Abstract
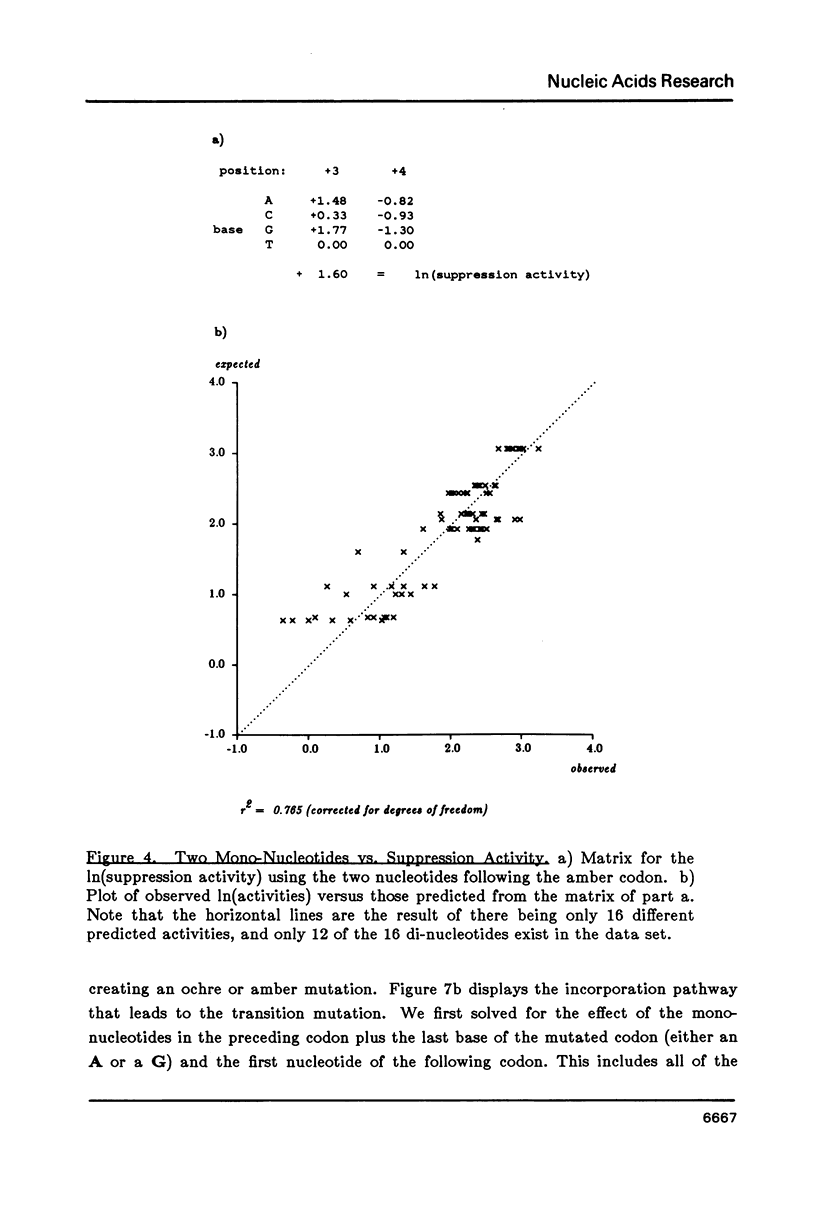
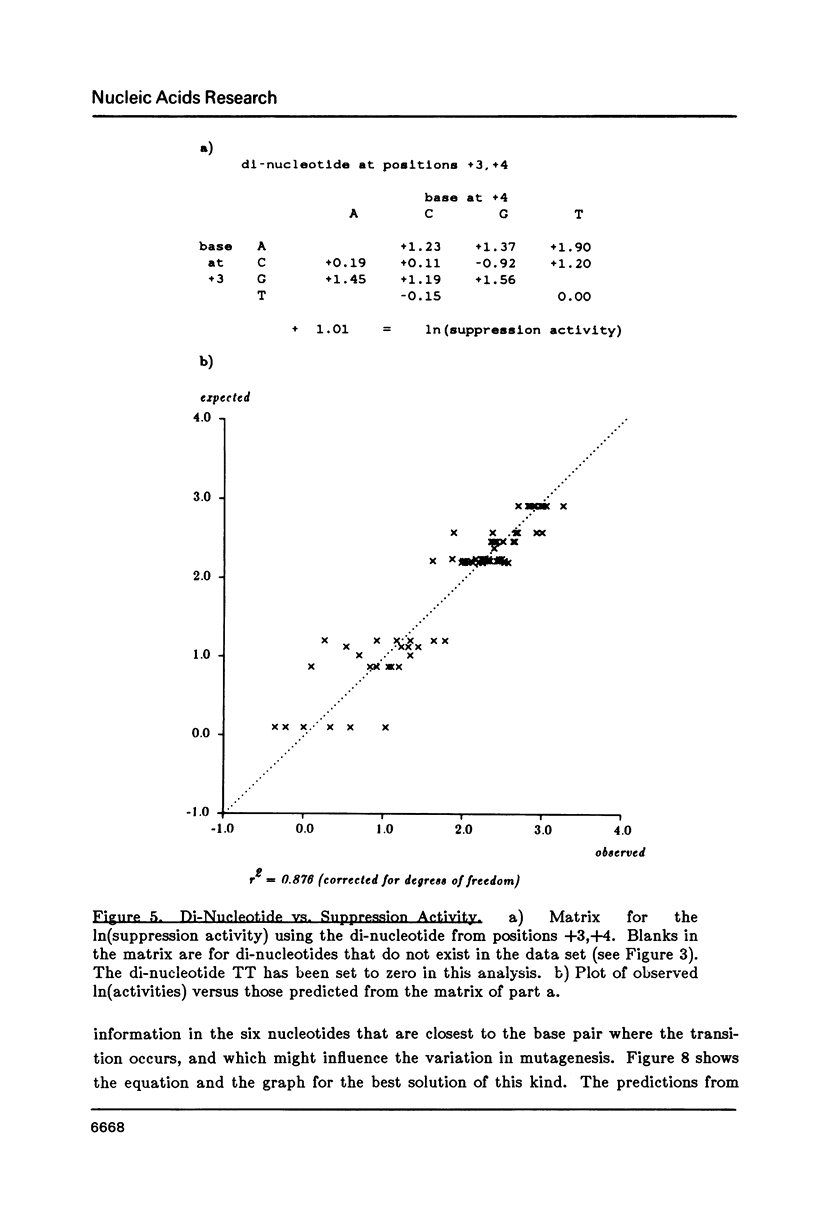
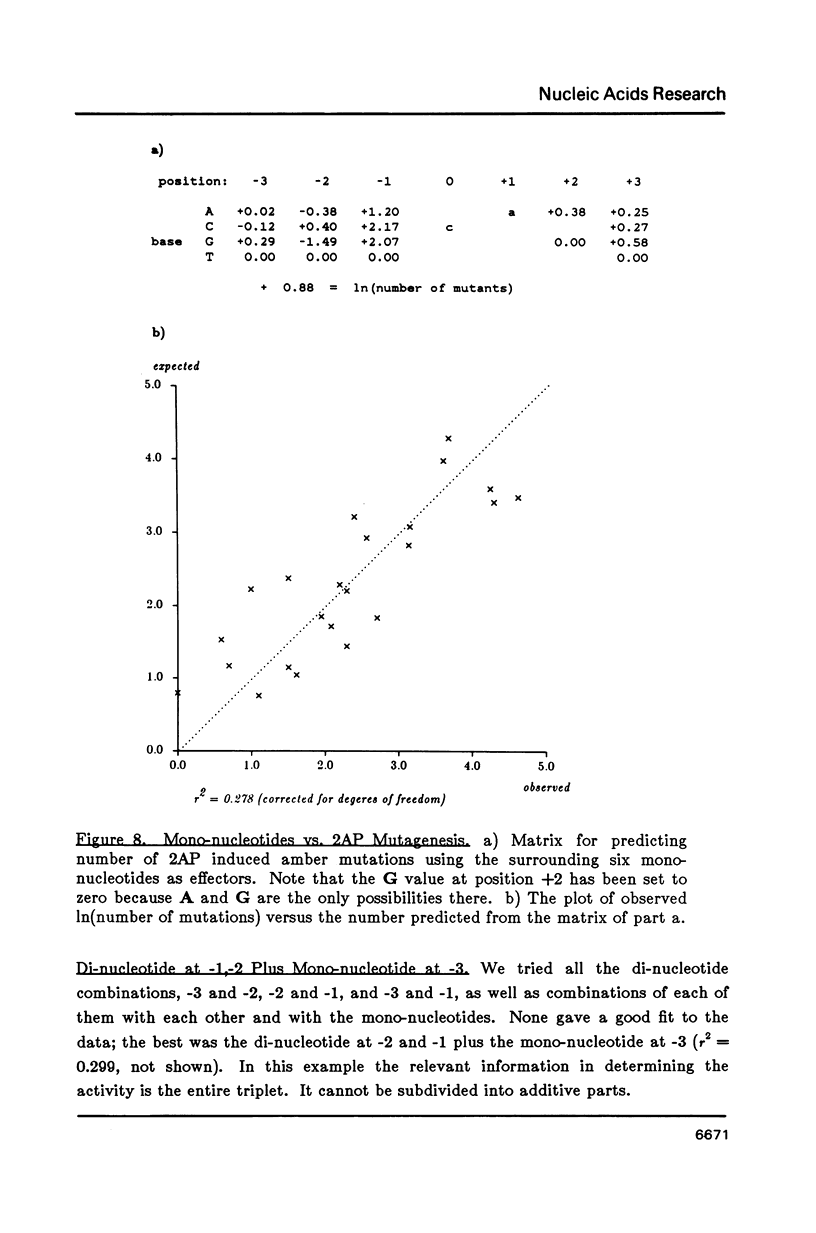
Matrices can be used to evaluate sequences for functional activity. Multiple regression can solve for the matrix that gives the best fit between sequence evaluations and quantitative activities. This analysis shows that the best model for context effects on suppression by su2 involves primarily the two nucleotides 3' to the amber codon, and that their contributions are independent and additive. Context effects on 2AP mutagenesis also involve the two nucleotides 3' to the 2AP insertion, but their effects are not independent. In a construct for producing beta-galactosidase, the effects on translational yields of the tri-nucleotide 5' to the initiation codon are dependent on the entire triplet. Models based on these quantitative results are presented for each of the examples.
Full text
PDF















Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ayer D., Yarus M. The context effect does not require a fourth base pair. Science. 1986 Jan 24;231(4736):393–395. doi: 10.1126/science.3510456. [DOI] [PubMed] [Google Scholar]
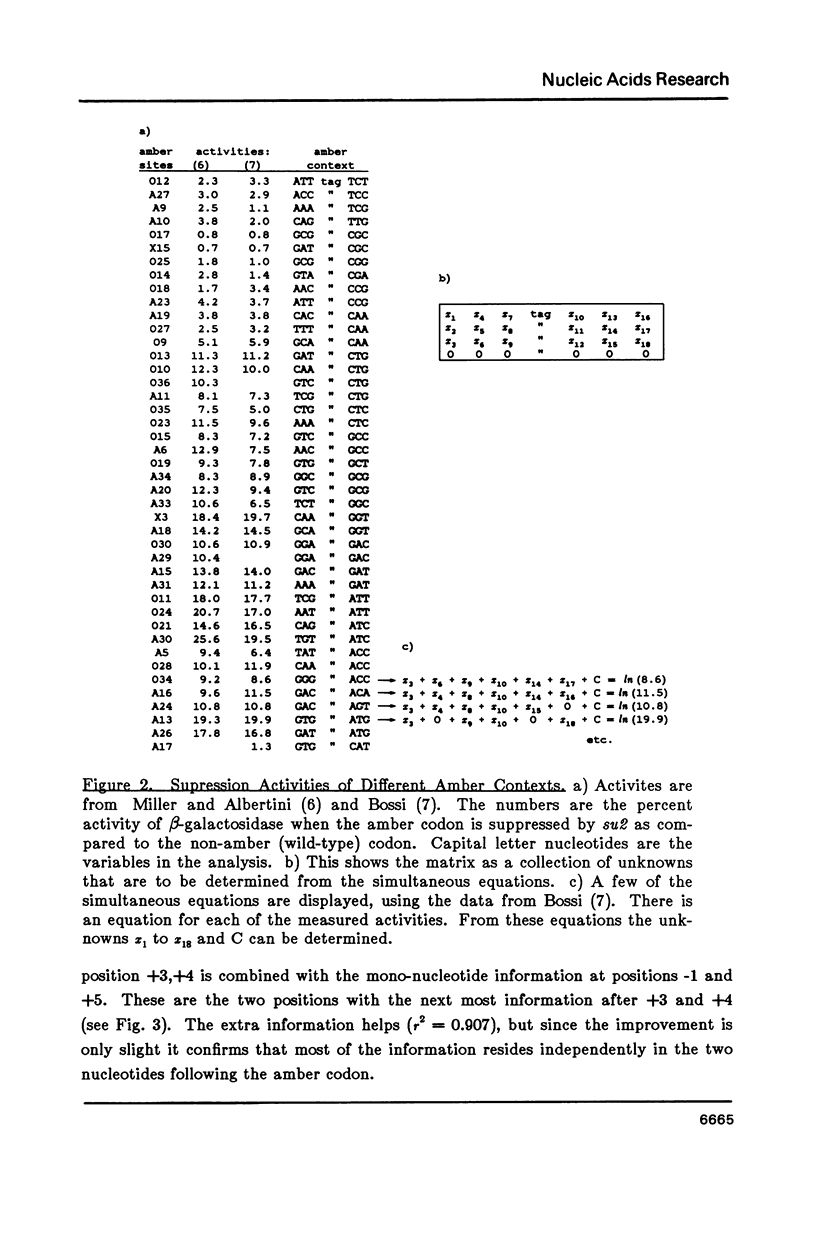
- Bossi L. Context effects: translation of UAG codon by suppressor tRNA is affected by the sequence following UAG in the message. J Mol Biol. 1983 Feb 15;164(1):73–87. doi: 10.1016/0022-2836(83)90088-8. [DOI] [PubMed] [Google Scholar]
- Brendel V., Trifonov E. N. A computer algorithm for testing potential prokaryotic terminators. Nucleic Acids Res. 1984 May 25;12(10):4411–4427. doi: 10.1093/nar/12.10.4411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coulondre C., Miller J. H., Farabaugh P. J., Gilbert W. Molecular basis of base substitution hotspots in Escherichia coli. Nature. 1978 Aug 24;274(5673):775–780. doi: 10.1038/274775a0. [DOI] [PubMed] [Google Scholar]
- Galas D. J., Eggert M., Waterman M. S. Rigorous pattern-recognition methods for DNA sequences. Analysis of promoter sequences from Escherichia coli. J Mol Biol. 1985 Nov 5;186(1):117–128. doi: 10.1016/0022-2836(85)90262-1. [DOI] [PubMed] [Google Scholar]
- Gold L., Pribnow D., Schneider T., Shinedling S., Singer B. S., Stormo G. Translational initiation in prokaryotes. Annu Rev Microbiol. 1981;35:365–403. doi: 10.1146/annurev.mi.35.100181.002053. [DOI] [PubMed] [Google Scholar]
- Harr R., Häggström M., Gustafsson P. Search algorithm for pattern match analysis of nucleic acid sequences. Nucleic Acids Res. 1983 May 11;11(9):2943–2957. doi: 10.1093/nar/11.9.2943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hui A., Hayflick J., Dinkelspiel K., de Boer H. A. Mutagenesis of the three bases preceding the start codon of the beta-galactosidase mRNA and its effect on translation in Escherichia coli. EMBO J. 1984 Mar;3(3):623–629. doi: 10.1002/j.1460-2075.1984.tb01858.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClure W. R. Mechanism and control of transcription initiation in prokaryotes. Annu Rev Biochem. 1985;54:171–204. doi: 10.1146/annurev.bi.54.070185.001131. [DOI] [PubMed] [Google Scholar]
- Mhaskar D. N., Goodman M. F. On the molecular basis of transition mutations. Frequency of forming 2-aminopurine-cytosine base mispairs in the G X C----A X T mutational pathway by T4 DNA polymerase in vitro. J Biol Chem. 1984 Oct 10;259(19):11713–11717. [PubMed] [Google Scholar]
- Miller J. H., Albertini A. M. Effects of surrounding sequence on the suppression of nonsense codons. J Mol Biol. 1983 Feb 15;164(1):59–71. doi: 10.1016/0022-2836(83)90087-6. [DOI] [PubMed] [Google Scholar]
- Mulligan M. E., Hawley D. K., Entriken R., McClure W. R. Escherichia coli promoter sequences predict in vitro RNA polymerase selectivity. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 2):789–800. doi: 10.1093/nar/12.1part2.789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ronen A. 2-Aminopurine. Mutat Res. 1980 Jan;75(1):1–47. doi: 10.1016/0165-1110(80)90026-3. [DOI] [PubMed] [Google Scholar]
- Schneider T. D., Stormo G. D., Haemer J. S., Gold L. A design for computer nucleic-acid-sequence storage, retrieval, and manipulation. Nucleic Acids Res. 1982 May 11;10(9):3013–3024. doi: 10.1093/nar/10.9.3013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schneider T. D., Stormo G. D., Yarus M. A., Gold L. Delila system tools. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):129–140. doi: 10.1093/nar/12.1part1.129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Staden R. Computer methods to locate signals in nucleic acid sequences. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 2):505–519. doi: 10.1093/nar/12.1part2.505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stormo G. D., Schneider T. D., Gold L. M. Characterization of translational initiation sites in E. coli. Nucleic Acids Res. 1982 May 11;10(9):2971–2996. doi: 10.1093/nar/10.9.2971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stormo G. D., Schneider T. D., Gold L., Ehrenfeucht A. Use of the 'Perceptron' algorithm to distinguish translational initiation sites in E. coli. Nucleic Acids Res. 1982 May 11;10(9):2997–3011. doi: 10.1093/nar/10.9.2997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tinoco I., Jr, Borer P. N., Dengler B., Levin M. D., Uhlenbeck O. C., Crothers D. M., Bralla J. Improved estimation of secondary structure in ribonucleic acids. Nat New Biol. 1973 Nov 14;246(150):40–41. doi: 10.1038/newbio246040a0. [DOI] [PubMed] [Google Scholar]
- Wulff D. L., Mahoney M., Shatzman A., Rosenberg M. Mutational analysis of a regulatory region in bacteriophage lambda that has overlapping signals for the initiation of transcription and translation. Proc Natl Acad Sci U S A. 1984 Jan;81(2):555–559. doi: 10.1073/pnas.81.2.555. [DOI] [PMC free article] [PubMed] [Google Scholar]


