,22 the function of the MORN domain is still unknown, despite its high conservation plants PIPKs. Alternatively, based on the findings of ARC3, ALS2 and AtPIP5K2,12,25,26 the function of the MORN domain possibly varies among PIPK isoforms and may thus have multifunctional roles. Therefore, it is necessary to identify interaction partners for the MORN domain of each plant PIPKs and to analyze phenotypes of transgenic plants carrying MORN domain-lacking PIPKs during developmental process and environmental stress responses.
Acknowledgements
This study was supported by The Swedish Research Council and the Swedish Foundation for Strategic Research.
Footnotes
Previously published online: www.landesbioscience.com/journals/psb/article/12922
References
- 1.Xue HW, Chen X, Mei Y. Function and regulation of phospholipid signalling in plants. Biochem J. 2009;421:145–156. doi: 10.1042/BJ20090300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ischebeck T, Seiler S, Heilmann I. At the poles across kingdoms: phosphoinositides and polar tip growth. Protoplasma. 2010;240:13–31. doi: 10.1007/s00709-009-0093-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Heck JN, Mellman DL, Ling K, Sun Y, Wagoner MP, Schill NJ, et al. A conspicuous connection: structure defines function for the phosphatidylinositol-phosphate kinase family. Crit Rev Biochem Mol Biol. 2007;42:15–39. doi: 10.1080/10409230601162752. [DOI] [PubMed] [Google Scholar]
- 4.Westergren T, Ekblad L, Jergil B, Sommarin M. Phosphatidylinositol 4-kinase associated with spinach plasma membranes. Isolation and characterization of two distinct forms. Plant Physiol. 1999;121:507–516. doi: 10.1104/pp.121.2.507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.DeWald DB, Torabinejad J, Jones CA, Shope JC, Cangelosi AR, Thompson JE, et al. Rapid accumulation of phosphatidylinositol 4,5-bisphosphate and inositol 1,4,5-trisphosphate correlates with calcium mobilization in salt-stressed Arabidopsis. Plant Physiol. 2001;126:759–769. doi: 10.1104/pp.126.2.759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Saavedra L, Balbi V, Dove SK, Hiwatashi Y, Mikami K, Sommarin M. Characterization of phosphatidylinositol kinases from the moss Physcomitrella patens: PpPIPK1 and PpPIPK2. Plant Cell Physiol. 2009;50:595–609. doi: 10.1093/pcp/pcp018. [DOI] [PubMed] [Google Scholar]
- 7.Lou Y, Gou JY, Xue HW. PIP5K9, an Arabidopsis phosphatidylinositol monophosphate kinase, interacts with a cytosolic invertase to negatively regulate sugar-mediated root growth. Plant Cell. 2007;19:163–181. doi: 10.1105/tpc.106.045658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ischebeck T, Stenzel I, Heilmann I. Type B phosphatidylinositol-4-phosphate 5-kinases mediate Arabidopsis and Nicotiana tabacum pollen tube growth by regulating apical pectin secretion. Plant Cell. 2008;20:3312–3330. doi: 10.1105/tpc.108.059568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kusano H, Testerink C, Vermeer JE, Tsuge T, Shimada H, Oka A, et al. The Arabidopsis phosphatidylinositol phosphate 5-kinase PIP5K3 is a key regulator of root hair tip growth. Plant Cell. 2008;20:367–380. doi: 10.1105/tpc.107.056119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Sous E, Kos B, Malhó R. Arabidopsis phosphatidylinositol-4-monophosphate 5-kinase 4 regulates pollen tube growth and polarity by modulating membrane recycling. Plant Cell. 2008;20:3050–3064. doi: 10.1105/tpc.108.058826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Stenzel I, Ischebeck T, König S, Hołubowska A, Sporysz M, Hause B, et al. The type B phosphatidylinositol-4-phosphate 5-kinase 3 is essential for root hair formation in Arabidopsis thaliana. Plant Cell. 2008;20:124–141. doi: 10.1105/tpc.107.052852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Camacho L, Smertenko AP, Pérez-Gómez J, Hussey PJ, Moore I. Arabidopsis Rab-E GTPases exhibit a novel interaction with a plasma-membrane phosphatidylinositol-4-phosphate 5-kinase. J Cell Sci. 2009;122:4383–4392. doi: 10.1242/jcs.053488. [DOI] [PubMed] [Google Scholar]
- 13.Mueller-Roeber B, Pical C. Inositol phospholipid metabolism in Arabidopsis. Characterized and putative isoforms of inositol phospholipid kinase and phosphoinositide-specific phospholipase C. Plant Physiol. 2002;130:22–46. doi: 10.1104/pp.004770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Takeshima H, Komazaki S, Nishi M, Iino M, Kangawa K. Junctophilins: a novel family of junctional membrane complex proteins. Mol Cell. 2000;6:11–22. doi: 10.1016/s1097-2765(00)00003-4. [DOI] [PubMed] [Google Scholar]
- 15.Im YJ, Davis AJ, Perera IY, Johannes E, Allen NS, Boss WF. The N-terminal membrane occupation and recognition nexus domain of Arabidopsis phosphatidylinositol phosphate kinase 1 regulates enzyme activity. J Biol Chem. 2007;282:5443–5452. doi: 10.1074/jbc.M611342200. [DOI] [PubMed] [Google Scholar]
- 16.Ma H, Lou Y, Lin WH, Xue HW. MORN motifs in plant PIPKs are involved in the regulation of subcellular localization and phospholipid binding. Cell Res. 2006;16:466–478. doi: 10.1038/sj.cr.7310058. [DOI] [PubMed] [Google Scholar]
- 17.Stace C, Manifava M, Delon C, Coadwell J, Cockcroft S, Ktistakis NT. PA binding of phosphatidylinositol 4-phosphate 5-kinase. Adv Enzyme Regul. 2008;48:55–72. doi: 10.1016/j.advenzreg.2007.11.008. [DOI] [PubMed] [Google Scholar]
- 18.Cockcroft S. Phosphatidic acid regulation of phosphatidylinositol 4-phosphate 5-kinases. Biochim Biophys Acta. 2009;1791:905–912. doi: 10.1016/j.bbalip.2009.03.007. [DOI] [PubMed] [Google Scholar]
- 19.Pical C, Westergren T, Dove SK, Larsson C, Sommarin M. Salinity and hyperosmotic stress induce rapid increases in phosphatidylinositol 4,5-bisphosphate, diacylglycerol pyrophosphate and phosphatidylcholine in Arabidopsis thaliana cells. J Biol Chem. 1999;274:38232–38240. doi: 10.1074/jbc.274.53.38232. [DOI] [PubMed] [Google Scholar]
- 20.Perera IY, Davis AJ, Galanopoulou D, Im YJ, Boss WF. Characterization and comparative analysis of Arabidopsis phosphatidylinositol phosphate 5-kinase 10 reveals differences in Arabidopsis and human phosphatidylinositol phosphate kinases. FEBS Lett. 2005;579:3427–3432. doi: 10.1016/j.febslet.2005.05.018. [DOI] [PubMed] [Google Scholar]
- 21.Jenkins GH, Fisette PL, Anderson RA. Type I phosphatidylinositol 4-phosphate 5-kinase isoforms are specifically stimulated by phosphatidic acid. J Biol Chem. 1994;269:11547–11554. [PubMed] [Google Scholar]
- 22.Mikami K, Saavedra L, Hiwatashi Y, Uji T, Hasebe M, Sommarin M. A dibasic amino acid pair conserved in the activation loop directs plasma membrane localization and is necessary for activity of plant type I/II phosphatidylinositol phosphate kinase. Plant Physiol. 2010;153:1004–1015. doi: 10.1104/pp.109.152686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Cooke FT, Dove SK, McEwen RK, Painter G, Holmes AB, Hall MN, et al. The stress-activated phosphatidylinositol 3-phosphate 5-kinase Fab1p is essential for vacuole function in S. cerevisiae. Curr Biol. 1998;8:1219–1222. doi: 10.1016/s0960-9822(07)00513-1. [DOI] [PubMed] [Google Scholar]
- 24.Shimada H, Koizumi M, Kuroki K, Mochizuki M, Fujimoto H, Ohta H, et al. ARC3, a chloroplast division factor, is a chimera of prokaryotic FtsZ and part of eukaryotic phosphatidylinositol-4-phosphate 5-kinase. Plant Cell Physiol. 2004;45:960–967. doi: 10.1093/pcp/pch130. [DOI] [PubMed] [Google Scholar]
- 25.Maple J, Vojta L, Soll J, Møller SG. ARC3 is a stromal Z-ring accessory protein essential for plastid division. EMBO Rep. 2007;8:293–299. doi: 10.1038/sj.embor.7400902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Otomo A, Hadano S, Okada T, Mizumura H, Kunita R, Nishijima H, et al. ALS2, a novel guanine nucleotide exchange factor for the small GTPase Rab5, is implicated in endosomal dynamics. Hum Mol Genet. 2003;12:1671–1687. doi: 10.1093/hmg/ddg184. [DOI] [PubMed] [Google Scholar]