Abstract
Circadian-regulated gene expression is predominantly controlled by a transcriptional negative feedback loop, and it is evident that chromatin modifications and chromatin remodeling are integral to this process in eukaryotes. We previously determined that multiple ATP–dependent chromatin-remodeling enzymes function at frequency (frq). In this report, we demonstrate that the Neurospora homologue of chd1 is required for normal remodeling of chromatin at frq and is required for normal frq expression and sustained rhythmicity. Surprisingly, our studies of CHD1 also revealed that DNA sequences within the frq promoter are methylated, and deletion of chd1 results in expansion of this methylated domain. DNA methylation of the frq locus is altered in strains bearing mutations in a variety of circadian clock genes, including frq, frh, wc-1, and the gene encoding the frq antisense transcript (qrf). Furthermore, frq methylation depends on the DNA methyltransferase, DIM-2. Phenotypic characterization of Δdim-2 strains revealed an approximate WT period length and a phase advance of approximately 2 hours, indicating that methylation plays only an ancillary role in clock-regulated gene expression. This suggests that DNA methylation, like the antisense transcript, is necessary to establish proper clock phasing but does not control overt rhythmicity. These data demonstrate that the epigenetic state of clock genes is dependent on normal regulation of clock components.
Author Summary
Circadian rhythms facilitate daily changes in gene expression via a transcriptional negative feedback loop. In eukaryotes, chromatin remodeling is an integral part of transcriptional regulation and is proving to be one of the major determinants for the proper timing and amplitude of clock-gene expression. We describe here the action of chromodomain helicase DNA–binding (CHD1), one of two ATP–dependent chromatin-remodeling enzymes required for normal circadian regulated gene expression of the central clock gene frequency (frq). Molecular analysis of strains lacking chd1 indicates that CHD1 is required for remodeling chromatin structure at the frq locus as a part of the daily clock cycle. Moreover, we discovered DNA methylation in the promoter of frq that diminishes over time in the absence of light/dark cycles and determined that normal DNA methylation appears to require a functional clock. The DNA methyltransferase DIM-2 is responsible for this DNA methylation, and the DNA methylation is required for proper phasing of clock gene expression. Collectively, these data demonstrate a close connection among chromatin remodeling, DNA methylation, and clock gene expression.
Introduction
Virtually all eukaryotes have the ability to control temporal gene expression as a function of time-of-day. The molecular mechanism of circadian rhythms is conserved among animals and fungi and involves a transcriptional negative feedback loop [1]–[5]. In Neurospora, the positive elements are the White Collar (WC) proteins, which drive rhythmic expression of the negative element frq. WC-1 and WC-2 are GATA-type zinc finger transcription factors that heterodimerize via PAS (Per-Arnt-Sim) domains to form the White Collar Complex (WCC) [6]–[9]. Rhythmic and light-activated WCC-driven expression of frq is regulated by two cis-acting sequences in the promoter, the Clock Box (C-box) and the proximal light regulated element (PLRE) [10], [11].
In the negative arm of the loop, translated FRQ protein associates with the FRQ-interacting RNA helicase, FRH [12], and undergoes regulated nuclear entry [13] modulated in part by the phosphorylation state of FRQ [14]. FRQ-FRH is believed to inhibit frq expression by promoting inactivation of the WCC through phosphorylation [15], [16] and via direct binding to the WCC [17]. In addition, circadian regulated frq expression is partially controlled by a chromatin-remodeling event that promotes accessibility to the C-box promoter element [18]. Once FRQ is synthesized, it undergoes progressive phosphorylation [19]–[21] while feeding back to inhibit its expression [22] and it is eventually turned over in a reaction that involves an F-box/WD40 protein FWD [23] and the SKP-cullin complex [24], thus releasing its negative effects on transcription. A plethora of data indicates FRQ phosphorylation and turnover establish period but less is known about the molecular mechanisms underlying phase determination.
Rhythmic changes in chromatin structure have been correlated with changes in transcriptional activity of clock-associated loci. In mammalian cells, rhythmic modifications include acetylation of histone H3 at Per1, Per2 and Cry, and acetylation of H4 at Per1, with the peak in acetylation occurring during the transcriptional activation phase [25], [26]. In contrast, during the repressive phase, histone H3 associated with Per1 and Per2 becomes methylated at K27 [27]. There are also rhythmic changes at the clock-controlled gene (ccg) encoding the D-element binding protein (Dbp), including H3K9me2 and HP1 binding, suggesting that at least one ccg is regulated by facultative heterochromatin formation [28]. In addition, the role of CLOCK as a histone acetyltransferase [29], and the acetylation of BMAL1 [30], combined with the observation that the NAD+-dependent histone deacetylase SIRT1 modulates the amplitude of circadian regulated gene expression [31], [32] has garnished attention. More recently, the mixed-lineage leukemia (MLL1) lysine methyltransferase (KMT2A) has been show to associate with CLOCK-BMAL1 and is required for rhythmic expression of clock genes [33]. Additionally, the histone demethylase JMJD5 (KDM8) is rhythmically expressed and is required for normal rhythms in both the Arabidopsis and mammalian clock suggesting that oscillations in H3K36 methylation are also an important component of clock function [34]. Current models of circadian gene regulation suggest that the clock controls its own phase-specific chromatin modifications, which help confer levels of expression appropriate for the time-of-day. How these “marks” are established after entrainment, transferred after replication, and ultimately interpreted so that rhythms are maintained and asynchronous expression is prevented are active areas of research.
It is clear that chromatin structure is instrumental for appropriate circadian regulation of transcription. In fact, circadian changes in nucleosome location are known to occur at both frq and mDbp [18], [35], indicating that the clock controls chromatin architecture at a subset of clock-regulated loci. Consistent with this idea, circadian regulated remodeling at frq involves the ATP-dependent chromatin-remodeling enzyme CLOCKSWITCH (CSW-1) [18]. Moreover, Drosophila KISMET protein, a member of the CHD subfamily of ATP-dependent remodeling enzymes, was shown to be a key regulator of photoresponses [36]. RNAi knockdowns of the kis gene resulted in activity rhythms in constant light similar to the cryb mutation [36].
In this report, we show that an additional remodeling enzyme CHD1, remodels chromatin at frq, is necessary for normal frq expression, and is involved in regulating chromatin structure at frq. Surprisingly, we report that DNA in the frq promoter is methylated and we observe an expanded domain of DNA methylation at frq and other light/clock regulated genes in Δchd1. This was unexpected because previously, only repeated DNA sequences were found to be methylated and presumably silenced (reviewed in [37]). We show transient and reversible methylation that is associated with regulatory and coding sequences of genes that are actively expressed. This normal DNA methylation at frq is altered in strains bearing mutations in circadian clock genes and the frq antisense transcript qrf, and it is catalyzed by the DNA methyltransferase DIM-2. Strains lacking dim-2 exhibit a small phase advance, suggesting that the methylation serves to limit the onset of circadian regulated transcription. These results demonstrate that transient, facultative, and presumably regulated DNA methylation also occurs in fungi and that chromatin remodeling and DNA methylation combine to fine turn circadian regulated gene expression.
Results
CHD1 is required for normal frq expression
Previously, we generated and screened 19 Neurospora knockout strains in which proposed ATP-dependent chromatin-remodeling enzymes were deleted, to identify genes necessary for remodeling chromatin structure at frq [18]. As is typical for circadian research in Neurospora, these knockouts were generated in a ras-1bd genetic background in order to facilitate subsequent screening of the overt rhythm in conidiation [38], [39]. In this genetic background, we found that two remodeling enzymes (orthologues of Sth1 and Chd1) appeared essential for viability and a third, CSW-1, whose closest homologues are Fun30 in Saccharomyces cerevisiae, ETL1 in mice, and SMARCAD in humans, remodels chromatin at frq [18]. Subsequent analysis indicated that Δchd1 is synthetically lethal with ras-1bd, explaining our inability to isolate viable ascospores that contain both Δchd1 and ras-1bd. We have since obtained a viable homokaryotic knockout of the chd1 homologue, NCU03060 in a WT background, from the Neurospora Genome Project knockout consortium [40]. Verification of the Δchd1 knockout strain and characterization of its associated phenotypes can be found in Figure S1.
Because chd1 is essential when combined with ras-1bd, we were unable to assay clock phenotypes with race tubes, the standard assay used to measure circadian regulated conidiation. Therefore, circadian rhythmicity was analyzed molecularly after a standard light to dark transfer by examining the normally rhythmic expression of frq mRNA transcript and protein. Northern blot analysis (Figure 1A) and quantitative RT PCR (Figure 1B) revealed that in the Δchd1 strain, frq expression was diminished and the amplitude of rhythmic RNA accumulation was significantly reduced and rapidly graded to arhythmicity. Western blot analysis (Figure 1C) confirmed that clock function was disrupted in the Δchd1 strain. FRQ was detected as multiple phosphorylated forms, indicating newly synthesized FRQ was continually replacing protein destined for turnover. Quantification of the western blot is shown in Figure 1D and is consistent with the RNA expression data showing that CHD1 is required for sustained rhythmicity. We also observed a striking defect in chromatin remodeling at the antisense promoter (see below) so we examined expression of the antisense transcript by quantitative RT-PCR using a primer specific for antisense in the reverse transcriptase reaction. We found no significant differences between WT and Δchd1 strains (Figure 1E). Although we did not see a rhythm in qrf in a true wild-type strain, we did confirm the low amplitude rhythm in the ras-1bd strain previously reported [41] (data not shown).
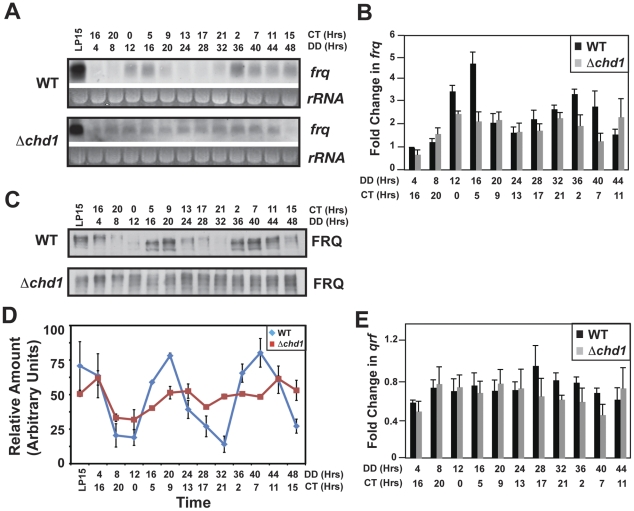
Figure 1. CHD1 is required for proper frq regulation.
(A) Northern blot and (B) Real-time RT-PCR analysis of frq RNA levels in Δchd1 and WT strains. (C) Immunoblot analysis on cell lysates using a FRQ antibody to compare expression in WT versus Δchd1 [19]. These and comparable data from three replicate experiments were quantified and are plotted in panel D which shows rapid loss of rhythmicity. (E) Real-Time RT-PCR using a qrf-specific primer in an RT reaction followed by quantitative PCR of the qrf transcript. DD indicates the length of time that samples were exposed to dark conditions and the corresponding circadian time (CT) is also shown. In this and subsequent figures LP15 identifies cultures that were kept in darkness for 24 hours and then exposed to saturating light treatment for 15 minutes prior to harvesting. Error bars represent the standard error.
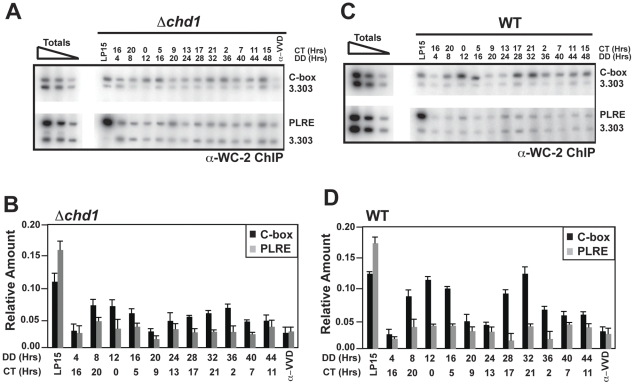
To further define frq expression, we used chromatin immunoprecipitation (ChIP) to assay the association of WC-2 with the frq promoter in Δchd1 (Figure 2A and 2B). We consistently detected a subtle, low-amplitude rhythm in WC-2 association with the C-box, but binding was reduced approximately 50% when compared with WT, consistent with the observed reduction in frq mRNA (Figure 2C and 2D). The low amplitude rhythms in frq message and rhythmic WC-2 binding to the C-box suggest a more supportive rather than immediately essential role for CHD1 in clock function; however, these analyses only monitor clock function for two circadian cycles.
Figure 2. Binding of WC-2 to the frq promoter in Δchd1.
ChIP analyses examining WC-2 binding at the C-box and PLRE in Δchd1 (A and B) and WT (C and D) strains. The panels show a dilution series of the total lysate and the 3.303 oligo pair is used as a measure of nonspecific background [18]. Error bars represent the standard error. The totals consisted of a 1∶100, 1∶1,000 and 1∶10,000 dilution of the starting DNA present in the cross-linked lysates.
To confirm the effects of Δchd1 on clock function and to examine frq expression over an extended circadian time course, we used an indirect method to measure rhythmicity of the core oscillator in real time using a frq promoter-luciferase reporter construct [42]. WT and Δchd1 strains containing the pVG110 construct integrated at the his-3 locus were grown on mini-race tubes and LUC activity was initially monitored over the entire tube and at region near the inoculation point (Figure S2). While clear robust rhythms were observed in WT, the Δchd1 strain typically displayed a single peak of frq-driven LUC activity with no evidence of sustained circadian regulation. Together, these data demonstrate that CHD1 is required to maintain normal high amplitude rhythms of WC-2 binding and frq mRNA accumulation, and is essential for persistent circadian expression of frq in synchronized Neurospora cultures.
CHD1 remodels chromatin at frq
Previously, we showed that chromatin in the frq promoter region, near the transcriptional start site (TSS) and the PLRE, is remodeled in response to light treatment but that this remodeling is independent of CSW-1 [18]. We suspected that this light-dependent remodeling might be catalyzed by CHD1 because of the failure to properly express frq in Δchd1 strains. Therefore, we asked if CHD1 is responsible for catalyzing the opening and closing of chromatin structure by comparing Δchd1 and WT nuclei using a limited MNase I digestion. Nuclei were isolated from 48-hour old cultures grown in the dark for 12 hours (CT0) and 24 hours (CT12), and treated with a 15 minute light-pulse (LP15) given after 24 hrs in darkness. Regions in frq were examined by partial MNaseI digestion followed by indirect end-labeling (Figure 3A). Within the region surrounding the PLRE and TSS, we consistently only observed a very minor remodeling defect in Δchd1 compared to WT (Figure S3). We extended our analysis to regions near the C-box promoter element, a region shown to be remodeled by CSW-1 [18], and did not observe any CHD1-dependent remodeling in that region (data not shown). In contrast to the subtle effect seen in the sense promoter, we detected unequivocal CHD1-dependent changes in chromatin structure in the antisense promoter (Figure 3B). A sequence near the 1250 site in the qrf promoter closely matches the imperfect repeat consensus sequence of the C-box and PLRE previously described [10]. WC-2 binding to the antisense light response element (aLRE) has been observed in WC-2 ChIP-seq experiments [43]. As indicated above, chromatin remodeling in the frq promoter persists in Δchd1 and may account for the low amplitude rhythms in frq mRNA and WC-2 binding observed in Δchd1 mutants (Figure 1A and Figure 2A). This observation is consistent with previous work and suggests that multiple chromatin remodeling enzymes regulate frq including the previously identified CSW-1 protein [18]. These data cumulatively indicate that chromatin structure at frq in a Δchd1 strain is different from that in WT and implicates CHD1 in maintaining chromatin architecture at the frq locus.
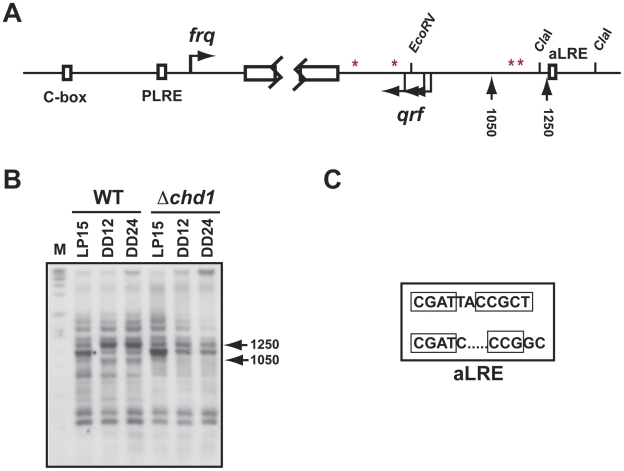
Figure 3. CHD1 is needed for remodeling at frq.
(A) Schematic representation of the frq locus with vertical arrows (1050 site and 1250 site) indicating the regions where remodeling is observed. Horizontal arrows indicate the major start sites of transcription for both frq and qrf. Red asterisks indicate the poly A sites for frq. (B) Chromatin structure was visualized by indirect end-labeling of DNA after partial MNaseI digestion. Nuclei were isolated from WT and Δchd1 strains grown in the dark (DD12 and DD24) and after a 15-minute light-pulse (LP15). The two arrows (1050 and 1250) identify specific MNase I sites in the region of the qrf promoter. (C) A consensus WCC binding site called the antisense light response element (aLRE) is located in the qrf promoter.
DNA methylation occurs at the frq promoter
Curiously, in the Southern blot data associated with the MNase I remodeling assay described in Figure S3, we routinely observed that a large percentage of the DNA isolated from Δchd1 was resistant to digestion with the restriction endonuclease NcoI. NcoI is sensitive to cytosine methylation, which suggested the possibility that frq DNA from Δchd1 is hypermethylated. Tests using other methylation-sensitive enzymes strongly supported this notion (Figure 4A–4D, and data not shown). This apparent DNA methylation at frq is surprising; in Neurospora, about 2% of cytosines are methylated, but in a variety of strains and under all conditions previously tested methylation has been shown to exist almost exclusively in relics of RIP (repeat-induced point mutation) and rDNA [37], [44], [45].
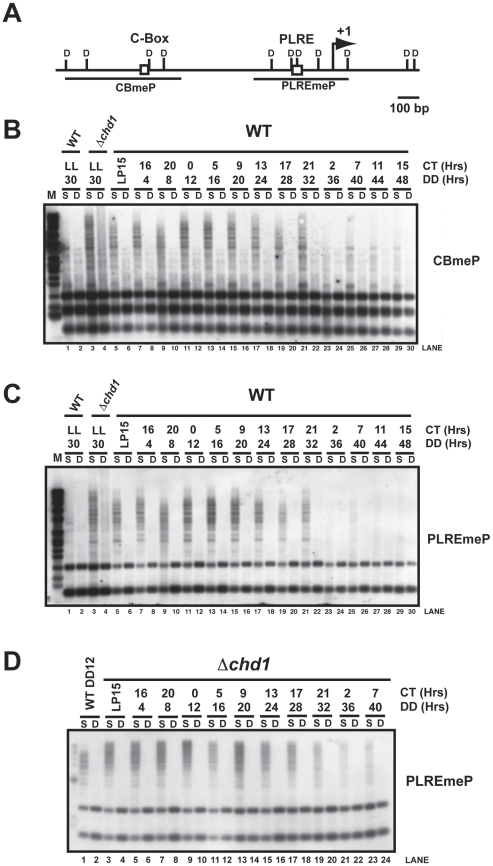
Figure 4. DNA methylation occurs at frq.
(A) Schematic diagram of the frq locus illustrating the location of the probes. (B) Genomic DNA was isolated from WT and Δchd1 strains grown in constant light (LL) at 30°C (Lanes 1–4) and compared to that from a WT strain grown in constant darkness (DD) for the indicated times over a 48-hrs time course. DNA was digested with Sau3AI (S) and DpnII (D) and probed with the C-box probe. (C) The blot represented in B was probed for the PLRE. (D) DNA was isolated from Δchd1 grown under standard circadian entrainment, handled as described and examined using the PLRE to detect DNA methylation.
A common assay for examining DNA methylation in Neurospora utilizes the methylation sensitive restriction endonuclease Sau3AI and its methylation insensitive isoschizomer, DpnII, followed by Southern blot analysis [46]. We first examined DNA methylation in WT and Δchd1 strains grown in constant light for 48 hrs at 30°C, which results in constitutive, non-rhythmic frq expression, and compared this with DNA methylation observed in WT and Δchd1 strains grown in constant darkness for two circadian cycles. Cultures were synchronized by a transfer from constant light at 25°C to constant darkness at 25°C and samples were collected every four hours (Figure 4). Southern blots were probed with DNA corresponding to the C-box (CBmeP) and PLRE (PLREmeP) to check for methylation at these sites (Figure 4A). The appearance of partially digested DNA in the Sau3AI digests suggested that the frq promoter is methylated (Figure 4B and 4C) under these conditions of growth in light followed by dark, and frq is hypermethylated in the Δchd1 strain relative to WT grown under the same conditions (Figure 4B and 4C, lanes 1–4). To ensure that inhibition of Sau3AI digestion at frq was due to DNA methylation, we routinely stripped and reprobed blots for a region of the mitochondrial genome (Figure S4 and data not shown). Mitochondrial DNA, which is not methylated, but is at much higher concentrations relative to nuclear DNA, was completely digested when compared with frq at similar time points (Figure S4). Interestingly, the DNA methylation at the frq promoter decreased with time in darkness, although not in an overtly circadian manner (Figure 4B and 4C, lanes 5–30). In addition, promoter methylation appeared greatly reduced in cultures grown in constant light for extended periods (48 hours in light). In a Δchd1 strain over the circadian time course frq promoter hypermethylation was seen at all time points examined (Figure 4D, compare lanes 1 and 2 with lanes 3–24).
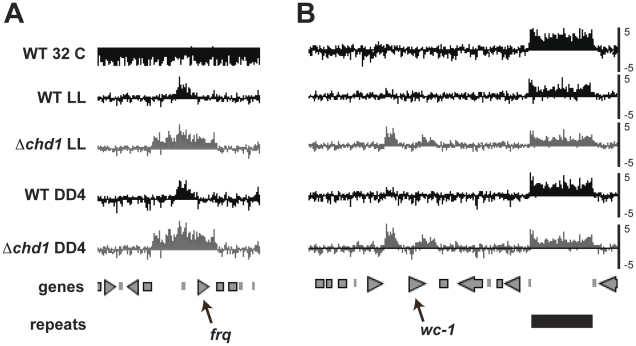
The unanticipated observation of DNA methylation at frq immediately suggested the possibility that other genetic loci were similarly modified and similarly affected by loss of chd1. To examine this we performed methylated DNA immunoprecipitation (MeDIP), followed by hybridization of differentially labeled input and MeDIP fractions to a high-density chromosome VII (LGVII) microarray (MeDIP-chip) [45]. MeDIP-chip experiments using genomic DNA from WT or Δchd1 cultures grown in constant light or in the dark for four hours revealed increased DNA methylation in the Δchd1 strain, consistent with Southern blot analyses. A peak of DNA methylation at the frq promoter was observed in the WT strain. In the Δchd1 strain, we observed an expanded domain of DNA methylation, which covered the entire frq promoter and open reading frame (Figure 5A). We next examined DNA methylation across the entire chromosome and found a peak of DNA methylation in the promoter of wc-1 in Δchd1 that was absent in the WT strain (Figure 5B). DNA methylation at wc-1 in Δchd1 was confirmed by a methyl-sensitive Southern blot, further supporting the validity of the MeDIP-Chip data (Figure S6). In contrast to frq and wc-1 where methylation is influenced by loss of CHD1, DNA methylation at RIP'd regions was similar in WT and Δchd1 (Figure 5B and data not shown), suggesting that CHD1 does not affect DNA methylation at constitutive heterochromatin. Because previous studies clearly showed a lack of methylation at the frq locus, it seems likely that particular growth conditions may be important to observe methylation and may indicate that it is dynamic [45]. Incidentally, when growth conditions consisting of elevated temperatures and extended light similar to previous work was used in this study, we failed to see significant methylation at frq (Figure 4B and 4C, lanes 1 and 2) supporting the notion that light-dark transitions or perhaps circadian entrainment are needed for methylation at frq.
Figure 5. DNA methylation at frq and wc-1.
Enrichment values for MeDIP experiments are shown at frq and wc-1 loci as log2 values (right). MeDIP data for WT and Δchd1 strains grown in constant light (LL) or for four hours in the dark (DD4) are shown below a plot of previously published MeDIP data obtained from WT grown at 32 degrees (WT 32) [45]. The positions of predicted genes are shown in grey and an annotated repeat near wc-1 is shown in black; frq and wc-1 are indicated.
We extended our analysis of the breadth of DNA methylation by examining more parts of the frq locus as well as other genes (Figure S5). Consistent with Figure 5, Southern hybridization confirmed that DNA methylation within the frq coding region is only seen in Δchd1, not WT. A third light and clock regulated gene, vvd [47] is similarly seen to be methylated in Δchd1 but not in WT (Figure S5E and S5F) but another light and clock controlled gene eas (ccg-2) appeared never to be methylated, even in Δchd1 (Figure S5C and S5D). Lastly, we examined the well-studied methylated Ψ63 region and found no discernable difference between WT and Δchd1 (Figure S5G and S5H), consistent with the microarray data, which revealed that CHD1 does not effect the strong methylation seen at relics of RIP.
Components of the clock impact normal methylation patterns
In considering common elements among the genes whose methylation is influenced by CHD1, it became clear that all are also regulated by light and by the components of the circadian clock. To examine this in more detail we followed DNA methylation in clock-defective strains (Figure 6, Figure S6). frq9 is a loss-of-function allele in which a frame-shift mutation results in premature termination; therefore, frq9 encodes a protein that is incapable of establishing circadian negative feedback [22]. Cultures containing frq9 were grown at 25°C in constant light and then transferred to darkness, and DNA was isolated and digested using the restriction endonucleases Sau3AI and DpnII followed by Southern blot analysis. We found a marked reduction of the slower migrating DNA in the Sau3AI digest compared with WT, indicative of hypomethylated DNA in frq9 (Figure 6A, compare lanes 1 and 2 with lanes 3–24). We next assayed how loss of FRH affected DNA methylation by using a strain that expresses an inducible hairpin RNA (dsfrh) that causes silencing of the frh gene [12]. Under conditions in which FRH expression was reduced, we found a noticeable decrease in DNA methylation at frq that was similar to that in the frq9 strain (Figure 6B). We conclude that normal levels of methylation require proper expression of core clock genes or at least FRQ-FRH dependent negative feedback, suggesting that both FRQ and FRH play an important, although perhaps indirect, role in regulating methylation at frq.
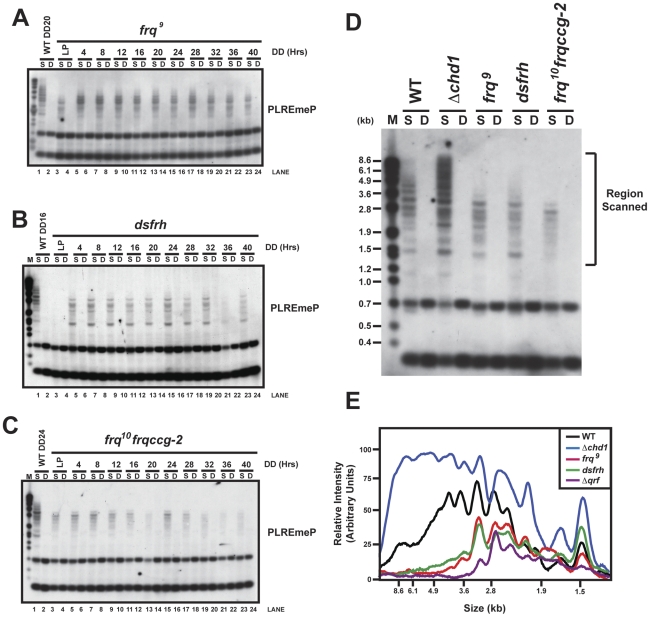
Figure 6. FRQ and the antisense transcript are required for normal DNA methylation.
(A)The methylation status of the frq promoter was examined in frq9, (B) a FRH knock-down strain dsfrh, and (C) a strain with reduced qrf expression, frq10frqccg-2. Strains were grown in light, transferred to darkness, samples collected at the indicated times, DNA was isolated and digested with Sau3AI (S) or DpnII (D) and processed for Southern hybridizations using a PLRE specific probe. WT DNA isolated from dark grown cultures was included as a control to compare differences in migration of methylated DNA. (D) To obtain a side-by-side comparison of the methylation pattern for the mutant strains used in this study, DNA obtained from the DD16 time point was taken for each strain and treated as in panel A, B and C. (E) The level of DNA methylation in the samples from (D) was quantified by performing densitometry on the region indicated in (D) and plotted versus the molecular weight marker. It is clear from this figure that Δchd1 is hypermethylated whereas frq9, dsfrh, and frq10frqccg-2 are all hypomethylated relative to WT.
An antisense frq transcript is also expressed from the frq locus [41]. Because antisense transcripts are known to be involved in X-chromosome inactivation (Xist/Tsix) [48], and because RNA-dependent DNA methylation (RdDM) requires formation of double stranded RNAs [49], we sought to examine the role of the frq antisense transcript in establishment of normal methylation. To test this, we used a strain frq10frqccg-2 [41] in which the 3′ region of frq, containing the promoter of this antisense transcript qrf, is replaced with the 3′ region of eas (ccg-2). This drastically reduces expression of the antisense transcript and results in a small phase lag in the oscillator [41]. In frq10frqccg-2, methylation is still present but is significantly reduced compared to WT (Figure 6C).
To further quantify the level of DNA methylation in WT and to allow direct comparison among Δchd1 and the clock mutant strains, we examined methylation at a single time point, DD16 (Figure 6D) and then performed a densitometric analysis on the methylated region (Figure 6E). We confirmed that the observed differences were not a result of the ras-1bd mutation that is present in frq9, dsfrh, and frq10frqccg-2, but not in WT or Δchd1. Because there was no significant difference in the methylation patterns of WT and ras-1bd, we conclude that ras-1bd has no apparent effect on the promoter methylation at frq (Figure S6B and S6C). The degree of methylation enhancement in Δchd1 and reduction in the strains bearing mutations in core clock components compared to WT is apparent. In the time course experiments, we routinely observed a reduction in the overall amount of methylated DNA at later time points (DD36 and beyond), which may suggest that the level of DNA methylation is reduced after prolonged incubations in the absence of light/dark cycles.
frq DNA is methylated by DIM-2
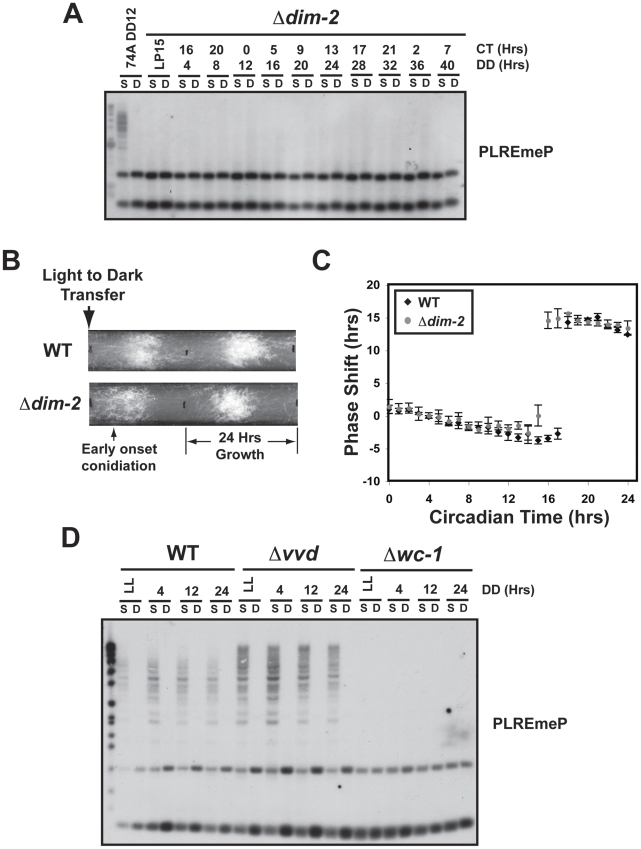
To better understand the possible role of DNA methylation in circadian regulated gene expression, we identified the methyltransferase responsible for this activity. Consistent with prior studies showing the DNA methyltransferase (DNMT) DIM-2 (defective in methylation) to be required for all observed DNA methylation in Neurospora [50], we found that Δdim-2 lacked all detectable methylation at frq (Figure 7A). This allowed us to examine the role of DNA methylation in circadian rhythmicity.
Figure 7. DIM-2 is required for frq methylation and effects the phase of the clock.
(A) DNA was isolated from a Δdim-2 strain and digested with Sau3AI (S) or DpnII (D) and probed with the PLRE probe. (B) Phenotype of Δdim-2 compared to WT grown on race tubes. Data are presented for two circadian cycles. (C) A phase-response curve comparing WT and Δdim-2 over one full circadian cycle. (D) DNA methylation analysis of Δvvd and Δwc-1 compared to WT grown under a limited circadian time course.
Circadian clock-regulated conidiation is still observed in Δdim-2, ras-1bd strains using race tubes, indicating that DNA methylation in not essential for rhythmicity per se (Figure 7B, Figure S7). There was, however, a slight phase-advance of approximately 2 hours in strains lacking DIM-2 (Figure 7B, Figure S7A). To confirm the phase-advance, we generated a Phase Response Curve (PRC) comparing Δdim-2 to WT (Figure 7C). Normally, a pulse of light will advance or delay the phase of the clock depending on the time when the pulse of light is given. In WT, the greatest shift in phase occurs around CT18; however, in Δdim-2 the largest shift was observed at CT15, further supporting the 2 hour phase-advance. A representative set of race tubes from the CT16 time point is shown in Figure S7B to highlight the phase shift difference between WT and Δdim-2. These results suggest that DNA methylation plays an ancillary but supportive role in clock regulation; namely, overt rhythms are not affected yet the onset of circadian clock-regulated expression is affected.
Further supporting the correlation between altered DNA methylation and altered circadian phase, we found that the vvd deletion strain, with its well-documented defects in clock phasing, also has an altered DNA methylation profile (Figure 7D). We note that the frq promoter is hypermethylated in Δvvd, further supporting the notion that normal DNA methylation is required for proper phasing. We also note that there is no DNA methylation in the Δwc-1 strain, consistent with a connection between normal expression of the frq/qrf locus and normal DNA methylation. The early onset of conidiation observed in Δdim-2 is consistent with the notion that DNA methylation is involved in transferring regulatory signals to pertinent clock genes thereby delaying the start of clock gene expression.
Discussion
We set out to understand how chromatin is remodeled at frq, and to investigate how this might help establish permissive and non-permissive states for rhythmic transcription. Our work led us to discover a link between CHD1-catalyzed chromatin remodeling, DNA methylation, and phase determination. We demonstrate that CHD1 contributes to apparent changes in chromatin structure at frq and is needed for normal frq expression. Surprisingly, we discovered DNA methylation at frq, providing the first example of promoter methylation in Neurospora, and we found that frq and other loci are hypermethylated in Δchd1 strains. We showed that DIM-2 is the DNMT responsible for methylating frq and that deletion of dim-2 results in a small phase defect.
Changes in DNA methylation at frq were also observed in Δvvd, and frq10frqccg-2, and both these strains have phase defects, providing a link between DNA methylation and phasing of clock gene expression. The apparent minor phase advance of approximately two hours in Δdim-2 is significant considering the circadian timescale. Under normal circadian conditions, the onset of frq expression occurs approximately 8 hours after the transition to dark conditions; therefore, a 2-hour advance represents a 25% shift. Moreover, there is probably a FRQ-induced refractory period after the transition to dark conditions where the WCC is inactive due to negative feedback caused by light-expressed FRQ-FRH. Thus WC-mediated transcription cannot occur until light-expressed FRQ is cleared by the proteasome, which takes roughly 4–6 hours. Based on expression profiles of frq in frq9 [22] and frh10 [51], we know expression of frq is suppressed upon transition to dark with a WT core-oscillator. Thus, under conditions where the negative elements are unaffected, the maximum observable phase advance should be approximately two hours, as found in the Δdim-2 strain.
In this report, we present data indicating that proper expression of both frq and its antisense counterpart, qrf, are essential for normal DNA methylation in this region. In fact, WC-1 dependent expression of frq, and its antisense noncoding counterpart qrf, clearly influences methylation. Interestingly, non-coding RNAs affect the methylation status of corresponding genes in other systems (i.e. mammals, plants) [52], [53], raising the possibility that the novel methylation that we report also depends on RNA. Further work is needed to explore this possibility but it is noteworthy that the mammalian clock gene mPer2 has promoter methylation [54], [55]. The hypermethylation phenotype observed in Δchd1 may result from improper expression of the sense/antisense pair or spurious transcripts, which would ultimately lead to defects in disiRNA (Dicer-independent small interfering RNA) production. A recent report indicates that in Neurospora sense/antisense transcripts produce disiRNA [56] and these may direct the DNA methylation through an unknown mechanism. Perhaps such RNAs could play a role in DNA methylation. An unrelated possibility is that CHD1 is involved in RNA processing of the qrf and/or 3′ end of the frq transcripts and this is coupled to a chromatin remodeling event whose misregulation leads in some way to the observed hypermethylation phenotype. A role for CHD1 in mRNA processing has been described in mammalian cell culture [57].
It is clear that CHD1 is needed for proper frq expression and does appear to remodel chromatin at frq. However, the role CHD1 plays in clock-regulated expression remains complicated by the residual remodeling that occurs in the absence of CHD1, and additional work is also needed to fully understand how DNA methylation influences frq expression. Regardless of the eventual role discovered for promoter methylation though, the simple fact that transient, apparently facultative, and presumably regulated DNA methylation occurs in the fungal kingdom is noteworthy. That this phenomenon went undiscovered for so long, but is so apparent in cultures grown at room temperature in light dark cycles, may serve to highlight the value of studying organisms under conditions that approximate the conditions in which they live in nature.
Materials and Methods
Strains and growth conditions
The wild-type (WT) Oak Ridge strain FGSC2489 (74A) strain or the clock WT strain, 328-4 (ras-1bd, A) were used as controls and parent strains for crosses. Strains generated in this study are shown in Table S1. The Δchd1 strain was generated by the knockout consortium [40], obtained from the Fungal Genetics Stock Center (FGSC, University of Missouri, Kansas City), and crossed to 74A generating XB99-1. Ascospores were germinated on complete media and genotyped by Southern blot for the hygromycin resistance cassette (Figure S1). The Δdim-2, ras-1bd strain, XB98-3, was generated by crossing FGSC8594 to 328-4 and spores were selected for hygromycin resistance and then screened on race tubes for the presence of the ras-1bd allele. For luciferase strains, the plasmid pVG110 [42], containing the full-length frq promoter fused to luciferase, was first transformed into rid− strains, FGSC9014 and FGSC9015, and targeted to the his-3 locus (Larrondo et al., in preparation). XB100-9 (Δchd1::hph, rid−, A) was crossed to FGSC9015 containing pVG110 and spores were selected on hygromycin, then screened for luminescence to obtain XB105-13. Growth media on race tubes consisted of 1X Vogel's salts, 0.1% glucose, 0.17% arginine [58]. Luciferase readings were measure using 12.5 µM luciferin as described [42].
The method used to generate the phase-response curve is outlined elsewhere [47]. Briefly, the phase response curve was generated using standard race tubes grown in the light for 12 or 24 hours and then transferred to dark conditions. After the third day of growth, individual race tubes were subjected to a 15 minute light pulse at the indicated circadian time and then placed back into a dark incubator for an additional 5–6 days (8 days of total growth). The phase on day 1 and on day 5 was determined using Chrono software and plotted as the difference of the two days.
The liquid culture assays were performed in media (2% LCM) containing 1X Vogel's salts, 0.17% arginine with 2% glucose and grown at 25°C. Conidia were used to seed mycelial mats in 75 mm Petri dishes and 0.5 cm plugs were cut from these and grown in 100 ml cultures. Mats were harvested by filtration, frozen in liquid nitrogen and ground in a mortar and pestle. Time course experiments have been described previously [22].
ChIP and chromatin-remodeling assays
The ChIP assays and oligonucleotides were identical to those previously described for WC-2 [18]. For MNaseI assays, nuclei were isolated in the dark as previously described [10], [13] and then subjected to limited MNaseI digestion following established protocols [59]. Equal amounts of nuclei were resuspended in MNaseI buffer A, treated with 1.0 ml 0.1 M CaCl2, incubated at 37°C for 1.5 minutes, and then varying amounts of MNaseI were added and the nuclei incubated for an additional 1.5 minutes. The enzyme was inactivated by the addition of Proteinase K (100 µg/ml) in TENS buffer (20 mM Tris-HCl, pH 7.4, 200 mM NaCl, 2 mM EDTA, 2% SDS), and the nuclei were incubated for a minimum of 2 hours to remove the chromatin. The DNA was purified by phenol chloroform extraction-EtOH precipitation and digested with NcoI as the secondary enzyme. DNA fragments were resolved on a 1.5% agarose gel, transferred to positively charged nitrocellulose, and probed with frqP4 specific probe. Remodeling at the antisense promoter was performed using the identical protocol except EcoRI was used as the restriction enzyme with frqP18. MeDIP-chip experiments were performed as previously described [45].
Northern, Southern, and Western blot
Standard Northern and Southern blots were performed using digoxigenin label DNA probes following Roche guidelines. All of the oligonucleotides used for probes are contain in Table S2. RNA was isolated from cells using a hot-phenol extraction or Trizol and 15 or 25 ug/ul of RNA were fractionated on a 1.3% agarose formaldehyde gel [22]. Quantitative RT-PCR was performed as described [60]. DNA was isolated using the Puragene Kit following the manufactures' protocol. Immunoblot analysis was as described [19].
Supporting Information
Circadian, growth, and molecular characterization of the Δchd1 strain. (A) WT 74A (FGSC 2489), ras-1bd (328-4) and Δchd1 were grown on race tubes. In addition to the genetic interaction with ras-1bd, the Δchd1 strain has an overall retarded growth phenotype. Confirmation of chd1 deletion strain by Southern blot using probes for chd1 (B) and hph (C). (D) Schematic of chd1 locus showing the relative locations of the probes.
(PDF)
CHD1 is required for normal WT circadian rhythms Expression of a frq-luc reporter construct was measured on race tubes in WT and Δchd1. (A) An actual race tube showing the areas used to obtain recordings. The luminescence was recorded for WT and Δchd1 isolates and plotted for the whole tube (B and C) and inoculation point (D and E). (F) Schematic representation of the sectional analysis used to monitor rhythms at the growth front. Light emitted from the area marked by each section was measured for the entire span of days required for strains to grow down the length of the tube. Bioluminescence levels are and plotted for WT (G) and Δchd1 (H). Each separate line traces the amount of light emitted by the culture within each small section. In G it is clear that cultures in each section remain rhythmic whereas in H, the Δchd1 strain shows no rhythm.
(PDF)
Analysis of remodeling in the frq promoter. We consistently observed subtle remodeling in the region surrounding the PLRE and TSS in a WT strain and this remodeling differed in Δchd1. (A) Schematic representation of the frq promoter highlighting the region around the PLRE and TSS. (B and D) Partial MNase I digestion probing the region around the PLRE and TSS. (C and E) Quantification of the remodeling observed in WT compared to Δchd1. The light-dependent remodeling that occurs at the TSS (Site 3) appears to be CHD1 independent. However, there is a slight defect in CHD1-dependent remodeling at the PLRE in the dark (Sites 1 and 2). Under normal WT conditions, frq is expressed in early subjective morning DD12 and the relative amount of site 1 is roughly half that of site 2 that we surmise is indicative of a more open chromatin state. At DD24 (and DD4, Panel D and E) in WT, a time when frq transcription is minimal, we see two predominant bands in the PLRE region, which are roughly at a 50∶50 ratio (site 1 and site 2) indicating a movement of nucleosomes to a more closed condensed state. In Δchd1 there does not appear to be a relative increase in Site 1 at the at the DD24 (or DD4, Panel D and E) time point and there is no discernable difference in Site 1 intensity relative to Site 2 between the DD12 and DD24 time points indicating a lack of chromatin remodeling at this site. Admittedly, this is difficult to observe because there is only a subtle change observed in this MNaseI assay, but the important point is that the relative intensity of site 1 to site 2 in Δchd1 is always lower compared to WT where the relative ratios of site 1 to site 2 are roughly equal at times when frq expression is low (DD4 and DD24); These data are further compounded by the large proportion of undigested DNA from Δchd1 nuclei (see methylation analysis). Yet, this experiment is very reproducible as the independent biological replicates show similar results.
(PDF)
A representative methylation Southern blot was probed for frq promoter DNA (PLREmeP) and then stripped and re-probed for a region of the mitochondrial genome (mitoP). WT DNA from two different time points (DD12 and DD16) were digested with Sau3AI and DpnII, resolved on a 1.0% agarose gel, transferred to nitrocellulose and probed as described in Material and Methods. To ensure complete cutting with Sau3A1, a 2X concentration is shown next to a standard amount.
(PDF)
DNA Methylation at other loci. WT and Δchd1 methylation southern blots were stripped and reprobed for the frq coding region (A and B), the ccg-2 promoter (C and D), the vvd promoter (E and F) and the ψ63 region (G and H).
(PDF)
DNA methylation at wc-1. (A) Data obtained from the MeDIP Chip experiment indicated that in addition to frq, wc-1 was also methylated in Δchd1. This was confirmed by a methylation sensitive Southern Blot using BfuCI (an isoschizomer of Sau3AI) and DpnII. (B and C) A side-by-side comparision was done to examine if there were any methylation difference in ras-1bd compared to an isogenic WT. It is clear there are no significant differences other than variations due to loading.
(PDF)
The Δdim-2 strain that has no DNA methylation retains a functional circadian clock but displays an altered phase. (A) Three independent isolates from a cross containing both Δdim-2 and ras-1bd were grown on race tubes. The phase on day 1 for each strain is shown at the right. (B) A sample racetube used to generate the phase response curve is shown. Note the large difference in phase shift on Day 6 between WT and Δdim-2.
(PDF)
Strains used in this study.
(PDF)
Oligonucleotides.
(PDF)
Acknowledgments
We thank Luis Larrondo for help with the luciferase experiments. We also thank members of the Dunlap, Loros, and Selker laboratories for interesting discussions throughout the course of this work.
Footnotes
The authors have declared that no competing interests exist.
This work was supported by NIH (www.NIH.gov): GM34985, GM68087 to JCD; GM083336 to JCD and JJL; M025690 to EUS; GM071223 to WJB; and by the Leukemia and Lymphoma Society (www.lls.org) 3295-10 to ZAL. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Bell-Pedersen D, Cassone VM, Earnest DJ, Golden SS, Hardin PE, et al. Circadian rhythms from multiple oscillators: lessons from diverse organisms. Nat Rev Genet. 2005;6:544–556. doi: 10.1038/nrg1633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Schibler U, Sassone-Corsi P. A web of circadian pacemakers. Cell. 2002;111:919–922. doi: 10.1016/s0092-8674(02)01225-4. [DOI] [PubMed] [Google Scholar]
- 3.Reppert SM, Weaver DR. Coordination of circadian timing in mammals. Nature. 2002;418:935–941. doi: 10.1038/nature00965. [DOI] [PubMed] [Google Scholar]
- 4.Heintzen C, Liu Y. The Neurospora crassa circadian clock. Adv Genet. 2007;58:25–66. doi: 10.1016/S0065-2660(06)58002-2. [DOI] [PubMed] [Google Scholar]
- 5.Brunner M, Schafmeier T. Transcriptional and post-transcriptional regulation of the circadian clock of cyanobacteria and Neurospora. Genes Dev. 2006;20:1061–1074. doi: 10.1101/gad.1410406. [DOI] [PubMed] [Google Scholar]
- 6.Cheng P, Yang Y, Gardner KH, Liu Y. PAS domain-mediated WC-1/WC-2 interaction is essential for maintaining the steady-state level of WC-1 and the function of both proteins in circadian clock and light responses of Neurospora. Mol Cell Biol. 2002;22:517–524. doi: 10.1128/MCB.22.2.517-524.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Crosthwaite SK, Dunlap JC, Loros JJ. Neurospora wc-1 and wc-2: transcription, photoresponses, and the origins of circadian rhythmicity. Science. 1997;276:763–769. doi: 10.1126/science.276.5313.763. [DOI] [PubMed] [Google Scholar]
- 8.Linden H, Macino G. White collar 2, a partner in blue-light signal transduction, controlling expression of light-regulated genes in Neurospora crassa. EMBO J. 1997;16:98–109. doi: 10.1093/emboj/16.1.98. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ballario P, Vittorioso P, Magrelli A, Talora C, Cabibbo A, et al. White collar-1, a central regulator of blue light responses in Neurospora, is a zinc finger protein. EMBO J. 1996;15:1650–1657. [PMC free article] [PubMed] [Google Scholar]
- 10.Froehlich AC, Liu Y, Loros JJ, Dunlap JC. White Collar-1, a circadian blue light photoreceptor, binding to the frequency promoter. Science. 2002;297:815–819. doi: 10.1126/science.1073681. [DOI] [PubMed] [Google Scholar]
- 11.Froehlich AC, Loros JJ, Dunlap JC. Rhythmic binding of a WHITE COLLAR-containing complex to the frequency promoter is inhibited by FREQUENCY. Proc Natl Acad Sci U S A. 2003;100:5914–5919. doi: 10.1073/pnas.1030057100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cheng P, He Q, Wang L, Liu Y. Regulation of the Neurospora circadian clock by an RNA helicase. Genes Dev. 2005;19:234–241. doi: 10.1101/gad.1266805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Luo C, Loros JJ, Dunlap JC. Nuclear localization is required for function of the essential clock protein FRQ. EMBO J. 1998;17:1228–1235. doi: 10.1093/emboj/17.5.1228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Diernfellner AC, Querfurth C, Salazar C, Hofer T, Brunner M. Phosphorylation modulates rapid nucleocytoplasmic shuttling and cytoplasmic accumulation of Neurospora clock protein FRQ on a circadian time scale. Genes Dev. 2009;23:2192–2200. doi: 10.1101/gad.538209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Schafmeier T, Haase A, Kaldi K, Scholz J, Fuchs M, et al. Transcriptional feedback of Neurospora circadian clock gene by phosphorylation-dependent inactivation of its transcription factor. Cell. 2005;122:235–246. doi: 10.1016/j.cell.2005.05.032. [DOI] [PubMed] [Google Scholar]
- 16.He Q, Cha J, He Q, Lee HC, Yang Y, et al. CKI and CKII mediate the FREQUENCY-dependent phosphorylation of the WHITE COLLAR complex to close the Neurospora circadian negative feedback loop. Genes Dev. 2006;20:2552–2565. doi: 10.1101/gad.1463506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Denault DL, Loros JJ, Dunlap JC. WC-2 mediates WC-1-FRQ interaction within the PAS protein-linked circadian feedback loop of Neurospora. EMBO J. 2001;20:109–117. doi: 10.1093/emboj/20.1.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Belden WJ, Loros JJ, Dunlap JC. Execution of the circadian negative feedback loop in Neurospora requires the ATP-dependent chromatin-remodeling enzyme CLOCKSWITCH. Mol Cell. 2007;25:587–600. doi: 10.1016/j.molcel.2007.01.010. [DOI] [PubMed] [Google Scholar]
- 19.Garceau NY, Liu Y, Loros JJ, Dunlap JC. Alternative initiation of translation and time-specific phosphorylation yield multiple forms of the essential clock protein FREQUENCY. Cell. 1997;89:469–476. doi: 10.1016/s0092-8674(00)80227-5. [DOI] [PubMed] [Google Scholar]
- 20.Baker CL, Kettenbach AN, Loros JJ, Gerber SA, Dunlap JC. Quantitative proteomics reveals a dynamic interactome and phase-specific phosphorylation in the Neurospora circadian clock. Mol Cell. 2009;34:354–363. doi: 10.1016/j.molcel.2009.04.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Tang CT, Li S, Long C, Cha J, Huang G, et al. Setting the pace of the Neurospora circadian clock by multiple independent FRQ phosphorylation events. Proc Natl Acad Sci U S A. 2009;106:10722–10727. doi: 10.1073/pnas.0904898106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Aronson BD, Johnson KA, Loros JJ, Dunlap JC. Negative feedback defining a circadian clock: autoregulation of the clock gene frequency. Science. 1994;263:1578–1584. doi: 10.1126/science.8128244. [DOI] [PubMed] [Google Scholar]
- 23.He Q, Cheng P, Yang Y, He Q, Yu H, et al. FWD1-mediated degradation of FREQUENCY in Neurospora establishes a conserved mechanism for circadian clock regulation. EMBOJ. 2003;22:4421–4430. doi: 10.1093/emboj/cdg425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.He Q, Cheng P, He Q, Liu Y. The COP9 signalosome regulates the Neurospora circadian clock by controlling the stability of the SCFFWD-1 complex. Genes Dev. 2005;19:1518–1531. doi: 10.1101/gad.1322205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Etchegaray JP, Lee C, Wade PA, Reppert SM. Rhythmic histone acetylation underlies transcription in the mammalian circadian clock. Nature. 2003;421:177–182. doi: 10.1038/nature01314. [DOI] [PubMed] [Google Scholar]
- 26.Curtis AM, Seo SB, Westgate EJ, Rudic RD, Smyth EM, et al. Histone acetyltransferase-dependent chromatin remodeling and the vascular clock. J Biol Chem. 2004;279:7091–7097. doi: 10.1074/jbc.M311973200. [DOI] [PubMed] [Google Scholar]
- 27.Etchegaray JP, Yang X, DeBruyne JP, Peters AH, Weaver DR, et al. The polycomb group protein EZH2 is required for mammalian circadian clock function. J Biol Chem. 2006;281:21209–21215. doi: 10.1074/jbc.M603722200. [DOI] [PubMed] [Google Scholar]
- 28.Ripperger JA, Schibler U. Rhythmic CLOCK-BMAL1 binding to multiple E-box motifs drives circadian Dbp transcription and chromatin transitions. Nat Genet. 2006;38:369–374. doi: 10.1038/ng1738. [DOI] [PubMed] [Google Scholar]
- 29.Doi M, Hirayama J, Sassone-Corsi P. Circadian regulator CLOCK is a histone acetyltransferase. Cell. 2006;125:497–508. doi: 10.1016/j.cell.2006.03.033. [DOI] [PubMed] [Google Scholar]
- 30.Hirayama J, Sahar S, Grimaldi B, Tamaru T, Takamatsu K, et al. CLOCK-mediated acetylation of BMAL1 controls circadian function. Nature. 2007;450:1086–1090. doi: 10.1038/nature06394. [DOI] [PubMed] [Google Scholar]
- 31.Asher G, Gatfield D, Stratmann M, Reinke H, Dibner C, et al. SIRT1 regulates circadian clock gene expression through PER2 deacetylation. Cell. 2008;134:317–328. doi: 10.1016/j.cell.2008.06.050. [DOI] [PubMed] [Google Scholar]
- 32.Nakahata Y, Kaluzova M, Grimaldi B, Sahar S, Hirayama J, et al. The NAD+-dependent deacetylase SIRT1 modulates CLOCK-mediated chromatin remodeling and circadian control. Cell. 2008;134:329–340. doi: 10.1016/j.cell.2008.07.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Katada S, Sassone-Corsi P. The histone methyltransferase MLL1 permits the oscillation of circadian gene expression. Nat Struct Mol Biol. 2010;17:1414–1421. doi: 10.1038/nsmb.1961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Jones MA, Covington MF, Ditacchio L, Vollmers C, Panda S, et al. Jumonji domain protein JMJD5 functions in both the plant and human circadian systems. Proc Natl Acad Sci U S A. 2010;107:21623–21628. doi: 10.1073/pnas.1014204108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ripperger JA, Shearman LP, Reppert SM, Schibler U. CLOCK, an essential pacemaker component, controls expression of the circadian transcription factor DBP. Genes Dev. 2000;14:679–689. [PMC free article] [PubMed] [Google Scholar]
- 36.Dubruille R, Murad A, Rosbash M, Emery P. A constant light-genetic screen identifies KISMET as a regulator of circadian photoresponses. PLoS Genet. 2009;5:e1000787. doi: 10.1371/journal.pgen.1000787. doi: 10.1371/journal.pgen.1000787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Suzuki MM, Bird A. DNA methylation landscapes: provocative insights from epigenomics. Nat Rev Genet. 2008;9:465–476. doi: 10.1038/nrg2341. [DOI] [PubMed] [Google Scholar]
- 38.Sargent ML, Woodward DO. Genetic determinants of circadian rhythmicity in Neurospora. J Bacteriol. 1969;97:861–866. doi: 10.1128/jb.97.2.861-866.1969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Belden WJ, Larrondo LF, Froehlich AC, Shi M, Chen CH, et al. The band mutation in Neurospora crassa is a dominant allele of ras-1 implicating RAS signaling in circadian output. Genes Dev. 2007;21:1494–1505. doi: 10.1101/gad.1551707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Colot HV, Park G, Turner GE, Ringelberg C, Crew CM, et al. A high-throughput gene knockout procedure for Neurospora reveals functions for multiple transcription factors. Proc Natl Acad Sci U S A. 2006;103:10352–10357. doi: 10.1073/pnas.0601456103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kramer C, Loros JJ, Dunlap JC, Crosthwaite SK. Role for antisense RNA in regulating circadian clock function in Neurospora crassa. Nature. 2003;421:948–952. doi: 10.1038/nature01427. [DOI] [PubMed] [Google Scholar]
- 42.Gooch VD, Mehra A, Larrondo LF, Fox J, Touroutoutoudis M, et al. Fully codon-optimized luciferase uncovers novel temperature characteristics of the Neurospora clock. Eukaryot Cell. 2008;7:28–37. doi: 10.1128/EC.00257-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Smith KM, Sancar G, Dekhang R, Sullivan CM, Li S, et al. Transcription factors in light and circadian clock signaling networks revealed by genomewide mapping of direct targets for Neurospora White Collar Complex. Eukaryot Cell. 2010;9:1549–1556. doi: 10.1128/EC.00154-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Selker EU, Tountas NA, Cross SH, Margolin BS, Murphy JG, et al. The methylated component of the Neurospora crassa genome. Nature. 2003;422:893–897. doi: 10.1038/nature01564. [DOI] [PubMed] [Google Scholar]
- 45.Lewis ZA, Honda S, Khlafallah TK, Jeffress JK, Freitag M, et al. Relics of repeat-induced point mutation direct heterochromatin formation in Neurospora crassa. Genome Res. 2009;19:427–437. doi: 10.1101/gr.086231.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Tamaru H, Selker EU. A histone H3 methyltransferase controls DNA methylation in Neurospora crassa. Nature. 2001;414:277–283. doi: 10.1038/35104508. [DOI] [PubMed] [Google Scholar]
- 47.Heintzen C, Loros JJ, Dunlap JC. The PAS protein VIVID defines a clock-associated feedback loop that represses light input, modulates gating, and regulates clock resetting. Cell. 2001;104:453–464. doi: 10.1016/s0092-8674(01)00232-x. [DOI] [PubMed] [Google Scholar]
- 48.Lee JT, Davidow LS, Warshawsky D. Tsix, a gene antisense to Xist at the X-inactivation centre. Nat Genet. 1999;21:400–404. doi: 10.1038/7734. [DOI] [PubMed] [Google Scholar]
- 49.Mette MF, Aufsatz W, van der Winden J, Matzke MA, Matzke AJ. Transcriptional silencing and promoter methylation triggered by double-stranded RNA. EMBO J. 2000;19:5194–5201. doi: 10.1093/emboj/19.19.5194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Kouzminova E, Selker EU. dim-2 encodes a DNA methyltransferase responsible for all known cytosine methylation in Neurospora. EMBO J. 2001;20:4309–4323. doi: 10.1093/emboj/20.15.4309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Shi M, Collett M, Loros JJ, Dunlap JC. FRQ-interacting RNA helicase mediates negative and positive feedback in the Neurospora circadian clock. Genetics. 2010;184:351–361. doi: 10.1534/genetics.109.111393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Ooi SK, O'Donnell AH, Bestor TH. Mammalian cytosine methylation at a glance. J Cell Sci. 2009;122:2787–2791. doi: 10.1242/jcs.015123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Feng S, Jacobsen SE, Reik W. Epigenetic reprogramming in plant and animal development. Science. 330:622–627. doi: 10.1126/science.1190614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Shih MC, Yeh KT, Tang KP, Chen JC, Chang JG. Promoter methylation in circadian genes of endometrial cancers detected by methylation-specific PCR. Mol Carcinog. 2006;45:732–740. doi: 10.1002/mc.20198. [DOI] [PubMed] [Google Scholar]
- 55.Ji Y, Qin Y, Shu H, Li X. Methylation analyses on promoters of mPer1, mPer2, and mCry1 during perinatal development. Biochem Biophys Res Commun. 391:1742–1747. doi: 10.1016/j.bbrc.2009.12.146. [DOI] [PubMed] [Google Scholar]
- 56.Lee HC, Li L, Gu W, Xue Z, Crosthwaite SK, et al. Diverse pathways generate microRNA-like RNAs and Dicer-independent small interfering RNAs in fungi. Mol Cell. 2010;38:803–814. doi: 10.1016/j.molcel.2010.04.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Sims RJ, 3rd, Millhouse S, Chen CF, Lewis BA, Erdjument-Bromage H, et al. Recognition of trimethylated histone H3 lysine 4 facilitates the recruitment of transcription postinitiation factors and pre-mRNA splicing. Mol Cell. 2007;28:665–676. doi: 10.1016/j.molcel.2007.11.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Dunlap JC, Loros JJ. Analysis of circadian rhythms in Neurospora: overview of assays and genetic and molecular biological manipulation. Methods Enzymol. 2005;393:3–22. doi: 10.1016/S0076-6879(05)93001-2. [DOI] [PubMed] [Google Scholar]
- 59.Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, et al. Chanda VB, editor. Current Protocols in Molecular Biology. 1994.
- 60.Chen CH, Ringelberg CS, Gross RH, Dunlap JC, Loros JJ. Genome-wide analysis of light-inducible responses reveals hierarchical light signalling in Neurospora. EMBO J. 2009;28:1029–1042. doi: 10.1038/emboj.2009.54. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Circadian, growth, and molecular characterization of the Δchd1 strain. (A) WT 74A (FGSC 2489), ras-1bd (328-4) and Δchd1 were grown on race tubes. In addition to the genetic interaction with ras-1bd, the Δchd1 strain has an overall retarded growth phenotype. Confirmation of chd1 deletion strain by Southern blot using probes for chd1 (B) and hph (C). (D) Schematic of chd1 locus showing the relative locations of the probes.
(PDF)
CHD1 is required for normal WT circadian rhythms Expression of a frq-luc reporter construct was measured on race tubes in WT and Δchd1. (A) An actual race tube showing the areas used to obtain recordings. The luminescence was recorded for WT and Δchd1 isolates and plotted for the whole tube (B and C) and inoculation point (D and E). (F) Schematic representation of the sectional analysis used to monitor rhythms at the growth front. Light emitted from the area marked by each section was measured for the entire span of days required for strains to grow down the length of the tube. Bioluminescence levels are and plotted for WT (G) and Δchd1 (H). Each separate line traces the amount of light emitted by the culture within each small section. In G it is clear that cultures in each section remain rhythmic whereas in H, the Δchd1 strain shows no rhythm.
(PDF)
Analysis of remodeling in the frq promoter. We consistently observed subtle remodeling in the region surrounding the PLRE and TSS in a WT strain and this remodeling differed in Δchd1. (A) Schematic representation of the frq promoter highlighting the region around the PLRE and TSS. (B and D) Partial MNase I digestion probing the region around the PLRE and TSS. (C and E) Quantification of the remodeling observed in WT compared to Δchd1. The light-dependent remodeling that occurs at the TSS (Site 3) appears to be CHD1 independent. However, there is a slight defect in CHD1-dependent remodeling at the PLRE in the dark (Sites 1 and 2). Under normal WT conditions, frq is expressed in early subjective morning DD12 and the relative amount of site 1 is roughly half that of site 2 that we surmise is indicative of a more open chromatin state. At DD24 (and DD4, Panel D and E) in WT, a time when frq transcription is minimal, we see two predominant bands in the PLRE region, which are roughly at a 50∶50 ratio (site 1 and site 2) indicating a movement of nucleosomes to a more closed condensed state. In Δchd1 there does not appear to be a relative increase in Site 1 at the at the DD24 (or DD4, Panel D and E) time point and there is no discernable difference in Site 1 intensity relative to Site 2 between the DD12 and DD24 time points indicating a lack of chromatin remodeling at this site. Admittedly, this is difficult to observe because there is only a subtle change observed in this MNaseI assay, but the important point is that the relative intensity of site 1 to site 2 in Δchd1 is always lower compared to WT where the relative ratios of site 1 to site 2 are roughly equal at times when frq expression is low (DD4 and DD24); These data are further compounded by the large proportion of undigested DNA from Δchd1 nuclei (see methylation analysis). Yet, this experiment is very reproducible as the independent biological replicates show similar results.
(PDF)
A representative methylation Southern blot was probed for frq promoter DNA (PLREmeP) and then stripped and re-probed for a region of the mitochondrial genome (mitoP). WT DNA from two different time points (DD12 and DD16) were digested with Sau3AI and DpnII, resolved on a 1.0% agarose gel, transferred to nitrocellulose and probed as described in Material and Methods. To ensure complete cutting with Sau3A1, a 2X concentration is shown next to a standard amount.
(PDF)
DNA Methylation at other loci. WT and Δchd1 methylation southern blots were stripped and reprobed for the frq coding region (A and B), the ccg-2 promoter (C and D), the vvd promoter (E and F) and the ψ63 region (G and H).
(PDF)
DNA methylation at wc-1. (A) Data obtained from the MeDIP Chip experiment indicated that in addition to frq, wc-1 was also methylated in Δchd1. This was confirmed by a methylation sensitive Southern Blot using BfuCI (an isoschizomer of Sau3AI) and DpnII. (B and C) A side-by-side comparision was done to examine if there were any methylation difference in ras-1bd compared to an isogenic WT. It is clear there are no significant differences other than variations due to loading.
(PDF)
The Δdim-2 strain that has no DNA methylation retains a functional circadian clock but displays an altered phase. (A) Three independent isolates from a cross containing both Δdim-2 and ras-1bd were grown on race tubes. The phase on day 1 for each strain is shown at the right. (B) A sample racetube used to generate the phase response curve is shown. Note the large difference in phase shift on Day 6 between WT and Δdim-2.
(PDF)
Strains used in this study.
(PDF)
Oligonucleotides.
(PDF)