Abstract
In the title compound, C22H20N+·Cl−, the anthracene system makes a dihedral angle of 72.65 (4)° with the benzene ring. The C—N—C—C torsion angles in the chain connecting the benzene ring and anthracene system are 52.24 (15) and −170.73 (11)°. The crystal structure is stabilized by intermolecular N—H⋯Cl and C—H⋯Cl hydrogen bonds, which link the molecules into tetramers about inversion centers.
Related literature
For the synthesis and structures of related compounds, see: Ashton et al. (1997 ▶). For formation of rotaxanes from sec-ammonium salts and crown ethers, see: Nakazono et al. (2008 ▶).
Experimental
Crystal data
C22H20N+·Cl−
M r = 333.84
Triclinic,

a = 6.7457 (13) Å
b = 10.761 (2) Å
c = 13.033 (3) Å
α = 94.45 (3)°
β = 104.84 (3)°
γ = 104.48 (3)°
V = 875.3 (3) Å3
Z = 2
Mo Kα radiation
μ = 0.22 mm−1
T = 113 K
0.28 × 0.24 × 0.20 mm
Data collection
Rigaku Saturn CCD area-detector diffractometer
Absorption correction: multi-scan (CrystalClear; Rigaku/MSC, 2005 ▶) T min = 0.941, T max = 0.957
5882 measured reflections
3048 independent reflections
2342 reflections with I > 2σ(I)
R int = 0.027
Refinement
R[F 2 > 2σ(F 2)] = 0.031
wR(F 2) = 0.088
S = 1.04
3048 reflections
226 parameters
3 restraints
H atoms treated by a mixture of independent and constrained refinement
Δρmax = 0.20 e Å−3
Δρmin = −0.21 e Å−3
Data collection: CrystalClear (Rigaku/MSC, 2005 ▶); cell refinement: CrystalClear; data reduction: CrystalClear; program(s) used to solve structure: SHELXS97 (Sheldrick, 2008 ▶); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008 ▶); molecular graphics: SHELXTL (Sheldrick, 2008 ▶); software used to prepare material for publication: SHELXTL.
Supplementary Material
Crystal structure: contains datablocks I, global. DOI: 10.1107/S1600536811016837/pv2409sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S1600536811016837/pv2409Isup2.hkl
Supplementary material file. DOI: 10.1107/S1600536811016837/pv2409Isup3.cml
Additional supplementary materials: crystallographic information; 3D view; checkCIF report
Table 1. Hydrogen-bond geometry (Å, °).
| D—H⋯A | D—H | H⋯A | D⋯A | D—H⋯A |
|---|---|---|---|---|
| N1—H1A⋯Cl1i | 0.92 (1) | 2.26 (1) | 3.0963 (13) | 152 (1) |
| N1—H1B⋯Cl1 | 0.92 (1) | 2.17 (1) | 3.0781 (16) | 170 (1) |
| C16—H16A⋯Cl1ii | 0.97 | 2.60 | 3.4824 (16) | 151 |
Symmetry codes: (i)  ; (ii)
; (ii)  .
.
Acknowledgments
We are grateful for financial support from the Hebei Natural Science Fundation (No. B2009000670) and the Foundation of the Education Department of Hebei Province (grant No. 2009117), People’s Republic of China.
supplementary crystallographic information
Comment
Sec-ammonium salts and crown ethers combine well to yield a stable pseudorotaxanes as a precursor of rotaxanes (Nakazono et al., 2008). (9-Anthracenyl) benzylammonium hexafluorophosphate and aromatic crown ethers give hydrogen-bonded complexes pseudorotaxane-like geometries (Ashton et al., 1997). In this paper we report the synthesis and crystal strucure of (9-anthracenyl) benzyl ammonium chloride.
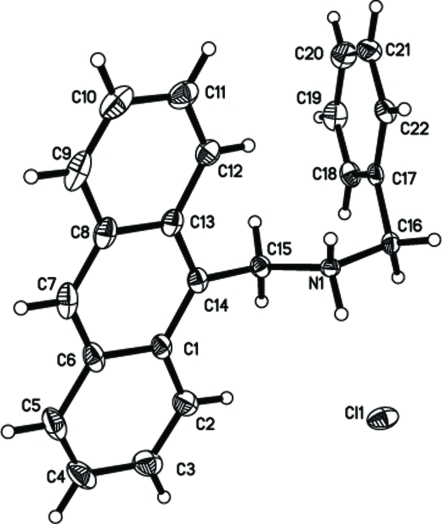
In the title compound (Fig. 1), anthracene ring makes a dihedral angle of 72.65 (4)° with benzene ring. The torsion angles in the chain connecting the benzene and the anthracene rings, (C15/N1—C16/C17 and (C16/N1—C15/C14) are 52.24 (15) ° and -170.73 (11)°, respectively. In the crystal structure, the crystal packing is stabilized by intermolecular N1—H1A···Cl1 and C16—H16A···Cl1 hydrogen bonds which link the molecules into tetramers about inversion centers (Table 1 and Fig. 2).
Experimental
A mixture of 9-anthracenealdehyde (6.18 g, 30 mmol) and benzylamine (3.86 g, 36 mmol) and molecular Sieve in toluene (200 mL) was heated under reflux with stirring in a water divider for 10 h. After the reaction mixture had cooled down to room temperature, the solvent was removed in vacuo to give the imine. The solid was dissolved in hot MeOH (150 mL), followed by drop-wise addition of NaBH4 (5.70 g, 150 mmol) and heating under reflux with stirring for 8 h. The reaction mixture was then allowed to cool down to room temperature, and concentrated HCl was added (pH<2). After evaporation of the solvent, the residue was suspended in H2O (70 mL) and extracted with CH2Cl2 (4 × 50 mL). The combined extracts were washed with 5% aqueous NaHCO3 (2× 70 mL) and H2O (70 mL) and then dried (MgSO4). Removal of the solvent in vacuo afforded the (9-anthracenyl)benzyl amine which was treated according to literature (Ashton et al., 1997) to prepare the title compound. Pale yellow single crystals of the title compound suitable for X-ray analysis were obtained by slow evaporation of an methnol solution.
Refinement
The H atoms were included at calculated positions with C—H = 0.93 and 0.97 Å for aryl and methylene type H-atoms, respectively, and refined in a riding mode with Uiso(H) = 1.2Ueq(C). The amino H-atoms were located from a difference Fourier map and were allowed to refine freely.
Figures
Fig. 1.
The molecular structure of the title complex, with 50% probablity ellipsoids.
Fig. 2.
Unit cell packing of the title complex, showing hydrogen bonded tetramers.
Crystal data
| C22H20N+·Cl− | Z = 2 |
| Mr = 333.84 | F(000) = 352 |
| Triclinic, P1 | Dx = 1.267 Mg m−3 |
| Hall symbol: -P 1 | Mo Kα radiation, λ = 0.71073 Å |
| a = 6.7457 (13) Å | Cell parameters from 2756 reflections |
| b = 10.761 (2) Å | θ = 2.4–27.9° |
| c = 13.033 (3) Å | µ = 0.22 mm−1 |
| α = 94.45 (3)° | T = 113 K |
| β = 104.84 (3)° | Block, yellow |
| γ = 104.48 (3)° | 0.28 × 0.24 × 0.20 mm |
| V = 875.3 (3) Å3 |
Data collection
| Rigaku Saturn CCD area-detector diffractometer | 3048 independent reflections |
| Radiation source: rotating anode | 2342 reflections with I > 2σ(I) |
| confocal | Rint = 0.027 |
| Detector resolution: 7.31 pixels mm-1 | θmax = 25.0°, θmin = 2.4° |
| ω and φ scans | h = −8→8 |
| Absorption correction: multi-scan (CrystalClear; Rigaku/MSC, 2005) | k = −12→10 |
| Tmin = 0.941, Tmax = 0.957 | l = −15→15 |
| 5882 measured reflections |
Refinement
| Refinement on F2 | Secondary atom site location: difference Fourier map |
| Least-squares matrix: full | Hydrogen site location: inferred from neighbouring sites |
| R[F2 > 2σ(F2)] = 0.031 | H atoms treated by a mixture of independent and constrained refinement |
| wR(F2) = 0.088 | w = 1/[σ2(Fo2) + (0.0502P)2] where P = (Fo2 + 2Fc2)/3 |
| S = 1.04 | (Δ/σ)max = 0.002 |
| 3048 reflections | Δρmax = 0.20 e Å−3 |
| 226 parameters | Δρmin = −0.21 e Å−3 |
| 3 restraints | Extinction correction: SHELXL97 (Sheldrick, 2008), Fc*=kFc[1+0.001xFc2λ3/sin(2θ)]-1/4 |
| Primary atom site location: structure-invariant direct methods | Extinction coefficient: 0.152 (9) |
Special details
| Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
| Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2, conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2 are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
| x | y | z | Uiso*/Ueq | ||
| Cl1 | 0.17351 (5) | 0.42040 (4) | 0.40032 (3) | 0.02288 (15) | |
| N1 | 0.38238 (18) | 0.52936 (11) | 0.64009 (9) | 0.0139 (3) | |
| C1 | 0.1631 (2) | 0.21072 (14) | 0.67206 (11) | 0.0184 (3) | |
| C2 | −0.0472 (2) | 0.21057 (16) | 0.61273 (12) | 0.0242 (4) | |
| H2 | −0.0774 | 0.2890 | 0.6018 | 0.029* | |
| C3 | −0.2034 (3) | 0.09727 (16) | 0.57217 (13) | 0.0305 (4) | |
| H3 | −0.3391 | 0.0995 | 0.5340 | 0.037* | |
| C4 | −0.1629 (3) | −0.02440 (17) | 0.58713 (13) | 0.0341 (4) | |
| H4 | −0.2721 | −0.1009 | 0.5597 | 0.041* | |
| C5 | 0.0344 (3) | −0.02903 (16) | 0.64127 (12) | 0.0305 (4) | |
| H5 | 0.0595 | −0.1091 | 0.6505 | 0.037* | |
| C6 | 0.2045 (3) | 0.08678 (15) | 0.68441 (12) | 0.0227 (4) | |
| C7 | 0.4101 (3) | 0.08322 (16) | 0.73768 (12) | 0.0262 (4) | |
| H7 | 0.4365 | 0.0029 | 0.7438 | 0.031* | |
| C8 | 0.5764 (2) | 0.19449 (15) | 0.78173 (11) | 0.0223 (4) | |
| C9 | 0.7883 (3) | 0.18991 (18) | 0.83401 (12) | 0.0299 (4) | |
| H9 | 0.8163 | 0.1098 | 0.8382 | 0.036* | |
| C10 | 0.9485 (3) | 0.29980 (18) | 0.87736 (12) | 0.0315 (4) | |
| H10 | 1.0853 | 0.2948 | 0.9103 | 0.038* | |
| C11 | 0.9084 (2) | 0.42185 (17) | 0.87262 (11) | 0.0273 (4) | |
| H11 | 1.0193 | 0.4969 | 0.9030 | 0.033* | |
| C12 | 0.7099 (2) | 0.43137 (16) | 0.82427 (11) | 0.0221 (4) | |
| H12 | 0.6871 | 0.5130 | 0.8231 | 0.026* | |
| C13 | 0.5357 (2) | 0.31906 (15) | 0.77525 (11) | 0.0188 (3) | |
| C14 | 0.3287 (2) | 0.32511 (14) | 0.72005 (11) | 0.0170 (3) | |
| C15 | 0.2884 (2) | 0.45631 (14) | 0.71774 (11) | 0.0167 (3) | |
| H15A | 0.1356 | 0.4454 | 0.6977 | 0.020* | |
| H15B | 0.3499 | 0.5065 | 0.7891 | 0.020* | |
| C16 | 0.3787 (2) | 0.66897 (14) | 0.64641 (11) | 0.0159 (3) | |
| H16A | 0.2322 | 0.6725 | 0.6209 | 0.019* | |
| H16B | 0.4571 | 0.7116 | 0.6000 | 0.019* | |
| C17 | 0.4765 (2) | 0.74042 (14) | 0.75999 (11) | 0.0160 (3) | |
| C18 | 0.3455 (2) | 0.75556 (14) | 0.82421 (12) | 0.0214 (4) | |
| H18 | 0.1978 | 0.7247 | 0.7963 | 0.026* | |
| C19 | 0.4352 (3) | 0.81680 (15) | 0.93001 (12) | 0.0277 (4) | |
| H19 | 0.3472 | 0.8261 | 0.9729 | 0.033* | |
| C20 | 0.6531 (3) | 0.86365 (16) | 0.97157 (13) | 0.0300 (4) | |
| H20 | 0.7122 | 0.9046 | 1.0424 | 0.036* | |
| C21 | 0.7851 (3) | 0.84988 (15) | 0.90801 (12) | 0.0254 (4) | |
| H21 | 0.9326 | 0.8817 | 0.9360 | 0.030* | |
| C22 | 0.6962 (2) | 0.78841 (14) | 0.80265 (11) | 0.0194 (3) | |
| H22 | 0.7848 | 0.7793 | 0.7601 | 0.023* | |
| H1A | 0.5219 (14) | 0.5298 (15) | 0.6487 (11) | 0.027 (4)* | |
| H1B | 0.3065 (17) | 0.4908 (14) | 0.5707 (8) | 0.018 (4)* |
Atomic displacement parameters (Å2)
| U11 | U22 | U33 | U12 | U13 | U23 | |
| Cl1 | 0.0145 (2) | 0.0350 (3) | 0.0170 (2) | 0.00489 (17) | 0.00408 (14) | −0.00094 (16) |
| N1 | 0.0128 (6) | 0.0149 (7) | 0.0127 (6) | 0.0023 (5) | 0.0032 (5) | 0.0016 (5) |
| C1 | 0.0249 (8) | 0.0171 (8) | 0.0153 (7) | 0.0035 (7) | 0.0110 (6) | 0.0039 (6) |
| C2 | 0.0265 (9) | 0.0237 (9) | 0.0219 (8) | 0.0031 (7) | 0.0102 (7) | 0.0024 (7) |
| C3 | 0.0249 (9) | 0.0339 (11) | 0.0270 (9) | −0.0032 (8) | 0.0090 (7) | 0.0034 (8) |
| C4 | 0.0413 (11) | 0.0230 (9) | 0.0302 (9) | −0.0108 (8) | 0.0172 (8) | 0.0002 (8) |
| C5 | 0.0479 (11) | 0.0167 (9) | 0.0285 (9) | 0.0017 (8) | 0.0208 (8) | 0.0045 (7) |
| C6 | 0.0365 (10) | 0.0161 (8) | 0.0194 (8) | 0.0044 (7) | 0.0171 (7) | 0.0039 (7) |
| C7 | 0.0434 (10) | 0.0212 (9) | 0.0259 (8) | 0.0176 (8) | 0.0201 (8) | 0.0109 (7) |
| C8 | 0.0321 (9) | 0.0257 (9) | 0.0171 (7) | 0.0140 (8) | 0.0138 (7) | 0.0085 (7) |
| C9 | 0.0413 (10) | 0.0403 (11) | 0.0247 (9) | 0.0288 (9) | 0.0174 (8) | 0.0174 (8) |
| C10 | 0.0261 (9) | 0.0532 (12) | 0.0221 (8) | 0.0187 (9) | 0.0089 (7) | 0.0148 (8) |
| C11 | 0.0264 (9) | 0.0391 (10) | 0.0164 (8) | 0.0077 (8) | 0.0065 (7) | 0.0068 (7) |
| C12 | 0.0253 (8) | 0.0254 (9) | 0.0164 (7) | 0.0073 (7) | 0.0069 (6) | 0.0048 (7) |
| C13 | 0.0251 (8) | 0.0225 (8) | 0.0138 (7) | 0.0095 (7) | 0.0107 (6) | 0.0056 (6) |
| C14 | 0.0226 (8) | 0.0167 (8) | 0.0157 (7) | 0.0061 (6) | 0.0111 (6) | 0.0039 (6) |
| C15 | 0.0176 (8) | 0.0164 (8) | 0.0173 (7) | 0.0041 (6) | 0.0075 (6) | 0.0033 (6) |
| C16 | 0.0159 (7) | 0.0143 (8) | 0.0189 (7) | 0.0049 (6) | 0.0056 (6) | 0.0059 (6) |
| C17 | 0.0197 (8) | 0.0097 (7) | 0.0197 (7) | 0.0051 (6) | 0.0056 (6) | 0.0053 (6) |
| C18 | 0.0231 (8) | 0.0166 (8) | 0.0277 (8) | 0.0078 (7) | 0.0099 (7) | 0.0057 (7) |
| C19 | 0.0395 (10) | 0.0256 (9) | 0.0255 (8) | 0.0147 (8) | 0.0168 (7) | 0.0038 (7) |
| C20 | 0.0439 (10) | 0.0219 (9) | 0.0211 (8) | 0.0095 (8) | 0.0049 (7) | −0.0015 (7) |
| C21 | 0.0251 (8) | 0.0180 (8) | 0.0269 (8) | 0.0029 (7) | 0.0006 (7) | 0.0008 (7) |
| C22 | 0.0216 (8) | 0.0148 (8) | 0.0224 (8) | 0.0046 (6) | 0.0079 (6) | 0.0031 (6) |
Geometric parameters (Å, °)
| N1—C15 | 1.4990 (17) | C10—H10 | 0.9300 |
| N1—C16 | 1.5051 (18) | C11—C12 | 1.359 (2) |
| N1—H1A | 0.917 (8) | C11—H11 | 0.9300 |
| N1—H1B | 0.922 (8) | C12—C13 | 1.426 (2) |
| C1—C14 | 1.410 (2) | C12—H12 | 0.9300 |
| C1—C2 | 1.431 (2) | C13—C14 | 1.418 (2) |
| C1—C6 | 1.442 (2) | C14—C15 | 1.504 (2) |
| C2—C3 | 1.360 (2) | C15—H15A | 0.9700 |
| C2—H2 | 0.9300 | C15—H15B | 0.9700 |
| C3—C4 | 1.421 (2) | C16—C17 | 1.512 (2) |
| C3—H3 | 0.9300 | C16—H16A | 0.9700 |
| C4—C5 | 1.352 (2) | C16—H16B | 0.9700 |
| C4—H4 | 0.9300 | C17—C22 | 1.386 (2) |
| C5—C6 | 1.425 (2) | C17—C18 | 1.3922 (19) |
| C5—H5 | 0.9300 | C18—C19 | 1.391 (2) |
| C6—C7 | 1.394 (2) | C18—H18 | 0.9300 |
| C7—C8 | 1.383 (2) | C19—C20 | 1.374 (2) |
| C7—H7 | 0.9300 | C19—H19 | 0.9300 |
| C8—C9 | 1.432 (2) | C20—C21 | 1.388 (2) |
| C8—C13 | 1.438 (2) | C20—H20 | 0.9300 |
| C9—C10 | 1.353 (2) | C21—C22 | 1.386 (2) |
| C9—H9 | 0.9300 | C21—H21 | 0.9300 |
| C10—C11 | 1.408 (2) | C22—H22 | 0.9300 |
| C15—N1—C16 | 114.67 (10) | C11—C12—C13 | 121.55 (16) |
| C15—N1—H1A | 112.3 (10) | C11—C12—H12 | 119.2 |
| C16—N1—H1A | 106.5 (10) | C13—C12—H12 | 119.2 |
| C15—N1—H1B | 109.8 (9) | C14—C13—C12 | 123.26 (14) |
| C16—N1—H1B | 106.0 (9) | C14—C13—C8 | 119.31 (14) |
| H1A—N1—H1B | 107.2 (9) | C12—C13—C8 | 117.42 (14) |
| C14—C1—C2 | 123.40 (14) | C1—C14—C13 | 120.77 (14) |
| C14—C1—C6 | 118.93 (14) | C1—C14—C15 | 120.96 (13) |
| C2—C1—C6 | 117.67 (14) | C13—C14—C15 | 118.23 (13) |
| C3—C2—C1 | 120.89 (16) | N1—C15—C14 | 112.01 (10) |
| C3—C2—H2 | 119.6 | N1—C15—H15A | 109.2 |
| C1—C2—H2 | 119.6 | C14—C15—H15A | 109.2 |
| C2—C3—C4 | 121.11 (16) | N1—C15—H15B | 109.2 |
| C2—C3—H3 | 119.4 | C14—C15—H15B | 109.2 |
| C4—C3—H3 | 119.4 | H15A—C15—H15B | 107.9 |
| C5—C4—C3 | 120.06 (16) | N1—C16—C17 | 111.53 (12) |
| C5—C4—H4 | 120.0 | N1—C16—H16A | 109.3 |
| C3—C4—H4 | 120.0 | C17—C16—H16A | 109.3 |
| C4—C5—C6 | 121.12 (16) | N1—C16—H16B | 109.3 |
| C4—C5—H5 | 119.4 | C17—C16—H16B | 109.3 |
| C6—C5—H5 | 119.4 | H16A—C16—H16B | 108.0 |
| C7—C6—C5 | 121.68 (15) | C22—C17—C18 | 119.11 (13) |
| C7—C6—C1 | 119.21 (15) | C22—C17—C16 | 120.99 (12) |
| C5—C6—C1 | 119.11 (15) | C18—C17—C16 | 119.88 (13) |
| C8—C7—C6 | 122.54 (15) | C19—C18—C17 | 120.05 (14) |
| C8—C7—H7 | 118.7 | C19—C18—H18 | 120.0 |
| C6—C7—H7 | 118.7 | C17—C18—H18 | 120.0 |
| C7—C8—C9 | 122.15 (15) | C20—C19—C18 | 120.37 (14) |
| C7—C8—C13 | 119.11 (14) | C20—C19—H19 | 119.8 |
| C9—C8—C13 | 118.74 (15) | C18—C19—H19 | 119.8 |
| C10—C9—C8 | 121.28 (16) | C19—C20—C21 | 120.05 (14) |
| C10—C9—H9 | 119.4 | C19—C20—H20 | 120.0 |
| C8—C9—H9 | 119.4 | C21—C20—H20 | 120.0 |
| C9—C10—C11 | 120.10 (15) | C22—C21—C20 | 119.72 (15) |
| C9—C10—H10 | 119.9 | C22—C21—H21 | 120.1 |
| C11—C10—H10 | 119.9 | C20—C21—H21 | 120.1 |
| C12—C11—C10 | 120.87 (16) | C17—C22—C21 | 120.71 (13) |
| C12—C11—H11 | 119.6 | C17—C22—H22 | 119.6 |
| C10—C11—H11 | 119.6 | C21—C22—H22 | 119.6 |
| C14—C1—C2—C3 | −177.07 (13) | C7—C8—C13—C12 | 178.28 (12) |
| C6—C1—C2—C3 | 1.8 (2) | C9—C8—C13—C12 | −1.57 (19) |
| C1—C2—C3—C4 | −0.1 (2) | C2—C1—C14—C13 | −178.15 (12) |
| C2—C3—C4—C5 | −0.9 (2) | C6—C1—C14—C13 | 3.01 (19) |
| C3—C4—C5—C6 | 0.2 (2) | C2—C1—C14—C15 | 4.2 (2) |
| C4—C5—C6—C7 | −178.11 (14) | C6—C1—C14—C15 | −174.63 (12) |
| C4—C5—C6—C1 | 1.5 (2) | C12—C13—C14—C1 | 179.28 (12) |
| C14—C1—C6—C7 | −3.89 (19) | C8—C13—C14—C1 | 0.21 (19) |
| C2—C1—C6—C7 | 177.20 (12) | C12—C13—C14—C15 | −3.02 (19) |
| C14—C1—C6—C5 | 176.47 (12) | C8—C13—C14—C15 | 177.92 (12) |
| C2—C1—C6—C5 | −2.44 (19) | C16—N1—C15—C14 | −170.73 (11) |
| C5—C6—C7—C8 | −178.83 (13) | C1—C14—C15—N1 | −106.13 (14) |
| C1—C6—C7—C8 | 1.5 (2) | C13—C14—C15—N1 | 76.17 (15) |
| C6—C7—C8—C9 | −178.44 (13) | C15—N1—C16—C17 | 52.24 (15) |
| C6—C7—C8—C13 | 1.7 (2) | N1—C16—C17—C22 | 81.05 (17) |
| C7—C8—C9—C10 | −179.48 (14) | N1—C16—C17—C18 | −97.18 (15) |
| C13—C8—C9—C10 | 0.4 (2) | C22—C17—C18—C19 | −0.7 (2) |
| C8—C9—C10—C11 | 0.6 (2) | C16—C17—C18—C19 | 177.52 (14) |
| C9—C10—C11—C12 | −0.4 (2) | C17—C18—C19—C20 | 0.6 (2) |
| C10—C11—C12—C13 | −0.9 (2) | C18—C19—C20—C21 | −0.1 (2) |
| C11—C12—C13—C14 | −177.20 (13) | C19—C20—C21—C22 | −0.2 (2) |
| C11—C12—C13—C8 | 1.9 (2) | C18—C17—C22—C21 | 0.5 (2) |
| C7—C8—C13—C14 | −2.6 (2) | C16—C17—C22—C21 | −177.76 (13) |
| C9—C8—C13—C14 | 177.55 (11) | C20—C21—C22—C17 | 0.0 (2) |
Hydrogen-bond geometry (Å, °)
| D—H···A | D—H | H···A | D···A | D—H···A |
| N1—H1A···Cl1i | 0.92 (1) | 2.26 (1) | 3.0963 (13) | 152 (1) |
| N1—H1B···Cl1 | 0.92 (1) | 2.17 (1) | 3.0781 (16) | 170 (1) |
| C16—H16A···Cl1ii | 0.97 | 2.60 | 3.4824 (16) | 151 |
Symmetry codes: (i) −x+1, −y+1, −z+1; (ii) −x, −y+1, −z+1.
Footnotes
Supplementary data and figures for this paper are available from the IUCr electronic archives (Reference: PV2409).
References
- Ashton, P. R., Ballardini, R., Balzani, V., Marcos, G.-L., Lawrence, S. M., Victoria, M.-D., Montalti, M., Piersanti, A., Prodi, L., Stoddart, J. F. & Williams, D. (1997). J. Am. Chem. Soc. 119, 10641–10651.
- Nakazono, K., Kuwata, S. & Takata, T. (2008). Tetrahedron Lett. 49, 2397–2400.
- Rigaku/MSC (2005). CrystalClear Rigaku/MSC Inc., The Woodlands, Texas, USA.
- Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. [DOI] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crystal structure: contains datablocks I, global. DOI: 10.1107/S1600536811016837/pv2409sup1.cif
Structure factors: contains datablocks I. DOI: 10.1107/S1600536811016837/pv2409Isup2.hkl
Supplementary material file. DOI: 10.1107/S1600536811016837/pv2409Isup3.cml
Additional supplementary materials: crystallographic information; 3D view; checkCIF report