Abstract
In the present study, clinical isolates of Bordetella pertussis collected in Poland from 1960 to 2005 were analyzed by pulsed-field gel electrophoresis (PFGE) according to protocols recommended in previous studies. Among the 110 isolates from 1995 to 2005, 59 PFGE patterns were found, most of which were different from those currently circulating in other European Union (EU) countries for which data are available. The PFGE patterns of currently disseminating B. pertussis clones were found within PFGE groups III and IV, as elsewhere in the EU, and in newly identified clusters A and C. Up to 70, 26, and 4%, respectively, of the currently isolated strains in Poland harbored ptxA1-prn1, ptxA1-prn2, and ptxA1-prn3 allele combinations, and most (82%) were found to be of the Fim2 phenotype. Differences in the extent of heterogeneity estimated by PFGE typing in B. pertussis populations circulating in Poland in comparison to other EU countries may be due to the different vaccine composition strategy, since routine pertussis vaccination was initiated in Poland in 1960.
INTRODUCTION
Similarly to other highly immunized communities in Europe, America, Asia, and Australia (9, 10, 20, 27, 35, 44), Poland has seen a resurgence of pertussis since the late 1990s (15). Children in Poland have been continuously immunized since 1960 with three primary doses and one booster dose at 2, 3 to 4, 5, and 16 to 18 months of life with a nationally produced whole-cell diphtheria-tetanus-pertussis (DTP) vaccine. An additional booster dose of acellular pertussis vaccine for 6-year-old children was introduced in 2004 to prevent the waning of immunity, which has been observed since the late 1990s in schoolchildren (39). An increased rates of pertussis was not found to be associated with decreased vaccination coverage, improved diagnostics, or the use of subpotent vaccine lots (15, 16). A previous study showed that B. pertussis strains circulating in Poland after the 1990s, as elsewhere in Europe, presented nonvaccine variants of pertussis toxin and pertactin gene alleles (17). A unique feature of the whole-cell pertussis (wP) component of this vaccine is that the vaccine strain composition has changed over time. An identical formulation of the wP component with respect to vaccine strains has been used, however, for 30 years.
Vaccine-driven evolution in the B. pertussis population measured by genetic and antigenic shifts is still hypothesized (5, 13, 19, 29, 32, 33, 34, 36, 42) since it was first suggested by Khattak et al. (23, 24). Epidemiological surveys of B. pertussis strains on the genome and gene levels have found recent isolates distinct from vaccine strains or isolates circulating before the 1990s in many countries using different typing methods (5, 8, 13, 14, 18, 19, 29, 30, 43). PFGE, multilocus sequence typing, and multilocus variable-number tandem repeat analysis have found some B. pertussis lineages to be more successful at disseminating and/or related to the increase of incidence in some countries (2, 3, 12, 19, 25, 38). Among many available methodologies capable of tracking genetic differences among isolates, PFGE has been the chosen reference method for epidemiological surveys (2, 31), since it is the most discriminatory (4).
A study published by the European research program for strain characterization and surveillance, the EUpertstrain II project, found B. pertussis strains from Poland isolated in 1999 to 2004 to be different from those observed in seven other European countries using the PFGE technique (19). A study on a more representative number of isolates from Poland has been suggested to confirm unique PFGE patterns observed in Poland. In the present study, we describe the B. pertussis strain population isolated in Poland from 1960 to 2005 with respect to vaccine strains used for production of the national DTP vaccine performed with the PFGE procedure according to EUpertstrain guidelines (www.eupertstrain.org/). The isolates were also characterized by serotyping and sequencing of PCR-amplified fragments of pertussis toxin ptxA subunit and pertactin genes. The goal of the study was to establish the extent of differences seen in B. pertussis strains circulating in Poland in order to contribute to the characterization of B. pertussis strains circulating in Europe.
MATERIALS AND METHODS
Collection of strains.
In all, 148 B. pertussis strains, including 139 clinical isolates, 6 historical vaccine strains (A/63, 21/60, 7/60, 25593/65, 1326/62, and 60623/67), and 3 current vaccine strains (606/67, 186/65, and 629/65), were used (Table 1). The reference B. pertussis strains were 18323, Tohama I, Bp134, B902, FR743, FIN12, and FR287, kindly supplied by Norman Fry (Health Protection Agency Centre for Infections, London, United Kingdom) and Reza Advani (Swedish Institute for Infectious Disease Control, Solna, Sweden). A EUpertstrain II culture collection of 140 strains from Finland, the Netherlands, France, Germany, Sweden, Denmark, and the United Kingdom was used as a control B. pertussis strain collection to ensure the level of validity and correctness of positioning isolates into previously described PFGE groups.
Table 1.
B. pertussis strains used for whole-cell pertussis vaccine production in Poland
| Strain, country, isolation yra | Allele combination |
|||
|---|---|---|---|---|
| 1960–1967 | 1967–1971 | 1971–1977 | Since 1978 | |
| 509, NL, <1966 | ptxA3-prn1 | ptxA3-prn1 | ||
| A/63, PL, 1963 | ptxA3-prn1 | ptxA3-prn1 | ptxA3-prn1 | |
| 21/60, PL, 1960 | ptxA1-prn1 | ptxA1-prn1 | ptxA1-prn1 | |
| 5/V, PL | NAb | |||
| 10/60, PL, 1960 | NA | |||
| 7/60, PL, 1960 | ptxA1-prn1 | |||
| 134, NL, <1966 | ptxA2-prn1 | ptxA2-prn1 | ptxA2-prn1 | |
| 25593/65, Y, <1965 | ptxA2-prn1 | |||
| 14/64, PL, 1964 | NA | |||
| 15/64, PL, 1964 | NA | |||
| 16/64, PL, 1964 | NA | |||
| 20/64, PL, 1964 | NA | |||
| 1326/62, PL, 1962 | ptxA2-prn1 | |||
| 60623/67, Y, <1967 | ptxA2-prn1 | |||
| 606/67, PL, 1967 | ptxA2-prn1 | ptxA2-prn1 | ||
| 186/65, Y, 1965 | ptxA2-prn1 | ptxA2-prn1 | ||
| 629/65, PL, <1965 | ptxA2-prn1 | ptxA2-prn1 | ptxA2-prn1 | |
Abbreviations: Y, former Yugoslavia; NL, Netherlands; PL, Poland.
NA, not available.
PFGE methodology.
PFGE was performed according to the standardized procedure for typing B. pertussis (31) with the modifications described (2). The obtained PFGE profiles were analyzed using GelCompar software version 5.1 (Applied Maths, Belgium). The unweighted pair group method using the arithmetic average (UPGMA) algorithm was used as the clustering method, with 2% band tolerance and 1.5% optimization settings with the Dice coefficient. All profiles were verified by visual comparison. PFGE profiles obtained for clinical isolates and vaccine strains were compared to B. pertussis PFGE reference strains representing the different profile groups: 18323 (PFGE group I), Tohama I (PFGE group II), Bp134 (PFGE group III), B902 (PFGE group IVα), FR743 (PFGE group IVβ), FIN12 (PFGE group IVγ), and FR 287 (PFGE group V), as previously described (7, 43). PFGE profiles were classified into groups based on a criterion of similarity higher than 86%.
Serotyping.
Serotype analysis was performed by using a microplate agglutination assay with monoclonal antibodies against fimbriae type 2 (Fim2; NIBSC, 04/154) and fimbriae type 3 (Fim3; NIBSC, 04/156) according to the EUpertstrain group recommendations (www.eupertstrain.org/). Strains expressing autoagglutination were designated as untypeable. Strains that do not express Fim2 and Fim3 were designated as Fim0.
Sequencing.
Parts of the genes encoding for the surface protein pertussis toxin A subunit (ptxA) and pertactin (prn) were sequenced as previously described (31, 42).
RESULTS
Of the 139 strains analyzed, 69 PFGE profiles within five previously described PFGE groups (III, IVα, IVβ, IVγ, and V) and three newly identified PFGE clusters (A, B, and C) were found (Fig. 1). See also Table S1 in the supplemental material.
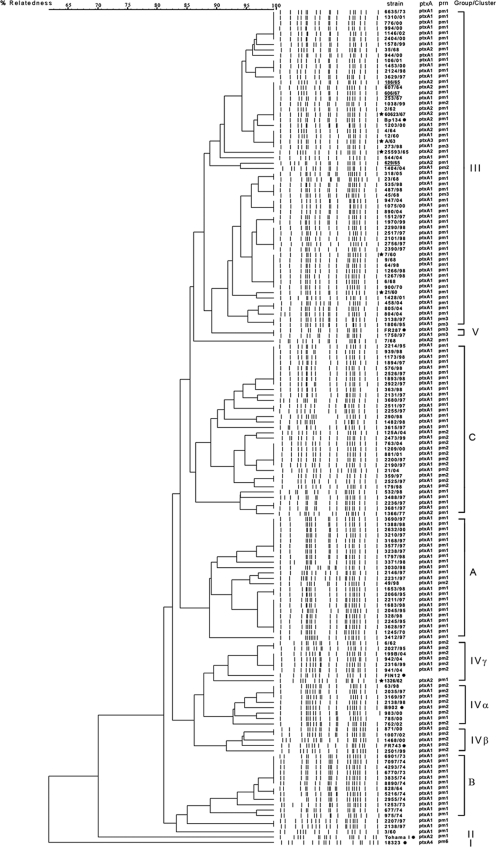
Fig. 1.
Dendrogram of 148 B. pertussis strains circulating in Poland from 1960 to 2005. Current vaccine strains are underlined, historical vaccine strains are indicated by an asterisk (*), and reference strains for different PFGE groups are indicated by a dot (●). The groups and clusters were identified to have an overall relatedness of ca. 86%. The classification of the pertactin allele (prn1, prn2, and prn3) and subunit A of the pertussis toxin allele (ptxA1, ptxA2, ptxA3, and ptxA4) is shown.
PFGE profiles typical for group I (represented by 18323) and group II (represented by Tohama I) were not found among the 139 clinical isolates and were related to other groups at the levels of 62 and 70%, respectively.
The relatedness levels among the strains classified into groups III, IVα, IVβ, IVγ, and V were found to be 88, 93, 92, 89, and 100%, respectively. For the newly identified clusters A, B, and C, relatedness achieved values of 89, 89, and 87%, respectively.
In group III, represented by reference strain Bp134, 21 different PFGE patterns were identified among 50 strains clustering together. Most of the isolates (38/50) found in group III were isolated after 1990. Among those isolated in the 1960s and 1970s, current (606/67, 186/65, and 629/65) and historical (60623/67, 25593/65, 7/60, 21/60, and A/63) vaccine strains were identified. Most of the strains within group III harbored the ptxA1-prn1 allele combination (40 of 50) (Table 2).
Table 2.
Distribution of PFGE groups and clusters versus ptxA-prn allele types among B. pertussis strains isolated in Poland within the periods 1960–77 and 1995–2005
| PFGE group/cluster | No. of strains isolated in: |
ptxA-prn allele combination | |
|---|---|---|---|
| 1960–1977 | 1995–2005 | ||
| III | 7 | 33 | ptxA1-prn1 |
| 0 | 2 | ptxA1-prn2 | |
| 1 | 3 | ptxA1-prn3 | |
| 4 | 0 | ptxA2-prn1 | |
| IVαβγ | 0 | 1 | ptxA1-prn1 |
| 1 | 15 | ptxA1-prn2 | |
| V | 0 | 1 | ptxA1-prn3 |
| A | 1 | 21 | ptxA1-prn1 |
| 0 | 1 | ptxA1-prn2 | |
| B | 12 | 0 | ptxA1-prn1 |
| C | 0 | 20 | ptxA1-prn1 |
| 0 | 11 | ptxA1-prn2 | |
| 1 | 0 | ptxA2-prn1 | |
PFGE groups IVα, IVβ, and IVγ, identified according to reference strains B902, FR743, and FIN12, contained five, three, and two distinct PFGE patterns found for seven, four, and six isolates, respectively. All isolates belonging to group IV were isolated in the 1990s and early 21st century, with the exception of a single isolate from the 1960s, identified in group IVγ. In group IVγ, the vaccine strain 1326/62 used for production up to 1967 was also found. Most of the strains found in PFGE groups IVα, IVβ, and IVγ harbored the ptxA1-prn2 (six, four, and six strains, respectively) allele combination.
Group V, represented by reference strain FR287, contained a single isolate (1758/97) from 1997 of the ptxA1-prn3 type. The position of group V in the dendrogram seemed to be biased since there were no more available strains clustered within group V.
The newly recognized clusters, C, A, and B contained 32, 23, and 12 isolates, respectively (Fig. 1). Clusters C and A contained strains isolated in the late 1990s with the exception of two isolates, 1366/77 and 1245/70. Most of the strains identified in clusters C and A presented the ptxA1-prn1 profile; however, 11 strains classified in the C cluster harbored ptxA1-prn2 alleles. Cluster B consisted of 11 strains from the 1970s and a single isolate from the 1960s, all presenting the ptxA1-prn1 allele type (Table 2). Unique PFGE patterns were found for isolate 7/68, related to group III and for three isolates (2207/97, 2138/97, and 3/60) more related to cluster B (Fig. 1).
The correctness of the assignment was verified according to the positioning of the European EUpertstrain collection of 140 strains from Finland, the Netherlands, France, Germany, Sweden, Denmark, and the United Kingdom. Most of the strains (127 of 141) from the EUpertstrain II culture collection (apart from the isolates from Poland) were classified into groups IVα and IVβ (data not shown).
Among the strains isolated in Poland from 1960 to 1977, 79 and 93% harbored the ptxA1 and prn1 alleles, respectively. All strains isolated in Poland in the period from 1995 to 2005 harbored the ptxA1 allele. Most of the isolates from 1995 to 2005 (70%) harbored the prn1 allele. The prn2 and prn3 alleles were found among 26 and 4% of the strains, respectively (Table 3).
Table 3.
Frequency of ptxA and prn alleles and Fim serotypes among B. pertussis strains isolated in Poland within the periods 1960–1977 and 1995–2005
| Period (yrs) | No. of strains | % (no. of strains) |
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
ptxA allele |
prn allele |
Serotype Fim |
||||||||||
| ptxA1 | ptxA2 | ptxA3 | prn1 | prn2 | prn3 | Fim2 | Fim3 | Fim2.3 | Fim0 | Untypeable | ||
| 1960–1977 | 29 | 79 (23) | 21 (6) | 93 (27) | 3.5 (1) | 3.5 (1) | 24 (7) | 38 (11) | 38 (11) | |||
| 1995–2005 | 110 | 100 (110) | 70 (77) | 26 (29) | 4 (4) | 82 (90) | 4.5 (5) | 12 (13) | 0.9 (1) | 0.9 (1) | ||
As many as 38, 38, and 24% of the strains isolated in the period from 1960 to 1977 expressed Fim2.3, Fim3, and Fim2, respectively (Table 3). Vaccine strains previously used for production were found to be of the Fim2 phenotype (A/63, 21/60, and 1326/62) and the Fim3 phenotype (509, 134, 25593/65, and 60623/67).
Currently used vaccine strains harbored the Fim2.3 phenotype, in contrast to the ones seen in the subsequent period from 1995 to 2005, since 12, 4.5, and 82% of the strains harbored Fim2.3, Fim3, and Fim2, respectively. A single isolate from 1995 to 2005 demonstrated autoagglutination, and another one was Fim0 (Table 3).
DISCUSSION
The extent of genetic heterogeneity of B. pertussis strains isolated from 1960 to 2005 in Poland, as found by application of the recommended PFGE standard procedure, appeared to be higher than described in other European countries. First, there were 59 PFGE patterns identified among 110 strains from 1995 to 2005 compared to 59 PFGE patterns found among 255 strains of the EUpertstrain I and II collections, respectively (19). Second, apart from previously described groups I to V, three new clusters designated in the study as A, B, and C were identified. In a study by Hallander et al. (19), Poland was found to be exceptional, since among 13 strains isolated from 1999 to 2004 and compared to strains from Finland, France, Germany, the Netherlands, Sweden, Denmark, and the United Kingdom, no isolate presented PFGE patterns typical for strains circulating in Europe. As much as 70% of the European isolates examined in the Hallander study (19) were assigned to 11 common PFGE profiles, the most prevalent one being BpSR11, which belongs to the IVβ PFGE group. BpSR11 comprised 10 to 50% of the isolates from Finland, France, Germany, the Netherlands, Sweden, Denmark, and the United Kingdom (19). In Sweden, the increased frequency of BpSR11 was to found to be related with more severe disease (3).
We found that only 47% of the Polish isolates studied belonged to PFGE groups III and IV. PFGE group III contained B. pertussis strains isolated from all periods studied, all three currently used vaccine strains and most of vaccine strains used historically for production of the national whole-cell pertussis vaccine. One vaccine strain (1326/62) used in previous periods for pertussis vaccine production was found in group IV. In contrast to the strains from other European countries (7, 19, 43), the B. pertussis strains we studied were not so commonly found in PFGE groups IVα, IVβ, and IVγ, since only seven, four, and six isolates were classified into groups IV α, IVβ, and IVγ, respectively. In the three newly recognized clusters, C, A, and B, 67 isolates (48%) were found. Cluster B, the most distinct one, was mainly represented by strains from the 1970s. Clusters C and A were comprised almost exclusively of strains isolated in late 1990s.
In the period from 1995 to 2005, compared to the period from 1960 to 1977, the frequency of strains belonging to group IV, cluster A, and cluster C greatly increased. The data we obtained may suggest that strains of group IV, as elsewhere in Europe, and strains belonging to newly identified clusters A and C are the clones most successfully disseminating in Poland. Moreover, most of the isolates (96%) found in cluster A harbored the ptxA1-prn1 allele. In cluster C, as much as 62.5 and 34% isolates harbored the ptxA1-prn1 and ptxA1-prn2 alleles, respectively. Generally speaking, after the 1990s, up to 70 and 26% of the strains were found to harbor vaccine-unrelated ptxA1-prn1 and ptxA1-prn2 alleles, respectively. Strains with ptxA1-prn1 alleles were, however, quite frequently isolated from 1960 to 1977 (72%) but slightly decreased in frequency in the following decades. In contrast, strains harboring ptxA1-prn2 were generally (except for a single isolate) not seen before the 1990s. Current vaccine strains harbor the ptxA2-prn1 alleles, but strains used in the past for wP production also harbored the ptxA3-prn1 and ptxA1-prn1 allele combinations (Table 1).
Changes in the distribution of ptxA1 and prn alleles in the period from 1995 to 2005, compared to 1960 to 1977, seemed to follow those observed in other highly immunized populations, where vaccine-unrelated variants actually predominate (22).
In our study, the shift observed in Fim serotypes did not match what has generally been observed in other highly vaccinated populations, where currently the Fim3 serotype has been the most frequent (1, 6, 21, 22, 26, 28, 40, 41, 45). In Poland, vaccine strains used for wP production harbor the Fim2.3 phenotype, contrasting with Fim2 among currently circulating isolates. This trend, however, does not seem to be unique among other vaccinated populations, as in Serbia, similarly to Poland, Fim2 is the prevalent serotype (1). Fim3 beside prn2, has been proposed as providing a survival advantage to B. pertussis isolates in wP-vaccinated population (28). However, shifts in serotype dominance might be attributed not only to vaccination status of a given population but also to vaccine strain-specific properties of the wP vaccine.
Weber et al. (43), Advani et al. (2), and Caro et al. (7) showed the linkage between group IV and the prn2 allele, between PFGE group V and the prn3 allele, and between PFGE group III and the prn1 allele. In our study, similar relationships between groups and alleles were confirmed.
The wider range of successfully disseminating clones in Poland compared to elsewhere in Europe as found by PFGE typing may be connected with wider vaccination selection pressure in the sense of the vaccine strains used. In Poland, awareness of pertussis has not increased among practitioners, and diagnostics have not improved. Vaccination coverage did not decrease over time and seems to be one of the highest in Europe, constantly oscillating ca. 98% (16). Analysis of the potency of the vaccine lots produced from 1972 to 2001 showed good product consistency, even if some changes have been introduced in the production and testing procedures (15). Waning immunity found among schoolchildren in the context of pathogen evolution was attributed to increases in pertussis rates (39). The exceptional history of the wP vaccine produced in Poland should definitely be taken into account as the force driving B. pertussis evolution, since several changes in vaccine strain composition have occurred in the past (Table 1). Until 1967, as many as 14 strains may have been used for vaccine production and they were differently allocated into particular vaccine lots. Although archival data are not fully complete, and not all of the strains used historically for production are currently available, some records clearly show that the strains used for vaccine production were changed periodically. From 1967 to 1971, 10 of the strains used for production were withdrawn, but one additional strain was added to the vaccine composition. In the subsequent period from 1971 to 1977, one vaccine strain was withdrawn and two additional strains were added. However, the reasons for these changes have remained unresolved. The composition of particular lots in the sense of isolates used for vaccine formulation was also found to be changeable. The vaccine strain composition of vaccine lots produced in 1976 is shown in Table 4 and indicates the level of formulation differences. The last vaccine strain composition change took place in 1978 and was aimed at eliminating strains expressing the highest toxicity and to lower the possibility of adverse events after vaccination (37). Since then, the same vaccine formulation has been used continuously for over 30 years.
Table 4.
Example of vaccine strain wP composition in the DTwP vaccine lots produced in 1976 in Poland
| Lot of DTwP vaccine | Composition (%) of final lot |
|||||
|---|---|---|---|---|---|---|
| 629/65 | 186/65 | 606/67 | A/63 | 134 | 21/60 | |
| S1/76 | 22 | 23 | 20 | 10 | 5 | 20 |
| S2/76 | 24 | 13 | 24 | 10 | 29 | |
| S3/76 | 18 | 11 | 16 | 15 | 5 | 5 |
| S4/76 | 25 | 17 | 33 | 5 | 15 | 5 |
| S5/76 | 21 | 17 | 20 | 15 | 27 | |
| S6/76 | 17 | 21 | 12 | 9 | 16 | 23 |
| S7/76 | 30 | 25 | 10 | 24 | 11 | |
| S8/76 | 31 | 28 | 10 | 10 | 11 | 10 |
| S9/76 | 24 | 24 | 21 | 10 | 10 | 11 |
| S10/76 | 28 | 30 | 16 | 20 | 6 | |
| S11/76 | 27 | 22 | 19 | 20 | 12 | |
| S12/76 | 20 | 21 | 17 | 19 | 12 | 11 |
The higher level of heterogeneity of B. pertussis strains observed in Poland as found by PFGE may have resulted from several different pressures induced by different vaccine strains capable of increasing B. pertussis population diversity. Thus, we suspect that the differences observed in B. pertussis populations circulating in Poland and other EU countries may have resulted from different vaccine composition strategies used since the initiation of routine pertussis vaccination.
Supplementary Material
ACKNOWLEDGMENTS
This study was supported by a grant for scientific research (N N404 167634) from the Ministry of Education and Science in Poland.
We thank Marta Balińska-Peroutková for help with the English version of the manuscript and Barbara Husejnow for excellent technical assistance.
Footnotes
Supplemental material for this article may be found at http://jcm.asm.org/.
Published ahead of print on 9 February 2011.
REFERENCES
- 1. Advani A., Donnelly D., Gustafsson L., Hallander H. O. 2007. Changes of the Swedish Bordetella pertussis population in incidence peaks during an acellular pertussis vaccine period between 1997 and 2004. APMIS 115:299–310 [DOI] [PubMed] [Google Scholar]
- 2. Advani A., Donnelly D., Hallander H. 2004. Reference system for characterization of Bordetella pertussis pulsed-field gel electrophoresis profiles. J. Clin. Microbiol. 42:2890–2897 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Advani A., Gustafsson L., Carlsson R.-M., Donnelly D., Hallander H. 2007. Clinical outcome of pertussis in Sweden: association with pulsed-field gel electrophoresis profiles and serotype. APMIS 115:736–742 [DOI] [PubMed] [Google Scholar]
- 4. Advani A., van der Heide H. G. J., Hallander H. O., Mooi F. R. 2009. Analysis of Swedish Bordetella pertussis isolates with three typing methods: characterization of an epidemic lineage. J. Microbiol. Methods 78:297–301 [DOI] [PubMed] [Google Scholar]
- 5. Borisova O., et al. 2007. Antigenic divergence between Bordetella pertussis clinical isolates from Moscow, Russia, and vaccine strains. Clin. Vaccine Immunol. 14:234–238 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Caro V., et al. 2006. Bordetella pertussis, Finland and France. Emerg. Infect. Dis. 12:987–989 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Caro V., et al. 2005. Pulsed-field gel electrophoresis analysis of Bordetella pertussis populations in various European countries with different vaccine policies. Microbes Infect. 7:976–982 [DOI] [PubMed] [Google Scholar]
- 8. Cassiday P., et al. 2000. Polymorphism in Bordetella pertussis pertactin and pertussis toxin virulence factors in the United States, 1935–1999. J. Infect. Dis. 182:1402–1408 [DOI] [PubMed] [Google Scholar]
- 9. Celentano L. P., Massari M., Paramatti D., Salmaso S., Tozzi A. E. 2005. Resurgence of pertussis in Europe. Pediatr. Infect. Dis. J. 24:761–765 [DOI] [PubMed] [Google Scholar]
- 10. Cordova S. P., Gilles M. T., Beers M. Y. 2000. The outbreak that had to happen: Bordetella pertussis in north-west Western Australia in 1999. Commun. Dis. Intell. 24:375–379 [DOI] [PubMed] [Google Scholar]
- 11. Dakic G., et al. 2010. Bordetella pertussis vaccine strains and circulating isolates in Serbia. Vaccine 28:1188–1192 [DOI] [PubMed] [Google Scholar]
- 12. Elomaa A., et al. 2005. Strain variation among Bordetella pertussis isolates in Finland, where the whole-cell pertussis vaccine has been used for 50 years. J. Clin. Microbiol. 43:3681–3687 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Fingermann M., et al. 2006. Differences of circulating Bordetella pertussis population in Argentina from the strain used in vaccine production. Vaccine 24:3513–3521 [DOI] [PubMed] [Google Scholar]
- 14. Fry N. K., et al. 2001. Genotypic variation in the Bordetella pertussis virulence factors pertactin and pertussis toxin in historical and recent clinical isolates in the United Kingdom. Infect. Immun. 69:5520–5528 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Gzyl A., Augustynowicz E., Gniadek G., Ślusarczyk J. 2004. Potency of pertussis component in the DTP vaccine: an overview of three decade study in Poland. Biologicals 32:129–137 [DOI] [PubMed] [Google Scholar]
- 16. Gzyl A., Augustynowicz E., Rabczenko D., Gniadek G., Slusarczyk J. 2004. Pertussis in Poland. Int. J. Epidemiol. 33:358–365 [DOI] [PubMed] [Google Scholar]
- 17. Gzyl A., Augustynowicz E., van Loo I., Ślusarczyk J. 2000. Temporal nucleotide changes in pertactin and pertussis toxin genes in Bordetella pertussis strains isolated from clinical cases in Poland. Vaccine 20:299–303 [DOI] [PubMed] [Google Scholar]
- 18. Hallander H. O., Advani A., Donnelly D., Gustafsson L., Carlsson R.-M. 2005. Shifts of Bordetella pertussis variants in Sweden from 1970 to 2003, during three periods marked by different vaccination programs. J. Clin. Microbiol. 43:2856–2865 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Hallander H., et al. 2007. Bordetella pertussis strains circulating in Europe in 1999 to 2004 as determined by pulsed-field gel electrophoresis. J. Clin. Microbiol. 45:3257–3262 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Han H.-J., et al. 2008. Antigenic variation in Bordetella pertussis isolates recovered from adults and children in Japan. Vaccine 26:1530–1534 [DOI] [PubMed] [Google Scholar]
- 21. Heikkinen E., et al. 2008. Bordetella pertussis isolates in Finland: serotype and fimbrial expression. BMC Microbiol. 8:162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Kallonen T., He Q. 2009. Bordetella pertussis strain variation and evolution postvaccination. Expert Rev. Vaccines 8:863–875 [DOI] [PubMed] [Google Scholar]
- 23. Khattak M. N., Matthews R. C., Burnie J. P. 1992. Is Bordetella pertussis clonal? BMJ 304:813–815 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Khattak M. N., Matthews R. C. 1993. Genetic relatedness of Bordetella species as determined by macrorestriction digests resolved by pulsed-field gel electrophoresis. Int. J. Syst. Bacteriol. 43:659–664 [DOI] [PubMed] [Google Scholar]
- 25. King A. J., et al. 2008. Comparative genomic profiling of Dutch clinical Bordetella pertussis isolates using DNA microarrays: identification of genes absent from epidemic strains. BMC Genomics 9:311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Kurova N., Njamkepo E., Brun D., Tseneva G., Guiso N. 2010. Monitoring of Bordetella isolates circulating in Saint Petersburg, Russia, between 2001 and 2009. Res. Microbiol. 161:810–815 [DOI] [PubMed] [Google Scholar]
- 27. Lee Y. S., Yang C. Y., Lu C. H., Tseng Y. H. 2003. Molecular epidemiology of Bordetella pertussis isolated in Taiwan, 1992–1997. Microbiol. Immunol. 47:903–909 [DOI] [PubMed] [Google Scholar]
- 28. Litt D. J., Neal S. E., Fry N. K. 2009. Changes in genetic diversity of the Bordetella pertussis population in the United Kingdom between 1920 and 2006 reflect vaccination coverage and emergence of a single dominant clonal type. J. Clin. Microbiol. 47:680–688 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Mooi F. R., et al. 1998. Polymorphism in the Bordetella pertussis virulence factors P. 69/pertactin and pertussis toxin in the Netherlands: temporal trends and evidence for vaccine-driven evolution. Infect. Immun. 66:670–675 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Mooi F. R., He Q., van Oirschot H., Mertsola J. 1999. Variation in the Bordetella pertussis virulence factors pertussis toxin and pertactin in vaccine strains and clinical isolates in Finland. Infect. Immun. 67:3133–3134 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Mooi F. R., Hallander H., Wirsing von Konig C. H., Hoet B., Guiso N. 2000. Epidemiological typing of Bordetella pertussis isolates: recommendations for a standard methodology. Eur. J. Clin. Microbiol. Infect. Dis. 19:174–181 [DOI] [PubMed] [Google Scholar]
- 32. Mooi F. R., van Loo I. H., King A. J. 2001. Adaptation of Bordetella pertussis to vaccination: a cause for its reemergence? Emerg. Infect. Dis. 7:526–528 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Mooi F. R. 2010. Bordetella pertussis and vaccination: the persistence of a genetically monomorphic pathogen. Infect. Genet. Evol. 10:36–49 [DOI] [PubMed] [Google Scholar]
- 34. Njamkepo E., Cantinelli T., Guigon G., Guiso N. 2008. Genomic analysis and comparison of Bordetella pertussis isolates circulating in low and high vaccine coverage areas. Microbes Infect. 10:1582–1586 [DOI] [PubMed] [Google Scholar]
- 35. Ntezayabo B., De Serres G., Duval B. 2003. Pertussis resurgence in Canada largely caused by a cohort effect. Pediatr. Infect. Dis. J. 22:22–27 [DOI] [PubMed] [Google Scholar]
- 36. Pereira A., Pereira A. S. P., Moreira-Filho C. A., Bando S. Y., Tambourgi D. V. 2005. Comparative analysis of Bordetella pertussis patient isolated strain and classical strains used in the pertussis vaccine. Vaccine 23:4353–4358 [DOI] [PubMed] [Google Scholar]
- 37. Schiller B., Rudnicka W. W., Robaczewska M. E. 1976. Stability of pertussis strains for vaccine production, abstr. 197. Proceedings of the Symposium on Stability and Effectiveness of Measles, Poliomyelitis and Pertussis vaccines Yugoslav Academy of Sciences and Arts, Zagreb, Poland [Google Scholar]
- 38. Schouls L. M., van der Heide H. G. J., Vauterin L., Vauterin P., Mooi F. R. 2004. Multiple-locus variable-number tandem repeat analysis of Dutch Bordetella pertussis strains reveals rapid genetic changes with clonal expansion during the late 1990s. J. Bacteriol. 186:5496–5505 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Ślusarczyk J., et al. 2002. Stan uodpornienia dzieci w wieku 6-8 lat przeciw krztuŚcowi, tę żcowi i błonicy. Przegl. Epidemiol. 56:39–48 [PubMed] [Google Scholar]
- 40. Tsang R. S. W., et al. 2004. Polymorphisms of the fimbria fim3 gene of Bordetella pertussis strains isolated in Canada. J. Clin. Microbiol. 42:5364–5367 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. van Amersfoorth S. C. M., et al. 2005. Analysis of Bordetella pertussis populations in European countries with different vaccination policies. J. Clin. Microbiol. 43:2837–2843 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. van Loo I. H., van der Heide H. G., Nagelkerke N. J., Verhoef J., Mooi F. R. 1999. Temporal trends in the population structure of Bordetella pertussis during 1949-1996 in a highly vaccinated population. J. Infect. Dis. 179:915–923 [DOI] [PubMed] [Google Scholar]
- 43. Weber C., Boursaux-Eude C., Coralie G., Caro V., Guiso N. 2001. Polymorphism of Bordetella pertussis isolates circulating for the last 10 years in France, where a single effective whole-cell vaccine has been used for more than 30 years. J. Clin. Microbiol. 39:4396–4403 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Zanardi L., Pascual F., Bisgard K., Murphy T., Wharton M. 2002. Pertussis–United States,1997-2000. MMWR Morb. Mortal. Wkly. Rep. 51:73–76 [PubMed] [Google Scholar]
- 45. Zhang L., et al. 2010. Effect of vaccination on Bordetella pertussis strains, China. Emerg. Infect. Dis. 16:1695–1701 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.