Abstract
Different strains of Listeria monocytogenes are well known to persist in individual food processing plants and to contaminate foods for many years; however, the specific genotypic and phenotypic mechanisms responsible for persistence of these unique strains remain largely unknown. Based on sequences in comK prophage junction fragments, different strains of epidemic clones (ECs), which included ECII, ECIII, and ECV, were identified and shown to be specific to individual meat and poultry processing plants. The comK prophage-containing strains showed significantly higher cell densities after incubation at 30°C for 48 h on meat and poultry food-conditioning films than did strains lacking the comK prophage (P < 0.05). Overall, the type of strain, the type of conditioning film, and the interaction between the two were all highly significant (P < 0.001). Recombination analysis indicated that the comK prophage junction fragments in these strains had evolved due to extensive recombination. Based on the results of the present study, we propose a novel model in which the concept of defective comK prophage was replaced with the rapid adaptation island (RAI). Genes within the RAI were recharacterized as “adaptons,” as these genes may allow L. monocytogenes to rapidly adapt to different food processing facilities and foods. If confirmed, the model presented would help explain Listeria's rapid niche adaptation, biofilm formation, persistence, and subsequent transmission to foods. Also, comK prophage junction fragment sequences may permit accurate tracking of persistent strains back to and within individual food processing operations and thus allow the design of more effective intervention strategies to reduce contamination and enhance food safety.
INTRODUCTION
Listeria monocytogenes is a unique food-borne pathogen that causes listeriosis, which ranges from febrile gastroenteritis to more severe life-threatening invasive diseases, especially for immunocompromised individuals (94). It is widely distributed in many wild and domestic animals and various natural environments and is resistant to a wide variety of environmental stresses (33). L. monocytogenes is considered a model organism for the study of host-pathogen interactions, especially as a model for intracellular pathogens of humans (47). However, L. monocytogenes may also be an excellent model for pathogen-environment interactions, because it is well known to cycle between being a pathogen in many wild and domestic animals and a saprophyte in diverse environments, including those found in various types of food processing facilities (41). However, while much is known about the pathogenic lifestyle of L. monocytogenes, much less is known about its saprophytic lifestyle, including the genetic and phenotypic factors affecting its colonization and persistence in food processing and retail environments and subsequent transmission to ready-to-eat (RTE) foods. This has resulted in numerous costly recalls, disease cases, and outbreaks, which are often associated with high mortality (94). As a result, a zero-tolerance policy for L. monocytogenes currently exists for RTE meat and poultry products manufactured in the United States.
Specific resident strains of L. monocytogenes have been found to persist for months to more than a decade (70, 78, 95, 105) in various types of food processing plants manufacturing meat products (5, 38, 40, 90, 105), poultry products (4–6, 32, 78, 85), dairy products (4, 5, 37, 48, 53, 101), seafood products (4, 5, 73, 86, 95), vegetables (5), and sandwiches (8) and also in retail environments (88). These persistent strains are genetically distinct from transient strains that are isolated sporadically (4, 5). Persistent strains of L. monocytogenes show greater adherence to food-contact surfaces than do nonpersistent strains (66, 74) and as a result contaminate foods more frequently than do sporadic strains (5, 48). Autio et al. (5) found that some pulsotypes of L. monocytogenes were repeatedly found in the same food product from the same producer while other pulsotypes were repeatedly found in products from multiple producers. While many reports have clarified the roles of numerous virulence genes and their coordinated expression in causing listeriosis, the genotypic and phenotypic mechanisms responsible for the persistence of different strains of L. monocytogenes in individual food processing and retail facilities remain to be elucidated.
Most bacteria, including L. monocytogenes, grow on surfaces and form biofilms (12, 93). Cells of L. monocytogenes within biofilms are more resistant to biocides (102), which increases the risk of food contamination. Initial adherence is critical for biofilm formation and depends on the physiochemical properties of the environmental surfaces as well as the biofilm-forming potential of the bacterial cells (18, 83, 102). L. monocytogenes can adhere to abiotic surfaces such as stainless steel, glass, plastic, polymers, and rubber that are present in the food processing environment (44, 102, 107). These bacteria also adhere to biotic surfaces such as other microorganisms and plant and animal tissues (44). Strains of L. monocytogenes show specificity in forming biofilms. For example, serotype 1/2a strains have been reported to adhere faster to surfaces and to produce larger biofilms than do other serotypes (35, 54). However, L. monocytogenes strains that are low biofilm producers have been reported to form dense biofilms in the presence of a strong biofilm producer (80). Various studies have demonstrated that specific genes in L. monocytogenes are required for different phases of biofilm formation. In L. monocytogenes, agr (accessory gene regulator) and ami (autolysin-adhesion gene) mutants were shown to be defective in the initial attachment to a surface (57, 84, 91). Also, genomic studies exploring gene expression during growth of L. monocytogenes on a surface demonstrated that the DegU orphan response regulator was important for biofilm formation (43, 57).
Conditioning films are well known to enhance the adherence of various microorganisms to numerous surfaces in natural environments (29, 39) and the surfaces of teeth (27, 56) and implanted medical devices (24). Bowden and Li (11) stressed the importance of nutritional influences of conditioning films on biofilm development. They indicated that conditioning films may act as important nutrient sources and that competition for such nutrients may serve as a strong selective force in the evolution of strains that form biofilms and colonize various surfaces. Many authors have speculated that biofilm formation is the reason for the persistence of specific strains in food processing plants. However, other explanations for this phenomenon have also been suggested, including continual reintroduction of the same strain over extended periods of time, random primary colonization, ability to grow and survive at low temperatures, competitiveness for nutrients, ability to mount a stringent response and undergo physiological adaptation to nutrient deprivation, resistance to sanitizers and/or heavy metals and antibiotics, ability to interact with other microorganisms to form a stable ecosystem within biofilms, or some combination of the above (70, 78, 83, 105).
In order to implement more effective intervention strategies to prevent persistent strains of L. monocytogenes from contaminating RTE foods, we must first identify their reservoirs and understand how they are transmitted from these reservoirs to foods. To accomplish this, it is critical to develop an accurate molecular subtyping method by selecting the right molecular subtyping marker(s) and applying it to a relevant collection of strains (36, 94). Multi-virulence-locus sequence typing (MVLST) of L. monocytogenes very accurately identified and differentiated the four epidemic clones (ECs) of L. monocytogenes (ECI, -II, -III, and -IV), outbreak clones that did not belong to a specific epidemic clone, and nonoutbreak strains (21, 65). However, MVLST was not able to differentiate outbreak clones within epidemic clones of L. monocytogenes (21, 65). To overcome this limitation, Chen and Knabel (20) targeted regions within the comK and PSA prophages of L. monocytogenes and designed PCR-based approaches that accurately differentiated different outbreak clones within epidemic clones. Orsi et al. (78) subsequently performed whole-genome sequencing on lineage II serotype 1/2a ECIII strains of L. monocytogenes from a 1988 sporadic case and the 2000 outbreak linked to RTE poultry products from the same processing plant in Texas. Their findings supported the conclusion of Chen and Knabel (20) that prophages in L. monocytogenes are excellent markers for differentiating outbreak clones within epidemic clones. The results of Orsi et al. (78) also revealed extensive recombination throughout the comK prophage in ECIII, which they suggested was due to recombination between the comK prophage in the ECIII strain in lineage II and the comK prophage in a serotype 1/2b strain in lineage I.
Most species of bacteria, including L. monocytogenes (49, 62), contain prophage DNA which can interact with infecting phage (49, 50). As a result, phage-prophage interactions are thought to be a dynamic force acting on bacterial populations and to account for a major portion of bacterial evolution that is occurring via lateral gene transfer (13, 15, 49, 50, 52). Therefore, temperate phages which integrate their genomes into host genomes could be a major source of novel genes that enhance the fitness of the bacterial host (34, 46, 49, 52, 103). Wang et al. (103) demonstrated that cryptic (defective) prophages in Escherichia coli K-12 enhance its resistance to osmotic, oxidative, and acid stresses and increase growth and biofilm formation. Prophages, including those in L. monocytogenes, constitute the major differences between genomes of closely related strains of bacteria (13). This is especially the case for epidemic clones and outbreak clones of L. monocytogenes, which are highly clonal in terms of their backbone genome (22, 72, 78, 81, 82). Extensive mosaicism is typical of prophages in many prokaryotes (71), including the comK prophage of L. monocytogenes (78), which is rendered inactive (defective) by gene deletions following insertion into the comK gene (16).
The Food Safety and Inspection Service (FSIS) is responsible for monitoring the safety of RTE meat and poultry products manufactured in the United States. As part of this responsibility, FSIS conducts risk-based L. monocytogenes testing programs in processing plants producing postlethality-exposed RTE meat and poultry products. As a result of its L. monocytogenes sampling program, FSIS has generated more than 500 L. monocytogenes isolates from different RTE meat processing facilities throughout the United States (104). Small fractions (∼4%) of these isolates were ECII based on pulsed-field gel electrophoresis (PFGE) profiles and multilocus genotyping (MLGT) (104); however, these methods could not prove conclusively that these isolates were ECII nor determine whether they represented different genotypes of ECII. Eifert et al. (32) also isolated L. monocytogenes strains that had ECII markers and PFGE patterns similar to those of the 1998 and 2002 L. monocytogenes outbreak clones (32) from two different turkey processing plants from nonadjoining states in the United States. L. monocytogenes serotype 1/2a epidemic clones, ECIII (76) and ECV (2008 Canadian outbreak; S. Knabel, A. Reimer, B. Verghese, M. Lok, J. Zergler, J. Farber, F. Pagotto, M. Graham, C. A. Nadon, Canadian Public Health Laboratory Network (CPHLN), and M. W. Gilmour, unpublished data), also appear to persist within RTE meat and poultry processing plants. Therefore, the purposes of the present study were (i) to confirm that the putative ECII isolates were ECII; (ii) to determine the genotypes of persistent strains of ECII, ECIII, and ECV by amplifying and sequencing comK prophage junction fragments; (iii) to assess whether recombination was occurring in these junction fragments; (iv) to determine the capacities of the different comK-prophage-containing genotypes to attach, grow, and form biofilms on RTE meat- and poultry-conditioning films, compared to those of strains lacking the comK prophage; and (v) to develop a genotypic/phenotypic model that might explain how sequence variations in the comK prophage might drive rapid niche-specific adaptation, biofilm formation, persistence, and transmission of L. monocytogenes.
MATERIALS AND METHODS
Bacterial isolates and DNA extraction.
Isolate identification numbers, sources, and dates of isolation are given in Table S1 in the supplemental material. Ten isolates were obtained from the Listeria collection at CDC, and 18 isolates were obtained from FSIS of the U.S. Department of Agriculture (USDA). For the 10 isolates obtained from CDC, six were associated with two outbreaks of listeriosis involving ECII and four isolates were ECIII isolates from the same plant in Texas (two isolates associated with one sporadic case in 1988 and two isolates associated with one outbreak of listeriosis in 2000). The 18 isolates obtained from FSIS were isolated by FSIS at different times from different meat processing facilities as part of their L. monocytogenes testing programs. FSIS had determined that these isolates had PFGE and MLGT subtypes similar to those of previously characterized ECII isolates (104). Bacterial isolates were streaked on tryptic soy agar with 0.6% yeast extract (TSAYE) (BD, Franklin Lakes, NJ) with incubation at 35°C for 24 h. For each strain, one colony on the plate was inoculated into 10 ml of tryptic soy broth with yeast extract (TSBYE) and then incubated at 35°C overnight. Cultures grown overnight were adjusted to an optical density of 0.2 at 650 nm, which is equivalent to approximately 107 CFU/ml. For all isolates, bacterial genomic DNAs were extracted using an UltraClean microbial DNA extraction kit (Mo Bio Laboratories, Solana Beach, CA) and stored at −20°C before use. Similar methods were used for genomic DNA extraction from 10 isolates from the work of Eifert et al. (32) in S. Kathariou's lab.
ECII PCR.
PCR amplifications were performed using PCR master mix (Qiagen Inc., Valencia, CA) and ECII-specific primers (see Table S2 in the supplemental material) with a Mastercycler gradient (Eppendorf Scientific Corp., Hamburg, Germany). Amplification conditions were 94°C for 15 min followed by 31 cycles of 94°C for 30 s, 59°C for 1.5 min, and 72°C for 1.5 min and a final extension at 72°C for 10 min.
MVLST.
Intragenic regions of the six virulence genes (prfA, inlB, inlC, dal, clpP, and lisR) were amplified as previously described by Chen et al. (21).
Outbreak clone PCR.
PCR amplification of the hypervariable comK prophage region in the 1998 and 2002 ECII outbreak strains (LMOh7858_2426) was performed as previously described by Chen and Knabel (20).
Amplifications of upstream and downstream junction fragments.
Amplification of the upstream and downstream junction fragments within comK was performed using methods modified from the work of Loessner et al. (64). These modifications included redesigned primers (see Table S2 in the supplemental material) and different cycling conditions. A single PCR program was used for both upstream and downstream junction fragment amplifications (initial denaturation at 95°C for 15 min followed by 40 cycles of 94°C for 30 s, 63°C for 1.5 min, and 72°C for 1.5 min and a final extension at 72°C for 10 min).
DNA sequencing.
All PCR products were purified using ExoSAP-IT (USB Corp., Cleveland, OH) prior to sequencing. DNA sequencing was performed at the Pennsylvania State University Genomics Core Facility. The sequences of two clinical isolates from the 2008 Canada outbreak (ECV) were downloaded from NCBI GenBank.
Sequence analysis.
Sequence analyses were performed using DNAStar software (v8.1) (DNAStar Inc.). Multiple sequence alignments were performed using MEGA (version 4.0.2) (97). The unweighted pair group method with arithmetic mean (UPGMA) implemented in MEGA was used to construct cluster diagrams using the upstream and downstream junction fragment sequences of L. monocytogenes isolates. One thousand random tree replications were performed, and bootstrap values were assigned to the nodes of the cluster diagram (see Fig. 2).
Fig. 2.
Cluster diagrams based on upstream and downstream junction fragment sequences in L. monocytogenes isolates described in Table S1 in the supplemental material; the plant information and prophage type are labeled in the figure. (A) Upstream junction fragment cluster diagram. (B) Downstream junction fragment cluster diagram. Nodes are labeled with bootstrap values. *, plant information is not available.
Recombination analysis.
The extent of recombination within the upstream and downstream junction fragments of L. monocytogenes was determined using the Recombination Detection Program (RDP), GENECONV, maximum chi-square, Chimera, and Sister Scan recombination detection methods implemented in RDP v.3.38 (68). Since no single program provided optimal performance under all conditions, a stringent criterion was applied, such that any event for detection of positive recombination breakpoints had to be supported by five or more methods with P values of ≤10−5.
Detection of spontaneous induction of the comK prophage.
Spontaneous induction of the comK prophage was detected by attP and attB PCR. Amplifications of the attP site within comK phage (Fig. 1 A) and the attB attachment site within comK in the L. monocytogenes genome (Fig. 1B) were performed using methods modified from the work of Loessner et al. (64). The modifications included redesigned primers (see Table S2 in the supplemental material) and cycling conditions as described above for amplification of the upstream and downstream junction fragments.
Fig. 1.
Schematic diagram of phage A118 integration into and excision out of the L. monocytogenes chromosome at comK. (A) Phage A118 with attP attachment site. Horizontal arrows indicate PCR priming sites for amplifying the fragment containing attP. (Modified from reference 64 with permission of John Wiley & Sons.) (B) L. monocytogenes comK gene containing attB attachment site. Horizontal arrows indicate priming sites for amplifying comK with attB attachment site. (C) Lysogenized L. monocytogenes showing the locations of the forward and reverse primers (horizontal arrows) for PCR amplification of upstream and downstream comK prophage junction fragments. The location of gene locus LMOh7858_2426 in the lysogenized strain (20) is shown with an upward-pointing arrow. (D) Lysogenized L. monocytogenes showing the PCR targets for the upstream and downstream comK prophage junction fragment PCR (RBS, ribosomal binding protein; NC, noncoding region; att, attachment site). The partial gene targets are underlined.
Detection of int gene in L. monocytogenes genomic DNA and comK phage DNA by PCR.
The phage integrase gene (int) in L. monocytogenes genomic DNA and comK phage DNA was amplified by PCR in strains F2365 (ECI), H7858 (ECII), N3-031 (ECIII), and 1001::A118 using primers described in Table S2 in the supplemental material. ECI strain F2365 was isolated from Mexican-style soft cheese during the 1985 California outbreak. The ECII and ECIII strains are described in Table 4. L. monocytogenes strain 1001::A118 is an artificially lysogenized serotype 1/2a strain. The cells were grown to early log phase (107 CFU/ml) at 35°C in tryptose broth (TB) supplemented with CaCl2 to a final concentration of 10 mM. Cells were centrifuged at 12,000 rpm for 1 min and resuspended in 5 ml of fresh TB. After overnight incubation in TB at 35°C, samples were centrifuged at 14,000 rpm for 10 min and the supernatant was filtered through a 0.2-μm sterile membrane filter (Thermo Scientific, Waltham, MA). Genomic DNA extraction was performed as described above. Phage DNA extraction was conducted as described by Asadulghani et al. (3). Briefly, phage DNA in the phage head was detected by PCR amplification of int after DNase (Rockland, Gilbertsville, PA), RNase (EMD, Gibbstown, NJ), and proteinase K (Rockland, Gilbertsville, PA) treatments. PCR amplification conditions were 94°C for 15 min followed by 30 cycles of 94°C for 30 s, 62°C for 1.5 min, and 72°C for 1.5 min and a final extension at 72°C for 10 min. Phage DNA amplification was repeated at least three times for each isolate, and int was confirmed by sequencing. The int primer sequences for detection of phage DNA are shown in Table S2 in the supplemental material.
Table 4.
Detection of spontaneous induction of the comK prophage in L. monocytogenes strains F2365, H7858, and N3-031 using attP and attB PCR, comK phage int PCR, and plaque formation
| Strain testeda | Serotype | Detection of spontaneous excision |
Amplification of int in: |
Plaque formationb | Interpretation | ||
|---|---|---|---|---|---|---|---|
| attP | attB | Lysogen | Phage | ||||
| F2365 (ECI) | 4b | − | − | − | − | − | comK prophage absent, no comK phage formed |
| H7858 (ECII) | 4b | + | + | + | − | − | comK prophage spontaneously induced, no or undetectable levels of defective comK phage |
| N3-031 (ECIII) | 1/2a | + | + | + | + | − | comK prophage spontaneously induced, defective comK phage |
| 1001::A118 | 1/2a | + | + | + | + | +c | comK prophage spontaneously induced, active comK phage |
Epidemic clone designation is indicated within parentheses after strain tested.
Indicator strains tested were ATCC 23074 and 1001 for serotypes 4b and 1/2a, respectively.
Approximately 3.00 log10 PFU/ml of filtrate as described in Materials and Methods.
Plaque assay.
To determine whether the prophages in strains H7858 (ECII) and N3-031 (ECIIII) were active or defective, a plaque assay was performed on these strains. All media used in the plaque assay were supplemented with CaCl2 to a final concentration of 10 mM. Cells were grown at 35°C in TB to early log phase (107 CFU/ml), centrifuged at 12,000 rpm for 1 min, and then resuspended in 1.8 ml of fresh TB. After overnight incubation in TB at 35°C, samples were centrifuged at 14,000 rpm for 10 min and the supernatant was filtered through a 0.2-μm sterile membrane filter (Thermo Scientific, Waltham, MA). One hundred microliters of filtrate was mixed with 100 μl of exponential-phase cells of the serotype 1/2a indicator strain 1001 (provided by Martin Loessner, ETH, Zurich, Switzerland) or 4b indicator strain ATCC 23074; the mixture was added to 5 ml of molten TSAYE containing 0.75% agar at 50°C and then poured onto solidified TSAYE containing 1.5% agar. Plaque formation was observed after overnight incubation at 30°C. The plaque assay was also performed on strains F2365 (ECI) and 1001::A118. Each plaque assay was replicated twice.
Epifluorescence microscopy.
Epifluorescence microscopy was used to study the effects of type of strain of L. monocytogenes, type of food-conditioning film, and their interaction on the density of L. monocytogenes cells on glass slides after incubation at 30°C for 48 h. Seven strains of L. monocytogenes were used in this study, including two strains that lacked the comK prophage (a lineage III strain, W1-111, and a lineage I ECI strain, F2365) and five strains that contained the comK prophage (three ECII strains [J1703, H7858, and OB020790], an ECIII strain [N3-031], and an ECV strain [08-5923]). Sterile TSBYE containing no cells served as the negative control for strains. Five large, unsliced, unopened cooked RTE foods from a local supermarket deli were used to make the food-conditioning films. These foods included Brie soft cheese, pork and beef hot dogs (containing salt, potassium lactate, sodium phosphate, sodium diacetate, sodium erythorbate, and sodium nitrite), fully cooked turkey breast (containing salt and sodium phosphate), pasteurized canned ham (containing salt, sodium phosphate, and sodium nitrite), and chicken breast (containing salt, potassium lactate, sodium phosphate, and sodium diacetate). Briefly, all foods (except the canned ham) were steamed for 60 min to destroy all vegetative cells that might be present on the surface of the foods and then cooled in an ice-water bath. Ten grams of each food was then aseptically sampled from the center of the food products and blended with 40 g of sterile water in a sterile blender jar using an Osterizer blender (Oster, Shelton, CT). Sixty microliters of the homogeneous food slurry of each food was transferred into each compartment of an eight-compartment CultureSlide (Falcon; Becton Dickinson, Franklin Lakes, NJ) (see Fig. S1 in the supplemental material). Food slurries in CultureSlide compartments were then air dried at 35°C for 3 h to form food-conditioning films on the surface of the glass CultureSlides. Sterile water containing no food served as the negative control for food-conditioning film.
The protocol for preparation of inoculates and fluorescent microscopy was adapted from the report by Kushwaha and Muriana (58). Briefly, all strains of L. monocytogenes were incubated in TSBYE at 35°C for 17 h and then diluted 10−5-fold with sterile TSBYE. Two hundred microliters of diluted culture of each strain was then added to each compartment of the CultureSlide (except for the no-L. monocytogenes control compartment), and then the CultureSlides were incubated at 30°C for 48 h (see Fig. S1 in the supplemental material for experimental design). After incubation, each compartment was rinsed three times using 200 μl of Tris buffer (pH 7.4; 0.05 M) and then stained using 200 μl of 5,6-carboxy-fluorescein diacetate (5,6-CFDA; Invitrogen, Carlsbad, CA) solution with incubation at 25°C for 15 min. After fluorescent staining, each compartment was rinsed three times with Tris buffer (pH 7.4; 0.05 M). The chambers of the CultureSlides were then removed, and the remaining slides were examined using a BX51 fluorescence microscope (excitation wavelength, ∼490 nm; detection wavelength, ∼510 nm) equipped with a DP20 camera (Olympus Optical, Tokyo, Japan). Pictures of each slide were taken at magnifications of ×100 and 400×, and the cell density of each strain on each food-conditioning film was analyzed visually and given cell density scores from 0 (no cells present) through 1 (very low), 2 (low), 3 (moderate), and 4 (high) to 5 (very high) by three separate individuals using visual scoring standards (see Fig. 3A). The experiment was replicated twice, and mean scores were calculated and analyzed statistically using analysis of variance (ANOVA) with Minitab software (version 16.0; Minitab, State College, PA). Pairwise comparisons were made by using Tukey's least significant difference test (α = 0.05).
Fig. 3.
(A) Epifluorescence photomicrographs showing different cell densities of L. monocytogenes on food-conditioning films. The number in the upper right corner of each picture indicates the cell density score, with 0 indicating absence of cells (turkey, no cells added) and 1 indicating very small (glass only, no food-conditioning film), 2 indicating small (hot dog with strain J1703), 3 indicating moderate (hot dog with strain OB020790), 4 indicating large (chicken with strain J1703), and 5 indicating very large (turkey with strain 08-5923) amounts of cells observed on the slides. Bars, 20 μm. (B) Epifluorescence photomicrographs showing the degradation of food-conditioning films and biofilm formation by the ECV strain (08-5923). Red arrows indicate undegraded food-conditioning films, and yellow arrows indicate biofilm formation. Bars, 40 μm.
Sequence analysis of comK prophage in 4b and 1/2a strains.
Sequence analyses were conducted on the comK prophage regions in the sequenced genomes of L. monocytogenes serotype 1/2a and 4b using BLAST, PSI_BLAST (1), and BLAST 2 (98). Homology searches were performed with both DNA and protein sequences with various levels of stringency to detect genes that have 100% sequence identity within an outbreak clone and unique genes within a serotype. BLAST searches for phage genes were also carried out on comK prophage sequences of ECII, ECIII, and ECV against the NCBI GenBank database.
Nucleotide sequence accession numbers.
Gene sequences were deposited into GenBank under accession numbers JF791254 to JF791319.
RESULTS
ECII PCR.
Positive amplifications of the ECII marker (19) indicated that all isolates from FSIS and the work of Eifert et al. (32) were ECII (Table 1).
Table 1.
PCR amplification results for ECII PCR, outbreak clone PCR, and junction fragment PCR of L. monocytogenes isolates from different processing plants
| Processing plant | ECII PCR | ECII outbreak clone PCR |
Junction fragment PCR |
||
|---|---|---|---|---|---|
| 2426_1998 primer pair | 2426_2002 primer pair | Upstream | Downstream | ||
| A | + | + | − | + | + |
| B | + | − | + | + | + |
| C | + | − | + | + | + |
| D | − | − | − | + | + |
| E | − | − | − | + | + |
| F | + | + | − | + | + |
| G | + | + | − | + | + |
| H | + | − | − | + | + |
| I | + | − | + | + | + |
| J | + | + | − | + | + |
| K | + | − | − | + | + |
| L | + | − | − | + | + |
| M | + | − | − | + | + |
| N | + | + | + | + | + |
| O | + | + | − | + | + |
MVLST.
Isolates from FSIS and the work of Eifert et al. (32) had the same MVLST sequence type (virulence type 19 [VT19]) as that of ECII (21). MVLST analysis of the two fully sequenced 2008 Canada outbreak isolates (ECV) revealed that they had an MVLST sequence type that was distinct from that of ECIII (VT1) or any other strain in our collection, and thus, they were classified into their own unique virulence type (VT59) in serotype 1/2a in lineage II (Knabel et al., unpublished).
Outbreak clone PCR.
Two primer pairs (2426_1998 and 2426_2002) (see Table S2 in the supplemental material) targeting the prophage region LMOh7858_2426 were able to differentiate between the 1998-1999 and 2002 outbreak clones of ECII based on the sizes of the amplified products (20). Isolates from plants F, G, J, and O had only the 1998-1999 ECII outbreak clone profile, the isolate from plant I had the 2002 ECII outbreak clone profile, and isolates from plant N had both the 1998-1999 and 2002 ECII outbreak clone profiles (Table 1). However, isolates from plant H (see Fig. S2) and plants K, L, and M (data not shown) showed no amplification with either of the above primer pairs.
Amplification of upstream and downstream junction fragments.
PCR experiments targeting the upstream and downstream junction fragments were conducted to determine whether or not the comK prophage was absent in isolates that failed to amplify using the above prophage PCR assay. Positive amplifications of the downstream junction fragment were seen in all isolates tested, including isolates from plant H (see Fig. S3 in the supplemental material), which as mentioned above had previously shown no amplification with prophage primer pair 2426_1998 or 2426_2002. Amplification of the upstream and downstream junction fragments was observed among all isolates analyzed in the present study using modified primer pairs. Sequencing results for ECII isolates from plants K and M (both from New York) and plant L (from New Jersey) indicated that the comK prophage was present. However, more than one product was amplified using these junction fragment primer pairs (data not shown), even when cycling conditions were modified and alternative primer pairs targeting upstream and downstream junction fragments were employed. Therefore, these isolates were not included in the present study.
Cluster diagrams were constructed based on the upstream and downstream junction fragment sequences (Fig. 2 A and B). Concatenated junction fragment sequences gave the same tree resolution as did the downstream junction fragment, and so isolates with identical downstream junction fragment sequences were assigned to the same prophage type (PT). The present study identified nine prophage types (Fig. 2), one each for ECIII and ECV and seven PTs associated with ECII isolates from meat and poultry processing plants in the United States. Of the seven ECII PTs, five PTs were unique to individual processing plants and two were found in multiple plants (Fig. 2; Table 2). In the latter cases, PT2 was found in three different processing plants and PT6 was found in two different processing plants (Fig. 2A and B). Both upstream and downstream junction fragments indicated that PT2 included isolates from plant B in Pennsylvania, plant C in New Jersey, and plant I in Indiana. Interestingly, plants B and C both produced the same type of food, RTE turkey deli meat. The upstream and downstream junction fragments indicated that PT6 contained isolates from plant G in Pennsylvania and one isolate from plant O (state information not available; Fig. 2). Similar to the situation above with PT2 isolates, both plant O (S. Kathariou, unpublished data) and plant G (FSIS, unpublished data) processed the same type of food; however, in this case it was raw turkey. Single nucleotide polymorphisms (SNPs) within the junction fragments that differentiated prophage types of L. monocytogenes were identified (Table 3; see also Table S3 in the supplemental material).
Table 2.
Virulence types, allelic profiles of upstream and downstream junction fragments, and prophage types (PTs) of isolates analyzed in this study
| Processing plant | Identification no. | MVLST virulence type | Junction fragment allelic profile |
Prophage type | |
|---|---|---|---|---|---|
| Upstream | Downstream | ||||
| A | H7858 | 19 | 1 | 1 | 1 |
| A | H7557 | 19 | 1 | 1 | 1 |
| B | J1816 | 19 | 2 | 2 | 2 |
| B | J1815 | 19 | 2 | 2 | 2 |
| C | J1925 | 19 | 2 | 2 | 2 |
| J1703 | 19 | 2 | 2 | 2 | |
| D | N3-031 | 1 | 3 | 3 | 3 |
| D | J1-101 | 1 | 3 | 3 | 3 |
| D | R2-603 | 1 | 3 | 3 | 3 |
| D | R2-499 | 1 | 3 | 3 | 3 |
| E | 08-5578 | 59 | 4 | 4 | 4 |
| E | 08-5923 | 59 | 4 | 4 | 4 |
| F | OB040119 | 19 | 5 | 5 | 5 |
| F | OB050272 | 19 | 5 | 5 | 5 |
| F | OB050273 | 19 | 5 | 5 | 5 |
| G | OB050226 | 19 | 6 | 6 | 6 |
| G | OB050347 | 19 | 6 | 6 | 6 |
| G | OB050350 | 19 | 6 | 6 | 6 |
| G | OB050351 | 19 | 6 | 6 | 6 |
| G | OB050355 | 19 | 6 | 6 | 6 |
| G | OB070122 | 19 | 6 | 6 | 6 |
| H | OB020621 | 19 | 5 | 7 | 7 |
| H | OB020790 | 19 | 5 | 7 | 7 |
| I | OB030029 | 19 | 2 | 2 | 2 |
| J | OB070181 | 19 | 7 | 8 | 8 |
| N | 1493 | 19 | 7 | 9 | 9 |
| N | 1495 | 19 | 7 | 9 | 9 |
| N | 1496 | 19 | 7 | 9 | 9 |
| N | 1498 | 19 | 7 | 9 | 9 |
| N | 1503 | 19 | 7 | 9 | 9 |
| N | 1506 | 19 | 7 | 9 | 9 |
| N | 1513 | 19 | 7 | 9 | 9 |
| N | 1514 | 19 | 7 | 9 | 9 |
| N | 1516 | 19 | 7 | 9 | 9 |
| N | 1117 | 19 | 6 | 6 | 6 |
Table 3.
Summary of SNPs within different regions of the upstream and downstream junction fragmentsa
| Region within junction fragment | No. of SNPs |
|
|---|---|---|
| Total | Nonsynonymous | |
| Upstream junction fragment | ||
| comK′ (RBS) | 1 | NA |
| comK′ | 2 | NA |
| Noncoding (prophage) | 52 | NA |
| HP1 (prophage) | 55 | 22 |
| HP2 (prophage) | 14 | 7 |
| Downstream junction fragment | ||
| Int (prophage) | 65 | 13 |
| Noncoding (prophage) | 9 | NA |
| ′comK | 3 | NA |
NA, not applicable because these are noncoding regions/pseudoregions. RBS, ribosomal binding site.
Recombination analysis.
Possible recombination events in the upstream and downstream fragments were tested with the RDP v.3.38 program. Three putative recombination events were identified in the downstream junction fragment. Among them, only two putative recombination events (prophage types PT5, PT6, and PT8) were supported by at least four methods (P ≤ 10−3) (see Fig. S4 in the supplemental material). Four putative recombination events were detected in the upstream junction fragment, and two of these recombination events in PT4 and PT6 were supported by five methods implemented in the RDP program with high statistical significance (P ≤ 10−5).
Detection of spontaneous induction of the comK prophage.
Positive amplifications with attB primer pairs were seen in all 12 isolates tested (see Fig. S4 in the supplemental material; also data not shown). Positive amplifications with attP primer pairs were seen in nine of the isolates tested but not in isolates OB070181, OB080398, and OB080567 (Fig. S4; also data not shown). Comparison of the attP and attB sequences from strains H7858 and N3-031 with the corresponding upstream and downstream junction fragment sequences from these same strains confirmed that the fragments amplified were the attP site within phage A118 and the attB attachment site within comK.
Detection of int in L. monocytogenes genomic DNA and comK phage DNA by PCR.
Amplification of int in genomic DNA was observed in L. monocytogenes strains H7858 (ECII), N3-031 (ECIII), and 1001::A118 but not in F2365 (ECI) (Table 4). Amplification of int in comK phage DNA was observed in L. monocytogenes strains N3-031 and 1001::A118 but not in F2365 or H7858 (Table 4). Amplicons were sequenced to confirm that int was amplified.
Plaque assays.
Phage plaques were observed on plates containing filtrate from L. monocytogenes strain 1001::A118 but were not observed with ECI strain F2365, ECII strain H7858, or ECIII strain N3-031 (Table 4).
Epifluorescence microscopy.
The effects of strain, type of RTE food-conditioning film, and their interaction on final cell density were all highly significant (P < 0.001). Tukey's multiple comparison revealed that strains lacking the comK prophage (ECI and lineage III) showed significantly lower cell densities than did 4b and 1/2a strains containing the comK propahge (P < 0.05). The two serotype 1/2a strains containing the comK prophage (ECIII and ECV) produced the highest cell densities (P < 0.05) and frequently formed biofilms on meat- and poultry-conditioning films (Fig. 3 A and B and data not shown). The ECV strain produced very high cell densities and mature biofilms when growing on conditioning films made from all four RTE muscle foods (hot dogs, turkey, ham, and chicken). Compared to other strains, the ECV strain formed unique biofilms, which contained high numbers of cells embedded in web-like extracellular polymeric substances (EPS) with empty spaces within the web-like network (Fig. 3A and B). In contrast, ECV showed very low cell density on conditioning films made from soft cheese. Among all RTE foods tested, chicken breast produced the highest average cell density across all strains (P < 0.05), followed by ham, turkey, hot dog, and soft cheese (Table 5). Food-conditioning films were always visible when cell densities were low or moderate; however, food-conditioning films were never observed at very high cell densities where biofilms were often seen (Fig. 3B). No biofilms were observed in the absence of food-conditioning films (Fig. 3A), even though cells had attached to the glass slides and nutrients were available in TSBYE.
Table 5.
Effects of type of strain, type of food-conditioning film, and their interaction on the cell density of L. monocytogenes on glass CultureSlides after incubation at 30°C for 48 h in TSBYEa
| Strain | PTb | Cell density of strain on FCF |
Strain means for all FCFse | |||||
|---|---|---|---|---|---|---|---|---|
| No-FCF control | Soft cheese | Hot dog | Turkey | Ham | Chicken | |||
| No-L. monocytogenes control | 0 | 0 | 0 | 0 | 0 | 0 | 0 A | |
| Lineage III (W1-111) | NAc | 1.0 | 1.0 | 1.0 | 1.5 | 2.5 | 1.0 | 1.4 B |
| ECI (F2365) | NA | 1.0 | 3.0 | 1.5 | 1.5 | 2.0 | 2.0 | 2.0 BC |
| ECII (J1703) | 2 | 1.0 | 1.5 | 2.5 | 1.0 | 3.0 | 4.0 | 2.4 BC |
| ECII (H7858) | 1 | 1.0 | 1.0 | 2.5 | 4.0 | 3.5 | 3.5 | 2.9 CD |
| ECII (OB020790) | 7 | 1.0 | 3.0 | 3.0 | 3.0 | 2.5 | 4.0 | 3.1 CDE |
| ECIII (N3-031) | 3 | 1.0 | 3.5 | 3.5 | 4.5 | 4.0 | 3.5 | 3.8 DE |
| ECV (08-5923) | 4 | 1.0 | 1.0 | 5.0 | 5.0 | 4.5 | 5.0 | 4.1 E |
| FCF means for all strainsd | 1.0 a | 2.0 ab | 2.7 bc | 2.9 bc | 3.1 bc | 3.3 c | ||
Data in the table are based on two replications of the experiment. The numbers in the table indicate cell densities of different strains present on different food-conditioning films (FCFs), with 0 indicating absence of cells and 1 indicating very small, 2 indicating small, 3 indicating moderate, 4 indicating large, and 5 indicating very large amounts of cells observed on the slides. Fractional numbers are calculated based on the results of two replications.
PT, prophage type.
NA, not applicable. The lineage III strain (W1-111) and ECI strain (F2365) do not contain the comK prophage.
Means of one FCF for all strains are calculated by averaging all the values in the column (except that for the no-L. monocytogenes control) corresponding to that FCF; means in this row that do not share a lowercase letter are significantly different (P < 0.05).
Means of one strain for all FCFs are calculated by averaging all the values in the row (except that for the no-FCF control) corresponding to that strain; means in the column that do not share an uppercase letter are significantly different (P < 0.05).
Sequence analysis of comK prophage of 4b and 1/2a strains.
Comparative genome analysis of L. monocytogenes revealed that the comK prophage from serotypes 1/2a and 4b have four open reading frames (ORFs) (LMOh7858_2410, LMOh7858_2411, LMOh7858_2421, and LMOh7858_2426) that show 100% sequence identity within the 1988 sporadic and 2000 outbreak strains of ECIII (77). comK prophage sequence comparison of serotype 1/2a epidemic clones (ECs) (ECIII and ECV) with genome-sequenced serotype 4b strain (ECII) revealed two ORFs unique to 1/2a ECs that also have 100% sequence identity within ECIII isolates (Fig. 4). Comparison of prophage protein sequences of ECII, ECIII, and ECV against the sequences available in NCBI GenBank revealed that while the majority of prophage genes in ECII (serotype 4b) and ECV (serotype1/2a) are from Listeria phage A006, most prophage genes in ECIII (serotype 1/2a) are from Listeria phage A118 (Fig. 5). In addition, comK prophage genes with homology to genes from Streptococcus and Enterococcus phages were also found in the comK prophage (data not shown).
Fig. 4.
Sequences of putative adaptons within comK prophages in epidemic clone strains of L. monocytogenes show 100% sequence identity within a processing plant over a 12-year period (plant D) but vary between processing plants (plants A and E). The putative adaptons ORFs HP1, HP2, gp15, gp13, and partial int (locus designations LMOh7858_2410, -2411, -2421, -2426, and -2475, respectively) are present in sequenced genomes of 1/2a and 4b serotypes of L. monocytogenes that contain the comK prophage and show 100% sequence identity within 1988 and 2000 ECIII isolates. The putative adaptons ORFs gp27 (LMOf6854_2338) and HP3 (LMOf6854_2375) (arrows shaded gray) indicate those comK prophage genes that are unique to serotype 1/2a strains but are not present in the ECII serotype 4b strain. comK′, N-terminal comK fragment; HP1, hypothetical protein 1; HP2, hypothetical protein 2; gp27, phage gp27 protein; gp15, phage gp15 protein; gp13, major tail protein; HP3, hypothetical protein 3; int, integrase; ′comK, C-terminal comK fragment. Arrows point to the corresponding positions of these loci in the comK prophage in ECII, ECIII, and ECV strains. Blocks (adaptons) with different shading indicate nonidentical sequences, and black blocks indicate 100% sequence identity within ECIII isolates.
Fig. 5.
Comparison of comK prophage genes in 1/2a and 4b serotypes of Listeria monocytogenes with sequenced bacteriophage genomes. Regions that correspond to different bacteriophages are shaded.
DISCUSSION
The backbone genome of L. monocytogenes is highly clonal (22, 72, 81, 82), which is consistent with the fact that L. monocytogenes has the highest degree of purifying selection observed in bacteria (75). Within individual epidemic clones, the backbone genomes are even more conserved (21, 25, 65). For example, Orsi et al. (78) used whole-genome sequence comparison to analyze two 1988 sporadic case isolates and two 2000 outbreak isolates of ECIII that were associated with the same processing plant over a 12-year period and were able to find only 11 total SNPs in the backbone genome between these strains. In contrast, the ∼40-kb comK prophage had 1,274 polymorphic sites that differentiated the 1988 isolates from the 2000 isolates (78). This very large number of polymorphisms in the comK prophage within apparently so short a period of time (12 years or less) was very likely due to extensive recombination (78).
The strong congruence between the upstream and downstream junction fragment sequences and their correlation with specific processing plants in the present study strongly suggested that these prophage types were not evolving randomly. Most meat and poultry processing plants in North America have been in existence for less than 100 years. Therefore, if the different persistent prophage types identified in the present study (Fig. 2) were due to rapid niche-specific adaptation in individual plants, only the comK prophage and not the backbone genome of L. monocytogenes would appear to provide enough sequence variation to account for this rapid adaptation. This is consistent with the concept of rapid evolution of microbial ecotypes (55) being driven by niche-specific selection (31). While it is difficult to predict the genes responsible for this ecological divergence, a DNA sequence-based approach which targets ecologically relevant markers could overcome these challenges. In the present study, ecologically distinct prophage types could be identified by sequencing the comK prophage junction fragments (Fig. 2). The fact that some prophage types were found in isolates from multiple processing plants may be due to a common food source in these plants. To achieve efficiency, many food processing plants run the same type of RTE food over the same packaging lines 16 h per day, 5 to 6 days per week, for months or years. This constant and prolonged exposure to the same or very closely related types of RTE foods may result in strains/ecotypes of L. monocytogenes that have evolved an increased ability to attach to, grow on, and form biofilms on these RTE food-conditioning films. Such strains would persist over other strains that are less fit (55). Initial bacterial cell attachment is critical for the formation of bacterial biofilms (14, 79). In food processing plants, food residues accumulate in enclosed areas that are difficult to clean and sanitize, e.g., slicers, etc., and thus represent harborage sites/reservoirs for L. monocytogenes. Because they contain food and water for extended periods of time, these harborage sites allow bacteria to become established and multiply (2, 23, 99). In summary, a common food in a food processing plant would likely produce a common food-conditioning film in harborage sites throughout the plant (10), which might select for attachment, growth, and biofilm formation by isolates with a common prophage type.
In order to test whether type of strain or type of food-conditioning film has an effect on cell attachment or biofilm formation, we utilized fluorescent microscopy to observe the effect of different food-conditioning films on growth and biofilm formation by different prophage types of L. monocytogenes and also strains of L. monocytogenes that lacked the comK prophage. The results demonstrated the critical role of food-conditioning films on attachment, growth, and biofilm formation by these different strains of L. monocytogenes. All EC strains, except ECII strain J1703, grew well on the respective foods from which they were originally isolated (see Table S1 in the supplemental material). However, it is not quite clear why the ECII strain J1703, which was originally isolated during the 2002 turkey deli outbreak, failed to grow on turkey-conditioning film in the present study. Research is under way in our lab (the Knabel lab) to test whether this ECII strain is capable of forming biofilms on other brands of RTE turkey products. A lineage I ECI strain and a lineage III strain, both of which lacked the comK prophage, showed significantly less growth on meat- and poultry-conditioning films in the present study than did ECII, ECIII, and ECV strains that contained the comK prophage (Fig. 3A; Table 5). This is consistent with a previous study (67) in which strain F2365 (ECI) was shown to be less capable of forming biofilms than were other 4b strains and also 1/2a strains. The different ECII isolates also showed variability in their attachment and growth on food-conditioning films, which might be due to their different comK prophage types. Similarly, Wang et al. (103) demonstrated that cryptic (defective) prophages in E. coli K-12 increase growth and biofilm formation. However, it cannot be excluded that other (currently unidentified) genetic differences may play a role in attachment, growth, and biofilm formation by both L. monocytogenes and E. coli. Interestingly, while ECII, ECIII, and especially ECV showed extensive growth and sometimes biofilm formation on muscle foods, they showed very little if any growth on soft cheese (Table 5). This is consistent with previous reports that RTE meat residues increased attachment and biofilm formation by L. monocytogenes (92), while milk proteins significantly reduced its attachment to various surfaces (106).
Many known bacterial cell surface-associated fitness determinants are located on genomic islands of prophage origin (42, 61). Interestingly, sequence analysis of the comK prophage revealed four open reading frames (ORFs) (Fig. 4) located in the “black block” regions (regions of 100% sequence identity in the comK prophage) of the work of Orsi et al. (78). These ORFs are present in all genome-sequenced 1/2a and 4b strains of L. monocytogenes that contain the comK prophage. In addition, 2 other ORFs, which are unique to genome-sequenced 1/2a strains and not present in the phage A118 genome or ECII, can also be found in these regions (Fig. 4). At present, it is unclear whether these genes or some other prophage genes are responsible for niche-specific adaption. We hypothesize that comK prophage genes might have been modified to help L. monocytogenes adapt to different food processing environments, and so we find it appropriate to tentatively term these genes “adaptons.” This is very similar to E. coli probiotic strain Nissle, where such determinants (similar to adaptons) are located on genomic islands that likely represent defective prophages and include adhesins thought to contribute to its survival and successful colonization of the human gut (42). Putative adaptons in the comK prophage may encode similar adhesins that mediate specific attachment to different food-conditioning films. While such adaptons would probably not fully explain attachment and biofilm formation by L. monocytogenes, as many adhesins are chromosomally encoded (7, 51), they may help to explain the persistence of specific L. monocytogenes prophage types in individual food processing plants that manufacture the same food product over an extended period of time (5, 70, 86) (Fig. 2B).
Given the evidence for extensive recombination within prophages in the present and other studies (3, 20, 78), we speculate that transduction of the entire comK prophage via defective comK phage particles might be mediating this rapid and extensive recombination. Phage-like elements that function solely for mediating gene exchange have been observed previously in the case of gene transfer agents (GTAs) in alphaproteobacteria (59) and Staphylococcus aureus pathogenicity island (SaPI) particles (87, 96). In the present study, the comK prophages in L. monocytogenes ECII and ECIII were shown to be spontaneously inducible, as positive amplifications of attP and attB were seen in all lysogenized isolates analyzed (see Fig. S5 in the supplemental material) (data not shown). The results in Table 4 demonstrate that the comK phage was spontaneously induced from ECIII strains N3-031 and 1001::A118 and that the comK phage genome was packaged within phage particles by these strains. These findings are consistent with those of Loessner et al. (63), who also revealed spontaneous induction of the comK prophage and subsequent phage formation. However, the prophages in L. monocytogenes are fairly stable, as only a small fraction of the population is spontaneously induced (63). Spontaneous induction and formation of defective transducing phages have been seen in various other microorganisms and are a mechanism by which various virulence genes (3) and genomic islands (49, 52), including pathogenicity islands (87), are mobilized into other strains. A recent study with E. coli K-12 showed that a spontaneously induced defective prophage, “e14,” is critical for biofilm formation (103).
Plaque-forming bacteriophages of L. monocytogenes have been reported to be serotype specific due to the different phage receptors on the surface of L. monocytogenes strains of different serotypes (30, 100). However, finding genes from various serotype-specific phages within defective comK prophages in different lineages of L. monocytogenes (Fig. 5) indicates that defective comK prophages recombine between phages/defective phages that infect different lineages and serotypes. In addition, standard phage susceptibility tests are based on plaque formation; however, Chen and Novick (17) in their studies reported that several staphylococcal phages transduced L. monocytogenes but could not form plaques. Spontaneous induction of defective comK phage from a donor cell of L. monocytogenes and subsequent transduction of the entire defective phage genome into a recipient cell that already contains either an infective or a defective comK prophage would likely result in frequent and large-scale homologous recombination. This phenomenon is known to produce rapid reshuffling of gene modules and/or gene sequences within bacterial populations (3, 13, 78) and is consistent with the hypothetical phage-mediated “life cycle” of genomic islands proposed by Dobrindt et al. (28) and the model proposed in this study (Fig. 6). Such a mechanism would be fundamentally different from generalized or specialized transduction, where only a small fraction (∼10−4) of the induced phage are actually transducing phages (62). The proposed mechanism would also be different than the lytic or lysogenic cycles of phage replication, because induction and recombination of defective comK phage would be occurring for the purpose of rapid evolution of L. monocytogenes in the absence of phage propagation and, if so, would likely be under the control of the bacterial host (13, 50).
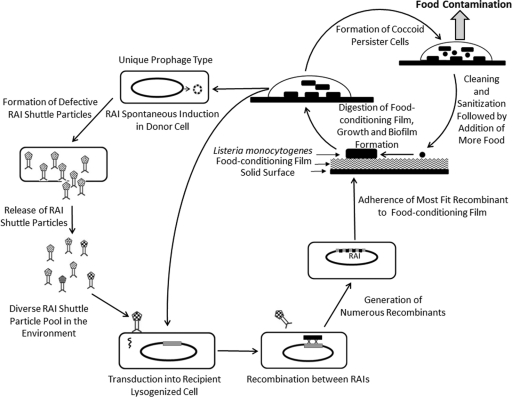
Fig. 6.
Proposed model for rapid niche-specific adaptation and persistence of L. monocytogenes. The cycle starts at the top with spontaneous induction of a rapid adaptation island (RAI) in the donor cell, followed by RAI phage formation and transduction of donor RAI into a recipient cell, which also contains a defective RAI or infective lysogenized phage integrated into its chromosome. Recombination between donor RAI and recipient RAI/comK prophage generates numerous RAI recombinants. Natural selection then acts on RAI recombinants to yield unique persistent prophage types that are adapted to individual processing plants or multiple plants manufacturing the same type of food product and thus produce the same type of food-conditioning film.
The proposed model for high-frequency recombination driving rapid niche-specific adaptation and persistence of L. monocytogenes in food processing plants (Fig. 6) is consistent with what is currently known about defective prophages and how they have been used by various bacteria to rapidly adapt to natural environments (45, 69). If this is also true for L. monocytogenes, we believe that defective comK prophages would be more accurately described as rapid adaptation islands (RAI) (Fig. 6). This RAI concept is consistent with the recent formulation of a novel concept of interphage interactions in defective prophage communities in E. coli O157:H7 (3). The finding that no comK phage DNA of ECII (H7858) was detected after PCR (Table 4) and the predominance of 1/2 phage genes in the comK prophage of a serotype 4 strain (Fig. 5) suggest that serotype 4b strains may act more like recipients than like donors (Fig. 6). Serotype 1/2a strains in lineage II are known to predominate and persist in food processing plants (77), possibly due to their enhanced ability to attach, grow, and form biofilms (9) (Table 5; Fig. 3B). Therefore, they are more likely to serve as donors of comK genes to other strains of L. monocytogenes in lineages I and II, thus allowing them to also adapt to these environments.
The results of the present study and the model presented help explain many of the previously well-known, but not well-understood, observations/phenomena associated with persistent strains of L. monocytogenes found in food processing plants, such as that (i) persistent strains in individual plants have unique genotypes/subtypes; (ii) persistent and prevalent strains are found in harborage sites containing RTE food residues; (iii) serotype 1/2a strains predominate over 4b strains in processing plants and foods; (iv) serotype 4b strains of ECII containing comK prophage are also known to be persistent in food processing plants; (v) ECI and lineage III strains of L. monocytogenes, which are of animal origin (6, 36, 60, 89) and lack the comK prophage (26; X. Deng, personal communication), are typically isolated from raw processing environments (11), raw foods, or RTE foods that have been cross contaminated (21, 24, 39) but not from postpasteurization environments or most pasteurized RTE foods (36); and (vi) RTE meat and poultry products are more frequently contaminated with L. monocytogenes than are most other RTE foods and thus are in the highest risk category for listeriosis. While the results of the present study support the model presented, further research is obviously needed to test the hypothesis that the comK prophage is playing a key role in attachment, growth, and biofilm formation on specific food-conditioning films. Specifically, the comK prophage should be cured (deleted) from those ECII, ECIII, and ECV strains that contain it and these isogenic strains should be compared to wild-type (WT) strains in a food-conditioning film model system to test this hypothesis. These studies are under way in our (Knabel's) laboratory.
The present study also confirms that biofilm formation is more likely to occur in difficult-to-clean harborage sites that contain food and water over extended periods of time (23). For this reason, it is critical to understand and eliminate harborage sites in order to prevent L. monocytogenes contamination of RTE foods. For example, the 2008 Canadian outbreak demonstrated that contaminated harborage sites deep within slicers likely provided long-standing reservoirs for L. monocytogenes, which was thought to have contributed significantly to this outbreak (2). Therefore, preventing L. monocytogenes contamination of RTE food products relies on detecting and eliminating harborage sites by first purchasing equipment that meets sanitary equipment design standards and then implementing a regular deep-cleaning and sanitizing program to remove all food residues to prevent biofilm formation and destroy all remaining L. monocytogenes bacteria.
Further research is needed to test the model proposed in the present study, especially RAI transduction and recombination, and the possible role(s) of putative adaptons in generating persistent prophage types of L. monocytogenes. If the model is confirmed to be accurate, it would lead to a more fundamental understanding of how L. monocytogenes rapidly adapts to and persists in individual food processing plants. It could also help identify the routes by which specific prophage types of L. monocytogenes are transmitted between and within processing plants and retail operations. This information would help food companies implement more effective intervention strategies to prevent persistent and dangerous strains of L. monocytogenes from colonizing food processing environments and thus prevent their transmission to RTE foods.
Supplementary Material
ACKNOWLEDGMENTS
We thank Peter Evans at FSIS, USDA, for identifying and providing PFGE profiles of putative ECII isolates in the FSIS culture collection; Martin Wiedmann at Cornell University for providing ECI and ECIII strains; Bala Swaminathan at CDC for providing ECII strains; and Martin Loessner at ETH, Zurich, Switzerland, for providing strains 1001 and 1001::A118.
S. Knabel was supported by a United States Department of Agriculture Special Grant on Milk Safety to the Pennsylvania State University, and S. Kathariou was supported by a grant from the American Meat Institute Foundation.
Footnotes
Supplemental material for this article may be found at http://aem.asm.org/.
Published ahead of print on 25 March 2011.
REFERENCES
- 1. Altschul S. F., et al. 1997. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25:3389–3402 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Anonymous. 17 April 2009, posting date. Lessons learned: the Canadian Food Inspection Agency's review of Est. 97B (Maple Leaf Consumer Foods Inc.). Canadian Food Inspection Agency, Ottawa, Ontario, Canada [Google Scholar]
- 3. Asadulghani M., et al. 2009. The defective prophage pool of Escherichia coli O157: prophage-prophage interactions potentiate horizontal transfer of virulence determinants. PLoS Pathog. 5:e1000408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Autio T., Keto-Timonen R., Lunden J., Bjorkroth J., Korkeala H. 2003. Characterisation of persistent and sporadic Listeria monocytogenes strains by pulsed-field gel electrophoresis (PFGE) and amplified fragment length polymorphism (AFLP). Syst. Appl. Microbiol. 26:539–545 [DOI] [PubMed] [Google Scholar]
- 5. Autio T., et al. 2002. Similar Listeria monocytogenes pulsotypes detected in several foods originating from different sources. Int. J. Food Microbiol. 77:83–90 [DOI] [PubMed] [Google Scholar]
- 6. Berrang M. E., Meinersmann R. J., Frank J. F., Smith D. P., Genzlinger L. L. 2005. Distribution of Listeria monocytogenes subtypes within a poultry further processing plant. J. Food Prot. 68:980–985 [DOI] [PubMed] [Google Scholar]
- 7. Bierne H., Cossart P. 2007. Listeria monocytogenes surface proteins: from genome predictions to function. Microbiol. Mol. Biol. Rev. 71:377–397 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Blatter S., Giezendanner N., Stephan R., Zweifel C. 2010. Phenotypic and molecular typing of Listeria monocytogenes isolated from the processing environment and products of a sandwich-producing plant. Food Control 21:1519–1523 [Google Scholar]
- 9. Borucki M. K., Peppin J. D., White D., Loge F., Call D. R. 2003. Variation in biofilm formation among strains of Listeria monocytogenes. Appl. Environ. Microbiol. 69:7336–7342 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Boulané-Petermann L. 1996. Processes of bioadhesion on stainless steel surfaces and cleanability: a review with special reference to the food industry. Biofouling 10:275–300 [DOI] [PubMed] [Google Scholar]
- 11. Bowden G. H., Li Y. H. 1997. Nutritional influences on biofilm development. Adv. Dent. Res. 11:81–99 [DOI] [PubMed] [Google Scholar]
- 12. Brooks J. D., Flint S. H. 2008. Biofilms in the food industry: problems and potential solutions. Int. J. Food Sci. Technol. 43:2163–2176 [Google Scholar]
- 13. Brussow H., Canchaya C., Hardt W. D. 2004. Phages and the evolution of bacterial pathogens: from genomic rearrangements to lysogenic conversion. Microbiol. Mol. Biol. Rev. 68:560–602 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Busscher H. J., Bos R., van der Mei H. C. 1995. Initial microbial adhesion is a determinant for the strength of biofilm formation. FEMS Microbiol. Lett. 128:229–234 [DOI] [PubMed] [Google Scholar]
- 15. Canchaya C., Proux C., Fournous G., Bruttin A., Brussow H. 2003. Prophage genomics. Microbiol. Mol. Biol. Rev. 67:238–276 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Chatterjee S. S., et al. 2006. Intracellular gene expression profile of Listeria monocytogenes. Infect. Immun. 74:1323–1338 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Chen J., Novick R. P. 2009. Phage-mediated intergeneric transfer of toxin genes. Science 323:139–141 [DOI] [PubMed] [Google Scholar]
- 18. Chen M.-Y. 2010. Towards real-time observation of conditioning film and early biofilm formation under laminar flow conditions using a quartz crystal microbalance. Biochem. Eng. J. 53:121–130 [Google Scholar]
- 19. Chen Y., Knabel S. J. 2007. Multiplex PCR for simultaneous detection of bacteria of the genus Listeria, Listeria monocytogenes, and major serotypes and epidemic clones of L. monocytogenes. Appl. Environ. Microbiol. 73:6299–6304 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Chen Y., Knabel S. J. 2008. Prophages in Listeria monocytogenes contain single-nucleotide polymorphisms that differentiate outbreak clones within epidemic clones. J. Clin. Microbiol. 46:1478–1484 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Chen Y., Zhang W., Knabel S. J. 2007. Multi-virulence-locus sequence typing identifies single nucleotide polymorphisms which differentiate epidemic clones and outbreak strains of Listeria monocytogenes. J. Clin. Microbiol. 45:835–846 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Cheng Y., Siletzky R. M., Kathariou S. 2008. Genomic divisions/lineages, epidemic clones, and population structure, p. 337–358 In Liu D. (ed.), Handbook of Listeria monocytogenes. CRC Press, New York, NY [Google Scholar]
- 23. Chmielewski R. A. N., Frank J. F. 2003. Biofilm formation and control in food processing facilities. Comp. Rev. Food Sci. Saf. 2:22–32 [DOI] [PubMed] [Google Scholar]
- 24. Costerton J. W., Montanaro L., Arciola C. R. 2005. Biofilm in implant infections: its production and regulation. Int. J. Artif. Organs 28:1062–1068 [DOI] [PubMed] [Google Scholar]
- 25. den Bakker H. C., Fortes E. D., Wiedmann M. 2010. Multilocus sequence typing of outbreak-associated Listeria monocytogenes isolates to identify epidemic clones. Foodborne Pathog. Dis. 7:257–265 [DOI] [PubMed] [Google Scholar]
- 26. Deng X., Phillippy A. M., Li Z., Salzberg S. L., Zhang W. 2010. Probing the pan-genome of Listeria monocytogenes: new insights into intraspecific niche expansion and genomic diversification. BMC Genomics 11:500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Diaz P. I., et al. 2006. Molecular characterization of subject-specific oral microflora during initial colonization of enamel. Appl. Environ. Microbiol. 72:2837–2848 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Dobrindt U., Hochhut B., Hentschel U., Hacker J. 2004. Genomic islands in pathogenic and environmental microorganisms. Nat. Rev. Microbiol. 2:414–424 [DOI] [PubMed] [Google Scholar]
- 29. Donlan R. M. 2002. Biofilms: microbial life on surfaces. Emerg. Infect. Dis. 8:881–890 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Dorscht J., et al. 2009. Comparative genome analysis of Listeria bacteriophages reveals extensive mosaicism, programmed translational frameshifting, and a novel prophage insertion site. J. Bacteriol. 191:7206–7215 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Dunn K. A., Bielawski J. P., Ward T. J., Urquhart C., Gu H. 2009. Reconciling ecological and genomic divergence among lineages of Listeria under an “extended mosaic genome concept.” Mol. Biol. Evol. 26:2605–2615 [DOI] [PubMed] [Google Scholar]
- 32. Eifert J. D., et al. 2005. Molecular characterization of Listeria monocytogenes of the serotype 4b complex (4b, 4d, 4e) from two turkey processing plants. Foodborne Pathog. Dis. 2:192–200 [DOI] [PubMed] [Google Scholar]
- 33. Farber J. M., Peterkin P. I. 1991. Listeria monocytogenes, a food-borne pathogen. Microbiol. Rev. 55:476–511 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Filee J., Forterre P., Laurent J. 2003. The role played by viruses in the evolution of their hosts: a view based on informational protein phylogenies. Res. Microbiol. 154:237–243 [DOI] [PubMed] [Google Scholar]
- 35. Folsom J. P., Siragusa G. R., Frank J. F. 2006. Formation of biofilm at different nutrient levels by various genotypes of Listeria monocytogenes. J. Food Prot. 69:826–834 [DOI] [PubMed] [Google Scholar]
- 36. Foxman B., Zhang L., Koopman J. S., Manning S. D., Marrs C. F. 2005. Choosing an appropriate bacterial typing technique for epidemiologic studies. Epidemiol. Perspect. Innov. 2:10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Fretz R., et al. 2010. Update: multinational listeriosis outbreak due to ‘Quargel’, a sour milk curd cheese, caused by two different L. monocytogenes serotype 1/2a strains, 2009-2010. Euro Surveill. 15(16):pii:19543. [PubMed] [Google Scholar]
- 38. Gilmour M. W., et al. 2010. High-throughput genome sequencing of two Listeria monocytogenes clinical isolates during a large foodborne outbreak. BMC Genomics 11:120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Goller C. C., Romeo T. 2008. Environmental influences on biofilm development. Curr. Top. Microbiol. Immunol. 322:37–66 [DOI] [PubMed] [Google Scholar]
- 40. Graves L. M., et al. 2005. Microbiological aspects of the investigation that traced the 1998 outbreak of listeriosis in the United States to contaminated hot dogs and establishment of molecular subtyping-based surveillance for Listeria monocytogenes in the PulseNet network. J. Clin. Microbiol. 43:2350–2355 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Gray M. J., Freitag N. E., Boor K. J. 2006. How the bacterial pathogen Listeria monocytogenes mediates the switch from environmental Dr. Jekyll to pathogenic Mr. Hyde. Infect. Immun. 74:2505–2512 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Grozdanov L., et al. 2004. Analysis of the genome structure of the nonpathogenic probiotic Escherichia coli strain Nissle 1917. J. Bacteriol. 186:5432–5441 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Gueriri I., et al. 2008. The DegU orphan response regulator of Listeria monocytogenes autorepresses its own synthesis and is required for bacterial motility, virulence and biofilm formation. Microbiology 154:2251–2264 [DOI] [PubMed] [Google Scholar]
- 44. Habimana O., Meyrand M., Meylheuc T., Kulakauskas S., Briandet R. 2009. Genetic features of resident biofilms determine attachment of Listeria monocytogenes. Appl. Environ. Microbiol. 75:7814–7821 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Hacker J., Carniel E. 2001. Ecological fitness, genomic islands and bacterial pathogenicity. A Darwinian view of the evolution of microbes. EMBO Rep. 2:376–381 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Hagens S., Loessner M. J. 2007. Bacteriophages of Listeria, p. 265–279 In H. Goldfine and H. Shen (ed.), Listeria monocytogenes: pathogenesis and host response. Springer, New York, NY [Google Scholar]
- 47. Hamon M., Bierne H., Cossart P. 2006. Listeria monocytogenes: a multifaceted model. Nat. Rev. Microbiol. 4:423–434 [DOI] [PubMed] [Google Scholar]
- 48. Harvey J., Gilmour A. 2001. Characterization of recurrent and sporadic Listeria monocytogenes isolates from raw milk and nondairy foods by pulsed-field gel electrophoresis, monocin typing, plasmid profiling, and cadmium and antibiotic resistance determination. Appl. Environ. Microbiol. 67:840–847 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Hatfull G. F. 2008. Bacteriophage genomics. Curr. Opin. Microbiol. 11:447–453 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Hendrix R. W. 2003. Bacteriophage genomics. Curr. Opin. Microbiol. 6:506–511 [DOI] [PubMed] [Google Scholar]
- 51. Jordan S. J., et al. 2008. Listeria monocytogenes biofilm-associated protein (bapL) may contribute to surface attachment of L. monocytogenes but is absent from many field isolates. Appl. Environ. Microbiol. 74:5451–5456 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Juhas M., et al. 2009. Genomic islands: tools of bacterial horizontal gene transfer and evolution. FEMS Microbiol. Rev. 33:376–393 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Kabuki D. Y., Kuaye A. Y., Wiedmann M., Boor K. J. 2004. Molecular subtyping and tracking of Listeria monocytogenes in Latin-style fresh-cheese processing plants. J. Dairy Sci. 87:2803–2812 [DOI] [PubMed] [Google Scholar]
- 54. Kalmokoff M. L., et al. 2001. Adsorption, attachment and biofilm formation among isolates of Listeria monocytogenes using model conditions. J. Appl. Microbiol. 91:725–734 [DOI] [PubMed] [Google Scholar]
- 55. Koeppel A., et al. 2008. Identifying the fundamental units of bacterial diversity: a paradigm shift to incorporate ecology into bacterial systematics. Proc. Natl. Acad. Sci. U. S. A. 105:2504–2509 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Kolenbrander P. E., Jakubovics N. S., Chalmers N. I., Palmer R. J., Jr 2007. Human oral multi-species biofilms: bacterial communities in health and disease, p. 175–193 In Kjelleberg S., Givskov M. (ed.), The biofilm mode of life: mechanisms and adaptation. Horizon Bioscience, Wymondham, Norfolk, United Kingdom [Google Scholar]
- 57. Kumar S., et al. 2009. A study on the effects of some laboratory-derived genetic mutations on biofilm formation by Listeria monocytogenes. World J. Microbiol. Biotechnol. 25:527–531 [Google Scholar]
- 58. Kushwaha K., Muriana P. M. 2009. Adherence characteristics of Listeria strains isolated from three ready-to-eat meat processing plants. J. Food Prot. 72:2125–2131 [DOI] [PubMed] [Google Scholar]
- 59. Lang A. S., Beatty J. T. 2007. Importance of widespread gene transfer agent genes in alpha-proteobacteria. Trends Microbiol. 15:54–62 [DOI] [PubMed] [Google Scholar]
- 60. Linnan M. J., et al. 1988. Epidemic listeriosis associated with Mexican-style cheese. N. Engl. J. Med. 319:823–828 [DOI] [PubMed] [Google Scholar]
- 61. Loeffler J. M., Fischetti V. A. 2006. Lysogeny of Streptococcus pneumoniae with MM1 phage: improved adherence and other phenotypic changes. Infect. Immun. 74:4486–4495 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Loessner M. J., Calendar R. 2006. The Listeria bacteriophages, p. 593–601 In Calendar R. (ed.), The bacteriophages. Oxford University Press Inc., New York, NY [Google Scholar]
- 63. Loessner M. J., Goeppl S., Busse M. 1991. Comparative inducibility of bacteriophage in naturally lysogenic and lysogenized strains of Listeria spp. by u.v. light and mitomycin C. Lett. Appl. Microbiol. 12:196–199 [Google Scholar]
- 64. Loessner M. J., Inman R. B., Lauer P., Calendar R. 2000. Complete nucleotide sequence, molecular analysis and genome structure of bacteriophage A118 of Listeria monocytogenes: implications for phage evolution. Mol. Microbiol. 35:324–340 [DOI] [PubMed] [Google Scholar]
- 65. Lomonaco S., Chen Y., Knabel S. J. 2008. Analysis of additional virulence genes and virulence gene regions in Listeria monocytogenes confirms the epidemiologic relevance of multi-virulence-locus sequence typing. J. Food Prot. 71:2559–2566 [DOI] [PubMed] [Google Scholar]
- 66. Lunden J. M., Miettinen M. K., Autio T. J., Korkeala H. J. 2000. Persistent Listeria monocytogenes strains show enhanced adherence to food contact surface after short contact times. J. Food Prot. 63:1204–1207 [DOI] [PubMed] [Google Scholar]
- 67. Marsh E. J., Luo H., Wang H. 2003. A three-tiered approach to differentiate Listeria monocytogenes biofilm-forming abilities. FEMS Microbiol. Lett. 228:203–210 [DOI] [PubMed] [Google Scholar]
- 68. Martin D., Rybicki E. 2000. RDP: detection of recombination amongst aligned sequences. Bioinformatics 16:562–563 [DOI] [PubMed] [Google Scholar]
- 69. McDaniel L. D., et al. 2010. High frequency of horizontal gene transfer in the oceans. Science 330:50. [DOI] [PubMed] [Google Scholar]
- 70. Møretrø T., Langsrud S. 2004. Listeria monocytogenes: biofilm formation and persistence in food-processing environments. Biofilms 1:107–121 [Google Scholar]
- 71. Morgan G. J., Pitts W. B. 2008. Evolution without species: the case of mosaic bacteriophages. Br. J. Philos. Sci. 59:745–765 [Google Scholar]
- 72. Nelson K. E., et al. 2004. Whole genome comparisons of serotype 4b and 1/2a strains of the food-borne pathogen Listeria monocytogenes reveal new insights into the core genome components of this species. Nucleic Acids Res. 32:2386–2395 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Norton D. M., et al. 2001. Molecular studies on the ecology of Listeria monocytogenes in the smoked fish processing industry. Appl. Environ. Microbiol. 67:198–205 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Norwood D. E., Gilmour A. 1999. Adherence of Listeria monocytogenes strains to stainless steel coupons. J. Appl. Microbiol. 86:576–582 [DOI] [PubMed] [Google Scholar]
- 75. Novichkov P. S., Wolf Y. I., Dubchak I., Koonin E. V. 2009. Trends in prokaryotic evolution revealed by comparison of closely related bacterial and archaeal genomes. J. Bacteriol. 191:65–73 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. Olsen S. J., et al. 2005. Multistate outbreak of Listeria monocytogenes infection linked to delicatessen turkey meat. Clin. Infect. Dis. 40:962–967 [DOI] [PubMed] [Google Scholar]
- 77. Orsi R. H., Bakker H. C., Wiedmann M. 2011. Listeria monocytogenes lineages: genomics, evolution, ecology, and phenotypic characteristics. Int. J. Med. Microbiol. 301:79–96 [DOI] [PubMed] [Google Scholar]
- 78. Orsi R. H., et al. 2008. Short-term genome evolution of Listeria monocytogenes in a non-controlled environment. BMC Genomics 9:539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Palmer J., Flint S., Brooks J. 2007. Bacterial cell attachment, the beginning of a biofilm. J. Ind. Microbiol. Biotechnol. 34:577–588 [DOI] [PubMed] [Google Scholar]
- 80. Pan Y. W., Breidt F., Kathariou S. 2009. Competition of Listeria monocytogenes serotype 1/2a and 4b strains in mixed-culture biofilms. Appl. Environ. Microbiol. 75:5846–5852 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81. Piffaretti J. C., et al. 1989. Genetic characterization of clones of the bacterium Listeria monocytogenes causing epidemic disease. Proc. Natl. Acad. Sci. U. S. A. 86:3818–3822 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Ragon M., et al. 2008. A new perspective on Listeria monocytogenes evolution. PLoS Pathog. 4:e1000146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Renier S., Hebraud M., Desvaux M. 18 November 2010. Molecular biology of surface colonization by Listeria monocytogenes: an additional facet of an opportunistic gram-positive foodborne pathogen. Environ. Microbiol. doi: 10.1111/j.1462-2920.2010.02378.x [DOI] [PubMed] [Google Scholar]
- 84. Rieu A., Weidmann S., Garmyn D., Piveteau P., Guzzo J. 2007. Agr system of Listeria monocytogenes EGD-e: role in adherence and differential expression pattern. Appl. Environ. Microbiol. 73:6125–6133 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Rorvik L. M., Aase B., Alvestad T., Caugant D. A. 2003. Molecular epidemiological survey of Listeria monocytogenes in broilers and poultry products. J. Appl. Microbiol. 94:633–640 [DOI] [PubMed] [Google Scholar]
- 86. Rorvik L. M., Aase B., Alvestad T., Caugant D. A. 2000. Molecular epidemiological survey of Listeria monocytogenes in seafoods and seafood-processing plants. Appl. Environ. Microbiol. 66:4779–4784 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87. Ruzin A., Lindsay J., Novick R. P. 2001. Molecular genetics of SaPI1—a mobile pathogenicity island in Staphylococcus aureus. Mol. Microbiol. 41:365–377 [DOI] [PubMed] [Google Scholar]
- 88. Sauders B. D., et al. 2004. Distribution of Listeria monocytogenes molecular subtypes among human and food isolates from New York State shows persistence of human disease-associated Listeria monocytogenes strains in retail environments. J. Food Prot. 67:1417–1428 [DOI] [PubMed] [Google Scholar]
- 89. Schlech W. F., et al. 1983. Epidemic listeriosis—evidence for transmission by food. N. Engl. J. Med. 308:203–206 [DOI] [PubMed] [Google Scholar]
- 90. Senczek D., Stephan R., Untermann F. 2000. Pulsed-field gel electrophoresis (PFGE) typing of Listeria strains isolated from a meat processing plant over a 2-year period. Int. J. Food Microbiol. 62:155–159 [DOI] [PubMed] [Google Scholar]
- 91. Shi X. M., Zhu X. N. 2009. Biofilm formation and food safety in food industries. Trends Food Sci. Technol. 20:407–413 [Google Scholar]
- 92. Somers E. B., Wong A. C. L. 2004. Efficacy of two cleaning and sanitizing combinations on Listeria monocytogenes biofilms formed at low temperature on a variety of materials in the presence of ready-to-eat meat residue. J. Food Prot. 67:2218–2229 [DOI] [PubMed] [Google Scholar]
- 93. Stoodley P., Sauer K., Davies D. G., Costerton J. W. 2002. Biofilms as complex differentiated communities. Annu. Rev. Microbiol. 56:187–209 [DOI] [PubMed] [Google Scholar]
- 94. Swaminathan B., Gerner-Smidt P. 2007. The epidemiology of human listeriosis. Microbes Infect. 9:1236–1243 [DOI] [PubMed] [Google Scholar]
- 95. Takahashi H., et al. 2009. Biofilm formation ability of Listeria monocytogenes isolates from raw ready-to-eat seafood. J. Food Prot. 72:1476–1480 [DOI] [PubMed] [Google Scholar]
- 96. Tallent S. M., Langston T. B., Moran R. G., Christie G. E. 2007. Transducing particles of Staphylococcus aureus pathogenicity island SaPI1 are comprised of helper phage-encoded proteins. J. Bacteriol. 189:7520–7524 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Tamura K., Dudley J., Nei M., Kumar S. 2007. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24:1596–1599 [DOI] [PubMed] [Google Scholar]
- 98. Tatusova T. A., Madden T. L. 1999. BLAST 2 Sequences, a new tool for comparing protein and nucleotide sequences. FEMS Microbiol. Lett. 174:247–250 [DOI] [PubMed] [Google Scholar]
- 99. Tompkin R. B. 2002. Control of Listeria monocytogenes in the food-processing environment. J. Food Prot. 65:709–725 [DOI] [PubMed] [Google Scholar]
- 100. Tran H. L., Fiedler F., Hodgson D. A., Kathariou S. 1999. Transposon-induced mutations in two loci of Listeria monocytogenes serotype 1/2a result in phage resistance and lack of N-acetylglucosamine in the teichoic acid of the cell wall. Appl. Environ. Microbiol. 65:4793–4798 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101. Unnerstad H., et al. 1996. Prolonged contamination of a dairy with Listeria monocytogenes. Neth. Milk Dairy J. 50:493–499 [Google Scholar]
- 102. Van Houdt R., Michiels C. 2010. Biofilm formation and the food industry, a focus on the bacterial outer surface. J. Appl. Microbiol. 109:1117–1131 [DOI] [PubMed] [Google Scholar]
- 103. Wang X., et al. 2010. Cryptic prophages help bacteria cope with adverse environments. Nat. Commun. 1:147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104. Ward T. J., et al. 2010. Molecular and phenotypic characterization of Listeria monocytogenes from FSIS surveillance of RTE foods and processing facilities. J. Food Prot. 73:861–869 [DOI] [PubMed] [Google Scholar]
- 105. Williams S. K. 2011. Molecular ecology of Listeria monocytogenes and other Listeria species in small and very small ready-to-eat meat processing plants. J. Food Prot. 74:63–77 [DOI] [PubMed] [Google Scholar]
- 106. Wong A. C. 1998. Biofilms in food processing environments. J. Dairy Sci. 81:2765–2770 [DOI] [PubMed] [Google Scholar]
- 107. Zameer F., Gopal S., Krohne G., Kreft J. 2010. Development of a biofilm model for Listeria monocytogenes EGD-e. World J. Microbiol. Biotechnol. 26:1143–1147 [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.