Abstract
MicroRNAs (miRNAs) are ~22-nucleotide-long noncoding RNAs that normally function by suppressing translation and destabilizing messenger RNAs bearing complementary target sequences. Some miRNAs are expressed in a cell- or tissue-specific manner and may contribute to the establishment and/or maintenance of cellular identity. Recent studies indicate that tissue-specific miRNAs may function at multiple hierarchical levels of gene regulatory networks, from targeting hundreds of effector genes incompatible with the differentiated state to controlling the levels of global regulators of transcription and alternative pre-mRNA splicing. This multilevel regulation may allow individual miRNAs to profoundly affect the gene expression program of differentiated cells.
There is a surprisingly small difference in the number of protein-encoding genes between organisms of vastly different morphological and behavioral complexity. A plausible explanation for this paradox may be in the increased elaboration of gene regulatory networks at the levels of transcription (1) and alternative pre-mRNA splicing (2). More recently, miRNAs have been proposed to play a role in the expansion of organismal complexity (3). Indeed, some miRNAs are expressed in a cell- or tissue-specific manner during embryonic development, suggestive of a role in cellular differentiation (4) [also see essay by O. Hobert in this issue (5)]. Certain of these tissue-specific miRNAs, such as miR-1 and miR-124, which are expressed in muscle cells and neurons, respectively, have been shown to stimulate differentiation of the corresponding cell types (6–8).
Targeting Gene Batteries
Several distinct molecular mechanisms may underlie the biological functions of miRNAs. Individual miRNAs can repress large sets of mRNAs that are not required at a particular developmental stage (9, 10). A number of these miRNA targets fall into the category of “gene batteries,” sets of functionally related effector genes that represent outputs of gene regulatory networks (11). Indeed, when miR-124 is introduced into nonneuronal mammalian cells a preferential reduction in the amounts of multiple nonneuronal mRNAs, for example, those encoding proteins required for cell proliferation or neural stem cell function, is observed (7, 9, 12). Conversely, depletion of miR-124 from primary neurons leads to the accumulation of a number of nonneuronal mRNA targets (13). A similar mode of action has been reported for miR-1 (9). Thus, in cells undergoing differentiation, miRNAs can effectively deplete unwanted mRNAs left over from progenitor cells. Furthermore, miRNAs may regulate a fraction of mRNA targets at the level of translation without affecting their stability (14, 15).
Targeting Regulators of Transcription
MiRNAs can also control the expression of critical transcriptional regulators, as first shown by the repression of transcription factor lin-14 by miRNA lin-4 (16, 17). Another example is miR-124, which targets mRNA of the small C-terminal domain phosphatase 1 (SCP1/CTDSP1) protein, a component of the RE1-silencing transcription repressor (REST/NRSF) (6) (Fig. 1). REST represses the transcription of a large number of neuron-specific genes in nonneural cells (13). Thus, by reducing SCP1 expression in differentiating neurons, miR-124 may facilitate the derepression of the neuronal transcription program (6). The REST complex may also repress miR-124 gene expression and thus establish a double-negative feedback loop between miR-124 and the REST complex (6, 13) (Fig. 1). Similarly, miR-1 and another muscle-specific miRNA, miR-133, regulate several transcription factors essential for muscle development (4, 8).
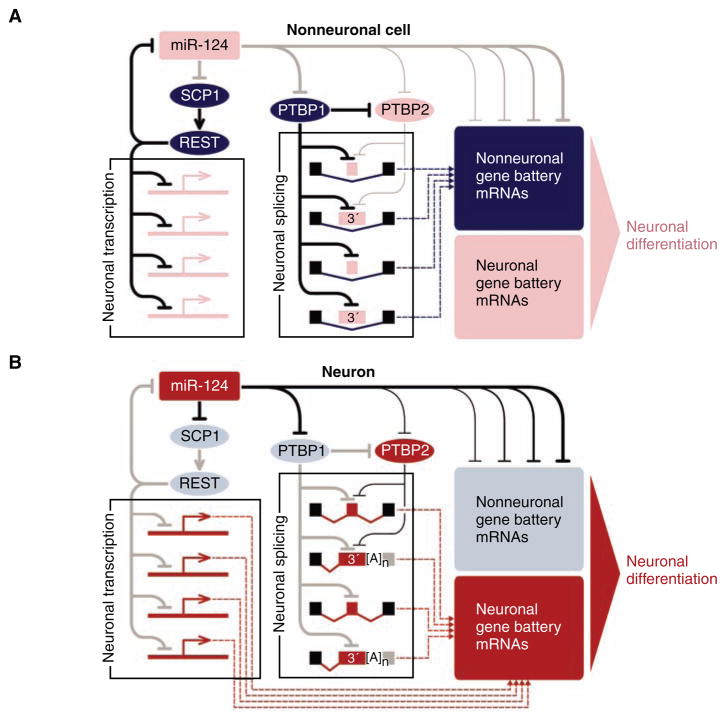
Fig. 1.
MiR-124 controls an extensive gene regulatory network. Active regulatory interactions are shown as solid black lines. Inactive interactions are in gray. Weaker regulatory interactions are rendered as thinner lines. Gene battery expression inputs to the cellular transcriptome are indicated as dashed lines. Nonneuronal elements are colored in dark blue (present) or light blue (absent); neuron-specific elements are colored in corresponding shades of red. This is a simplified diagram that does not account for potential inputs from other neuron-specific miRNAs, as well as the recently identified miRNA function in translational activation (15). Furthermore, PTBP1 and PTBP2 may also activate subsets of alternative exons by using a yet-to-be understood mechanism (19). (A) In nonneuronal cells or neural precursors, miR-124 is either absent or present at a low amount, which allows efficient expression of global repressors of neuronal transcription and alternative pre-mRNA splicing, as well as nonneuronal gene batteries. (B) In differentiating neurons, miR-124 quantities increase, leading to the down-regulation of corresponding repressors, thus allowing the production of the corresponding neuronal proteins. Inversely, the nonneuronal battery genes are directly down-regulated by miR-124.
Targeting Regulators of Alternative Splicing
Recent studies have identified yet another activity of miRNAs: the induction of large-scale changes in gene expression by targeting global regulators of alternative pre-mRNA splicing (7, 18). In early muscle cell precursors, a repressor of alternative splicing called polypyrymidine tract-binding protein 1 (PTBP1/PTB/hnRNP-I) and its homolog, PTBP2 (nPTB/brPTB/PTBLP), repress the inclusion of a number of muscle-specific exons into mature mRNAs. However, upon myotube differentiation, PTBP1 and PTBP2 protein quantities are reduced, leading to increased inclusion of their target exons during splicing. This splicing switch is regulated, at least in part, by miRNAs: The aforementioned miR-133 strongly suppresses PTBP2 production, whereas miR-1 and its sequence homolog miR-206 may also contribute to down-regulation of the PTBP1 and PTBP2 quantities during muscle development (18).
The amounts of PTBP1 and PTBP2 are also regulated by miRNAs during the development of the nervous system (7, 19). PTBP1 functions in this context as a repressor of neuron-specific alternative exon inclusion, and it is expressed in neural precursors as well as many other types of nonneuronal cells. However, in differentiating and mature neurons, PTBP1 quantities decrease, leading to the inclusion of a number of neuron-specific alternative exons in mature mRNA (19). Similar to the regulation in muscle cells, the reduced PTBP1 expression in neurons is mediated by miR-124, which interacts with conserved and nonconserved cognate target sites in the 3′ untranslated region (3′UTR) of the PTBP1 mRNA (7) (Fig. 1).
In addition to other alternative exons, PTBP1 represses the inclusion of exon 10 of PTBP2 pre-mRNA (7, 19, 20). When this exon is skipped (i.e., excluded from mature mRNA), PTBP2 mRNA acquires a premature termination codon and as a result is degraded by the nonsense-mediated decay machinery (7, 19, 20). During neuronal differentiation, miR-124 quantities increase, thus reducing PTBP1 amounts and resulting in the accumulation of correctly spliced PTBP2 mRNA, which in turn leads to a substantial increase in PTBP2 protein. Notably, although PTBP1 and PTBP2 are closely related homologs, PTBP1 appears to be a much stronger repressor of neuron-specific alternative exons (7, 19). Thus, the reduction in PTBP1 and increase in PTBP2 results in a global switch from nonneuronal to neuron-specific alternative splicing patterns, leading to the production of neuron-specific protein isoforms. Interestingly, PTBP2 mRNA contains conserved miR-124 binding sites that allow miR-124 to suppress PTBP2 expression, albeit less efficiently than PTBP1 (7). This regulation may provide a mechanism that dampens PTBP2 expression (Fig. 1).
The increasing diversity of cellular differentiation in metazoans, accompanied by an increase in the complexity of gene regulatory networks, must have required a mechanism to prevent interference between spatially or temporally adjacent gene expression programs. The above examples argue that at least some miRNAs play an important role in this mechanism by effectively rewiring the cell-specific networks at all levels of the regulatory hierarchy, from the gene battery to global regulators of transcription and alternative splicing. It seems likely that other examples of this multilevel regulation will be revealed as target repertoires of other miRNAs are determined.
Acknowledgments
We thank S. Oliferenko, S. Buchanan, B. Friedman, E. Morimoto, and B. McCallum for critical comments. This effort was supported by NIH (grant 2R01NS043915-27; T.M.) and Leukemia and Lymphoma Society (E.V.M.).
Contributor Information
Eugene V. Makeyev, Email: makeyev@mcb.harvard.edu.
Tom Maniatis, Email: maniatis@mcb.harvard.edu.
References and Notes
- 1.Levine M, Tjian R. Nature. 2003;424:147. doi: 10.1038/nature01763. [DOI] [PubMed] [Google Scholar]
- 2.Maniatis T, Tasic B. Nature. 2002;418:236. doi: 10.1038/418236a. [DOI] [PubMed] [Google Scholar]
- 3.Niwa R, Slack FJ. Curr Opin Genet Dev. 2007;17:145. doi: 10.1016/j.gde.2007.02.004. [DOI] [PubMed] [Google Scholar]
- 4.Zhao Y, Srivastava D. Trends Biochem Sci. 2007;32:189. doi: 10.1016/j.tibs.2007.02.006. [DOI] [PubMed] [Google Scholar]
- 5.Hobert O. Science. 2008;319:1785. doi: 10.1126/science.1151651. [DOI] [PubMed] [Google Scholar]
- 6.Visvanathan J, Lee S, Lee B, Lee JW, Lee SK. Genes Dev. 2007;21:744. doi: 10.1101/gad.1519107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Makeyev EV, Zhang J, Carrasco MA, Maniatis T. Mol Cell. 2007;27:435. doi: 10.1016/j.molcel.2007.07.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chen JF, et al. Nat Genet. 2006;38:228. doi: 10.1038/ng1725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lim LP, et al. Nature. 2005;433:769. doi: 10.1038/nature03315. [DOI] [PubMed] [Google Scholar]
- 10.Giraldez AJ, et al. Science. 2006;312(75) doi: 10.1126/science.1122689. published online 15 February 2006. [DOI] [PubMed] [Google Scholar]
- 11.Davidson EH. The Regulatory Genome: Gene Regulatory Networks in Development and Evolution. Academic Press; San Diego, CA: 2006. [Google Scholar]
- 12.Cao X, Pfaff SL, Gage FH. Genes Dev. 2007;21:531. doi: 10.1101/gad.1519207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Conaco C, Otto S, Han JJ, Mandel G. Proc Natl Acad Sci USA. 2006;103:2422. doi: 10.1073/pnas.0511041103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Karginov FV, et al. Proc Natl Acad Sci USA. 2007;104:1929. [Google Scholar]
- 15.Vasudevan S, Tong Y, Steitz JA. Science. 2007;318:1931. doi: 10.1126/science.1149460. published online 29 November 2007. [DOI] [PubMed] [Google Scholar]
- 16.Lee RC, Feinbaum RL, Ambros V. Cell. 1993;75:843. doi: 10.1016/0092-8674(93)90529-y. [DOI] [PubMed] [Google Scholar]
- 17.Wightman B, Ha I, Ruvkun G. Cell. 1993;75:855. doi: 10.1016/0092-8674(93)90530-4. [DOI] [PubMed] [Google Scholar]
- 18.Boutz PL, Chawla G, Stoilov P, Black DL. Genes Dev. 2007;21:71. doi: 10.1101/gad.1500707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Boutz PL, et al. Genes Dev. 2007;21:1636. doi: 10.1101/gad.1558107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Spellman R, Llorian M, Smith CW. Mol Cell. 2007;27:420. doi: 10.1016/j.molcel.2007.06.016. [DOI] [PMC free article] [PubMed] [Google Scholar]