SUMMARY
Lipin family members (lipin 1, 2, 3) are bi-functional proteins that dephosphorylate phosphatidic acid (PA) to produce diacylglycerol (DAG) and act in the nucleus to regulate gene expression. Although other components of the triglyceride synthesis pathway can form oligomeric complexes, it is unknown whether lipin proteins also exist as oligomers. In this study, by using various approaches, we revealed that lipin 1 formed stable homo-oligomers with itself and hetero-oligomers with lipin 2/3. Both the N- and C-terminal regions of lipin 1 mediate its oligomerization in a head-to-head/tail-to-tail manner. We also show that lipin 1 subcellular localization can be influenced through oligomerization, and the individual lipin 1 monomers in the oligomer function independently in catalyzing dephosphorylation of PA. This study provides evidence that lipin proteins function as oligomeric complexes and that the three mammalian lipin isoforms can form combinatorial units.
Keywords: lipin, oligomer, FRET, phosphatidic acid phosphatase
INTRODUCTION
Lipin proteins are a novel family of Mg2+-dependent phosphatidic acid phosphatases (PAPs), which catalyze the dephosphorylation of PA into DAG [1]. They are evolutionally conserved and show similar primary organization [2]. Lipin proteins consist of a N-terminal domain (N-LIP) and a C-terminal domain (C-LIP), with the latter containing a conserved DXDXT motif that is required for the catalytic activity [1–3]. Distinct from fungi, nematodes and insects that only express one lipin isoform, mammals express three lipin isoforms named lipin 1, 2, and 3 [2]. In addition, three lipin 1 protein isoforms are produced by alternative mRNA splicing [4–6]. Mice lacking lipin 1 have a reduction in adipose tissue mass and are insulin resistance [7]. Targeted overexpression of lipin 1 in mouse adipose tissue results in obesity, yet the mice maintain insulin sensitivity [8]. In humans, point mutations in the LPIN2 gene encoding lipin 2 cause Majeed syndrome [9], an autoinflammatory disorder, which may result from ablation of the PAP activity of the enzyme [10].
The lipin proteins primarily localize to the cytosol and are found in both the soluble and cytosolic membrane compartments, but can also be found in the nucleus [2,3,5,11]. Biochemical studies of PAP activities conducted prior to the identification of the gene encoding lipin 1 lead to the hypothesis that this enzymatic activity transiently associates with membranes to dephosphorylate PA (reviewed in [12]). Subsequent work has suggested that phosphorylation controls lipin 1 movement between cellular compartments [3,11]. The yeast lipin Pah1p has been reported to associate with the promoters of genes involved in phospholipid biosynthesis [13]. In mammalian cells, lipin 1 can positively or negatively regulate gene expression via interaction with transcription factors, such as the peroxisome proliferator-activated receptor α (PPARα) and NFATc4, as well as transcriptional coregulators such as PPARγ-coactivator 1α (PGC-1α) and histone deacetylases [14,15]. The PAP catalytic activity of lipin 1 appears to be separable from its activity in the regulation of transcription.
In the course of studies designed to identify lipin1 interacting proteins by immunoprecipitation of epitope tagged lipin isoforms we made the surprising discovery that lipin 1 can self associate. This study was conducted to investigate potential interactions between lipin isoforms. We show that lipin 1 forms both stable homo-oligomers and can also form hetero-oligomers with lipin 2 and lipin 3, suggesting that the function of these enzymes may be linked in a previously unappreciated manner.
Experimental
Chemicals
DAG (1–2-dioleoyl-sn-glycerol Dioleoyl) and PA (1,2-dioleoyl-sn-glycero-3-phosphate) were purchased from Avanti Polar Lipids. N-Ethylmaleimide (NEM) was purchased from Sigma. Lambda-Phosphatase (λPPase) was purchased from New England Biolabs.
Plasmids
V5-tagged lipin 1a, lipin 1b, lipin 2 and lipin 3 expression vectors were kind gifts from Dr. Karen Reue (University of California, Los Angeles) [16]. HA-Lipin 1a, HA-lipin 1b (G84R), HA-lipin 1b (Lmut), HA-lipin 1b (D712E), GST-lipin 1b (1–641), and GST-lipin 1b (642–924) were described previously [14]. To construct V5-lipin 1b (N1, 1–171), V5-lipin 1b (N2, 1–641), V5-lipin 1b (C1, 121–924), and V5-lipin 1b (C2, 469–924), corresponding cDNA fragments were PCR amplified from mouse lipin 1b and ligated into the pcDNA 3.1 D/V5-His-TOPO vector (Invitrogen). V5-LPP3 was described previously [17]. EYFP (Venus) and ECFP (Cerulean)-tagged lipin 1b constructs were constructed by subcloning full-length lipin 1b upstream of the fluorescent tags. Both Cerulean and Venus contained the monomeric A206K mutation. CAAX-lipin 1b was generated by PCR using the reverse primer 5'-GAGCGGCCGCctatgagagcacgcacttgcatgacatgcactgacggtcctctggctgcttagctgaggctgaatgc -3’. This removed the endogenous stop codon from lipin 1 and added the amino acid sequence KQPEDRQCMSCKCVLS*, with the ras-derived CAAX box at the carboxy terminus underlined. All of the clones were confirmed by DNA sequencing. Adenoviral constructs expressing HA-tagged lipin 1 and 2 have been previously described [14,18]
Antibodies
Anti-lipin 1 (LAb1 and LAb2) and anti-lipin 2 polyclonal antibodies were described previously [4,18]. The following antibodies were obtained commercially: anti-GAPDH (Abcam); anti-V5 (Invitrogen); anti-HA (Santa Cruz Biotechnology, Inc); anti-lipin 1 (Santa Cruz (C-15)); anti-V5 agarose (Sigma); anti-HA agrose (Sigma); Alexa Fluor488™, Alexa Fluor555™, and Alexa Fluor568™ anti-IgG antibody (Invitrogen); HRP-conjugated anti-IgG antibodies (Pierce); AP-conjugated anti-IgG antibodies (Sigma).
Cell culture and transfection
COS-7 (ATCC), HEK293T (American Type Culture Collection) and HEK293A (Invitrogen) cells were maintained in DMEM supplemented with 10% fetal bovine serum (FBS) (Invitrogen) and antibiotics. Cell transfections were performed with Lipofectamine™ 2000 (Invitrogen). 3T3-L1 cells were differentiated and maintained as previously described [3].
Generation of lipin baculovirus
Recombinant baculoviruses for expression of lipin 1 b with an N-terminal tandem His6 and HA tag were generated using the Bac-to-Bac baculovirus expression system (Invitrogen) according to the manufacturer’s protocol. Viral stocks were obtained by transfection of Sf9 insect cells cultured Grace’s Insect Medium with recombinant bacmids.
Purification of recombinant lipin protein
Recombinant lipin 1b protein tagged with both HA and 6xHis was extracted from a soluble fraction obtained from lysates of Sf9 cells that were infected with the recombinant lipin 1b baculovirus and cultured for 48h post-infection. Lipin protein was purified with Ni-NTA Purification System (Invitrogen). Briefly, pelleted cells were suspended in lysis buffer (50 mM NaH2PO4, 0.5 M NaCl, pH 8.0) and disrupted by three freeze-thaw cycles using liquid nitrogen and a 37°C water bath. After centrifugation at 6,000 rpm for 15 min, the supernatant was collected and mixed with the Ni-NTA resin for 1h at 4°C with gentle agitation. The resin was allowed to settle under gravity, and washed with 8ml of washing buffer (50 mM NaH2PO4, 0.5 M NaCl, 0.02 M imidazole, pH 8.0) 5 times. Purified lipin protein was eluted with 6ml of elution buffer (50 mM NaH2PO4, 0.5 M NaCl, 0.25 M imidazole, pH 8.0). The eluates were combined and concentrated to about 100 µl using Millipore Amicon ultra-4 centrifugal filter units.
Immunoprecipitations and immunoblotting
Cells were lysed by brief sonication in lysis buffer (50 mM Tris pH 7.5, 250 mM NaCl, 1 mM EDTA, 1 mM EGTA, 0.5% Triton X-100, 10% glycerol, complete Protease Inhibitor Cocktail (Roche Diagnostics). Lysates were subjected to immunoprecipitation with anti-V5 or anti-HA agarose. Immune complexes were washed five times with lysis buffer and subjected to immunoblotting with proper antibodies. Immunoblotting analysis was performed as previously described [3,19]. 3T3-L1 cells were homogenized with glass/Teflon in buffer A (50 mM β-glycerophosphate, 50 mM NaCl, 1 mM EDTA, 1 mM EGTA, and 10 mM sodium phosphate (pH 7.4), and 10 mM Triton X-100) supplemented with protease inhibitors (1 mM phenylmethylsulfonyl fluoride, 10 µg/ml leupeptin, 10 µg/ml aprotinin, 10 µg/ml pepstatin, 0.25 µM microcystin) and cleared by centrifugation at 16,000 g for 10 minutes. [3]. Lysates were subjected to immunoprecipitation using non-immune IgG, LAb2 [4], or lipin 2 antibodies [18].
Immunofluorescence microscopy
Immunofluorescent staining was performed as previously described [20]. In brief, cells were fixed with 4% paraformaldehyde in PBS for 30 minutes. Following fixation, cells were treated with 0.2% Triton X-100 in PBS for 5 minutes. After blocking with 10% FBS in PBS for 20 minutes, cells were incubated for 1 hour with primary antibody, followed by washing in PBS and incubation for 1 hour with the relevant secondary antibody. Nuclei were stained with Hoechst 33342 (Invitrogen) or DAPI (Vector Laboratories). Pictures of cells were taken with a laser-scanning confocal microscope (Olympus FV500, Tokyo, Japan) or a Zeiss Axiovert 200 fluorescent microscope. Images were pseudo-colorized and merged with ImageJ or Adobe Photoshop.
Forster Resonance Energy Transfer (FRET) Imaging
COS-7 cells on coverslips in 6-well plates were transfected with 6 µg of plasmids. Forty-eight hours after transfection cells were fixed at room temperature in 4% paraformaldehyde for 15 minutes. The system used for spectral imaging was a Zeiss Axiovert 200M epifluorescent motorized microscope coupled to a Zeiss 510 confocal-multiphoton imaging system at the University of Virginia’s W. M. Keck Center for Cellular Imaging. Details of imaging and the computational analysis used to separate donor spectral bleed-through (DSBT) and acceptor spectral bleed-through (ASBT) from the signal acquired during excitation of the FRET channel have been published [21]. Briefly, reference spectra for untagged, monomeric Cerulean or Venus fluorescent proteins were subtracted from cells expressing both lipin-Venus and Cerulean fusion proteins using linear unmixing and the algorithm previously described [21–23]. All Fluorescence Lifetime Imaging Microscopy (FLIM) imaging was performed on live HEK-293T cells that were transfected 36–48 hours previously with 3 µg lipin 1b-Cerulean, or lipin 1b-Cerulean + lipin 1b-Venus. FLIM data was acquired using a multiphoton system consisting of a Nikon TE300 epifluorescent microscope coupled to a Biorad Radiance2100 confocal and multiphoton system. A 10W Verdi tunable (700 to 1000nm) pulsed Ti:sapphire laser tuned to 820 nm was used for 2-photon excitation of the Cerulean fluorophore at 13 nsec pulse intervals. Photons were collected using a photomultiplier tube (PMT) over a period of 120 seconds for each measurement. Data collection and analysis was performed using SPCImage software (Becker & Hickl GmbH) as well as ImageJ.
GST-pull down
GST-pull down experiment was performed as previously described [24], with major modifications. In brief, HEK293A cells transiently expressing various V5-tagged lipin 1b fragments were lysed in ice cold lysis buffer (50 mM Tris pH 7.5, 250 mM NaCl, 1 mM EDTA, 1 mM EGTA, 0.5% Triton X-100, 10% glycerol, complete Protease Inhibitor Cocktail (Roche Diagnostics). Samples were briefly sonicated and centrifuged at 16,000g at 4°C for 30 minutes. The supernatant was incubated with anti-V5 agarose at 4 °C overnight. The beads were extensively washed with lysis buffer, and the recombinant lipin peptide was eluted by the incubation with 70 µL V5-peptide (100 µg/ml) in lysis buffer at 4°C for 1 h. After brief centrifugation, the supernatants were mixed with GST, GST-lipin 1b (1–641), or GST-lipin 1b (642–924) prebound to Glutathione Sepharose beads in 1 ml of binding buffer (50 mM Tris pH 7.5, 250 mM NaCl, 1 mM EDTA, 1 mM EGTA, 0.5% Triton X-100, 10% glycerol, complete Protease Inhibitor Cocktail (Roche Diagnostics), and incubated with rotation for 4 hours at 4 °C. The beads were washed four times with 1 ml binding buffer, separated by SDS–PAGE and analyzed by immunoblotting.
PAP activity assay
Ten cm plates of HEK-293T cells were transfected (Lipofectamine 2000) with 0.5 µg of myc-lipin 1b, and 0.25, 0.5 or 1.0 µg of HA-lipin 1b (D712E) plasmids. Forty-eight hours after transfection cells were homogenized in glass/Teflon at 1,000 rpm in 1 ml buffer A. Immunoprecipitations were performed as previously described [3] and divided into separate aliquots for electrophoresis and PAP activity measurement. PAP activity was determined by measurement of soluble [32P]-orthophosphate release from [32P]-PA using substrates presented in Triton X-100/phosphatidic acid mixed micelles as previously described [18].
Gel filtration chromatography of 3T3-L1 adipocyte extracts
3T3-L1 adipocytes were homogenized by glass/Teflon in a minimal volume of TES buffer (255 mM sucrose, 1 mM EDTA, 0.1 mM DTT, 1 mM Tris, pH 7.4) supplemented with protease inhibitors. Extracts were centrifuged at 16,000 g for ten minutes and the resulting supernatants subjugated to 175,000 g centrifugation for 60 minutes. The soluble fraction was filtered through 0.45 µm cellulose acetate syringe filter and 400 µl representing 1.0 to 1.6 mg total protein was loaded on a Superose 6 10/300 GL column equilibrated with buffer A. The sample was eluted at a flow rate of 0.1 ml/minute and 1 ml fractions collected. The columns were calibrated with molecular weight standards obtained from GE/Pharmacia. PAP activity measurements were performed on 40 µl aliquots for 20 minutes at 37 °C. In some cases samples were pre-incubated in 10 mM Triton X-100 or 1% fatty acid poor bovine serum albumin for 15 minutes at room temperature with continuous mixing before loading on column.
Gel Filtration Chromatography of recombinant lipin 1b protein
50 µl of concentrated protein representing approximately 10 µg of purified lipin 1b analysed by gel filtration chromatography on a Superose 6 10/300 GL column. The flow rate used in the separation was 0.5 ml/minutes with phosphate buffered saline (137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.76 mM KH2PO4, pH 7.4). The columns were calibrated with molecular weight standards obtained from GE/Pharmacia. After 14 min, 30 fractions of 0.5ml of each were collected. PAP activity measurements of gel filtration fractions of recombinant lipin 1 were performed essentially as described [16]. Briefly, 65 µl of column eluates were assayed for times up to 60 minutes (<20% substrate hydrolysis) at 30 °C in 100 µl volumes of reaction buffer (50 mM Tris, pH 7.4, 1 mM MgCl2, 2 mM Triton X-100, 2 mg/ml Fatty Acid Free BSA, 0.1 mM PA (with ~20,000 dpm of 32P labeled PA made by phosphorylation of DG with bacterial DG kinase)) for the release of soluble [32P]-orthophosphate from Triton X-100/[32P]-phosphatidic acid mixed micelles.
Blue Native Polyacrylamide Gel Electrophoresis (BN-PAGE)
HEK293A cells cultured in 35mm plates were transfected with pcDNA3 (vector), lipin 1b-V5 or lipin 1b-V5 (D712E). 36 hours later, cells were lysed in 100 µl lysis buffer (50 mM Tris pH 7.5, 50 mM NaCl, 1% digitonin, complete Protease Inhibitor Cocktail (Roche Diagnostics)). Following centrifugation, 15 µl of samples was loaded to NativePAGE™ Novex Bis-Tris Gel sytem (Invitrogen), transferred to PVDF membrane, according to manufacturer’s manual. The proteins on the membrane were fixed with 8% acetic acid and probed with anti-V5 antibody and HRP-coupled secondary antibody.
RESULTS
Lipin proteins self-associate in mammalian cells
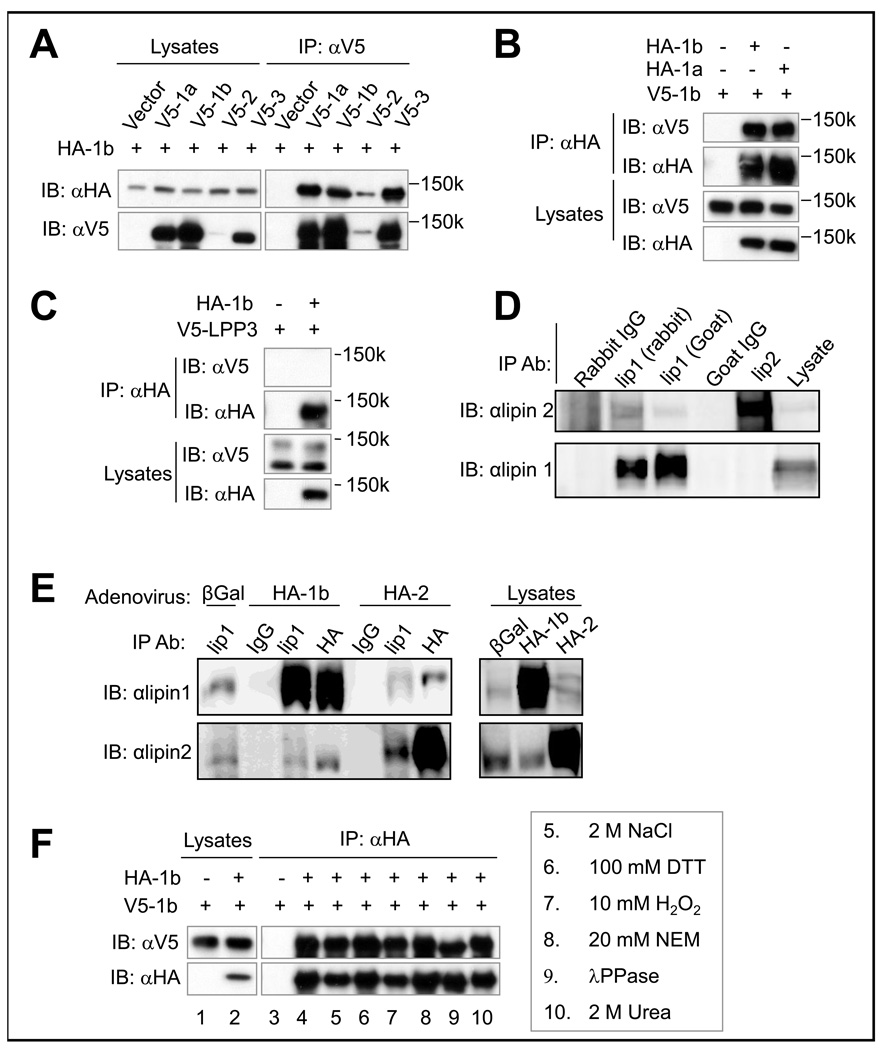
COS-7 cells were transiently transfected with plasmid constructs encoding N-terminally HA-tagged lipin 1b, together with or without C-terminally V5-tagged lipin 1a, 1b, 2, or 3, respectively. Thirty-six hours later, the cells were homogenized in stringent lysis buffer (0.5% Triton X-100 and 250 mM NaCl) and epitope tagged proteins were precipitated with anti-V5 agarose. As shown in Figure 1A, HA-lipin 1b was co-immunoprecipitated with all of the V5-tagged lipin isoforms. In a reciprocal co-IP experiment, HA-tagged lipin 1a or lipin 1b also associated with lipin 1b-V5 (Figure 1B). As reported previously [16], the V5-tagged lipin 2 was expressed at lower levels than V5-tagged lipin 1 or 3 (Figure 1A). However, despite decreased expression V5-tagged lipin 2 still co-immunoprecipitated lipin 1 at levels proportionate to its expression. Next, since LPP family proteins are also PA phosphatases and have been reported to form homo- or hetero-oligomers [25], we examined the possible interaction between lipin 1b and LPP3, a representative LPP. As shown in Figure 1C, HA-lipin 1b did not co-immunoprecipitate with V5-tagged LPP3, arguing that lipin 1 associates with itself and other lipin isoforms but not with LPPs. In the subsequent experiment we found that endogenous lipin 2 was immunoprecipitated with anti-lipin 1 antibodies from the lysates of differentiated 3T3-L1 cells. This indicates that the lipin-lipin associations are not a result of ectopic expression of the proteins (Figure 1D). Whereas we failed to detect lipin 1 in anti-lipin 2 immunoprecipitates from 3T3-L1 lysates, this could be explained by the possibility that association with lipin 1 might prevent the accessibility of the lipin 2 epitope to the anti-lipin 2 antibody. Finally, immunoprecipitation of adenovirally delivered HA-tagged lipin 1 or lipin 2 in 3T3-L1 adipocytes revealed that endogenous lipin 1 and 2 were able to interact with recombinant lipin 1 and 2 (Figure 1E). Together, these co-immunoprecipitation experiments reveal that lipin proteins can form homo- or hetero-oligomers in vivo.
Figure 1. Lipin 1 coimmunoprecipitates with itself and lipin 2, and 3, but not LPP3.
(A) COS-7 cells were transfected with constructs for expression of HA-lipin 1b and various V5-tagged lipin expression vectors as indicated. 36 hours after transfection, cell lysates were subjected to immunoprecipitation with an anti-V5 antibody, followed by immunoblotting with anti-HA or anti-V5 antibodies. (B) COS-7 cells were cotransfected with vectors for expression of V5-lipin 1b and HA-tagged lipin 1a or lipin 1b as indicated. 36 hours after transfection, lysates were subjected to immunoprecipitation with anti-HA antibody, followed by immunoblotting with anti-HA or anti-V5 antibodies. (C) COS-7 cells were cotransfected with vectors for expression of V5-LPP3 and HA-tagged lipin 1b. 36 hours after transfection, lysates were subjected to immunoprecipitation with anti-HA antibody, followed by immunoblotting with anti-HA or anti-V5 antibodies. (D) 3T3-L1 adipocytes were homogenized in buffer A containing 10 mM Triton X-100 and equal amounts of lysates were used for immunoprecipitation with rabbit or goat non-immune IgG, lipin 1 (LAb2 or Santa Cruz C-15), or lipin 2 antibodies, followed by immunoblotting with anti-lipin 1 (LAb1) or 2 antibodies. (E) 3T3-L1 adipocytes were infected for two days with adenovirus vectors expressing HA-tagged lipin 1 or 2. Cells were homogenized and immunoprecipitations were performed as in 1D, but with anti-lipin 1 (LAb2) or HA (12CA5) antibodies. (F) COS-7 cells were transfected with vectors for expression of V5-lipin 1b, with or without HA-lipin 1b, and after 36 hours cell lysates were prepared and were subjected to immunoprecipitation with anti-HA antibody. After 3 washes with lysis buffer, the immunoprecipitates were washed another time with either Tris buffer (50 mM Tris-HCl pH 7.5, 50 mM NaCl) (lanes 4), or Tris buffer containing indicated materials (lanes 5–10). The beads were subjected to another round of washing with lysis buffer and then subjected to immunoblot analysis.
Lipin 1 forms stable oligomers
Next, we examined the biochemical properties of lipin oligomers by identifying experimental conditions that would allow us to reversibly assemble and disassemble these complexes in vitro. Coordination of Mg2+ in the active site is necessary for the PAP activity of lipin 1 [1]. However, lipin oligomerization was observed in the presence of both EDTA and EGTA indicating that these associations are divalent cation-independent. PAP activity of lipin 1 is dependent on NEM-sensitive cysteines [16,26]. However, as shown in Figure 1F, washing immunoprecipitates with a buffer containing either oxidants NEM or H2O2, or the reducing agent DTT, did not affect lipin 1b self-association (Figure 1F, lane 6,7,8). This suggests that lipin 1 oligomerization is redox-insensitive, and does not involve formation of intermolecular disulphide bonds. Lipin 1 is a phosphoprotein and phosphorylation of lipin 1 may play a regulatory role in mediated oligomerization. However, we found that lipin 1 oligomerization appears to be independent of its phosphorylation status, because incubation of the immunoprecipitates with λPPase, which effectively dephosphorylated the protein as indicated by altered mobility on SDS-PAGE (Figure 1F), did not impair the lipin-lipin interaction (Figure 1F, lane 9). Finally, the interaction between V5-lipin 1b and HA-lipin 1b was resistant to high concentration of salt (2 M NaCl) and moderate concentration (2 M) of urea (Figure 1F, lane 5, 10), suggesting that lipin 1 oligomers are stable even at increased ionic strength or in the presence of a denaturant.
Mapping the domains of lipin 1 in mediating oligomerization
We next investigated the regions of lipin 1 that are necessary for self association. A series of N- or C-terminally truncated variants of lipin 1b with C-terminal V5 tag was constructed and co-expressed with HA-lipin 1b in COS-7 cells. The interactions between the full-length and the truncated mutants of lipin 1b were determined by co-immunoprecipitation. As shown in Figure 2A and 2B, full length lipin 1b interacted with all of the N- and C-terminal fragments of lipin 1b employed, suggesting that the N-terminal (1–171) and C-terminal (469–924) regions of lipin 1b simultaneously mediate lipin 1 oligomerization. Using dimerization, the simplest organization form of lipin 1, as an example, there are two possible models for lipin-lipin association: (1) “head-to-head/tail-to-tail” (2) “head-to-tail/tail-to-head”. To test these two possibilities, glutathione sepharose-bound GST-lipin 1b (1–641) or lipin 1b (642–924) from bacteria was incubated with V5-lipin 1b (1–171) or V5-lipin 1b (469–924) immunopurified from HEK293A cells and interacting proteins were evaluated by immunoblotting with anti-V5 antibody. As shown in Figure 2C, the N-terminal part of lipin 1b only interacted with the N-terminus, while the C-terminal part preferred to associate with the C-terminus. Our data thus suggest a model that lipin 1 oligomerizes through “head-to-head/tail-to-tail” interactions.
Figure 2. Mapping the lipin 1b domains in mediating its oligomerization.
(A) Schematic representation of the V5-tagged lipin 1b truncation mutant constructs used in co-IP and GST-pull down experiments. Their abilities to interact with HA-lipin 1b or GST-fused lipin 1b fragments are indicated as “+” (positive), “−” (negative) or “N/D” (not determined). NLIP, CLIP, and NLS (nuclear localization sequence) are shown. (B) COS-7 cells were co-transfected with HA-lipin 1b and various V5-tagged lipin 1b fragments as indicated. 36 hours after transfection, the lysates were subjected to immunoprecipitation with anti-V5 antibody, followed by immunoblotting with anti-HA or anti-V5 antibody. (C) In vitro interaction between GST-lipin 1b fragments and immunopurified V5-tagged lipin 1b fragments.
Co-localization of lipin family members in cells
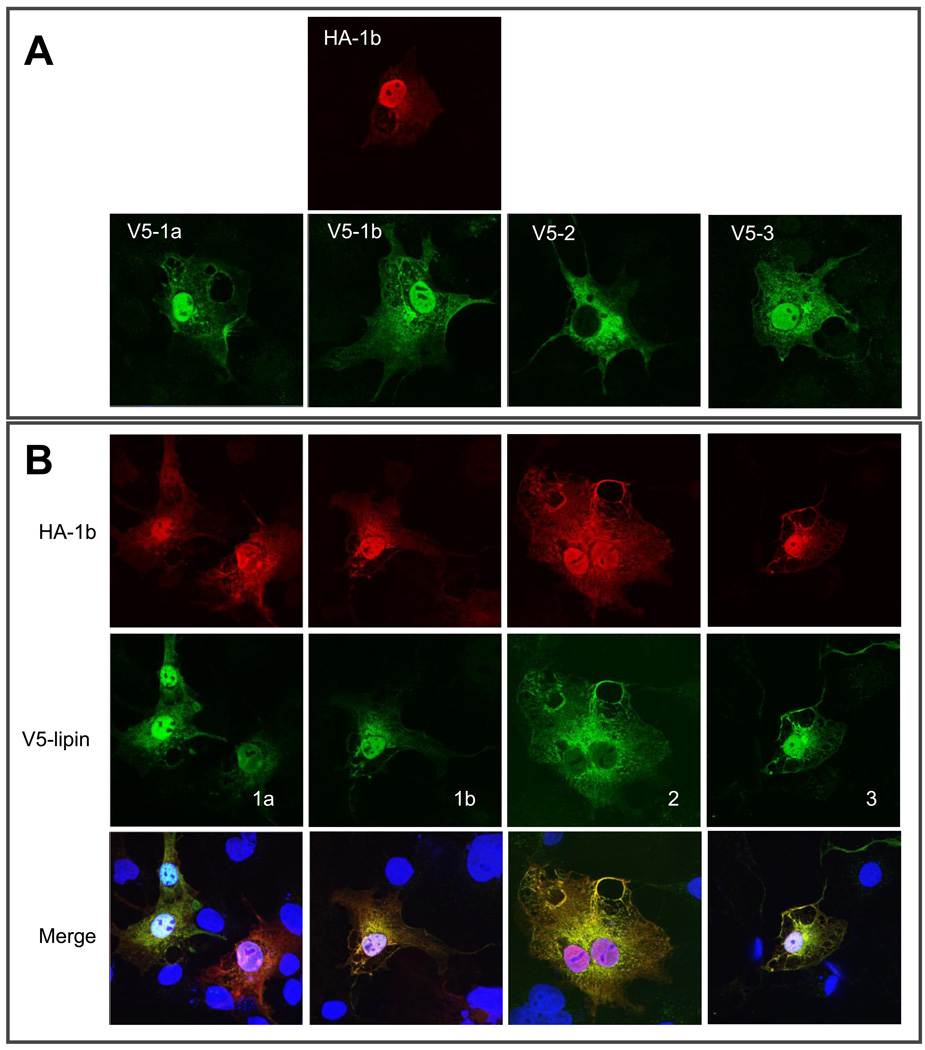
Lipins exhibit variable subcellular localization patterns. As shown in Figure 3A, except for lipin 2, which is mainly excluded from nucleus, other lipin isoforms including lipin 1a, 1b, and 3 are targeted to both cytosol and nucleus in the COS-7 cells. Coexpression of HA-tagged lipin 1b with individual V5-tagged lipin isoforms revealed substantial co-localization (as shown by yellow immunofluorescence in the merged panels of Figure 3B). Lipin 1 also showed a co-localization with lipin 2 outside of the nucleus (Figure 3B). Co-localization of these lipin isoforms is consistent with their ability to self associate as revealed by the experiments shown in Figure 1.
Figure 3. Colocalization of lipin 1 and lipin 1, 2, and 3 in cells.
(A) COS-7 cells were transfected with vectors for expression of V5-tagged lipin 1a, 1b, 2 or 3, or HA-tagged lipin 1b. 36 hours after transfection, the cells were fixed and subjected to immunostaining with anti-HA or anti-V5 antibody. (B) COS-7 cells were cotransfected with HA-tagged lipin 1b and V5-tagged lipin 1a, 1b, 2 or 3. 36 hours after transfection, the cells were subjected to immunostaining with anti-HA and anti-V5 antibody. Red: HA-lipin 1b; Green: V5-tagged lipin isoforms; Blue: Hoechst 33342.
Demonstration of intermolecular interactions between lipin monomers in cells by FRET
The data shown in Figure 3 indicate that lipin isoforms are co-localized in cells but provide no information about the proximity of these proteins. To address this issue lipin 1 was tagged at the carboxy-terminus with either Venus fluorescent protein (YFP) or Cerulean fluorescent protein (CFP) and energy transfer from the donor (Cerulean-lipin 1) to the acceptor (Venus-lipin 1) was determined by spectral imaging. Venus-lipin 1 and Cerulean-lipin 1 showed extensive co-localization as expected from the immunofluorescent imaging results (Figure 4A). In the event of close opposition of the fluorescent tags, such as occurs upon interaction of the tagged proteins, energy from the donor molecule can be transferred to the acceptor molecule by a process termed Förster Resonance Energy Transfer, or FRET. Because FRET is dependent on proximity, fluorophores must be within ~1–10 nm of each other for energy transfer to occur, a distance that also typically occurs during protein-protein interaction. Excitation at 458 nm in cells where Venus-lipin 1 and Cerulean-lipin 1 were co-expressed led to an increase in emission at 540 nm when compared to either alone (Figure 4B). To quantitate the amount of energy transferred by excitation of the donor to emission of the acceptor requires subtraction of the contaminating donor spectral bleed-through (DSBT) and acceptor spectral bleed-through (ASBT). For example, excitation of either Cerulean or Cerulean-lipin 1 alone at 458 nm gives a peak emission at 500 nm, however, DSBT can be seen in the emission at 540 nm in double labeled cells (Figure 4B). While excitation of YFP or Venus-lipin 1 at 514 nm gives a peak emission at 540 nm, excitation at 458 nm also leads to ASBT emission at 540 nm, albeit at a reduced level compared to excitation at 514 nm. In order to subtract DSBT and ASBT from the actual FRET signal, linear unmixing was used to remove the DSBT from the FRET channel, and a computer algorithm was employed to identify and remove contaminating ASBT from the FRET signal [21,22]. The resulting values, termed processed FRET (PFRET) showed that the efficiency of energy transferred from the donor to the acceptor (E%) was 21.7% (SD +/−4.66, n=5) which is similar in magnitude to the energy transfer efficiency observed for a number of other well characterized protein-protein interactions such as CFP- and YFP-tagged C/EBPα homodimers at 23% (SD +/−5%) [21]. That the energy transfer E% values are not dependent on acceptor concentration is shown in Figure 4C to the right where the calculated E% values from a single cell are plotted against the acceptor intensity (intensity/pixels). Non-specific transfer of energy (E%) that is dependent on acceptor intensity gives a correlation coefficient approaching 1, while the correlation coefficient of the fluorescently-tagged lipin 1 protein(s) in this representative example was −0.284, showing that the calculated E% was independent of acceptor concentration. Taken together, these results indicate that the association between lipin isoforms observed using biochemical approaches in cell lysates reflects close interactions between these proteins in live cells.
Figure 4. FRET analysis of fluorescent-tagged lipin 1.
(A) Co-localization of Venus-lipin 1b and Cerulean-lipin 1b. Images merged in Image J. (B) Single cell example of uncorrected emission spectra from co-transfected Venus-lipin 1b and Cerulean-lipin 1b (Co-transfection) at 514 or 458nm, and single transfected Venus-lipin 1b (Acceptor alone) or Cerulean-lipin 1b (Donor Alone) excited at 458nm. (C) (Left) Intensity map showing E% of a representative cell where spectral bleed through of both donor and acceptor have been removed by linear unmixing and a computer algorithm (PFRET). (Right) Graph of E% versus acceptor intensity of individual cell shown on left. (D) Decay curve for cells transfected with either Cerulean-lipin 1b (Cer-lip) alone or Cerulean-lipin 1b and Venus-lipin 1b (Cer-lip + Ven-lip). (E) Average decay time (nsec) per cell for Cerulean-lipin 1b (Cer-lip) and Cerulean-lipin 1b + Venus-lipin 1b (Cer-lip + Ven-lip), * indicates statistical significance (P < 0.00001).
As a second measure of FRET we employed Fluorescence Lifetime Imaging Microscopy (FLIM) to measure changes in fluorescence decay of the donor in the presence of an acceptor. We measured the lifetime of the donor, Cerulean-lipin 1b, with and without the acceptor, Venus-lipin 1b, in live HEK-293T cells. In order to do so, the fluorophores were excited by a single laser pulse at 820 nm and the photons released during the return of the fluorophore to the ground state from the excited state were captured and displayed as data points. SPCImage software generated a fit curve based on this data that verified an exponential decay of the fluorophore. Figure 4D compares the fluorescence decay of Cerulean-lipin 1 alone with Cerulean-lipin 1 plus Venus-lipin 1. In this example the presence of Venus-lipin 1 decreases the slope of the decay curve of Cerulean-lipin 1. Quantitation of multiple cells reveals this is significant with the average decay time of Cerulean-lipin 1 decreasing from 2.226 nsec (SD +/− 0.08, n=14) to 2.101 nsec (SD +/− 0.09, n=16, P < 0.00001) in the presence of Venus-lipin 1, providing additional support for the Venus and Cerulean fluorophores being in close enough apposition for energy to be transferred from the donor to the acceptor.
Forced relocalization of lipin 1 in cells
Subcellular fractionation studies indicate that lipin 1 translocates between the cytosol and the endoplasmic reticulum/microsomal membranes in a dynamic manner dependent on its phosphorylation state and fatty acid availability [3,27]. However, when visualized by immunofluorescence intracytosolic changes in lipin 1 localization are subtle (data not shown). Therefore to test the hypothesis that lipin self association could lead to redistribution of lipin isoforms in cells we determined whether co-expression of a plasma membrane-targeted lipin 1b variant (see below) could redirect cytosolic lipin 1b to the plasma membrane through oligomerization. We employed a myc-tagged lipin 1, which is localized diffusely in transfected COS-7 cells, with undetectable plasma membrane staining (Figure 5, upper panels). In addition, we developed a plasma membrane-targeted lipin 1b (lipin 1b-CAAX) construct, which upon overexpression showed clear plasma membrane localization in addition to the whole cell distribution (Figure 3, middle panels). The plasma membrane-targeted lipin 1b was obtained by fusing a CAAX-box motif to the carboxy-terminal tail of lipin 1b, to direct prenylation of lipin 1 [28]. Co-transfection of both constructs resulted in the easily observable relocalization of myc-lipin 1 to the plasma membrane. A portion of the images are shown at a larger size at the bottom. Similar results were seen with lipin 2 co-transfected with lipin 1-CAAX (data not shown). These results demonstrate that oligomerization between lipin isoforms can direct intracellular localization.
Figure 5. Forced plasma membrane-targeting of lipin 1 via a CAAX domain can recruit soluble lipin 1 to the plasma membrane.
COS-7 cells were transfected with plasmids for expression of CAAX-tagged lipin 1b and/or myc-tagged lipin 1b. 48 hours after transfection cells were fixed and subjected to immunostaining with anti-HA and/or anti-Myc antibodies. Transfections indicated to the left, and immunostaining shown at the top. Red: HA-lipin 1b-CAAX; Green: Myc-lipin 1b, Blue: DAPI.
Estimation of the native size of endogenous and recombinantly expressed lipin 1
Taken together, the data presented indicate that lipin proteins can self associate to form oligomeric complexes that can be isolated biochemically and visualized by fluorescence imaging. The NEM-sensitive PAP activity (a hallmark feature of the lipin enzymes) of rat liver cytosol was reported to elute with an Mr of 540,000 during gel filtration chromatography [29]. Because 3T3-L1 adipocytes express high levels of lipin 1 protein, we analyzed soluble fractions of these cells by Superose 6 gel filtration chromatography to determine the size of native lipin 1. As detected by Western blotting, lipin 1 and lipin 2 were eluted from the column with an Mr of 500,000–600,000; the same size observed for the majority of the PAP activity (Figure 6A and B). Although the predicted molecular weight of the lipin1, 2 and 3 polypeptides is ~90–100 kDa, on SDS-PAGE these proteins migrate aberrantly with an apparent Mr of ~140–150 kDa. In the case of lipin 1, phosphorylation accounts for part, but not all, of the unexpected decrease in mobility on SDS-PAGE [3]. Gel filtration measures the hydrodynamic radius of a molecule, not the molecular weight, and the tertiary structure of lipin molecules could partially account for the discrepancy in its elution from the column. Alternatively, unidentified post-translational modifications other than phosphorylation could produce changes in lipin migration on SDS-PAGE and elution by size exclusion chromatography. Indeed, a proportion of lipin 1 has been recently reported to be SUMOylated, a covalent modification of ~17 kDa [30]. Nevertheless, it is highly unlikely that post-translational modification or the shape of individual lipin molecules would change the elution profile from Mr of 100 kDa to an Mr of 500–600 kDa. Based purely on the predicted molecular weight of lipin 1, tetramerization of lipin 1 would produce an Mr of approximately 400 kDa if the lipin tetramer adopted a mostly spherical shape. Therefore a combination of oligomerization, probably tetramerization, molecular shape, and possibly post-translational modifications likely account for size of lipin 1 as measured by size exclusion chromatography. Addition of 10 mM Triton X-100 to cytosolic extracts before loading on the column caused a partial shift in the lipin 1 elution profile consistent with the addition of the predicted Mr of a Triton X-100 micelle, whereas treatment of extracts with 1% BSA was without effect (Figure 6A and data not shown).
Figure 6. Lipin 1 exists as an oligomer in 3T3-L1 adipocytes.
(A) Column fractions from Superose 6 gel filtration chromatography of 3T3-L1 adipocte cytosolic extract immunoblotted for lipin 1 and 2. Bottom panel is same extracts incubated with 10mM TX-100 before loading on column. (B) Lipin 1 immunoreactivity (arbitrary units) in column fractions versus Mg2+-dependent PAP activity (nmol/min/ml).
We also examined the molecular weight of recombinant lipin 1 that was expressed in insect cells and purified by metal ion affinity chromatography. Similar to our observations with endogenously expressed lipin 1 in 3T3-L1 cells, purified lipin 1 eluted as a discrete peak of Mr ~600,000, coincident with the elution pattern of PAP activity from the column (Figure 7A and B). Isolation of mono/di/tetrameric lipin 1 would allow functional studies to determine whether the different oligomeric forms of lipin 1 display differing enzymatic activities. In an attempt to obtain fractions containing lipin 1 species with molecular weights that are consistent with those of a monomer or dimer we employed a Superdex 200 column to increase the resolution of recombinant lipin eluting at Mr 50–500,000 (data not shown). However, recombinant lipin appears to be largely, if not exclusively, tetrameric and so we were unable to address the question of oligomer effects on PAP activity.
Figure 7. Recombinant lipin 1 forms dimers and tetramers.
(A) Column fractions from Superose 6 gel filtration chromatography of purified recombinant lipin 1b were collected and analyzed for PAP activity. (B) Immunoblotting of column fractions. (C) HEK293A cells transiently expressing V5-lipin 1b or V5-lipin 1b (D712E) were homogenized in digitonin-containing lysis buffer, the lysates separated by BN-PAGE and subjected to immunoblotting with anti-V5 antibody.
As an alternative method to gel filtration chromatography, extracts from HEK293A cells expressing V5-tagged lipin 1b were subjected to Blue Native PAGE gel electrophoresis in an attempt to preserve the native molecular structure. As shown in Figure 7C, immunoblotting with anti-V5 antibody clearly showed that native, non-denatured lipin 1b existed in two forms with a molecular weight slightly more than two and four times the predicted lipin 1b-V5 monomer (compared to the MW with SDS-PAGE in Figure 1A). Like gel filtration, native PAGE measures hydrodynamic radius and the more the divergence from a spherical shape the larger the molecule or molecular complex will appear by these chromatographic methods. These experiments suggest that both endogenous and recombinant lipin 1 exists primarily as tetrameric complexes both in cells and when purified in vitro. While more detailed analyses will be required to rule out the presence of other non-lipin proteins in these complexes, our determination of the molecular weight of recombinant lipin 1 by gel filtration chromatography and by non denaturing electrophoresis argue that the major lipin 1 species is a tetramer.
The relationship between oligomerization and the catalytic activity of lipin 1
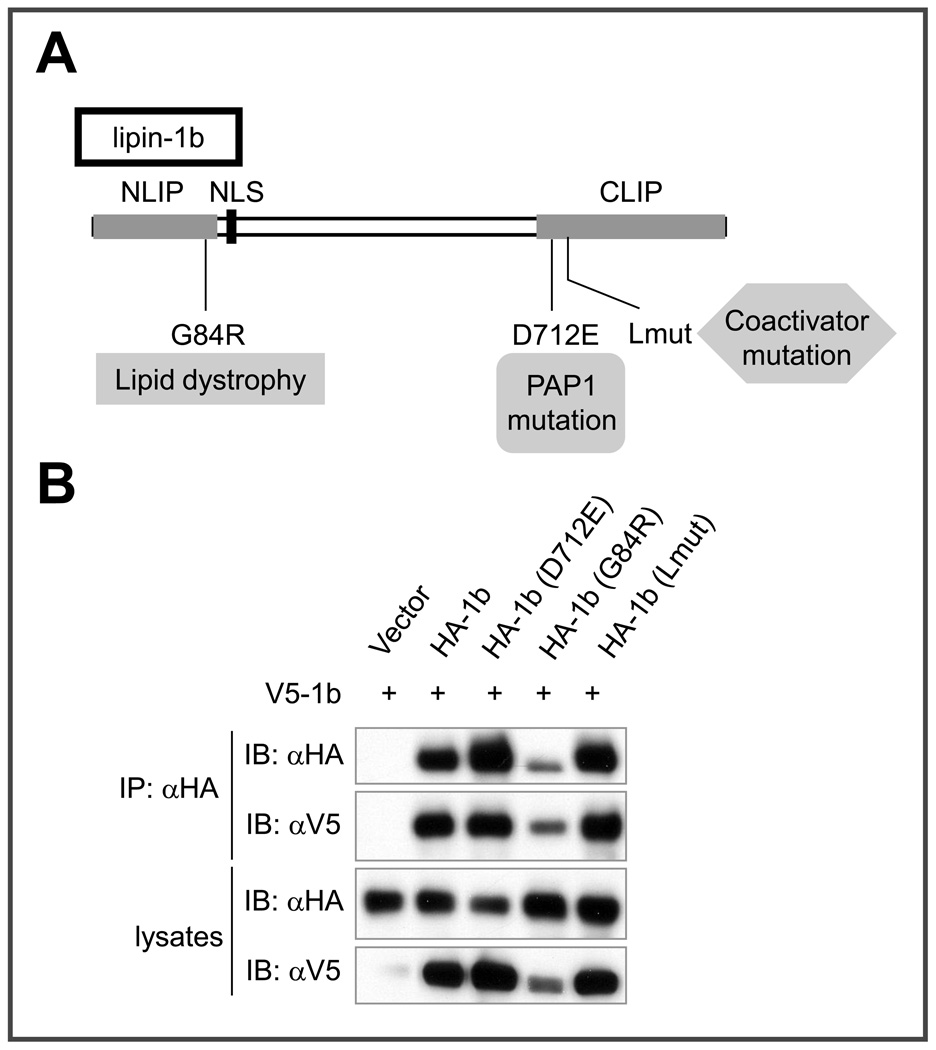
Clearly lipin 1 variants with altered self association would be valuable reagents to probe the regulation and biological importance of this interaction. We therefore examined the self association of a series of previously described lipin 1 mutants with defects in enzymatic activity of phosphorylation. As shown in Figure 8, lipin 1b (G84R), a natural mutation that results in mouse fatty liver dystrophy phenotype and reduces lipin enzymatic activity [2,3], lipin 1b (D712E), a catalytic site mutation that eliminates PAP activity [3], and the coactivator-deficient mutant lipin 1b (Lmut) [14], retained the ability to associate with V5-lipin 1b. Similar results were obtained with other single (or double)-site (s) lipin 1 mutants we generated (e.g., sumoylation-deficient lipin 1 mutant [30], lipin phosphorylation site mutants, and others), which are predicted to be deficient in various post-translational modifications (data not shown). Together with the data shown in Figures 1F and 2, these findings suggest that lipin 1 oligomers are stable and difficult to disrupt by single (or double) amino acid mutations. This could result from a “double security” mechanism for oligomerization that utilizes both amino- and carboxy-terminal interactions. Our results also indicate that the loss of function phenotypes associated with the synthetic and naturally occurring lipin 1 mutations noted above do not result from defects in self association.
Figure 8. Co-immunoprecipitation between lipin 1b and various lipin 1b mutants.
(A) Schematic representation of the functional effects of the lipin 1b point mutant constructs used in co-IP experiments. (B) COS-7 cells were co-transfected with V5-lipin 1b and various HA-tagged lipin 1b mutants as indicated. 36 hours after transfection, the lysates were subjected to immunoprecipitation with anti-HA antibody, followed by immunoblotting with anti-HA or anti-V5 antibody.
We next determined how the lipin 1 monomers function as PAP enzymes in the context of oligomers. Since our findings suggest lipin 1 self associates in a ‘head-to-head/tail-to-tail’ orientation, we postulated that the PAP activity might require two functional catalytic sites in physical proximity to function. To test this we co-expressed catalytically-deficient HA-tagged lipin 1b (D712E) with myc-tagged lipin 1b in HEK293T cells. The PAP activity in lysates of myc-tagged lipin 1b transfected cells was unaffected by increasing amounts of the HA-tagged D712E lipin 1b mutant (data not shown), demonstrating that PAP-null lipin 1 does not act in a dominant manner to inhibit the PAP activity of its oligomeric partner(s). We also verified that the measured PAP activity came from HA-lipin1b-D712E:myc-lipin 1b oligomers by immunoprecipitation (Figure 9A). Although tetrameric combinations preclude a simple 1:1 stoichiometry within each lipin 1 oligomer at equimolar amounts, the selective use of epitope tags suggests that there will be at least one mutant or wild-type lipin 1b within each immunoprecipitated oligomer. We speculated that if a catalytically inactive monomer of lipin 1b negatively interferes with the PAP activity of its wild-type lipin 1b oligomeric partner, then HA-lipin 1 (D712E) immune complexes would not contain detectable PAP activity. In fact, we detected a linear increase in the PAP activity contained in HA-lipin 1(D712E) immune complexes with the increasing amount of transfected wild-type lipin 1b (Figure 9B). As a control, immunoprecipitated HA-lipin 1b (D712E) itself showed no detectable PAP activity and anti-HA antibodies did not immunoprecipitate catalytically active myc-tagged lipin 1b (Figure 9B). These observations suggest that lipin 1 monomers function independently in catalyzing dephosphorylation of PA (Figure 9C).
Figure 9. Individual lipin 1b monomers in oligomers dephosphorylate PA independently.
(A and B) HEK-293T cells were transfected with 0.5 µg of plasmids for expression of myc-lipin 1b (+), and either 0.5 µg (+), or 0.25, 0.5 or 1.0 µg of plasmids for expression of HA-lipin 1b (D712E). The cells were lysed and after immunoprecipitating HA-lipin 1b (D712E) the immunoprecipitate was split into separate fractions, with one fraction used for Western blotting (A) and the other fraction divided into multiple aliquots and measured for PAP activity in the presence and absence of Mg2+ (B). (C) Simplified model depicting the putative dimeric arrangement and activity of individual lipin 1b monomers in a lipin 1b dimer. Black: catalytically inactive monomer; Gray: catalytically active monomer.
DISCUSSION
We demonstrate that lipin proteins self-associate and appear to exist predominately as stable homo- and hetero- dimers/ tretramers in vitro and in vivo. Our evidence for this includes observations from co-immunoprecipitation studies in which both overexpressed and endogenous lipin 1 robustly interacts with itself and other lipin isoforms. Gel filtration and native protein electrophoresis suggests that both native and recombinant lipin 1 exists as a dimer and tetramer with little if any monomeric protein present. The three lipin isoforms are extensively co-localized in transfected cells, and the spatial distance between lipin monomers is close enough to allow for efficient fluorescence resonance energy transfer. Mutational studies combined with co-immunoprecipitation and in vitro GST-pulldown experiments support a model in which lipin monomers self associate in a “head-to-head/tail-to-tail” orientation. Catalytically-inactive lipin 1 (D712E) mutant was able to efficiently pulldown PAP activity when co-expressed with wild-type lipin 1 in cells, indicating that the active sites of individual lipin 1 monomers operate independently in the oligomer.
Our observations raise the interesting possibility that oligomerization plays an important role in lipin 1 function. It is conceivable that the lipin 1 oligomers may be more stable or trafficked differentially to the appropriate cellular locale. Alternatively, oligomerization could directly affect some of the functions of lipin 1. For example, oligomerization could facilitate the ability of lipin 1 to function as an adapter protein to regulate gene transcription. Lipin 1 is predicted to simultaneously bind PPARα, PGC-1α, and p300 [14]. In this model, it is feasible that the dimerization/tretramerization of lipin 1 allows it to interact with each component of the transcriptional machinery to exert its “scaffold” functions. The observation that lipin 1, lipin 2 and lipin 3 all can interact with PPARα ([14] and our unpublished data)) suggests a possibility that they could be recruited to the gene promoter and activate transcription in a hetero-oligomer form, perhaps in specific physiological contexts.
Our results also indicate that the catalytic activity of lipin 1 oligomerization is not necessary for its oligomerization. This conclusion is supported by several observations, including; that lipin 1 oligomerization does not require Mg2+; NEM, a thiol-targeting agent that abrogates the PAP activity of lipin 1, has no effect on oligomerization; enzymatically inactive lipin 1 mutants can still form oligomers; and wild-type lipin 1 and enzymatically inactive lipin 1 function independently wildtype lipin 1 in oligomers.
In lower organisms, such as yeast and worms, there is only one lipin isoform [2]. The present study suggests that in higher organisms that express more than one lipin isoform hetero-oligomerization could produce lipin complexes with unique functions. For example, mammalian lipin 2 and 3 show a lower Vmax for catalytic activity compared to lipin 1 [16]. Given that each lipin monomer functions independently as a phosphatase within the oligomeric complex, hetero-oligomers containing lipin 1 would be predicted to have a higher PAP activity than lipin 2 or 3 homo-oligomers. In liver, lipin 2 is expressed at much higher levels than lipin 1 [18]. However, lipin 1 is robustly induced by stimuli that increase hepatic PAP activity (e.g., fasting and glucocorticoids; [14]), which likely serves to increase lipin 1 content or proportion in hetero-oligomers. In addition, lipin 2 demonstrates a different intracellular localization when compared to lipin 1, and may be enriched in the nuclear envelope [31], where the lipin phosphatase Dullard/NET56 is localized [32,33]. Thus, hetero-oligomerization between lipin 2 and lipin 1 in the tissues where they are both expressed (such as liver and adipocytes) could allow a lipin 2-mediated recruitment of lipin 1 onto nuclear envelope in order to obtain a high level of PAP activity at the nuclear periphery. As expression of the lipin isoforms approach parity, it seems reasonable to conclude there will be a great likelihood of forming heterodimers with resulting changes in specific activity as well as subcellular localization.
Homo-oligomerization may be related to substrate-induced lipin 1 activation. Allosterically activated enzymes are frequently multisubunit protein complexes. For example, acyl-coenzyme A:cholesterol acyltransferase 1 (ACAT1) is thought to have separate dimerization domains in the amino- and carboxy-termini, similar to what we have observed with lipin 1 [34]. In a mixed micelle assay system, purified ACAT1 exhibits allosteric activation by its substrate, cholesterol [35]. Purified Pah1p, the yeast lipin homolog, shows positive cooperative kinetics in its activation by its substrate PA, indicative of two PA binding sites and suggestive of allosteric activation [36,37], although it remains to be demonstrated that Pah1p forms oligomers. Mammalian lipin 1 also shows positive cooperative kinetics with PA, although it is unclear whether this is due to cooperative interaction between lipin and its substrate or allosteric activation of lipin enzymatic activity by the substrate [6,16]. Interestingly, the inhibition of rat lipin 1 by acyl-CoAs and fatty acids has been described as occurring through a negative allosteric interaction [38]. Whereas the lipin 1 monomers function independently in dephosphorylating PA within the oligomeric complex, it is possible that full lipin catalytic activity, particularly with regards to substrate cooperativity, requires oligomerization. To definitively determine whether lipin 1 oligomerization regulates allosteric activation by substrate will require identification of lipin mutants that fail to oligomerize and/or the isolation and characterization of lipin monomers. Unfortunately our efforts to generate such mutants and to isolate monomeric lipin 1 have been unsuccessful, so further work will be required to test these interesting possibilities.
The last two enzymatic reactions in triacylglycerol synthesis are lipin-mediated dephosphorylation of phosphatidic acid to generate diacylglycerol, followed by the acylation of diacylglycerol to form triacylglycerol (reviewed in [39]). There are two diacylglycerol acyltransferases, DGAT1 and DGAT2. DGAT1 is structurally similar to ACAT1 while DGAT2 is unrelated [40]. Interestingly, DGAT1 also forms homo- and hetero-oligomers [41]. Although allosteric activation of DGAT1 by diacylglycerol or acyl-CoAs has not been shown, an amino-terminal fragment of DGAT1 can bind to acyl-CoAs in a cooperative fashion suggestive of allosteric activation [42,43]. Like lipin family members, dimerization of DGAT1 is not required for enzymatic activity, as heterodimerization with a truncated and inactive isoform of DGAT1 did not eliminate catalytic activity of the full-length form [41]. It is curious that the two proteins that catalyze the final two steps in TAG synthesis, lipin 1 and DGAT1, both form homo- and hetero-oligomers. Future questions should address whether oligomerization of these enzymes has any functional role in channeling lipid intermediates towards esterification.
In summary, this study reveals for the first time that lipins can form homo- and hetero-oligomers and suggests that the three mammalian lipin isoforms are tightly linked with each other and serve as an integrated unit.
ACKNOWLEDGMENTS
We are indebted to Dr. Larry Gerace and Dr. Karen Reue for kindly providing reagents used in this study. We also thank Dr. Ammasi Periasamy and Dr. Yuansheng Sun for valuable assistance in acquiring and interpreting FRET and FLIM data.
FUNDING
The work was supported by the grant from the National Basic Research Program of China 2006CB911001 (CC), NIH grants R01-DK078187 (BF), R01-GM50388 and NCRR P20RR021954 (AM). TEH is supported by R01-DK052753. HR is an American Heart Association Post Doctoral Research Fellow.
Abbreviations used
- ACAT1
acyl-coenzyme A:cholesterol acyltransferase 1
- ASBT
acceptor spectral bleed through
- BN-PAGE
Blue Native-PAGE
- C-LIP
carboxy-terminal lipin domain
- C/EBPα
CCAAT/enhancer binding protein α
- DAG
diacylglycerol
- DSBT
donor spectral bleed-through
- DGAT
diacylglycerol acyltransferase
- DMEM
Dulbecco’s Modified Eagle Medium
- ECFP
enhanced cyan fluorescent protein
- EYFP
enhanced yellow fluorescent protein
- FLIM
Fluorescence Lifetime Imaging Microscopy
- FRET
Forster Resonance Imaging Transfer
- GAPDH
glyceraldehyde-3-phosphate dehydrogenase
- LPP
lipid phosphate phosphatases
- N-LIP
amino-terminal lipin domain
- NEM
N-ethylmaleimide
- PA
phosphatidic acid
- PAP
Mg2+-dependent phosphatidic acid phosphatase
- PFRET
Processed FRET
- PGC-1α
PPARγ-coactivator 1α
- PPARα
peroxisome proliferator-activated receptor α
REFERENCES
- 1.Han GS, Wu WI, Carman GM. The Saccharomyces cerevisiae Lipin homolog is a Mg2+-dependent phosphatidate phosphatase enzyme. J. Biol. Chem. 2006;281:9210–9218. doi: 10.1074/jbc.M600425200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Peterfy M, Phan J, Xu P, Reue K. Lipodystrophy in the fld mouse results from mutation of a new gene encoding a nuclear protein, lipin. Nat. Genet. 2001;27:121–124. doi: 10.1038/83685. [DOI] [PubMed] [Google Scholar]
- 3.Harris TE, Huffman TA, Chi A, Shabanowitz J, Hunt DF, Kumar A, Lawrence JC., Jr. Insulin controls subcellular localization and multisite phosphorylation of the phosphatidic acid phosphatase, lipin 1. J. Biol. Chem. 2007;282:277–286. doi: 10.1074/jbc.M609537200. [DOI] [PubMed] [Google Scholar]
- 4.Huffman TA, Mothe-Satney I, Lawrence JC., Jr. Insulin-stimulated phosphorylation of lipin mediated by the mammalian target of rapamycin. Proc. Natl. Acad. Sci. U. S. A. 2002;99:1047–1052. doi: 10.1073/pnas.022634399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Peterfy M, Phan J, Reue K. Alternatively spliced lipin isoforms exhibit distinct expression pattern, subcellular localization, and role in adipogenesis. J. Biol. Chem. 2005;280:32883–32889. doi: 10.1074/jbc.M503885200. [DOI] [PubMed] [Google Scholar]
- 6.Han GS, Carman GM. Characterization of the human LPIN1-encoded phosphatidate phosphatase isoforms. J. Biol. Chem. 2010;285:14628–14638. doi: 10.1074/jbc.M110.117747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Reue K, Xu P, Wang XP, Slavin BG. Adipose tissue deficiency, glucose intolerance, and increased atherosclerosis result from mutation in the mouse fatty liver dystrophy (fld) gene. J. Lipid Res. 2000;41:1067–1076. [PubMed] [Google Scholar]
- 8.Phan J, Reue K. Lipin, a lipodystrophy and obesity gene. Cell Metab. 2005;1:73–83. doi: 10.1016/j.cmet.2004.12.002. [DOI] [PubMed] [Google Scholar]
- 9.Ferguson PJ, Chen S, Tayeh MK, Ochoa L, Leal SM, Pelet A, Munnich A, Lyonnet S, Majeed HA, El Shanti H. Homozygous mutations in LPIN2 are responsible for the syndrome of chronic recurrent multifocal osteomyelitis and congenital dyserythropoietic anaemia (Majeed syndrome) J. Med. Genet. 2005;42:551–557. doi: 10.1136/jmg.2005.030759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Donkor J, Zhang P, Wong S, O'Loughlin L, Dewald J, Kok BP, Brindley DN, Reue K. A conserved serine residue is required for the phosphatidate phosphatase activity but not transcriptional coactivator functions of lipin-1 and lipin-2. J. Biol. Chem. 2009 doi: 10.1074/jbc.M109.023663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Peterfy M, Harris TE, Fujita N, Reue K. Insulin-stimulated interaction with 14-3-3 promotes cytoplasmic localization of lipin-1 in adipocytes. J. Biol. Chem. 2010;285:3857–3864. doi: 10.1074/jbc.M109.072488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Brindley DN, Pilquil C, Sariahmetoglu M, Reue K. Phosphatidate degradation: phosphatidate phosphatases (lipins) and lipid phosphate phosphatases. Biochim. Biophys. Acta. 2009;1791:956–961. doi: 10.1016/j.bbalip.2009.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Santos-Rosa H, Leung J, Grimsey N, Peak-Chew S, Siniossoglou S. The yeast lipin Smp2 couples phospholipid biosynthesis to nuclear membrane growth. EMBO J. 2005;24:1931–1941. doi: 10.1038/sj.emboj.7600672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Finck BN, Gropler MC, Chen Z, Leone TC, Croce MA, Harris TE, Lawrence JC, Jr., Kelly DP. Lipin 1 is an inducible amplifier of the hepatic PGC-1alpha/PPARalpha regulatory pathway. Cell Metab. 2006;4:199–210. doi: 10.1016/j.cmet.2006.08.005. [DOI] [PubMed] [Google Scholar]
- 15.Kim HB, Kumar A, Wang L, Liu GH, Keller SR, Lawrence JC, Jr., Finck BN, Harris TE. Lipin 1 represses NFATc4 transcriptional activity in adipocytes to inhibit secretion of inflammatory factors. Mol. Cell Biol. 2010;30:3126–3139. doi: 10.1128/MCB.01671-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Donkor J, Sariahmetoglu M, Dewald J, Brindley DN, Reue K. Three mammalian lipins act as phosphatidate phosphatases with distinct tissue expression patterns. J. Biol. Chem. 2007;282:3450–3457. doi: 10.1074/jbc.M610745200. [DOI] [PubMed] [Google Scholar]
- 17.Liu GH, Guan T, Datta K, Coppinger J, Yates J, III, Gerace L. Regulation of Myoblast Differentiation by the Nuclear Envelope Protein NET39. Mol. Cell Biol. 2009 doi: 10.1128/MCB.00684-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Gropler MC, Harris TE, Hall AM, Wolins NE, Gross RW, Han X, Chen Z, Finck BN. Lipin 2 is a liver-enriched phosphatidate phosphohydrolase enzyme that is dynamically regulated by fasting and obesity in mice. J. Biol. Chem. 2009;284:6763–6772. doi: 10.1074/jbc.M807882200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Liu GH, Qu J, Shen X. Thioredoxin-mediated negative autoregulation of peroxisome proliferator-activated receptor alpha transcriptional activity. Mol. Biol. Cell. 2006;17:1822–1833. doi: 10.1091/mbc.E05-10-0979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Qu J, Liu GH, Wu K, Han P, Wang P, Li J, Zhang X, Chen C. Nitric oxide destabilizes Pias3 and regulates sumoylation. PLoS. One. 2007;2:e1085. doi: 10.1371/journal.pone.0001085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Chen Y, Mauldin JP, Day RN, Periasamy A. Characterization of spectral FRET imaging microscopy for monitoring nuclear protein interactions. J. Microsc. 2007;228:139–152. doi: 10.1111/j.1365-2818.2007.01838.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Chen Y, Periasamy A. Intensity range based quantitative FRET data analysis to localize protein molecules in live cell nuclei. J. Fluoresc. 2006;16:95–104. doi: 10.1007/s10895-005-0024-1. [DOI] [PubMed] [Google Scholar]
- 23.Day RN, Booker CF, Periasamy A. Characterization of an improved donor fluorescent protein for Forster resonance energy transfer microscopy. J. Biomed. Opt. 2008;13:031203. doi: 10.1117/1.2939094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Liu GH, Qu J, Shen X. NF-kappaB/p65 antagonizes Nrf2-ARE pathway by depriving CBP from Nrf2 and facilitating recruitment of HDAC3 to MafK. Biochim. Biophys. Acta. 2008;1783:713–727. doi: 10.1016/j.bbamcr.2008.01.002. [DOI] [PubMed] [Google Scholar]
- 25.Long JS, Pyne NJ, Pyne S. Lipid phosphate phosphatases form homo-and hetero-oligomers: catalytic competency, subcellular distribution and function. Biochem. J. 2008;411:371–377. doi: 10.1042/BJ20071607. [DOI] [PubMed] [Google Scholar]
- 26.Butterwith SC, Hopewell R, Brindley DN. Partial purification and characterization of the soluble phosphatidate phosphohydrolase of rat liver. Biochem. J. 1984;220:825–833. doi: 10.1042/bj2200825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Cascales C, Mangiapane EH, Brindley DN. Oleic acid promotes the activation and translocation of phosphatidate phosphohydrolase from the cytosol to particulate fractions of isolated rat hepatocytes. Biochem. J. 1984;219:911–916. doi: 10.1042/bj2190911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wright LP, Philips MR. Thematic review series: lipid posttranslational modifications. CAAX modification and membrane targeting of Ras. J. Lipid Res. 2006;47:883–891. doi: 10.1194/jlr.R600004-JLR200. [DOI] [PubMed] [Google Scholar]
- 29.Day CP, Yeaman SJ. Physical evidence for the presence of two forms of phosphatidate phosphohydrolase in rat liver. Biochim. Biophys. Acta. 1992;1127:87–94. doi: 10.1016/0005-2760(92)90205-a. [DOI] [PubMed] [Google Scholar]
- 30.Liu GH, Gerace L. Sumoylation regulates nuclear localization of lipin-1alpha in neuronal cells. PLoS. One. 2009;4:e7031. doi: 10.1371/journal.pone.0007031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Grimsey N, Han GS, O'Hara L, Rochford JJ, Carman GM, Siniossoglou S. Temporal and spatial regulation of the phosphatidate phosphatases lipin 1 and 2. J. Biol. Chem. 2008;283:29166–29174. doi: 10.1074/jbc.M804278200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Kim Y, Gentry MS, Harris TE, Wiley SE, Lawrence JC, Jr., Dixon JE. A conserved phosphatase cascade that regulates nuclear membrane biogenesis. Proc. Natl. Acad. Sci. U. S. A. 2007;104:6596–6601. doi: 10.1073/pnas.0702099104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Schirmer EC, Florens L, Guan T, Yates JR, III, Gerace L. Nuclear membrane proteins with potential disease links found by subtractive proteomics. Science. 2003;301:1380–1382. doi: 10.1126/science.1088176. [DOI] [PubMed] [Google Scholar]
- 34.Yu C, Zhang Y, Lu X, Chen J, Chang CC, Chang TY. Role of the N-terminal hydrophilic domain of acyl-coenzyme A:cholesterol acyltransferase 1 on the enzyme's quaternary structure and catalytic efficiency. Biochemistry. 2002;41:3762–3769. doi: 10.1021/bi0120188. [DOI] [PubMed] [Google Scholar]
- 35.Chang CC, Lee CY, Chang ET, Cruz JC, Levesque MC, Chang TY. Recombinant acyl-CoA:cholesterol acyltransferase-1 (ACAT-1) purified to essential homogeneity utilizes cholesterol in mixed micelles or in vesicles in a highly cooperative manner. J. Biol. Chem. 1998;273:35132–35141. doi: 10.1074/jbc.273.52.35132. [DOI] [PubMed] [Google Scholar]
- 36.Wu WI, Lin YP, Wang E, Merrill AH, Jr., Carman GM. Regulation of phosphatidate phosphatase activity from the yeast Saccharomyces cerevisiae by sphingoid bases. J. Biol. Chem. 1993;268:13830–13837. [PubMed] [Google Scholar]
- 37.Wu WI, Carman GM. Regulation of phosphatidate phosphatase activity from the yeast Saccharomyces cerevisiae by phospholipids. Biochemistry. 1996;35:3790–3796. doi: 10.1021/bi952808f. [DOI] [PubMed] [Google Scholar]
- 38.Elabbadi N, Day CP, Gamouh A, Zyad A, Yeaman SJ. Relationship between the inhibition of phosphatidic acid phosphohydrolase-1 by oleate and oleoyl-CoA ester and its apparent translocation. Biochimie. 2005;87:437–443. doi: 10.1016/j.biochi.2005.01.003. [DOI] [PubMed] [Google Scholar]
- 39.Coleman RA, Lee DP. Enzymes of triacylglycerol synthesis and their regulation. Prog. Lipid Res. 2004;43:134–176. doi: 10.1016/s0163-7827(03)00051-1. [DOI] [PubMed] [Google Scholar]
- 40.Cases S, Smith SJ, Zheng YW, Myers HM, Lear SR, Sande E, Novak S, Collins C, Welch CB, Lusis AJ, Erickson SK, Farese RV., Jr. Identification of a gene encoding an acyl CoA:diacylglycerol acyltransferase, a key enzyme in triacylglycerol synthesis. Proc. Natl. Acad. Sci. U. S. A. 1998;95:13018–13023. doi: 10.1073/pnas.95.22.13018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Cheng D, Meegalla RL, He B, Cromley DA, Billheimer JT, Young PR. Human acyl-CoA:diacylglycerol acyltransferase is a tetrameric protein. Biochem. J. 2001;359:707–714. doi: 10.1042/0264-6021:3590707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Siloto RM, Madhavji M, Wiehler WB, Burton TL, Boora PS, Laroche A, Weselake RJ. An N-terminal fragment of mouse DGAT1 binds different acyl-CoAs with varying affinity. Biochem. Biophys. Res. Commun. 2008;373:350–354. doi: 10.1016/j.bbrc.2008.06.031. [DOI] [PubMed] [Google Scholar]
- 43.Weselake RJ, Madhavji M, Szarka SJ, Patterson NA, Wiehler WB, Nykiforuk CL, Burton TL, Boora PS, Mosimann SC, Foroud NA, Thibault BJ, Moloney MM, Laroche A, Furukawa-Stoffer TL. Acyl-CoA-binding and self-associating properties of a recombinant 13.3 kDa N-terminal fragment of diacylglycerol acyltransferase-1 from oilseed rape. BMC. Biochem. 2006;7:24. doi: 10.1186/1471-2091-7-24. [DOI] [PMC free article] [PubMed] [Google Scholar]