Abstract
The current consensus model for the circadian clock in mammals is based on a transcription-translation feedback loop. In this model, CRY and PER proteins repress their own transcription by suppressing the transactivator function of the CLOCK:BMAL1 heterodimer directly (physical model) and by facilitating post-translational modifications (chemical model). Most of the data for this model come from genetic and cell biological experiments. Here, we have purified all of the core clock proteins and performed in vitro and in vivo biochemical experiments to test the physical model. We find that CLOCK:BMAL1 binds to an E-box sequence in DNA and that CRY binds stably to the CLOCK:BMAL1:E-box ternary complex independently of PER. Both CRY and PER bind to CLOCK and BMAL1 off DNA but, in contrast to CRY, PER does not bind to the CLOCK:BMAL1:E-box complex. Unexpectedly, PER actually interferes with the binding of CRY to the CLOCK:BMAL1:E-box ternary complex. CRY likely destabilizes the CLOCK:BMAL1 heterodimer on DNA by a post-translational mechanism after binding to the complex. These findings support some aspects of the canonical model, but also suggest that some key features of the model need to be revised.
Keywords: Chromatin Immunoprecipitation (ChiP), DNA-Protein Interaction, Protein-Protein Interactions, Transcription Factors, Transcription Repressor, Circadian, Cryptochrome, Period
Introduction
The circadian clock is the internal timekeeping system that generates a daily rhythm in physiology and behavior of an organism (1–3). In mice and humans this rhythmic behavior is generated by a molecular clock with a periodicity of about 24 h that consists of a transcription-translation feedback loop (TTFL).2 The core mammalian molecular clock is made up of 4 genes/proteins and their paralogs: Clock (Npas2), Bmal1, Cry (Cry1 and Cry2), and Per (Per1 and Per2), and according to the current consensus model, operates as follows: the BMAL1 and CLOCK proteins, which are members of the bHLH (basic helix-loop-helix) family of transcription factors, make a heterodimer, which binds to the E-box element in the promoters of Per and Cry genes and activates their transcription (positive arm). The CRY and PER proteins make heterodimeric complexes, which, after a time lag enter the nucleus and inhibit the CLOCK:BMAL1-activated transcription of their own genes (negative arm). This core circuitry is consolidated by additional interlocking transcriptional circuits as well as post-translational modifications and proteolytic degradation of the core clock proteins (1, 3, 4).
The TTFL model is largely based on genetic data from mice with mutations in the so-called core clock genes, on protein-protein interaction data, reporter gene assays, and ChIP analysis (5–10). Various studies have led to several models for how CRY and PER repress their own transcription as well as the transcription of output genes controlled by the transactivation function of CLOCK:BMAL1. These, not necessarily mutually exclusive models, include: 1) physical (steric) models and 2) chemical (catalytic) models. In the steric model, binding of CRY, PER, or the PER:CRY complex to CLOCK and BMAL1 interferes with their transcription function (10). In the catalytic model, the “repressor” is actually a protein that recruits enzymes for post-translational modification of the activator complex and eventualy changes its activity (11–13). There are numerous versions of both the chemical and physical models, and quite likely both mechanisms contribute to the generation of robust circadian rhythmicity at the transcriptional level.
Here we have used a biochemical approach to investigate the mechanisms by which the two proteins in the negative arm of the mammalian TTFL model, CRY and PER, may interfere with the transactivator function of CLOCK and BMAL1. Specifically, we have addressed the following questions: (a) do CRY or PER interfere with the formation of the CLOCK:BMAL1 complex as has been suggested for zebrafish CRY1a (14) or do they inhibit CLOCK:BMAL1 from binding to its target sequence by shifting the equilibrium to the unbound form as has been suggested for Drosophila PER (15, 16); (b) do both CRY and PER interact with CLOCK and BMAL1 on and off DNA; (c) during repression of CLOCK-BMAL1, does PER bring CRY to the CLOCK:BMAL1:E-box complex (17) or is it the other way around? We find that both CRY and PER bind to CLOCK-BMAL1 off DNA. Unexpectedly, and not predicted by any of the current models, only CRY binds to the CLOCK:BMAL1:DNA ternary complex. Neither the PER:CRY heterodimer nor PER alone can bind to CLOCK:BMAL1:DNA and, in fact, at a sufficiently high concentration PER interferes with the binding of CRY to this complex. These in vitro data were supported by ChIP analysis with various mouse clock gene knock-out cell lines, which revealed that binding of CRY to the cognate promoters is dependent on BMAL1 but independent of PER. In light of these findings we propose a revised TTFL model for the mammalian circadian clock.
EXPERIMENTAL PROCEDURES
Cells and Antibodies
The Cry1−/− Cry2−/− mouse skin fibroblasts have been described previously (18). The isogenic Bmal1+/+ and Bma1−/− and Clock+/+ and Clock−/− mouse embryonic fibroblast cell lines (19) were kindly provided by Dr. Marina Antoch (Roswell Park Cancer Institute). The Per1−/− Per2−/− skin fibroblasts were prepared from a male double knock-out mouse (20) kindly provided by Dr. Cheng Chi Lee (University of Texas, Houston Medical School). Mouse embryonic fibroblasts expressing mCry1-FLAG and mPer2-FLAG were made by retrovirus infection. Briefly, mCry1-Flag-pBABEpuro and mPer2-Flag-pBABEpuro constructs were made and co-transfected together with pVSVG and pCIHPZ into HEK293T to produce retrovirus particles. Then mouse embryonic fibroblast cells were infected with the retrovirus and transfectants were selected in medium containing puromycin for 2 weeks. Single colonies were picked and cultured for subsequent analysis.
Anti-mCRY1 (IgM type monoclonal) and anti-CRY2 antibodies were described previously (18, 21). Anti-CLOCK and anti-BMAL1 antibodies were purchased from Santa Cruz and Bethyl Laboratories. Rabbit anti-PER2 antibodies were generated by using the N-terminal 200 amino acids of mPER2 protein as immunogen as described previously (22) and immunopurified by antigen that was covalently coupled to AminoLink resin (Pierce). Anti-Myc(9E10), anti-V5, anti-FLAG, and anti-His antibodies were purchased from Santa Cruz, Invitrogen, Sigma, and Abgent, respectively.
Plasmids
Plasmids used in this work were obtained from Addgene. Detailed construct cloning information is described under supplemental “Methods”. The hCry1-His baculovirus construct was described previously (23).
Clock Proteins
Proteins were purified either from baculovirus-infected Sf21 insect cells or from transfected HEK293T cells. Most of the proteins used in our experiments were purified with the baculovirus system, except Myc-CRY1 and Myc-CRY2, which were purified from the mammalian expression system. Proteins purified using the two systems showed similar properties. Detailed purification procedures are described under supplemental “Methods”. MAX (MYC-associated factor X) protein used in some gel mobility shift assays as a control was purchased from ProteinOne, USA. Drosophila CRY was generated as described elsewhere (24).
Electrophoretic Mobility Shift Assay
Two DNA duplexes were used as DNA binding substrates: a 14-bp duplex “M34” (25) and a 60-bp duplex “P2GS.” Sequences of these oligos are listed in supplemental Table S1. To prepare radiolabeled probes, 40 pmol of M34TOP or P2GSA were labeled with [γ-32P]ATP using polynucleotide kinase (New England Biolabs). DNA was then extracted with phenol/chloroform, and the labeled oligonucleotides were annealed with complementary strands (M34BTM and P2GS). Mini-quick spin oligo columns (Roche Applied Science) were used to remove the free ATP from labeled DNA. In EMSA experiments, the binding buffer consisted of 50 mm Tris-HCl, pH 8.0, 100 mm KCl, 1 mm EDTA, 6 mm dithiothreitol, and 20 μg/ml of poly(dI-dC) (Sigma). Radiolabeled DNA at ∼1 nm was mixed in 15 μl with clock proteins in the following order depending on the experiments: 1) CLOCK:BMAL1 for 10 min, 2) CLOCK:BMAL1 for 5 min, then CRY, PER, or hCRY1:PER2 protein for 8 min, or 3) CLOCK:BMAL1 for 5 min, then CRY for 8 min, and PER for 8 min. All reactions were incubated at 22 °C followed by 10 min at 4 °C. In antibody supershift experiments, about 1 μg of antibody was added for an additional 5 min after the original incubation at 22 °C. The reactions were loaded onto 4% nondenaturing polyacrylamide gels, and products were resolved at 100 V for 3 h at 4 °C. We note that under these reaction conditions CRY in isolation does not exhibit any DNA binding activity (Fig. 1D). Gels were dried and exposed to x-ray film.
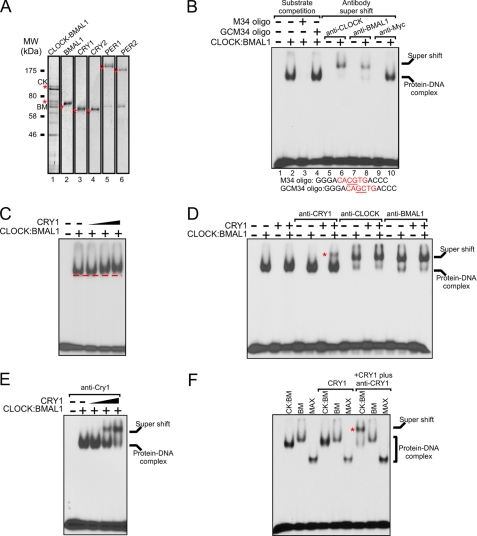
FIGURE 1.
Ternary complex of CRY:CLOCK:BMAL1 on an E-box. A, Coomassie Blue staining of SDS-polyacrylamide gels of mouse core clock proteins. The locations of the clock proteins are marked by asterisks to differentiate from nonspecific contaminants present in some of the preparations. When co-expressed, CLOCK and BMAL1 both become partially phosphorylated, and the phosphorylated and non-phosphorylated forms of each protein were run in the gel as doublets. The phosphorylated form of each can be converted to the non-phosphorylated form by incubation with λ-phosphatase (data not shown). B, electrophoretic mobility shift assay with the CLOCK:BMAL1 heterodimer (0.2 nm). The target was a 32P-labeled 14-bp duplex M34 with the E-box sequence (1 nm). The specificity of the complexes was confirmed by competition with a 100-fold excess of non-radioactive wild-type (M34) or mutant duplexes (GCM34) and by supershifting the DNA-protein complexes with the appropriate antibodies. Substrate sequences (one strand of each duplex) are listed under the gel. C, effect of CRY on the mobility of the CLOCK:BMAL1:E-box complex. The E-box duplex (1 nm) was incubated with fixed amounts of CLOCK:BMAL1 dimer (0.2 nm) and increasing amounts of CRY1 (0.1, 0.3, and 1 nm) as indicated. A line parallel to the leading edge of the CLOCK:BMAL1:E-box complex is drawn to indicate the subtle change in mobility of the protein-DNA band upon addition of CRY1. D, antibody supershift in the presence of various combinations of CRY1 (0.2 nm) and CLOCK:BMAL1 dimer (0.2 nm). The mobility of the supershifted CRY:CLOCK:BMAL1:E-box complex is indicated with an asterisk. E, effect of CRY1 concentration on the extent of the CRY1:CLOCK:BMAL1:E-box complex formation. CLOCK and BMAL1 (0.2 nm) were first incubated with the E-box duplex (1 nm). Then increasing amounts of CRY1 (0.1, 0.3, and 1 nm) were added, followed by addition of a constant amount of anti-CRY1 antibody (1 μg) to supershift protein-DNA complexes containing CRY1. F, binding of CRY1 to the CLOCK:BMAL1:E-box is dependent on the presence of CLOCK in the complex. CLOCK:BMAL1 (0.2 nm), BMAL1(2 nm), or MAX (0.5 nm) E-box-binding proteins were incubated with the E-box oligomer followed by CRY1 (1 nm) or CRY1 plus anti-CRY1 antibody. Note that only the CLOCK:BMAL1:E-box complex is supershifted upon addition of CRY1 plus anti-CRY1 (indicated by an asterisk) CK-BM (CLOCK:BMAL1), BM (BMAL1), and MAX.
DNase I Footprinting
Protein/DNA mixtures were prepared as in the electrophoretic mobility shift assay using P2GS DNA. Reactions, in addition to DNA, included: 1) no protein, 2) BMAL1:CLOCK (0.2 nm) alone, or 3) CLOCK:BMAL1 (0.2 nm) plus an amount of CRY (1 nm) that binds all of the CLOCK:BMAL1:E-box complexes as determined by gel shift analysis. Reactions in 15 μl were incubated at 22 °C for 20 min. Then 0.5 units of DNase I (Promega) and MgSO4 (10 mm), CaCl2 (1 mm) were added to each reaction, and after 5 min at 22 °C, digestion was stopped by adding EDTA to 25 mm. Then the reactions were loaded onto a 4% nondenaturing polyacrylamide gel to separate protein-bound and free DNA. Bands representing free DNA were from reaction 1, the CLOCK:BMAL1:E-box complex from reaction 2, and the CRY:CLOCK:BMAL1:E-box complex from reaction 3 were excised from the gel, and DNA was eluted from the gel slices and run on a 10% denaturing polyacrylamide gel that was visualized by autoradiography.
Pulldown Assay for Protein-Protein Interactions
Purified recombinant proteins at 0.1 nm were mixed in 200 μl of binding buffer (50 mm Tris-HCl, pH 7.5, 150 mm NaCl, 100 μg/ml of BSA, 0.05% Nonidet P-40) and incubated at 22 °C for 15 min. Then 20 μl of either FLAG- or V5-agarose (Sigma) beads were added and the mixture was incubated at 4 °C for 90 min. Then, the beads were washed with 1 ml of ice-cold binding buffer 4 times and after the final wash resuspended in 25 μl of SDS loading buffer and heated at 95 °C for 5 min. The proteins were then separated on SDS-PAGE and visualized by immunoblotting.
Chromatin Immunoprecipitation
We used a previously described dual cross-link ChIP procedure (26) with some modifications. Detailed procedures are described under supplemental “Methods”.
siRNA Treatment, RNA Extraction, and Reverse Transcription
siRNAs specific for mCry1, mCry2 (SMARTpool siRNA, L-040485-01-0005, and L-014151-01-0005) and CYCP-B control siRNA (D-001820-02) were purchased from Dharmacon. Transfections were performed with 50 pmol of siRNAs using the transfection reagent RNAi Max (Invitrogen). Following a 48-h transfection the cells were collected and total RNA was extracted using the RNeasy Mini Kit (Qiagen). Total RNA (500 ng) was reverse transcribed with oligo(dT)20 using SuperScript III (Invitrogen) according to the manufacturer's instructions.
Quantitative Real-time RT-PCR
Real-time PCR assays were performed by using an ABI 7300 system (Applied Biosystems) and MaximaTM SYBR Green/ROX qPCR Master Mix (Fermentas). Quantitation of ChIP was performed with the primer sets in supplemental Table S1. Each sample was analyzed in triplicate. To compare ChIP data from different chromatin samples, signals obtained from each chromatin sample were divided by signals obtained from the corresponding input sample and are presented as “% input.”
Quantitation of Dbp mRNA level was performed with the primer sets (mDbpF and mDbpR). Each sample was analyzed in triplicate, and cycle thresholds of individual genes were normalized to the corresponding Gapdh mRNA expression values obtained with the primer set (mGapdhF and mGapdhR). Relative mRNA levels were assessed by defining the value of CYCP-B control RNA treatment as one in each experiment. Sequences of the primers sets used in this study are listed under supplemental Table S1.
RESULTS
Purification of Mouse Clock Proteins
To test the conventional TTFL model that posits that the CLOCK:BMAL1 heterodimer drives the transcription of target genes and that the PER:CRY heterodimer binds to and interferes with the transactivator function of CLOCK:BMAL1, we purified these proteins using baculovirus or mammalian expression systems. Fig. 1A shows the purified full-length proteins analyzed by SDS-PAGE and Coomassie Blue staining. We found that BMAL1, CRY1/2, and PER1/2 could be expressed individually and purified in reasonable quantities using immunoaffinity chromatography. However, when CLOCK was expressed alone, it was somewhat unstable and insoluble as reported previously (27). When coexpressed with BMAL1, CLOCK could be purified in a reasonable yield and with one-to-one stoichiometry, as shown in lane 1. We used both mouse CRY1 and CRY2 and mouse PER1 and PER2 in our experiments as dictated by availability and practical considerations. In addition, we used human CRY1 for the CRY1:PER2 heterodimer and human CRY1 alone as a reference cryptochrome in experiments with this heterodimer. Because most of the experiments were performed with mouse clock proteins we will omit the “m” designation for the sake of simplicity.
Formation of CLOCK:BMAL1:CRY1 Ternary Complex on E-box DNA
Both CRYs and PERs bind to the CLOCK:BMAL1 heterodimer as evidenced by yeast two-hybrid assays, protein-protein pulldown, and co-immunoprecipitation assays (5–7, 9). These findings led to the original TTFL model that proposed that the PER:CRY heterodimer (or a higher order oligomer) binds to the CLOCK:BMAL1 complex on E-boxes of the cognate promoters and suppresses the transactivation function of the complex. In addition, reporter gene assays have led to the conclusion that CRY is the major repressor of CLOCK:BMAL1 by direct interaction with either CLOCK alone or with both CLOCK and BMAL1 in a multiprotein complex (5, 7, 28). However, a recent study that, in addition to reporter gene assays, employed co-immunoprecipitation assay and behavioral analysis of transgenic mice overexpressing PER2 or CRY1 concluded that PER2 directly binds to CLOCK:BMAL1 and brings CRY1 to the complex (17). We wished to test these and similar models by conducting experiments with purified proteins.
First, we tested the binding of CLOCK:BMAL1 to a 14-bp duplex containing an E-box by gel mobility shift assay (25). As seen in Fig. 1B, CLOCK:BMAL1 binds to the E-box specifically and this binding is competed out with 100-fold excess oligomer of the same sequence but not with an oligomer having a mutated sequence (lanes 3 and 4), as reported previously (25). Importantly, the retarded band is supershifted both with anti-CLOCK and anti-BMAL1 antibodies (anti-Myc as negative control), indicating that the protein-DNA complex contains both the CLOCK and BMAL1 proteins (lanes 6 and 8).
Next, we investigated the effect of CRY1 on the CLOCK:BMAL1:E-box ternary complex. In Fig. 1C the complex was incubated with increasing concentrations of CRY1 and then analyzed by gel electrophoresis. Two conclusions emerge from this figure. First, CRY1 does not disrupt the ternary complex under our experimental conditions as evidenced by the intensity of the CLOCK:BMAL1:DNA band in the absence and presence of CRY. Second, in the presence of CRY, in particular at the highest CRY concentration used in this experiment, there is a slight supershift of the protein-DNA complex, suggesting that CRY associates with CLOCK:BMAL1 on DNA. To further examine this association, we attempted to supershift the presumed CRY:CLOCK:BMAL1:DNA complex with antibodies against each of the 3 proteins. As seen in Fig. 1D the complex is nearly completely supershifted by anti-CLOCK and anti-BMAL1 antibodies and a significant fraction of it was supershifted with anti-CRY antibodies as well. In Fig. 1E we show that with high concentrations of CRY protein, most of the protein-DNA complex is supershifted, indicating that under appropriate assay conditions nearly all of the CLOCK:BMAL1:E-box complex is bound to CRY1. In addition, CRY2 protein, but not Drosophila CRY binds to the CLOCK:BMAL1:E-box complex in our assay (supplemental Fig. S1A). The CLOCK:BMAL1:CRY1 complex co-purified from mammalian cells also showed clear binding activity to the E-box as a ternary complex (supplemental Fig. S1, B and C). To ascertain the specificity of CRY1 binding to CLOCK:BMAL1 on an E-box we performed additional experiments to test effects of CRY on other E-box-binding proteins including the BMAL1 homodimer and the MAX homodimer (29, 30). Fig. 1F shows that the association of CRY1 with the CLOCK:BMAL1:E-box complex is specific, because the mobility of the BMAL1:E-box or the MAX:E-box complexes are not affected by CRY1 or CRY1 plus anti-CRY1 antibody under conditions where the entire CRY1:CLOCK:BMAL1:E-box is supershifted by the anti-CRY1 antibody. These data unambiguously show that the CRY protein does not dissociate CLOCK:BMAL1 from the E-box by physical interaction; in contrast, it binds to the CLOCK:BMAL1:E-box complex with high specificity and without the intermediacy of another protein.
Footprint of CLOCK:BMAL1
The data in Fig. 1 show that under appropriate reaction conditions nearly all of the BMAL1:CLOCK:E-box complexes are associated with CRY1. Yet, even under conditions of saturating concentration of CRY1 only a minor CRY1-induced supershift is detected in the DNA-protein complex. This could be due either to the relatively small molecular mass of CRY1 (≈65 kDa) compared with that of the CLOCK:BMAL1 heterodimer (≈180 kDa) or to the shape and electrostatic charge of the proteins involved. It is well known that the degree of mobility shift of DNA-protein complexes is not linear with the size of the binding protein. Indeed, as seen in Fig. 1F the BMAL1 homodimer:DNA complex migrates slower than the CLOCK:BMAL1-DNA complex even though the latter complex possesses a molecular mass ≈25 kDa more than the former. Hence, we considered the possibility that the CRY1:CLOCK:BMAL1 complex on an E-box, which is clearly detectable only by anti-CRY1 antibody induced supershift in gel retardation experiments, may be more readily detectable by DNase I footprinting, which has higher resolution than the gel retardation assay. For DNase I footprinting, we used a 60-bp fragment P2GS encompassing an E-box in the mouse Per1 promoter, which was shown to be functional in a reporter gene assay (9) (Fig. 2A). In the gel mobility shift assay using this fragment we obtained results similar to those obtained with the 14-bp E-box fragment used in Fig. 1 (Fig. 2B and supplemental Fig. S1D). Importantly, in the presence of anti-CRY1 antibodies, most of the DNA-protein complex was supershifted in the reaction that included CRY1 in the reaction mixture. Interestingly, when these complexes were probed by DNase I footprinting the CLOCK:BMAL1 heterodimer produced a rather small footprint of 8-bp centered around the E-box (Fig. 2C), and this footprint was not affected by CRY1 under conditions known to produce the CRY1:CLOCK:BMAL1 complex on the E-box. Even though this result did not provide further insight on the anatomy of the CLOCK:BMAL1:CRY complex, the footprint is consistent with the known properties of interactions of bHLH family protein with DNA (30). In the complex of BMAL1 and CLOCK, both of which are bHLH-PAS domain proteins, contact with DNA is made with the bHLH domains, which are located at the extreme N terminus of each member of the heterodimer. The extended structure of this family of proteins explains the small footprint and also explains the lack of effect of CRY1 on the footprint because CRYs bind to the PAS domains and the C-terminal domains distant from the DNA contact region (31, 32).
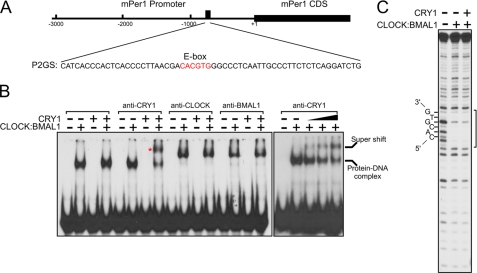
FIGURE 2.
DNase I footprint of clock proteins on an E-box. A, schematic representation of the mPer1 promoter illustrating the relative position and the sequence of one strand of the 60-bp duplex encompassing the mPer1 E-box2. CDS indicates coding DNA sequence. B, EMSA with the 60-bp duplex and supershift of the protein-DNA (0.4 nm CRY and 0.2 nm CLOCK:BMAL1 dimer) complexes with the indicated antibodies. The asterisk identifies the supershifted CRY:CLOCK:BMAL1:E-box complex. C, DNase I footprint of the clock protein-DNA complexes. The position of the E-box is indicated.
Effect of PER on CLOCK:BMAL1:E-box and CRY:CLOCK:BMAL1:E-box Complexes
Although CRY is the dominant repressor of the CLOCK:BMAL1 transactivation function as shown by reporter gene assays (7) and transcription assays with Cry1−/− Cry2−/− knock-out mice (33), numerous studies suggest that CRY works as a heterodimer with PER in the TTFL (3, 22). The coordinated inhibitor function of PER and CRY can be classified by the following different steric models: (a) PER:CRY complex binds to CLOCK:BMAL1 off DNA and disrupts the heterodimer or interferes with its binding to E-boxes in the cognate promoters. (b) PER:CRY binds to CLOCK:BMAL1 through CRY and blocks the transactivation function of the transcription factor (22). (c) The PER:CRY complex binds to CLOCK:BMAL1 by direct binding of PER to the complex, and not the other way around, even though CRY is considered the nominal repressor (17). To differentiate among these models, we investigated the combined effects of PER and CRY on the CLOCK:BMAL1:E-box complex. Because PER2 appears to be more important than PER1 for normal clock function in mice (17, 34), we used PER2 in most of our experiments.
The various models were tested by gel mobility shift assays. Fig. 3A and supplemental Fig. S1E show that, in contrast to CRY1 and CRY2, PER1 and PER2 do not affect the mobility of the CLOCK:BMAL1: E-box complex. However, because even with CRYs the mobility shift is subtle, this finding alone does not prove that PER2 does not associate with the complex. In Fig. 3B we employed the more stringent test of antibody-mediated band supershifting. When Myc-tagged CRY1 and PER2 were added to the CLOCK:BMAL1:E-box reaction mixture, Myc-CRY1 slightly retarded the DNA-protein band, which, importantly, upon the addition of Myc antibodies was nearly completely supershifted. In contrast, Myc-PER2 at the same concentration as Myc-CRY1 had no effect on the mobility of the CLOCK:BMAL1:E-box complex, and most importantly, the addition of anti-Myc antibodies to the Myc-PER2 plus CLOCK:BMAL1:E-box reaction mixture did not affect the mobility of the protein-DNA band. We conclude that under our reaction conditions PER2 does not directly bind to the CLOCK:BMAL1:E-box complex.
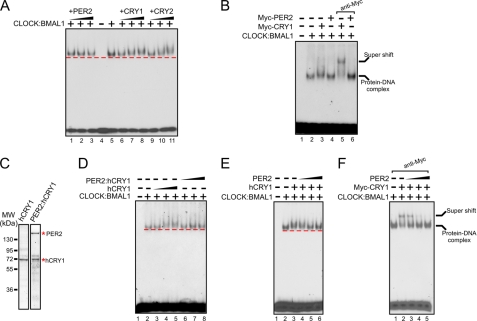
FIGURE 3.
Effect of PER on clock protein-DNA complexes. A, effect of PER2 on the CLOCK:BMAL1:E-box complex. The 14-bp E-box duplex (1 nm) was incubated with CLOCK:BMAL1 (0.2 nm) and then increasing concentrations (0.2, 0.6, and 1.5 nm) of PER2, CRY1, or CRY2 were added, and the protein-DNA complexes were analyzed by EMSA. The dashed line indicates the leading edge of migration of the CLOCK:BMAL1:E-box complex. Note the lack of effect of PER2 on the mobility of this complex (lanes 1–3). In contrast both CRY1 (lanes 6–8) and CRY2 (lanes 9–11) retard the DNA-protein complex. B, analysis of the composition of the protein-DNA bands by antibody supershift. In addition to CLOCK:BMAL1 (0.2 nm) equal concentrations of either Myc-PER2 (1 nm) or Myc-CRY1 (1 nm) were included in the reactions followed by anti-Myc antibodies in the last two lanes. Note the supershift of the protein-DNA band formed in the presence of Myc-CRY1 (lane 5) but not Myc-PER2 (lane 6). C, Coomassie Blue staining of SDS-PAGE showing the hCRY1 and PER2:hCRY1 used in the electrophoretic mobility shift assay. D, effects of hCRY1 and PER2:hCRY1 on CLOCK:BMAL1:E-box complexes. Increasing concentrations (0.1, 0.3, and 1 nm) of hCRY1 or the PER2:hCRY1 complex were added to the reactions as indicated before electrophoresis. Note the supershift caused by hCRY1 (lanes 4 and 5) but not by the PER2:hCRY1 complex (lanes 6–8) containing the same concentrations of hCRY1 as in lanes with hCRY1 alone. E and F, PER2 removes CRY1 from the CRY1:CLOCK:BMAL1:E-box complex. The E-box duplex was incubated with the indicated proteins: 0.2 nm CLOCK:BMAL1 dimer and 0.3 nm hCRY1 (in panel E) or 0.3 nm Myc-CRY1 (in panel F) before separating on a native gel. Note that the presence of hCRY1 causes a slight retardation of the CLOCK:BMAL1:E-box complex (panel E, lane 3), which is reversed by inclusion of increasing concentrations (0.2, 0.6, and 1.2 nm) of PER2 (panel E, lanes 4–6). The effect of PER2 on removing CRY1 from the protein-DNA complex is more clearly shown when the anti-Myc antibody supershifted Myc-CRY1:CLOCK:BMAL1:E-box complex is incubated with increasing concentrations of PER2 (panel F, lanes 3–5).
Failure of PER2 to bind to the CLOCK:BMAL1:E-box was unexpected. We considered the possibility that solitary PER2 may not be properly folded, and that in vivo PER2 might be mostly associated with CRY and hence the PER:CRY complex may represent the functionally relevant form of the protein. With these considerations, we purified the PER2:hCRY1 complex and the hCRY1 as a control (Fig. 3C) and tested them in gel mobility shift assays. The results are shown in Fig. 3D. As expected, CRY1 alone supershifts the CLOCK:BMAL1:E-box complex. Surprisingly, however, the PER2:CRY1 heterodimer does not affect migration of the CLOCK:BMAL1:E-box even though CRY1 alone at a concentration equivalent to its concentration in the PER2:CRY1 heterodimer clearly retards the protein-DNA band. The most parsimonious explanation of these data is that PER2 interferes with binding of CRY1 to the CLOCK:BMAL1:E-box complex. However, because such a conclusion is contrary to the generally held view that PER either mediates the binding of CRY to CLOCK:BMAL1, or is in a complex with CRY that binds to the CLOCK:BMAL1:DNA complex, we were concerned that the lack of binding by PER2:CRY1 might have been caused by an inactive protein preparation. Therefore, to test if indeed PER2 interfered with binding of CRY to the CLOCK:BMAL1:E-box complex we performed gel mobility shift assays to test the effects of PER on a pre-assembled CRY:CLOCK:BMAL1:E-box complex. The results in Fig. 3, E and F, show that supershifting of the CLOCK:BMAL1:E-box complex by CRY is abolished when PER2 is added to the complex as evidenced by the migration of the protein-DNA band without (Fig. 3E) and with (Fig. 3F) antibodies against the affinity tag on CRY1. Based on these findings, we conclude that PER2 does not recruit CRY to CLOCK:BMAL1 bound to the promoter of the cognate genes, but it actually impairs its binding.
In Vitro Interactions Among Purified CLOCK Proteins
Failure to detect an interaction of PER2 with CLOCK:BMAL1 on DNA was surprising because the interactions of PER2 with all clock proteins has been demonstrated by numerous studies (17). Hence, to ascertain whether the PER2 that was used in mobility shift experiments was active, we tested it for protein-protein interactions with the other clock proteins by co-immunoprecipitation of purified proteins. In Fig. 4 we compare the binding of PER2 and CRY1 to each other, and to BMAL1 and CLOCK. As expected, individually purified PER2 and CRY1 interact with one another (Fig. 4A). Significantly, PER2 binds to both BMAL1 and CLOCK with higher affinity than CRY1 (Fig. 4, B and C). Although quantitative binding experiments with recombinant proteins must be considered with some caution, these data nevertheless do show that PER2 used in our mobility shift assays is at least as active as CRY1 in protein-protein interactions with CLOCK and BMAL1. Hence, we conclude that the failure to detect binding of either PER2 or the PER2:CRY1 complex to the CLOCK:BMAL1:E-box ternary complex is not due to the misfolding or an otherwise inactive form of PER2; rather, it is a manifestation of an intrinsic property of the core clock system.
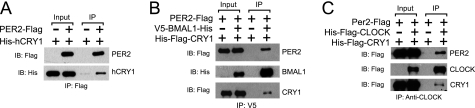
FIGURE 4.
Protein-protein interactions among purified clock proteins in vitro. CLOCK, BMAL1, CRY, and PER protein were individually purified from insect cells. Proteins (0.1 nm) were mixed as indicated, one of the clock proteins was immunoprecipitated and immunoprecipitates were tested for the other interacting proteins by immunoblotting (IB). A, PER2 and CRY1 interaction. B, interaction of BMAL1 with CRY1 and PER2. C, interaction of CLOCK with CRY1 and PER2. The input lanes contain 20% of the proteins used in the immunoprecipitation (IP) reactions.
Effect of Mutations in Core Clock Genes on Binding of Cryptochrome to Target Promoters in Vivo
The in vitro data we have presented indicate that CRY may be recruited to target promoters by direct protein-protein interactions with CLOCK:BMAL1 bound to E-boxes in cognate promoters. To test this prediction in vivo we carried out ChIP assays with CRY and CLOCK antibodies in cell lines from mice in which clock genes have been knocked out (supplemental Fig. S2, B and C). ChIP results are shown in Fig. 5 and supplemental Figs. S3 and S4. Several noteworthy conclusions emerge from ChIP assays with anti-CRY1 antibodies, shown in orange in Fig. 5A. First, in agreement with the in vitro data, CRY1 binds independently of PER1 and PER2 proteins to E-box elements in the promoter regions of both Per1 and Per2 genes. Second, in contrast to the in vitro data, which show that CRY1 binds to the CLOCK:BMAL1:E-box complex but not to the BMAL1:E-box complex (Fig. 1F), the binding of CRY1 to E-box in vivo is independent of CLOCK. However, considering that BMAL1 is capable of making heterodimers with other PAS domain proteins such as NPAS2, this was to be expected (35–37). Indeed, ChIP reveals that binding of CRY1 to both Per1 and Per2 E-boxes is absolutely dependent on BMAL1 (Fig. 5A, right column). These data confirm the conclusion from the in vitro experiments with purified proteins that CRY binds directly to CLOCK:BMAL1 situated on an E-box.
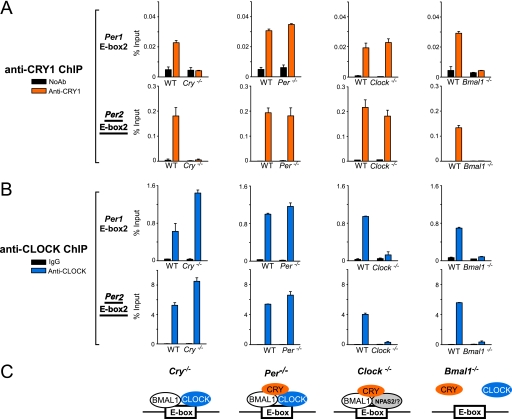
FIGURE 5.
CRY1 binds E-boxes in Per1 and Per2 promoters in the absence of PER protein and affects CLOCK-BMAL1 dynamics at cognate promoters in mouse fibroblasts. Wild-type or clock mutant mouse fibroblasts were used in ChIP experiments to determine the effects of null mutations of various clock genes on binding of CRY1 and CLOCK to target promoters. ChIP was analyzed by quantitative PCR and the values represent the average of three experiments ± S.E. From left to right, 4 columns of bar graphs show the ChIP data from different knock-out cell lines. From top to bottom, the four rows of graphs show binding to the Per1 Ebox2 or the Per2 Ebox2 (22, 49), as indicated on the left. A, CRY1 ChIP (represented by orange color). Dual cross-linked nuclear extracts were isolated from fibroblasts of the indicated genotypes and subjected to ChIP. The immunoprecipitation was performed with anti-CRY1 or no antibody control. B, CLOCK ChIP (represented by blue color). The immunoprecipitation was performed with anti-CLOCK polyclonal antibodies or IgG control. C, schematic diagram summarizing how CLOCK, BMAL1, and CRY1 occupy E-boxes in Per1 and Per2 promoters in fibroblasts with various genetic backgrounds.
Next, we investigated the binding of CLOCK to the same E-boxes (Fig. 5B, blue). The results reveal several interesting patterns regarding the interactions of the core clock proteins with clock gene promoters and with one another. First, the absence of CRY enhances the occupancy by CLOCK of E-boxes in both Per1 and Per2 promoters even though it has been reported that in mouse liver, in the absence of CRY, the level of CLOCK is reduced (22). It is conceivable that CRY promotes post-translational modifications of CLOCK that facilitates its removal from the cognate promoters (11). Second, in contrast to the absence of Cry genes, the absence of Per genes has only a modest stimulatory effect on CLOCK occupancy of Per1 and Per2 E-boxes. This is possibly because in Per1−/− Per2−/− cells, both CRY1 and CRY2 levels are lower in the nucleus (22), resulting in only a modest effect on the CLOCK protein level and its occupancy of E-boxes. Finally, in the Bmal1−/− background, there is no significant binding of CLOCK to the Per1 and Per2 promoters, consistent with the canonical clock model that BMAL1 is an irreplaceable component of the positive arm of the TTFL.
Overall, the ChIP experiments support the conclusions from the gel mobility shift assays with purified proteins that CLOCK:BMAL1 binds to E-boxes in cognate promoters, that recruitment of CRY to the CLOCK:BMAL1:E-box ternary complex is independent of PER and support previous reports that BMAL1, in the absence of CLOCK, can make heterodimers with other PAS domain proteins such as NPAS2 to regulate transcription (38) (Fig. 5C). Hence, in vitro experiments should be considered as essential tools for testing any molecular model for the mammalian circadian clock.
Probing for PER2 Protein Binding in Vivo to E-boxes in Promoters of CLOCK:BMAL1 Controlled Genes
The in vitro experiments showed that PER protein does not bind to the CLOCK:BMAL1:E-box complex. To examine this issue in vivo, we conducted ChIP assays with polyclonal anti-PER2 antibodies and wild-type mouse skin fibroblast cells. However, our experiments failed to reveal any measurable binding of PER2 protein to either Per1 or Per2 promoters (data not shown). Because ChIP assay results may be dramatically affected by the quality of the antibodies, we carried out ChIP assays using anti-FLAG antibodies in fibroblast cell lines that stably express CRY1-FLAG (control) or PER2-FLAG proteins. The immunoprecipitation assay results in Fig. 6A show that PER2-FLAG and CRY1-FLAG were precipitated at comparable levels from the two stable lines. CLOCK and BMAL1 bound to both FLAG-tagged proteins, more strongly to PER2-FLAG. Therefore, both FLAG-tagged proteins are active. However, the ChIP assay results with anti-FLAG antibodies in Fig. 6B show that CRY1-FLAG, but not PER2-FLAG, can bind to Per1 and Per2 promoters. FLAG-tagged PER2 failed to bind these regions when the FLAG tag was at either the C terminus (Fig. 6B) or N terminus (data not shown). These results indicate that at least in skin fibroblast, PER2 does not bind to these E-boxes. The strong interaction of PER2 with CLOCK and BMAL1 probably occurs only when these proteins are off DNA.
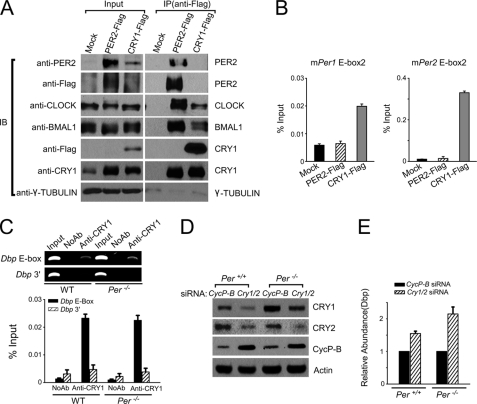
FIGURE 6.
CRY1 represses Dbp transcription independently of PER1 and PER2 in mouse fibroblasts. A, expression of clock proteins in mouse fibroblasts stably expressing CRY1-FLAG or PER2-FLAG. The cell lines stably expressing the tagged proteins are identified above each lane. Nuclear extracts prepared from a control cell line, Mock, and the two cell lines expressing PER2-FLAG or CRY1-FLAG, were subjected to immunoprecipitation (IP), and the immunoprecipitates were probed by immunoblotting (IB) for the clock proteins using antibodies listed on the left side. Blots were cut to envision the proteins listed on the right. Note the PER and CRY proteins were double immunoblotted with anti-FLAG antibodies, and with anti-CRY or anti-PER antibodies. The input lanes contain 10% of the material used in the IP. Note that the PER2-FLAG protein pulls down CLOCK, BMAL1, and CRY1 proteins, and the CRY1-FLAG protein pulls down CLOCK and BMAL1 but not PER2 presumably because the poor sensitivity of the anti-PER2 antibody (first row) does not permit detection of the endogenous PER2 signal. B, ectopic CRY1 binds to the E-boxes of Per1 and Per2, but ectopically expressed PER2 does not. ChIP assays were performed using anti-FLAG antibodies with the cell lines indicated on the x-axis, which express comparable levels of PER2-FLAG or CRY1-FLAG. Mock is a normal mouse fibroblast line as in A. C, binding of CRY1, independently of PER, to the E-box in the Dbp promoter. Wild-type or Per1−/− Per2−/− mouse skin fibroblasts were subjected to ChIP after dual cross-linking with either CRY1 antibodies or no antibody control. Top panel, semi-quantitative analysis of the ChIP data. Bottom panel, ChIP results of the Dbp E-box and a control non-regulatory sequence (Dbp 3′) were analyzed by quantitative PCR. D, down-regulation of CRY1 and CRY2 by siRNA in wild-type (Per+/+) and Per1−/−Per2−/− mouse skin fibroblasts (Per−/−) using antibodies against the proteins listed on the right. Whole cell extracts from cells treated with the indicated siRNAs were analyzed by immunoblotting; Cyclophilin-B (CycP-B) was used as a control for siRNA specificity and Actin was used as a loading control. E, down-regulation of CRY both in wild-type and Per null cells leads to up-regulation of Dbp transcription. The relative abundance of Dbp mRNA was determined by quantitative RT-PCR. Results obtained with Cry1/2 siRNA were normalized to results obtained with CycP-B siRNA, which were given a value of 1.
Effect of Cryptochrome on Transcription of a Clock-controlled Gene in the Presence and Absence of PER
The data presented so far show that CRY can bind to CLOCK:BMAL1 situated at E-boxes of target promoters without the help of PER. To determine whether this binding affects transcription we used a CLOCK:BMAL1-controlled gene with high amplitude circadian oscillation, Dbp, as a readout (8). First, we confirmed that as in the case of the Per1 and Per2 promoters, CRY1 binds to the E-box in the Dbp gene independently of PER (Fig. 6C). Then to determine the effects of CRY binding on Dbp transcription in the absence and presence of PER, we down-regulated CRY1 and CRY2 in the wild-type and Per mutant background and analyzed the cells for Dbp transcription as measured by quantitative RT-PCR. Clearly, the reduction in CRY level (Fig. 6D) is accompanied by a proportional increase in the level of Dbp mRNA (Fig. 6E), consistent with the consensus clock model whereby the binding of CRY to CLOCK:BMAL1 at the E-boxes of the target genes represses the transcription of these genes. Importantly, both in wild-type and Per1−/− Per2−/− cell lines, the Dbp mRNA level goes up similarly upon down-regulation of CRY. These findings suggest that CRY is the predominant repressor of Dbp, which is regulated with a robust rhythmicity by the CLOCK:BMAL1 transcriptional activator.
DISCUSSION
The TTFL, which was proposed over a decade ago as the mechanistic model for the mammalian circadian clock, remains as the starting point for the more detailed models that have been elaborated subsequently as new levels of regulation were discovered. However, despite the extensive genetic and cell biological data that exist in support of the canonical model, a rigorous biochemical test of the model has been lacking. Clearly, biochemical experiments with purified proteins are necessary to test some key points of the canonical model. Here, we attempted to address some key features of the model by conducting experiments with purified proteins in vitro and ChIP assays with cell lines mutated in various clock genes. Our findings support some aspects of the conventional model, but also suggest that some features of the canonical model need to be revised. A revised model consistent with our data is shown in Fig. 7 and is elaborated below.
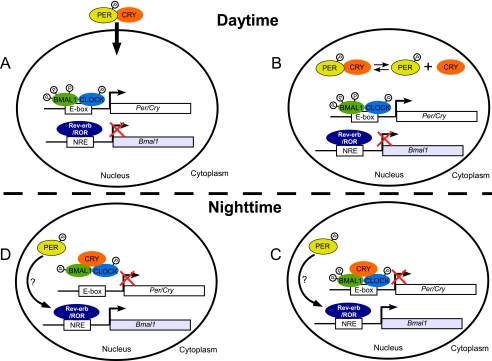
FIGURE 7.
Model for the roles of CRY and PER in the TTFL of the mammalian circadian clock. A and B, during the day (or subjective day) CLOCK:BMAL1 bind to the Per and Cry promoters and activate their transcription. In addition, REV-ERB/ROR regulates Bmal1 transcription. The PER and CRY proteins make heterodimers and enter the nucleus. Within the nucleus there is a dynamic equilibrium of the PER:CRY complex and solitary PER and CRY (most likely in complexes with other proteins); C and D, during the night (or subjective night), solitary CRY binds to the CLOCK:BMAL1:E-box ternary complex and inhibits the transcription of Per and Cry (as well as other CLOCK:BMAL1 controlled genes) both by physically interfering with the transactivation function of CLOCK:BMAL1, as well as by interfering with the post-translational modification of these proteins, which seems to be important for their transactivation functions. Data from mouse liver ChIP shows that PER2 can bind to the promoter of Bmal1 and control its transcription independently of CRY (42). This is a highly simplified model to emphasize that CRY can directly interact with CLOCK:BMAL1 on DNA to repress its transactivator function and that similarly, based on data from other studies (5, 42), PER2 can regulate the transcription of genes in the “stabilizing loop” of the clock independently of CRY to affect their transcription. NRE, nuclear responsive element.
Mechanism of CRY Repression of CLOCK:BMAL1 Transcriptional Activation
Our in vitro and in vivo findings are in agreement with previous reports in showing that bHLH-PAS domain proteins CLOCK and BMAL1 make a heterodimer, which binds to E-boxes in the promoters of cognate genes and functions as a transcriptional activator. Furthermore, our reconsitituted system and ChIP assays clearly demonstrate that CRY interacts strongly with CLOCK:BMAL1:E-box complexes. Binding was BMAL1 dependent both in vitro and in vivo, indicating that CRY is directly recruited to the E-box by a (CLOCK/NPAS2):BMAL1 heterodimer. Also of considerable importance is our finding that this binding of CRY is independent of PER: ChIP assays clearly demonstrated that CRY binds to regulatory E-boxes even in Per knock-out cells, and in vitro, PER was not only unnecessary for, but actually impaired CRY binding to the BMAL1:CLOCK:E-box DNA complex. Binding of CRY to the transactivator complex in the absence of PER is biologically meaningful, because we found that in Per1−/− Per2−/− knock-out cells, CRY down-regulated BMAL1:CLOCK-activated transcription of the Dbp gene. All of these results are in agreement with a reporter gene assay that found the inhibition of transactivation by CRY depends on the interaction of CRY with either CLOCK or both CLOCK and BMAL1 bound to DNA in S2 cells, which express no other mammalian clock proteins (5). Thus, independently of PER, CRY directly interacts with the BMAL1:CLOCK:E-box complex in a BMAL1-dependent manner to form a stable complex on DNA in which the transactivator function of CLOCK:BMAL1 is inhibited by CRY.
In addition to blocking transactivation, CRY may mediate an enzymatic function. We found that CLOCK occupied the Per1 E-box2, and to a lesser extent Per2 E-box2, at higher frequency in Cry null cells than wild-type cells, suggesting that CRY may destabilize the CLOCK:BMAL1:E-box complex in vivo (Fig. 7D). Relevant to this is a report showing that CLOCK:BMAL1 binds rhythmically to the Dbp promoter (8). Because under all experimental conditions tested, CRY did not destabilize the ternary complex in vitro, we ascribe the discrepancy between the in vivo and in vitro data to the effect of CRY on the phosphorylation status of other clock proteins including PER, CLOCK, and BMAL1 in vivo as it is known that the phosphorylation status of CLOCK affects its protein stability and DNA binding activity (12, 19, 39).
Regulation of the CRY:CLOCK:BMAL1:E-box Complex by PER
Although it is clear that PER and CRY bind to one another with high affinity, the significance of this interaction for their respective functions has been a matter of considerable debate. Some reports suggest that CRY is required for nuclear transport of PER, and others indicate that, in fact, CRY depends on PER for nuclear entry (40, 41). In addition, it has been reported that PER and CRY are required for the stability of one another, which somewhat complicates the interpretation of the results obtained with CRY or PER mutant mice and cell lines.
Similarly, even though yeast two-hybrid assays and co-immunoprecipitation experiments have revealed that in pairwise tests both CRY and PER directly interact with both CLOCK and BMAL1, these tests do not address the question of whether these interactions occur both off and on DNA. Some studies have concluded that CRY is the bona fide repressor that binds to CLOCK:BMAL1 and brings along PER to E-boxes in target promoters (22). However, a recent study that employed co-immunoprecipition and behavioral analysis of CRY1- or PER2- overproducing mice suggested that PER2 bridges CRY and CLOCK:BMAL1 bound to DNA to drive the circadian negative feedback loop (17).
In our study, we focused on the question of CRY/PER interactions with CLOCK and BMAL1 both on and off DNA in a defined system. These in vitro data were further verified by ChIP experiments in mouse fibroblasts to ascertain that the results we observed with the in vitro system were not an artifact of the reaction conditions. Contrary to our expectations, based on numerous reports indicating an essential role for PER in repressing CLOCK:BMAL1 in vivo, and based on in vitro data, including our own, which show strong PER/CLOCK/BMAL1 interaction off DNA, we were unable to detect the formation of a PER:CLOCK:BMAL1:E-box complex under a variety of experimental conditions. Furthermore, the presence of CRY in the reaction mixture does not promote the recruitment of PER to the CLOCK:BMAL1:DNA complex. In contrast, ChIP assays failed to reveal measurable binding of PER2 protein to either Per1 or Per2 promoters in wild-type fibroblasts. This is consistent with our model that CRY binds directly to the BMAL1:CLOCK:E-box complex (Fig. 7C). However, the lack of PER2 binding to cognate promoters in fibroblasts differs from the findings with ChIP using liver nuclei (42). Most likely, liver cells express proteins, which are not expressed in fibroblasts and interact with PER2 and recruit it to the target promoters in a tissue-specific manner.
Most surprisingly, perhaps, was our finding that even though solitary CRY binds to a CLOCK:BMAL1:E-box in vitro, the CRY:PER heterodimer does not, suggesting that PER interferes with binding of CRY to the transactivator on DNA. In support of this conclusion, we found that when PER was added to the CRY:CLOCK:BMAL1:E-box complex, CRY was removed from the complex. This is consistent with a model whereby CRY is in a dynamic equilibrium with CLOCK:BMAL1:E-box, and in the presence of PER at a sufficiently high concentration in the nucleus, all CRY becomes trapped in the form of a PER:CRY heterodimer, which cannot bind to the CLOCK:BMAL1:E-box (Fig. 7B). This conclusion may explain the severely disrupted rhythm in PER2 overexpressing fibroblasts or mice (17), but is counterintuitive to the genetically defined role of PER as a repressor in the core clock circuitry, and calls for re-evaluation of the core molecular clock model or revision of at least some aspects of the model.
Quite conceivably, PER functions as a repressor of CLOCK:BMAL1 not in the conventional sense by physical interaction with the transactivator complex, but by participating in stabilization and nuclear transport of CRY, which is the actual repressor, as well as by regulating transcription through interactions with some nuclear receptors as reported previously (42, 43) (Fig. 7C). We note, however, that this model does not incorporate the well known and rather strong interactions of PER with CLOCK and BMAL1. It is conceivable that PER interacts with these proteins off DNA, and in doing so, prevents their association with DNA although this is not consistent with the reporter gene assay results that show that the PER protein alone is a weak inhibitor of CLOCK:BMAL1 activity. It is, however, possible that PER, in addition to aiding CRY repressor function in its capacity as chaperone. It may, to a limited degree, aid in the formation of the CRY:CLOCK:BMAL1:E-box complex in its capacity as a molecular matchmaker by transient interaction with CLOCK:BMAL1 on DNA. Clearly, additional work is needed to clarify the mechanism by which PER participates in the negative arm of the transcription-translation feedback loop, and, if there is some tissue specificity (such as the suprachiasmatic nucleus versus peripheral tissue) in the core clock circuitry.
Enzymatic (Catalytic) Control of Circadian TTFL
It has been shown that the core clock proteins are subject to phosphorylation and ubiquitin-proteasome system-mediated degradation (1). In both Neurospora and Drosophila, the negative elements inhibit the DNA binding of the positive elements by recruiting kinases to phosphorylate the positive elements (44, 45). Undoubtedly, these and other post-translational events contribute significantly to the generation of a robust TTFL with circadian periodicity (21, 39, 46–48). Here, we have presented some basic biochemical facts regarding the contributions of the interactions of the proteins in the positive arm (CLOCK and BMAL1) and negative arm (CRY and PER) of the clock with one another on and off DNA to the generation of the circadian TTFL. We believe quantitative in vitro studies of the post-translational modifications of clock proteins are needed for a comprehensive description of the mammalian circadian clock.
Supplementary Material
Acknowledgments
We thank J. S. Takahashi, S. M. Reppert, C. C. Lee, and M. P. Antoch for reagents and L. A. Lindsey-Boltz and S. Adar for critical comments on the manuscript.
This work was supported, in whole or in part, by National Institutes of Health Grant GM31082.

The on-line version of this article (available at http://www.jbc.org) contains supplemental “Methods,” Table S1, and Figs. S1–S4.
- TTFL
- transcription-translation feedback loop
- MAX
- MYC-associated factor X.
REFERENCES
- 1. Takahashi J. S., Hong H. K., Ko C. H., McDearmon E. L. (2008) Nat. Rev. Genet. 9, 764–775 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Hastings M. H., Reddy A. B., Maywood E. S. (2003) Nat. Rev. Neurosci. 4, 649–661 [DOI] [PubMed] [Google Scholar]
- 3. Reppert S. M., Weaver D. R. (2002) Nature 418, 935–941 [DOI] [PubMed] [Google Scholar]
- 4. Lowrey P. L., Takahashi J. S. (2004) Annu. Rev. Genomics Hum. Genet. 5, 407–441 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Shearman L. P., Sriram S., Weaver D. R., Maywood E. S., Chaves I., Zheng B., Kume K., Lee C. C., van der Horst G. T., Hastings M. H., Reppert S. M. (2000) Science 288, 1013–1019 [DOI] [PubMed] [Google Scholar]
- 6. Griffin E. A., Jr., Staknis D., Weitz C. J. (1999) Science 286, 768–771 [DOI] [PubMed] [Google Scholar]
- 7. Kume K., Zylka M. J., Sriram S., Shearman L. P., Weaver D. R., Jin X., Maywood E. S., Hastings M. H., Reppert S. M. (1999) Cell 98, 193–205 [DOI] [PubMed] [Google Scholar]
- 8. Ripperger J. A., Schibler U. (2006) Nat. Genet. 38, 369–374 [DOI] [PubMed] [Google Scholar]
- 9. Gekakis N., Staknis D., Nguyen H. B., Davis F. C., Wilsbacher L. D., King D. P., Takahashi J. S., Weitz C. J. (1998) Science 280, 1564–1569 [DOI] [PubMed] [Google Scholar]
- 10. Etchegaray J. P., Lee C., Wade P. A., Reppert S. M. (2003) Nature 421, 177–182 [DOI] [PubMed] [Google Scholar]
- 11. Kondratov R. V., Kondratova A. A., Lee C., Gorbacheva V. Y., Chernov M. V., Antoch M. P. (2006) Cell Cycle 5, 890–895 [DOI] [PubMed] [Google Scholar]
- 12. Yoshitane H., Takao T., Satomi Y., Du N. H., Okano T., Fukada Y. (2009) Mol. Cell. Biol. 29, 3675–3686 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Dardente H., Fortier E. E., Martineau V., Cermakian N. (2007) Biochem. J. 402, 525–536 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Ishikawa T., Hirayama J., Kobayashi Y., Todo T. (2002) Genes Cells 7, 1073–1086 [DOI] [PubMed] [Google Scholar]
- 15. Menet J. S., Abruzzi K. C., Desrochers J., Rodriguez J., Rosbash M. (2010) Genes Dev. 24, 358–367 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Lee C., Bae K., Edery I. (1999) Mol. Cell. Biol. 19, 5316–5325 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Chen R., Schirmer A., Lee Y., Lee H., Kumar V., Yoo S. H., Takahashi J. S., Lee C. (2009) Mol. Cell 36, 417–430 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Gauger M. A., Sancar A. (2005) Cancer Res. 65, 6828–6834 [DOI] [PubMed] [Google Scholar]
- 19. Spengler M. L., Kuropatwinski K. K., Schumer M., Antoch M. P. (2009) Cell Cycle 8, 4138–4146 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Fu L., Pelicano H., Liu J., Huang P., Lee C. (2002) Cell 111, 41–50 [DOI] [PubMed] [Google Scholar]
- 21. Partch C. L., Shields K. F., Thompson C. L., Selby C. P., Sancar A. (2006) Proc. Natl. Acad. Sci. U.S.A. 103, 10467–10472 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Lee C., Etchegaray J. P., Cagampang F. R., Loudon A. S., Reppert S. M. (2001) Cell 107, 855–867 [DOI] [PubMed] [Google Scholar]
- 23. Ozgür S., Sancar A. (2006) Biochemistry 45, 13369–13374 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Ozturk N., Selby C. P., Annayev Y., Zhong D., Sancar A. (2011) Proc. Natl. Acad. Sci. U.S.A. 108, 516–521 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Hogenesch J. B., Gu Y. Z., Jain S., Bradfield C. A. (1998) Proc. Natl. Acad. Sci. U.S.A. 95, 5474–5479 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Nakahata Y., Kaluzova M., Grimaldi B., Sahar S., Hirayama J., Chen D., Guarente L. P., Sassone-Corsi P. (2008) Cell 134, 329–340 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Doi M., Hirayama J., Sassone-Corsi P. (2006) Cell 125, 497–508 [DOI] [PubMed] [Google Scholar]
- 28. Sato T. K., Yamada R. G., Ukai H., Baggs J. E., Miraglia L. J., Kobayashi T. J., Welsh D. K., Kay S. A., Ueda H. R., Hogenesch J. B. (2006) Nat. Genet. 38, 312–319 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Rutter J., Reick M., Wu L. C., McKnight S. L. (2001) Science 293, 510–514 [DOI] [PubMed] [Google Scholar]
- 30. Walhout A. J., Gubbels J. M., Bernards R., van der Vliet P. C., Timmers H. T. (1997) Nucleic Acids Res. 25, 1493–1501 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Kiyohara Y. B., Tagao S., Tamanini F., Morita A., Sugisawa Y., Yasuda M., Yamanaka I., Ueda H. R., van der Horst G. T., Kondo T., Yagita K. (2006) Proc. Natl. Acad. Sci. U.S.A. 103, 10074–10079 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Langmesser S., Tallone T., Bordon A., Rusconi S., Albrecht U. (2008) BMC Mol. Biol. 9, 41 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Vitaterna M. H., Selby C. P., Todo T., Niwa H., Thompson C., Fruechte E. M., Hitomi K., Thresher R. J., Ishikawa T., Miyazaki J., Takahashi J. S., Sancar A. (1999) Proc. Natl. Acad. Sci. U.S.A. 96, 12114–12119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Zheng B., Albrecht U., Kaasik K., Sage M., Lu W., Vaishnav S., Li Q., Sun Z. S., Eichele G., Bradley A., Lee C. C. (2001) Cell 105, 683–694 [DOI] [PubMed] [Google Scholar]
- 35. DeBruyne J. P., Weaver D. R., Reppert S. M. (2007) Nat. Neurosci. 10, 543–545 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Reick M., Garcia J. A., Dudley C., McKnight S. L. (2001) Science 293, 506–509 [DOI] [PubMed] [Google Scholar]
- 37. Debruyne J. P., Noton E., Lambert C. M., Maywood E. S., Weaver D. R., Reppert S. M. (2006) Neuron 50, 465–477 [DOI] [PubMed] [Google Scholar]
- 38. DeBruyne J. P., Weaver D. R., Reppert S. M. (2007) Curr. Biol. 17, R538–539 [DOI] [PubMed] [Google Scholar]
- 39. Kondratov R. V., Chernov M. V., Kondratova A. A., Gorbacheva V. Y., Gudkov A. V., Antoch M. P. (2003) Genes Dev. 17, 1921–1932 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Yagita K., Tamanini F., Yasuda M., Hoeijmakers J. H., van der Horst G. T., Okamura H. (2002) EMBO J. 21, 1301–1314 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Miyazaki K., Mesaki M., Ishida N. (2001) Mol. Cell. Biol. 21, 6651–6659 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Schmutz I., Ripperger J. A., Baeriswyl-Aebischer S., Albrecht U. (2010) Genes Dev. 24, 345–357 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Grimaldi B., Bellet M. M., Katada S., Astarita G., Hirayama J., Amin R. H., Granneman J. G., Piomelli D., Leff T., Sassone-Corsi P. (2010) Cell Metab. 12, 509–520 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. He Q., Cha J., He Q., Lee H. C., Yang Y., Liu Y. (2006) Genes Dev. 20, 2552–2565 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Yu W., Zheng H., Houl J. H., Dauwalder B., Hardin P. E. (2006) Genes Dev. 20, 723–733 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Asher G., Gatfield D., Stratmann M., Reinke H., Dibner C., Kreppel F., Mostoslavsky R., Alt F. W., Schibler U. (2008) Cell 134, 317–328 [DOI] [PubMed] [Google Scholar]
- 47. Asher G., Reinke H., Altmeyer M., Gutierrez-Arcelus M., Hottiger M. O., Schibler U. (2010) Cell 142, 943–953 [DOI] [PubMed] [Google Scholar]
- 48. Hirayama J., Sahar S., Grimaldi B., Tamaru T., Takamatsu K., Nakahata Y., Sassone-Corsi P. (2007) Nature 450, 1086–1090 [DOI] [PubMed] [Google Scholar]
- 49. Yoo S. H., Ko C. H., Lowrey P. L., Buhr E. D., Song E. J., Chang S., Yoo O. J., Yamazaki S., Lee C., Takahashi J. S. (2005) Proc. Natl. Acad. Sci. U.S.A. 102, 2608–2613 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.